Abstract
BCR-ABL-positive acute myeloid leukemia (AML) is a rare subtype of AML that is now included as a provisional entity in the 2016 revised WHO classification of myeloid malignancies. Since a clear distinction between de novo BCR-ABL+ AML and chronic myeloid leukemia (CML) blast crisis is challenging in many cases, the existence of de novo BCR-ABL+ AML has been a matter of debate for a long time. However, there is increasing evidence suggesting that BCR-ABL+ AML is in fact a distinct subgroup of AML. In this study, we analyzed all published cases since 1975 as well as cases from our institution in order to present common clinical and molecular features of this rare disease. Our analysis shows that BCR-ABL predominantly occurs in AML-NOS, CBF leukemia, and AML with myelodysplasia-related changes. The most common BCR-ABL transcripts (p190 and p210) are nearly equally distributed. Based on the analysis of published data, we provide a clinical algorithm for the initial differential diagnosis of BCR-ABL+ AML. The prognosis of BCR-ABL+ AML seems to depend on the cytogenetic and/or molecular background rather than on BCR-ABL itself. A therapy with tyrosine kinase inhibitors (TKIs) such as imatinib, dasatinib, or nilotinib is reasonable, but—due to a lack of systematic clinical data—their use cannot be routinely recommended in first-line therapy. Beyond first-line treatment of AML, the use of TKI remains an individual decision, both in combination with intensive chemotherapy and/or as a bridge to allogeneic stem cell transplantation. In each single case, potential benefits have to be weighed against potential risks.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The Philadelphia chromosome, corresponding to the BCR-ABL rearrangement, is the cytogenetic hallmark of chronic myelogenous leukemia (CML) and is frequently found in high-risk acute lymphoblastic leukemia (ALL). In addition, 0.5–3 % of all acute myelogenous leukemia (AML) cases also carry this fusion gene [1–5].
The distinction between this rarely occurring BCR-ABL+ AML and CML in (primary) blast crisis can be difficult, and the diagnostic challenges have led to an ongoing debate as to whether BCR-ABL+ AML does actually exist as a distinctive subgroup of AML. During the past, numerous case reports on BCR-ABL+ AML have been published, diminishing further doubts on its existence.
In this study, all published cases of BCR-ABL+ AML since 1975 as well as cases from our institution were analyzed with respect to their clinical and molecular features. Furthermore, a diagnostic algorithm for this rare but challenging problem in clinical practice will be suggested.
Patients and methods
References were primarily obtained by a PubMed literature search. Cross-references were used to obtain additional cases.
Cases from the literature were collected starting in 1975; cases from our own institution were available in a database that begins in 1998. In total, 164 cases of de novo BCR-ABL+ acute myeloid leukemia and 14 cases of relapsed BCR-ABL+ AML were analyzed. For a better comparability between the cases over this long period of time, published cases were re-classified (if necessary and possible) according to the latest WHO classification [6] or, in the case of acute undifferentiated leukemia (AUL)/mixed phenotype acute leukemia (MPAL), according to EGIL and/or WHO classification. Since classification was based on the EGIL criteria in the majority of cases, a re-classification according to WHO 2016 could not always be made due to a lack of information. Cases re-classified to AUL according to EGIL or to MPAL according to WHO were excluded from the analysis, since MPAL with BCR-ABL re-arrangement is an own entity within MPAL in WHO classification. Eight cases from the literature and two cases from our institution were excluded after being re-classified as AUL/MPAL according to the immunophenotype reported. Fifteen additional cases from the literature were excluded since they were BCR-ABL+ AUL according to the authors’ own statements.
Eight cases with 100 % BCR-ABL-positive metaphases, published before 1990, were excluded from the analysis because no additional features were described helping to distinguish AML from CML. One additional case was excluded because the patient had an antecedent myeloproliferative disorder, and four cases were excluded since no information was given that helped to distinguish de novo BCR-ABL+ AML from CML blast crisis. Cases describing secondary AML with an antecedent MDS are included and analyzed as AML with myelodysplasia-related changes. Finally, data on epidemiology, molecular pathogenesis, and phenotype of BCR-ABL+ AML are based on 126 cases of de novo BCR-ABL + AML, fulfilling the above-mentioned criteria. A detailed summary of all cases is given in the supplementary tables.
Results
Epidemiology
The Philadelphia chromosome is supposed to be a rare event in acute myeloid leukemia. Its reported incidence ranges from 0.5 to 3 % [1–5]. Most cases are described as de novo AML, few authors report on a new acquisition of the Philadelphia chromosome upon disease relapse or transformation of an antecedent myelodysplastic syndrome into AML. In contrast, BCR-ABL is much more common in acute leukemia of ambiguous lineage and thus was classified as an own subgroup in WHO classification [6].
So far, the overall incidence is mainly estimated from larger case series without any standardized definition. The accuracy will soon improve since several large study groups are acquiring these data in a joint effort to improve the molecular characterization of AML. Currently, a precise distinction between AML and CML in blast crisis is still limited by the lack of a clear definition; however, CML in primary blast crisis can be considered an extremely rare event and patients with secondary blast crisis can usually be identified by their medical history.
Molecular pathogenesis and phenotype
By definition, AML is a clonal disease. In comparison to most solid cancers, AML shows a limited spectrum of cytogenetic aberrations and mutations with different roles in pathogenesis [7–9].
Basic features of leukemogenesis according to the model of Gilliland and Griffin [10] are inhibition of differentiation and apoptosis (caused by a class II mutation as the leukemia initiating event) and the subsequent acquisition of a proliferative advantage (due to class I mutations). Typical examples are PML-RARA as a class II event and FLT3 or KIT mutations as class I events [7, 10]. Applying this model, BCR-ABL belongs to the group of class I mutations, conferring a proliferative advantage to the affected cells [11].
There is increasing evidence that the presence of age-dependent clonal hematopoiesis is a risk factor for the development of hematological malignancies [12, 13]. It is well known that BCR-ABL may also temporarily occur at very low levels in the blood of healthy individuals [14–17]. The co-incidence of spontaneously occurring BCR-ABL within clonal hematopoiesis could possibly explain the development of a BCR-ABL+ AML and the underlying “mutational background”; the maturation stage of the affected cell might decide whether the additional acquisition of BCR-ABL leads to AML, CML, or just a very small and only temporarily detectable BCR-ABL+ subclone.
BCR-ABL1 has been described in AML together with different class II aberrations such as CBFB-MYH11, RUNX1-RUNX1T1, PML-RARA, and NPM1 [1, 11, 18–35].
Based on the aforementioned model of AML pathogenesis, possible interplays of BCR-ABL with co-occurring genetic events are conceivable but the proof-of-principle experiment in an appropriate pre-clinical model is missing due to the lack of BCR-ABL+ AML models. The generation of a BCR-ABL+ AML cell line or the overexpression of various known genes together with BCR-ABL in hematopoietic precursor cells could be the first step to improve our understanding of molecular interactions during pathogenesis.
Recently, two major clonal evolution patterns during AML relapse were defined based on a genome-wide sequencing approach of several AML cases [36]. The first pattern implicates that the founding clone of the disease gains mutations and evolves into the relapse clone(s). The second suggests that a subclone of the initial clone survives therapy, gains additional mutations, and expands at relapse. In both models, a founding mutation is followed by a cooperating mutation able to convey a proliferative advantage [36]. Interestingly, mutations in genes involved in epigenetic regulation appear to occur early in the evolution of AML [9]. In view of this model, the acquisition of BCR-ABL is an interesting option to explain leukemia relapse and will be discussed later.
Interestingly, Nacheva EP et al. [37] recently showed that BCR-ABL+ AML displays characteristics of lymphoid disease as it shares deletions of IKZF and CDKN2A/B as well as the concomitant loss within the immunoglobulin and T cell receptor gene complexes that are all also found in BCR-ABL+ ALL and CML. Remarkably, all these aberrations have been found to be absent in myeloid blast crisis of CML [38], possibly adding a helpful tool for differential diagnosis in difficult cases.
However, larger data sets are needed before these biomarkers can be integrated into routine differential diagnosis. Confirmation of this intriguing finding would provide further evidence for a different and possibly unique biology of BCR-ABL+ AML, namely the appearance of lymphoid genomics in the guise of a myeloid phenotype.
A possible role of BCR-ABL within AML pathogenesis is depicted in Fig. 1.
Characteristic features of BCR-ABL+ AML
Regarding the WHO 2016 classification, most cases of BCR-ABL+ AML belong to the subgroup AML-NOS (38 %, corresponding to 48/126 of de novo BCR-ABL+ AML cases), followed by AML with myelodysplasia-related changes (32.5 %, 41/126 cases) and AML with recurrent genetic aberrations (in particular CBF leukemia with 16.7 %, corresponding to 21/126 cases). It seems very likely that the accumulation of cases in the subgroup AML-NOS may be caused by incomplete cytogenetic and/or molecular analyses, particularly in older case reports. Considering coinciding cytogenetic/molecular events, CBF leukemia with inv(16) or t(8;21) and NPM1 mutations are reported by several authors [1, 11, 18–35]:
-
1)
BCR-ABL in CBF-AML
Interestingly, 30 cases (23.8 %) of the 126 collected de novo BCR-ABL+ AML cases were AML with recurrent genetic alterations, mainly CBF leukemias in 21/126 cases. The most common recurrent genetic alteration was inv(16) in 17 cases; only one case was reported as promyelocytic leukemia carrying the t(15;17) translocation and three cases with the translocation t(8;21). This accumulation of “good risk” phenotypes (irrespective of BCR-ABL) is remarkable, since the vast majority of cases with BCR-ABL+ AML belongs to the intermediate or high-risk group according to ELN. On the one hand, this may explain why BCR-ABL+ AML is perceived as a “high risk” AML; on the other hand, it clearly shows that BCR-ABL may also arise in a “good risk” molecular background (see Fig. 2c).
However, the occurrence of inv(16) is not restricted to AML. It can also be found in blast crisis of CML [reviewed in 18, 39]. Therefore, a clear distinction of de novo AML and CML blast crisis remains challenging even in these inv(16)+ cases. The distribution of BCR-ABL p190 and p210 in inv(16)-CBF-AML was notably different from CML blast crisis: 53 % (9 cases) carried the p190 transcript in AML, whereas p190 was only found in one of 19 published cases of CML-AP/BC with inv(16) [18, 19, 39–48]. Therefore, the detection of the minor BCR-ABL transcript in AML with inv(16) might be helpful in identifying de novo BCR-ABL+ AML. Furthermore, inv(16) is a characteristic class II event and therefore present in every leukemic clone in AML, whereas it is mostly not in blast crisis of CML.
In AML with t(8;21), two thirds of cases were also p190 positive underlining the aforementioned hypothesis [11, 31].
Whether the overall prognosis of this favorable AML subgroup is worsened by BCR-ABL cannot be determined due to the small number of patients, but several long-term survivors are reported in the cohort of BCR-ABL+ CBF leukemia.
-
2)
BCR-ABL+ AML with concomitant mutations
Considering the 126 cases of de novo BCR-ABL+ AML, 6 had an additional NPM1 mutation at primary diagnosis [1, 11, 32–34]. In total, information on NPM1 mutational status was given in 15 cases. Notably, 3 of these 6 NPM1+ AML patients were long-term survivors, and for the remaining 3 cases, no survival data were available. This may again show that the overall prognosis of AML is not determined by BCR-ABL alone. However, the small number of cases does not allow any final conclusion.
The FLT3 mutational status was reported in 13 cases with two of them being positive (one FLT3-ITD, one FLT3-TKD) [1, 49]. This is remarkable, since it shows that two class I mutations, each conferring a proliferative advantage, can be acquired subsequently.
-
3)
Acute myeloid leukemia with myelodysplasia-related changes
Eighteen cases of MDS that became BCR-ABL positive upon transformation into secondary AML were reported [3, 11, 50–60]. In nine cases, information regarding the type of the BCR-ABL transcript was available. In 17 % of all AML cases (3/18), expression of p190 was reported; another 22 % (4/18) carried the p210 transcript. In 11 % (2/18) both transcripts were detectable. Fifty percent (9/18) of these patients had a complex-aberrant karyotype at the time of BCR-ABL acquisition. In most cases, BCR-ABL was documented for the first time upon transformation into AML (the same finding as reported by [61]).
The WHO classification of 2008 and its 2016 revision include cytogenetic criteria for the diagnosis of “acute myeloid leukemia with myelodysplasia-related changes” [6]. According to these criteria, 23 additional cases can be classified as AML with myelodysplasia-related changes although a history of documented MDS is not reported [1, 4, 30, 34, 61–66]. Therefore, AML with myelodysplasia-related changes represents the second largest subgroup with 41 cases in total after AML-NOS.
-
4)
Therapy-related BCR-ABL+ AML
Five cases of therapy-related BCR-ABL+ AML were described, two more cases can possibly be considered as “therapy-related”, but a lack of information prevents a definitive classification [31, 35, 60, 67–70]. No correlation with radiation or a specific therapy-regimen can be established due to the low number of cases.
-
5)
BCR-ABL at AML relapse
In 14 published cases, BCR-ABL was acquired upon AML relapse [11, 55, 71–77]. In 7 of these 14 patients, the morphologic classification was given. Four cases were classified as AML with maturation (FAB-M2), one case as AML with minimal differentiation (AML-FAB M0), and another case as myelomonocytic AML (AML-FAB M4). In these cases of BCR-ABL+ AML, the p190 transcript was reported in seven cases (one case expressed p210; no information was given in six cases). These data underline p190 as the predominant BCR-ABL transcript in AML.
Ten of these 14 patients succumbed to their disease or infectious complications; no outcome was reported for the remaining cases. At relapse, most patients acquired further, mostly high-risk cytogenetic aberrations such as deletions of chromosome seven. Therefore, the unfavorable outcome is not necessarily due to BCR-ABL alone; however, BCR-ABL may contribute to an increased genetic instability.
-
6)
BCR-ABL+ mixed phenotype acute leukemia
The occurrence of a Philadelphia chromosome is a frequent event in acute leukemia of ambiguous lineage as defined by the WHO classification [6].
In a larger case series, 16 % of the evaluated MPAL patients had a BCR-ABL rearrangement [78]. In a detailed analysis by the German ALL group, 30 % of their biphenotypic acute leukemia (BAL) patients were BCR-ABL positive [79]. In another case series, Afty et al. [80] found a shorter survival in the BCR-ABL+ BAL as compared to BCR-ABL+ AML. Furthermore, patients with BAL were older than patients with AML. None of the patients of this case series was treated with a tyrosine kinase inhibitor.
Clinical features and diagnosis
Little is known about characteristic clinical features of BCR-ABL+ AML. In general, its nature as a high-risk leukemia is assumed although there are no systematic data on outcome. Thus, the collected cases were analyzed regarding their clinical features.
As mentioned above, the distinction between BCR-ABL+ AML and CML in primary myeloid blast crisis can be difficult. Although both diseases are rare events, differential diagnosis is of major importance in order to choose the most appropriate therapy (intensive induction chemotherapy vs. tyrosine kinase inhibitor followed by an early allogeneic stem cell transplant).
The first step of differential diagnosis is a thorough history and physical examination: Has the patient ever presented before with unexplained leukocytosis? Does the patient have an antecedent blood disorder or a hepatosplenomegaly?
Splenomegaly is one of the clinical hallmarks of CML. Among our 126 cases of de novo BCR-ABL+ AML, we only found 16.7 % of cases with splenomegaly (corresponding to 21/126 cases); the absence of splenomegaly was explicitly stated in 28.6 % of all cases (36/126), whereas no definitive statement was given in 54.7 % (69/126) suggesting that in most of these cases, splenomegaly was at least not a prominent clinical feature. Thus, the absence of relevant splenomegaly seems to support the diagnosis “de novo BCR-ABL+ AML” rather than “CML blast crisis.” This is underlined by a recent publication by Soupir CP et al. [4].
Another hallmark of CML is the concomitant basophilia of ≥2 % WBC. This feature was only present in 5.6 % (7/126 cases) of BCR-ABL+ AML cases. 57.2 % (4/7) of these cases with basophilia were accompanied by splenomegaly and 85.7 % (6/7) by 100 % BCR-ABL-positive metaphases on cytogenetic analysis. Thus, cases with basophilia may occur in both, de novo AML and CML-BC, but are definitely rare. Of note, some authors had already excluded questionable cases with basophilia (e.g. [1]) so that our data might underrepresent cases with basophilia. Nevertheless, it seems appropriate to conclude that the absence of basophilia supports the diagnosis of de novo BCR-ABL+ AML.
In 32.5 % (41/126) of all cases, the immunophenotype was reported. Remarkably, co-expression of lymphoid markers was reported in 34.2 % of these cases (14/41), but was not sufficient for a classification as MPAL or BAL according to WHO or EGIL respectively. This is in line with a recent finding by Atfy et al. reporting an aberrant lymphoid co-expression in seven out of nine BCR-ABL+ AML cases [80].
For the collected 126 patients, the FAB classification was reported in 79 cases. The most common subtypes were M4eo, M0, and M1, followed by M7, M4, and M2; M3 was only seen once. For further information regarding FAB and WHO classification, see Fig. 2a, b.
Interestingly, megakaryoblastic AML (FAB M7) was reported in approximately 6.3 % of AML-NOS cases in contrast to less than 1 % in a population of AML-NOS [81].
Bacher and colleagues [11] as well as Berger [5] also suggested that the presence of the Philadelphia chromosome in less than 100 % of metaphases during conventional karyotyping is the major criterion for the diagnosis of BCR-ABL+ AML, since CML-BC/AP is characterized by the presence of the Philadelphia chromosome in virtually all metaphases. This is due to the fact that CML is an early progenitor disease carrying the key cytogenetic abnormality in all cells.
Since 70 % of all cases with convincing de novo BCR-ABL+ AML in our collection express the Philadelphia-chromosome in 100 % of all metaphases, we suggest a modified use of this criterion: In the absence of basophilia, typical AML-like co-aberrations such as a complex-aberrant karyotype or a deletion of chromosome 7 may nevertheless lead to the diagnosis of AML.
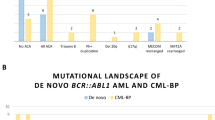
In our case series, the p190 transcript was found in 27.8 % (35/126) of all cases, p210 in 30.9 % (39/126), both transcripts in 2.4 % (3/126), e6a2 in 1.6 % (2/126), and no information was given in 37.3 % (47/126) of all cases (see Fig. 3). Since p190 is very rare in CML, the presentation with a p190 transcript is in favor of the diagnosis “AML” rather than “CML.” Thus, the type of transcript can sometimes help to distinguish AML from CML as suggested [5, 11, 82].
The overall prognosis of BCR-ABL+ AML is generally assumed to be unfavorable. Thus, the NCCN Clinical Practice Guidelines in Oncology for AML [83] include this entity into the poor-risk group, comparable with complex-aberrant karyotype AML. The ELN risk classification [84] does not include BCR-ABL+ AML so far. Hence, we determined the prognostic subgroup of all collected cases according to ELN criteria [84] independently of BCR-ABL: 21.4 % (27/126) were classified as favorable, 39.7 % (50/126) belonged to the intermediate-I and intermediate-II risk groups, 30.2 % of the patients (38/126) had an adverse prognosis, and for 8.7 % (11/126) of the patients, no information was available (see Fig. 2c). Therefore, the majority of cases—irrespective of BCR-ABL—already belonged to the less favorable ELN risk groups. Thus, the influence of BCR-ABL itself can hardly be determined. The same accounts for the favorable risk group, but it is at least remarkable that a considerable part of these patients showed a good response to therapy with long-term survival without allogeneic stem-cell transplantation being reported (e.g., in CBF leukemia/inv(16): 5/17 patients survived without stem cell transplantation, 4/17 patients who did not receive a SCT died, 3/17 patients who received SCT died, 3/17 patients survived after SCT, and for 2 patients, no survival data were available). In conclusion, BCR-ABL alone is not a valuable prognostic parameter so far. It seems more likely that the prognosis of BCR-ABL+ AML rather depends on the genetic background (concurrent aberrations) than on BCR-ABL itself [25].
Treatment of BCR-ABL+ AML
Currently, no standardized treatment for BCR-ABL+ AML exists. Considering the small number of cases, a randomized trial testing different treatment regimens can hardly ever be expected. Furthermore, an evaluation of the treatment response to tyrosine kinase inhibitors (TKIs) is complicated by the fact that the patients concomitantly received chemotherapy and allogeneic bone marrow transplants. In addition, many cases were published before TKIs were integrated into clinical practice. For the future, a world-wide registry would be helpful to address the efficacy of imatinib and other second- or third-generation TKIs in this rare disease.
In our case collection, 24.6 % of de novo BCR-ABL+ AML patients received a therapy with a TKI (imatinib: 29/31 TKI-treated cases; dasatinib: 4 cases; nilotinib: 1 case; some of the patients received a sequential therapy with more than one TKI). Only in two patients, a severe adverse event was reported leading to a withdrawal of the TKI (hepatotoxicity and diarrhea CTC grade 3 upon imatinib treatment [85]).
In relapsed BCR-ABL+ AML, seven patients received a tyrosine kinase inhibitor (imatinib: five cases, dasatinib: one case and nilotinib: two cases; some of the patients received a sequential therapy with two different TKIs). One patient achieved a CCyR, but died of a brain abscess, two patients had a transient response [73, 76], and two patients had no response [66, 75]. Interestingly, in three patients, the TKI treatment led to an eradication of the Ph-positive clone despite an overall refractory disease [65, 74, 77]. This clearly shows the clonal diversity of BCR-ABL+ AML in contrast to chronic phase CML, which is strictly addicted to BCR-ABL. In summary, therapy with a TKI alone does not induce sustained responses in BCR-ABL+ AML, although such a response was occasionally reported [e.g., 86]. This may be due to a very rapid clonal evolution, leading to resistance in a much higher proportion of patients and in a substantially shorter time than in CML. Furthermore, in contrast to CML, BCR-ABL is not supposed to be the key driver of the disease, but nevertheless might temporarily provide a proliferative advantage to a particular BCR-ABL+ subclone. However, as most reports show the eradication of the BCR-ABL+ subclone in AML, TKI therapy alone does not seem to control the disease, possibly owing to the fact that BCR-ABL is not a driving mutation in the founder clone. These aspects emphasize the importance of a differentiation between BCR-ABL+ AML and CML blast crisis: Whereas TKI therapy without chemotherapy is at least an option for remission induction in CML blast crisis before allogeneic stem cell transplantation (SCT), intensive induction chemotherapy remains the cornerstone of curative AML therapy and is usually followed by consolidation with chemotherapy or allogeneic stem cell transplantation according to ELN risk stratification, age, comorbidities, performance status, and donor availability [9, 87, 88]. Furthermore, allogeneic SCT is the standard of care for CML blast crisis after remission induction in eligible patients, whereas in the light of our current study, the need for SCT can at least be discussed in all cases of BCR-ABL+ AML belonging to the favorable ELN risk group.
Although the addition of a TKI to standard chemotherapy for BCR-ABL+ AML is reasonable and probably safe, we believe that there are no sufficient clinical data in support of a routine use in addition to chemotherapy during first-line therapy. The use of a TKI alone or in combination with chemotherapy in relapsed AML patients remains an individual decision, but it should not defer standard salvage therapy including allogeneic stem cell transplantation in eligible patients. Treatment of BCR-ABL+ AML is also reviewed elsewhere [4, 34, 85].
Conclusions and discussion
During the past years, an increasing number of data has accumulated which argues in favor of the existence of BCR-ABL+ AML. Therefore, BCR-ABL+ AML is now included as a new provisional entity in the 2016 revised WHO classification of hematopoietic malignancies [6]. In AML, BCR-ABL seems to cooperate with several AML-specific aberrations such as inv(16) and myelodysplasia-related cytogenetic aberrations; however, the molecular mechanism of both disease initiation and cooperation with other aberrations remains largely unknown. A better molecular characterization of BCR-ABL+ AML will always be hampered by its very low incidence. Furthermore, routine cytogenetic testing might lead to an underestimation of its real incidence since less dominant small BCR-ABL+ subclones could be missed and PCR for BCR-ABL is not routinely performed in AML. Thus, a BCR-ABL rearrangement might not be found upon routine analysis.
In clinical routine diagnostics, the distinction between de novo BCR-ABL+ AML and CML in blast crisis remains challenging in every single case. Therefore, we have generated the following algorithm for primary differential diagnosis which relies on a thorough analysis of all published cases (see Fig. 4).
Briefly, after exclusion of acute leukemia of ambiguous lineage by flow cytometry (which constitutes a separate entity according to WHO), the patient’s medical history with regard to an antecedent hematologic disease is of major importance. In particular, unexplained leukocytosis and/or splenomegaly rather point towards the diagnosis of CML blast crisis than AML. In contrast, prior signs of MDS in peripheral blood or bone marrow may support the diagnosis of (secondary) AML. Furthermore, basophilia or significant splenomegaly at diagnosis strongly diminishes the probability of de novo AML. Detection of p190-transcript and occurrence of any BCR-ABL transcript in less than 100 % of metaphases supports the diagnosis of AML rather than CML. Finally, persistent CCyR after conventional chemotherapy is extremely unusual for CML in blast crisis and supports the diagnosis of AML.
In summary, the following basic requirements for diagnosis of BCR-ABL+ AML are recommended:
-
1)
No antecedent hematological anomaly
-
2)
No basophilia upon disease onset
Our proposed diagnostic algorithm is depicted in Fig. 4.
With regard to therapy of BCR-ABL+ AML, we feel that TKI therapy cannot be routinely recommended as a part of first-line therapy. However, TKI as a part of salvage therapy is a reasonable approach, although any prognostic benefit remains unproven. Furthermore, it is not clear whether to use imatinib, dasatinib, or nilotinib. For bosutinib and ponatinib, no data are available. If indicated, we would choose the most appropriate TKI by taking into account the patient’s comorbidities in analogy to CML treatment. Whether the TKI should be used concomitantly with salvage chemotherapy, after chemotherapy or as a maintenance therapy or a “bridging” to transplant after a remission has been achieved, cannot be answered so far. In our clinical practice, we would use a TKI primarily in relapsed BCR-ABL+ AML, starting after a standard salvage regimen has been administered. However, possible benefits have always to be weighed against the risk of clonal selection in favor of a BCR-ABL-negative subclone. In case of BCR-ABL+ MPAL, we would prefer a therapy protocol in analogy to BCR-ABL+ ALL since these protocols are well established [89] and there is no clear evidence supporting the use of AML protocols in this particular situation.
In prospect of the recently revised WHO classification, we are looking forward to emerging new aspects on this interesting new entity, both in basic research and clinical management.
References
Konoplev S, Yin CC, Kornblau SM, Kantarjian HM, Konopleva M, Andreeff M, Lu G, Zuo Z, Luthra R, Medeiros LJ, Bueso-Ramos CE (2013) Molecular characterization of de novo Philadelphia chromosome-positive acute myeloid leukemia. Leuk Lymphoma. doi:10.3109/10428194.2012.701739
Grimwade D, Hills RK, Moorman AV, Walker H, Chatters S, Goldstone AH, Wheatley K, Harrison CJ, Burnett AK, National Cancer Research Institute Adult Leukaemia Working Group (2010) Refinement of cytogenetic classification in acute myeloid leukemia: determination of prognostic significance of rare recurring chromosomal abnormalities among 5876 younger adult patients treated in the United Kingdom Medical Research Council trials. Blood. doi:10.1182/blood-2009-11-254441
Keung YK, Beaty M, Powell BL, Molnar I, Buss D, Pettenati M (2004) Philadelphia chromosome positive myelodysplastic syndrome and acute myeloid leukemia-retrospective study and review of literature. Leuk Res 28(6):579–586
Soupir CP, Vergilio JA, Dal Cin P, Muzikansky A, Kantarjian H, Jones D, Hasserjian RP (2007) Philadelphia chromosome-positive acute myeloid leukemia: a rare aggressive leukemia with clinicopathologic features distinct from chronic myeloid leukemia in myeloid blast crisis. Am J Clin Pathol 127(4):642–650
Berger R (1993) Differences between blastic chronic myeloid leukemia and Ph-positive acute leukemia. Leuk Lymphoma 11(Suppl 1):235–237
Arber DA, Orazi A, Hasserjian R, Thiele J, Borowitz MJ, Le Beau MM, Bloomfield CD, Cazzola M, Vardiman JW (2016) The 2016 revision to the World Health Organization (WHO) classification of myeloid neoplasms and acute leukemia. Blood.
Grove CS, Vassiliou GS (2014) Acute myeloid leukaemia: a paradigm for the clonal evolution of cancer? Dis Model Mech. doi:10.1242/dmm.015974
Lawrence MS, Stojanov P, Polak P, Kryukov GV, Cibulskis K, Sivachenko A, Carter SL, Stewart C, Mermel CH, Roberts SA, Kiezun A, Hammerman PS, McKenna A, Drier Y, Zou L, Ramos AH, Pugh TJ, Stransky N, Helman E, Kim J, Sougnez C, Ambrogio L, Nickerson E, Shefler E, Cortés ML, Auclair D, Saksena G, Voet D, Noble M, DiCara D, Lin P, Lichtenstein L, Heiman DI, Fennell T, Imielinski M, Hernandez B, Hodis E, Baca S, Dulak AM, Lohr J, Landau DA, Wu CJ, Melendez-Zajgla J, Hidalgo-Miranda A, Koren A, McCarroll SA, Mora J, Lee RS, Crompton B, Onofrio R, Parkin M, Winckler W, Ardlie K, Gabriel SB, Roberts CW, Biegel JA, Stegmaier K, Bass AJ, Garraway LA, Meyerson M, Golub TR, Gordenin DA, Sunyaev S, Lander ES, Getz G (2013) Mutational heterogeneity in cancer and the search for new cancer-associated genes. Nature. doi:10.1038/nature12213
Döhner H, Weisdorf DJ, Bloomfield CD (2015) Acute myeloid leukemia. N Engl J Med. doi:10.1056/NEJMra1406184
Gilliland DG (2002) Molecular genetics of human leukemias: new insights into therapy. Semin Hematol 39(4 Suppl 3):6–11
Bacher U, Haferlach T, Alpermann T, Zenger M, Hochhaus A, Beelen DW, Uppenkamp M, Rummel M, Kern W, Schnittger S, Haferlach C (2011) Subclones with the t(9;22)/BCR-ABL1 rearrangement occur in AML and seem to cooperate with distinct genetic alterations. Br J Haematol. doi:10.1111/j.1365-2141.2010.08472.x
Jaiswal S, Fontanillas P, Flannick J, Manning A, Grauman PV, Mar BG, Lindsley RC, Mermel CH, Burtt N, Chavez A, Higgins JM, Moltchanov V, Kuo FC, Kluk MJ, Henderson B, Kinnunen L, Koistinen HA, Ladenvall C, Getz G, Correa A, Banahan BF, Gabriel S, Kathiresan S, Stringham HM, McCarthy MI, Boehnke M, Tuomilehto J, Haiman C, Groop L, Atzmon G, Wilson JG, Neuberg D, Altshuler D, Ebert BL (2014) Age-related clonal hematopoiesis associated with adverse outcomes. N Engl J Med. doi:10.1056/NEJMoa1408617
Genovese G, Kähler AK, Handsaker RE, Lindberg J, Rose SA, Bakhoum SF, Chambert K, Mick E, Neale BM, Fromer M, Purcell SM, Svantesson O, Landén M, Höglund M, Lehmann S, Gabriel SB, Moran JL, Lander ES, Sullivan PF, Sklar P, Grönberg H, Hultman CM, McCarroll SA (2014) Clonal hematopoiesis and blood-cancer risk inferred from blood DNA sequence. N Engl J Med. doi:10.1056/NEJMoa1409405
Biernaux C, Loos M, Sels A, Huez G, Stryckmans P (1995) Detection of major bcr-abl gene expression at a very low level in blood cells of some healthy individuals. Blood 86(8):3118–3122
Ismail SI, Naffa RG, Yousef AM, Ghanim MT (2014) Incidence of bcr-abl fusion transcripts in healthy individuals. Mol Med Rep. doi:10.3892/mmr.2014.1951
Song J, Mercer D, Hu X, Liu H, Li MM (2011) Common leukemia- and lymphoma-associated genetic aberrations in healthy individuals. J Mol Diagn. doi:10.1016/j.jmoldx.2010.10.009
Bose S, Deininger M, Gora-Tybor J, Goldman JM, Melo JV (1998) The presence of typical and atypical BCR-ABL fusion genes in leukocytes of normal individuals: biologic significance and implications for the assessment of minimal residual disease. Blood 92(9):3362–3367
Wu Y, Slovak ML, Snyder DS, Arber DA (2006) Coexistence of inversion 16 and the Philadelphia chromosome in acute and chronic myeloid leukemias: report of six cases and review of literature. Am J Clin Pathol 125(2):260–266
Ninomiya S, Kanemura N, Tsurumi H, Kasahara S, Hara T, Yamada T, Moriwaki H (2011) Coexistence of inversion 16 and the Philadelphia chromosome comprising P190 BCR-ABL in chronic myeloid leukemia blast crisis. Int J Hematol. doi:10.1007/s12185-011-0854-3
Secker-Walker LM, Morgan GJ, Min T, Swansbury GJ, Craig J, Yamada T, Desalvo L, Medina JW, Chowdhury V, Donahue RP (1992) Inversion of chromosome 16 with the Philadelphia chromosome in acute myelomonocytic leukemia with eosinophilia. Report of two cases. Cancer Genet Cytogenet 58(1):29–34
Miura I, Takatsu H, Yamaguchi A, Hashimoto K, Nimura T, Nishinari T, Niitsu H, Miura AB (1994) Standard Ph chromosome, t(9;22)(q34;q11), as an additional change in a patient with acute myelomonocytic leukemia (M4Eo) associated with inv(16)(p13q22). Am J Hematol 45(1):94–96
Siddiqui AD, Sheikh ZS, Liu D, Seiter K (2002) Coexistence of inversion 16 and the Philadelphia chromosome in patients with acute myelogenous leukemia. Leuk Lymphoma 43(5):1137–1140
Tirado CA, Valdez F, Klesse L, Karandikar NJ, Uddin N, Arbini A, Fustino N, Collins R, Patel S, Smart RL, Garcia R, Doolittle J, Chen W (2010) Acute myeloid leukemia with inv(16) with CBFB-MYH11, 3'CBFB deletion, variant t(9;22) with BCR-ABL1, and del(7)(q22q32) in a pediatric patient: case report and literature review. Cancer Genet Cytogenet. doi:10.1016/j.cancergencyto.2010.03.001
Roth CG, Contis L, Gupta S, Agha M, Safyan E (2011) De novo acute myeloid leukemia with Philadelphia chromosome (BCR-ABL) and inversion 16 (CBFB-MYH11): report of two cases and review of the literature. Leuk Lymphoma. doi:10.3109/10428194.2010.538941
Dai HP, Xue YQ, Wu LL, Pan JL, Gong YL, Wu YF, Zhang J, Wu DP, Chen SN (2012) p210 BCR-ABL1 as a secondary change in a patient with acute myelomonocytic leukemia (M4Eo) with inv(16). Int J Hematol. doi:10.1007/s12185-012-1190-y
Svaldi M, Lanthaler A, Venturi R, Coser P, Mitterer M (2001) Simultaneous occurrence of bcr-abl and inv16 in a case of M1 acute myeloid leukemia. Leukemia 15(4):695
Mecucci C, Noens L, Aventin A, Testoni N, Van den Berghe H (1988) Philadelphia-positive acute myelomonocytic leukemia with inversion of chromosome 16 and eosinobasophils. Am J Hematol 27(1):69–71
Cividin M, Brizard F, Sorel N, Renaud M, Guilhot F, Brizard A (2004) p190(BCR-ABL) rearrangement as a secondary change in a case of acute myelo-monocytic leukemia with inv(16)(p13q22). Leuk Res 28(1):97–99
Preudhomme C, Lai JL, Plantier I, Demory JL, Zandecki M, Fenaux P (1992) Cytogenetic and molecular remission in a case of acute myeloid leukaemia(AML) with inversion of chromosome 16 (inv(16)) and Philadelphia chromosome (Ph). Br J Haematol 82(3):623–626
Cho BS, Kim HJ, Lee S, Eom KS, Min WS, Lee JW, Kim CC (2007) Successful interim therapy with imatinib prior to allogeneic stem cell transplantation in Philadelphia chromosome-positive acute myeloid leukemia. Eur J Haematol 79(2):170–173
Dallorso S, Sessarego M, Garré ML, Haupt R, Pasino M, Sansone R (1990) Secondary acute promyelocytic leukemia with t(8;21) and t(9;22) at onset and loss of the Philadelphia chromosome at relapse. Cancer Genet Cytogenet 47(1):41–46
Verhaak RG, Goudswaard CS, van Putten W, Bijl MA, Sanders MA, Hugens W, Uitterlinden AG, Erpelinck CA, Delwel R, Löwenberg B, Valk PJ (2005) Mutations in nucleophosmin (NPM1) in acute myeloid leukemia (AML): association with other gene abnormalities and previously established gene expression signatures and their favorable prognostic significance. Blood 106(12):3747–3754
Suzuki T, Kiyoi H, Ozeki K, Tomita A, Yamaji S, Suzuki R, Kodera Y, Miyawaki S, Asou N, Kuriyama K, Yagasaki F, Shimazaki C, Akiyama H, Nishimura M, Motoji T, Shinagawa K, Takeshita A, Ueda R, Kinoshita T, Emi N, Naoe T (2005) Clinical characteristics and prognostic implications of NPM1 mutations in acute myeloid leukemia. Blood 106(8):2854–2861
Reboursiere E, Chantepie S, Gac AC, Reman O (2015) Rare but authentic Philadelphia-positive acute myeloblastic leukemia: two case reports and a literature review of characteristics, treatment and outcome. Hematol Oncol Stem Cell Ther. doi:10.1016/j.hemonc.2014.09.002
Mozziconacci MJ, Sainty D, Gabert J, Arnoulet C, Simonetti J, Toiron Y, Costello R, Hagemeijer A, Lafage-Pochitaloff M (1998) The Philadelphia chromosome as a secondary abnormality in two cases of acute myeloid leukemia. Br J Haematol 102(3):873–875
Ding L, Ley TJ, Larson DE, Miller CA, Koboldt DC, Welch JS, Ritchey JK, Young MA, Lamprecht T, McLellan MD, McMichael JF, Wallis JW, Lu C, Shen D, Harris CC, Dooling DJ, Fulton RS, Fulton LL, Chen K, Schmidt H, Kalicki-Veizer J, Magrini VJ, Cook L, McGrath SD, Vickery TL, Wendl MC, Heath S, Watson MA, Link DC, Tomasson MH, Shannon WD, Payton JE, Kulkarni S, Westervelt P, Walter MJ, Graubert TA, Mardis ER, Wilson RK, DiPersio JF (2012) Clonal evolution in relapsed acute myeloid leukaemia revealed by whole-genome sequencing. Nature. doi:10.1038/nature10738
Nacheva EP, Grace CD, Brazma D, Gancheva K, Howard-Reeves J, Rai L, Gale RE, Linch DC, Hills RK, Russell N, Burnett AK, Kottaridis PD (2013) Does BCR-ABL1 positive acute myeloid leukaemia exist? Br J Haematol. doi:10.1111/bjh.12301
Nacheva EP, Brazma D, Virgili A, Howard-Reeves J, Chanalaris A, Gancheva K, Apostolova M, Valgañon M, Mazzullo H, Grace C (2010) Deletions of immunoglobulin heavy chain and T cell receptor gene regions are uniquely associated with lymphoid blast transformation of chronic myeloid leukemia. BMC Genomics. doi:10.1186/1471-2164-11-41
Merzianu M, Medeiros LJ, Cortes J, Yin C, Lin P, Jones D, Glassman A, Kantarjian H, Huh Y (2005) inv(16)(p13q22) in chronic myelogenous leukemia in blast phase: a clinicopathologic, cytogenetic, and molecular study of five cases. Am J Clin Pathol 124(5):807–814
Tsuboi K, Komatsu H, Miwa H, Iida S, Banno S, Wakita A, Nitta M, Ueda R (2002) Lymphoid blastic crisis of chronic myelogenous leukaemia with inv(16)(p13;q22). Leuk Res 26(8):771–774
Evers JP, Bagg A, Himoe E, Zwiebel JA, Jacobson RJ (1992) Temporal association of marrow eosinophilia with inversion of chromosome 16 in recurrent blast crises of chronic myelogenous leukemia. Cancer Genet Cytogenet 62(2):134–139
Asou N, Sanada I, Tanaka K, Hidaka M, Suzushima H, Matsuzaki H, Kawano F, Takatsuki K (1992) Inversion of chromosome 16 and bone marrow eosinophilia in a myelomonocytic transformation of chronic myeloid leukemia. Cancer Genet Cytogenet 61(2):197–200
Heim S, Christensen BE, Fioretos T, Sørensen AG, Pedersen NT (1992) Acute myelomonocytic leukemia with inv(16)(p13q22) complicating Philadelphia chromosome positive chronic myeloid leukemia. Cancer Genet Cytogenet 59(1):35–38
Patel BB, Mohamed AN, Schiffer CA (2006) "Acute myelogenous leukemia like" translocations in CML blast crisis: two new cases of inv(16)/t(16;16) and a review of the literature. Leuk Res 30(2):225–232
Myint H, Ross FM, Hall JL, Hamblin TJ (1997) Early transformation to acute myeloblastic leukaemia with the acquisition of inv(16) in Ph positive chronic granulocytic leukaemia. Leuk Res 21(5):473–474
Mohamed AN, Pemberton P, Zonder J, Schiffer CA (2003) The effect of imatinib mesylate on patients with Philadelphia chromosome-positive chronic myeloid leukemia with secondary chromosomal aberrations. Clin Cancer Res 9(4):1333–1337
Silva PM, Lourenço GJ, Bognone RA, Delamain MT, Pinto-Junior W, Lima CS (2006) Inherited pericentric inversion of chromosome 16 in chronic phase of chronic myeloid leukaemia. Leuk Res 30(1):115–117
Colović M, Janković G, Bila J, Djordjević V, Wiernik PH (1998) Inversion of chromosome 16 in accelerated phase of chronic myeloid leukaemia: report of a case and review of the literature. Med Oncol 15(3):199–201
Palmisano M, Grafone T, Ottaviani E, Testoni N, Baccarani M, Martinelli G (2007) NPM1 mutations are more stable than FLT3 mutations during the course of disease in patients with acute myeloid leukemia. Haematologica 92(9):1268–1269
Kondo T, Tasaka T, Sano F, Matsuda K, Kubo Y, Matsuhashi Y, Nakanishi H, Sadahira Y, Wada H, Sugihara T, Tohyama K (2009) Philadelphia chromosome-positive acute myeloid leukemia (Ph + AML) treated with imatinib mesylate (IM): a report with IM plasma concentration and bcr-abl transcripts. Leuk Res. doi:10.1016/j.leukres.2009.03.017
Paietta E, Racevskis J, Bennett JM, Neuberg D, Cassileth PA, Rowe JM, Wiernik PH (1998) Biologic heterogeneity in Philadelphia chromosome-positive acute leukemia with myeloid morphology: the Eastern Cooperative Oncology Group experience. Leukemia 12(12):1881–1885
Sindt A, Deau B, Brahim W, Staal A, Visanica S, Villarese P, Rault JP, Macintyre E, Delabesse E (2006) Acute monocytic leukemia with coexpression of minor BCR-ABL1 and PICALM-MLLT10 fusion genes along with overexpression of HOXA9. Genes Chromosom Cancer 45(6):575–582
Smadja N, Krulik M, De Gramont A, Brissaud P, Debray J (1985) Acquisition of a Philadelphia chromosome concomitant with transformation of a refractory anemia into an acute leukemia. Cancer 55(7):1477–1481
Kohn G, Manny N, Eldor A, Cohen MM (1975) De novo appearance of the ph-1 chromosome in a previously monosomic bone marrow (45,XX,-6): conversion of a myeloproliferative disorder to acute myelogenous leukemia. Blood 45(5):653–657
Katsuno M, Yamashita S, Sadamura S, Umemura T, Hirata J, Nishimura J, Nawata H (1994) Late-appearing Philadelphia chromosome in a patient with acute nonlymphocytic leukaemia derived from myelodysplastic syndrome: detection of P210- and P190-type BCR-ABL fusion gene transcripts at the leukaemic stage. Br J Haematol 87(1):51–56
Hirsch-Ginsberg C, Childs C, Chang KS, Beran M, Cork A, Reuben J, Freireich EJ, Chang LC, Bollum FJ, Trujillo J, et al. (1988) Phenotypic and molecular heterogeneity in Philadelphia chromosome-positive acute leukemia. Blood 71(1):186–195
Maddox AM, Keating MJ, Trujillo J, Cork A, Youness E, Ahearn MJ, McCredie KB, Freireich EJ (1983) Philadelphia chromosome-positive adult acute leukemia with monosomy of chromosome number seven: a subgroup with poor response to therapy. Leuk Res 7(4):509–522
Yamashita S, Umemura T, Sadamura S, Takahira H, Nishimura J, Nawata H, Katsuno M, Okamura J, Horibe K (1996) Acute leukemias expressing p210-and p 190-type BCR-ABL mRNAs: report of two cases and review of the literature. Acta Haematol 96(2):99–104
Ohyashiki K, Kocova M, Ryan DH, Rowe JM, Sandberg AA (1986) Secondary acute myeloblastic leukemia with a Ph translocation in a treated Wegener's granulomatosis. Cancer Genet Cytogenet 19(3–4):331–333
Ohyashiki K, Ohyashiki JH, Raza A, Preisler HD, Sandberg AA (1987) Phenylbutazone-induced myelodysplastic syndrome with Philadelphia translocation. Cancer Genet Cytogenet 26(2):213–216
Lesesve JF, Troussard X, Bastard C, Hurst JP, Nouet D, Callat MP, Lenormand B, Piguet H, Flandrin G, Macintyre E (1996) p190BCR-ABL rearrangement in myelodysplastic syndromes: two reports and review of the literature. Br J Haematol 95(2):372–375
Larripa I, Gutiérrez M, Giere I, Acevedo S, Bengió R, Slavutsky I (1992) Complex karyotype with PH1 chromosome in myelodysplasia: cytogenetic and molecular studies. Leuk Lymphoma 6:401–406
Papageorgiou SG, Pappa V, Economopoulou C, Tsirigotis P, Konsioti F, Ionnidou ED, Chondropoulos S, Vasilatou D, Papageorgiou E, Economopoulos T, Dervenoulas J (2010) Dasatinib induces long-term remission in imatinib-resistant Philadelphia chromosome-positive acute megakaryoblastic leukemia but fails to prevent development of central nervous system progression. Leuk Res. doi:10.1016/j.leukres.2010.03.032
Fukunaga A, Sakoda H, Iwamoto Y, Inano S, Sueki Y, Yanagida S, Arima N (2013) Abrupt evolution of Philadelphia chromosome-positive acute myeloid leukemia in myelodysplastic syndrome. Eur J Haematol. doi:10.1111/ejh.12056
Isoda A, Nakahashi H, Hoshino T, Mitsui T, Yoshida Y (2007) Insufficient outcomes with imatinib mesylate: case report of Ph-positive acute myeloid leukemia evolving from myelodysplastic syndrome. Am J Hematol 82(6):501–502
Kelemen K, Galani K, Conley CR, Greipp PT (2014) Secondary Philadelphia chromosome and erythrophagocytosis in a relapsed acute myeloid leukemia after hematopoietic cell transplantation. Cancer Genet. doi:10.1016/j.cancergen.2014.05.013
Nakase K, Yamamoto Y, Morita K, Yamaguchi T, Nishii K, Shiku H (2006) Haunting appearance of BCR-ABL fusion gene products in a patient with therapy related leukaemia. Leuk Res 30(1):106–108
Kurzrock R, Shtalrid M, Talpaz M, Kloetzer WS, Gutterman JU (1987) Expression of c-abl in Philadelphia-positive acute myelogenous leukemia. Blood 70(5):1584–1588
Alimena G, Cedrone M, Nanni M, De Cuia MR, Lo Coco F, De Sanctis V, Cimino G, Mancini M (1995) Acute leukemia presenting a variant Ph chromosome with p190 expression, dup 3q and -7, developed after malignant lymphoma treated with alkylating agents and topoisomerase II inhibitors. Leukemia 9(9):1483–1486
Han JY, Theil KS (2006) The Philadelphia chromosome as a secondary abnormality in inv(3)(q21q26) acute myeloid leukemia at diagnosis: confirmation of p190 BCR-ABL mRNA by real-time quantitative polymerase chain reaction. Cancer Genet Cytogenet 165(1):70–74
Najfeld V, Geller M, Troy K, Scalise A (1998) Acquisition of the Ph chromosome and BCR-ABL fusion product in AML-M2 and t(8;21) leukemia: cytogenetic and FISH evidence for a late event. Leukemia 12(4):517–519
Jacobsen RJ, Himoe E, Sacher RA, Shashaty GG (1986) Late appearance of Philadelphia chromosome. Br J Haematol 63(2):392–394
Aoki J, Kakihana K, Kobayashi T, Hirashima Y, Akiyama H, Ohashi K, Sakamaki H (2012) Tyrosine kinase inhibitor therapy for acute myeloid leukemia with late-appearing Philadelphia chromosome. Leuk Res. doi:10.1016/j.leukres.2011.10.008
Yagyu S, Morimoto A, Kakazu N, Tamura S, Fujiki A, Nakase Y, Iehara T, Hosoi H, Kuroda H (2008) Late appearance of a Philadelphia chromosome in a patient with therapy-related acute myeloid leukemia and high expression of EVI1. Cancer Genet Cytogenet. doi:10.1016/j.cancergencyto.2007.09.023
Shah N, Leaker MT, Teshima I, Baruchel S, Abdelhaleem M, Ye CC (2008) Late-appearing Philadelphia chromosome in childhood acute myeloid leukemia. Pediatr Blood Cancer. doi:10.1002/pbc.21317
Prebet T, Michallet AS, Charrin C, Hayette S, Magaud JP, Thiébaut A, Michallet M, Nicolini FE (2004) Secondary Philadelphia chromosome after non-myeloablative peripheral blood stem cell transplantation for a myelodysplastic syndrome in transformation. Bone Marrow Transplant 33(2):247–249
Neuendorff NR, Schwarz M, Hemmati P, Türkmen S, Bommer C, Burmeister T, Dörken B, le Coutre P, Arnold R, Westermann J (2015) BCR-ABL1(+) acute myeloid leukemia: clonal selection of a BCR-ABL1(−) subclone as a cause of refractory disease with nilotinib treatment. Acta Haematol. doi:10.1159/000368176
Weinberg OK, Seetharam M, Ren L, Alizadeh A, Arber DA (2014) Mixed phenotype acute leukemia: a study of 61 cases using World Health Organization and European Group for the Immunological Classification of Leukaemias criteria. Am J Clin Pathol. doi:10.1309/AJCPPVUPOTUVOIB5
Heesch S, Neumann M, Schwartz S, Bartram I, Schlee C, Burmeister T, Hänel M, Ganser A, Heuser M, Wendtner CM, Berdel WE, Gökbuget N, Hoelzer D, Hofmann WK, Thiel E, Baldus CD (2013) Acute leukemias of ambiguous lineage in adults: molecular and clinical characterization. Ann Hematol. doi:10.1007/s00277-013-1694-4
Atfy M, Al Azizi NM, Elnaggar AM (2011) Incidence of Philadelphia-chromosome in acute myelogenous leukemia and biphenotypic acute leukemia patients: and its role in their outcome. Leuk Res. doi:10.1016/j.leukres.2011.04.011
Walter RB, Othus M, Burnett AK, Löwenberg B, Kantarjian HM, Ossenkoppele GJ, Hills RK, van Montfort KG, Ravandi F, Evans A, Pierce SR, Appelbaum FR, Estey EH (2013) Significance of FAB subclassification of "acute myeloid leukemia, NOS" in the 2008 WHO classification: analysis of 5848 newly diagnosed patients. Blood. doi:10.1182/blood-2012-10-462440
Chen SJ, Flandrin G, Daniel MT, Valensi F, Baranger L, Grausz D, Bernheim A, Chen Z, Sigaux F, Berger R (1988) Philadelphia-positive acute leukemia: lineage promiscuity and inconsistently rearranged breakpoint cluster region. Leukemia 2(5):261–273
O’Donnell MR, Tallman MS et al (2016) NCCN Clinical Practise Guidelines in Oncology: AML. Version 1. Available at: NCCN.org
Röllig C, Bornhäuser M, Thiede C, Taube F, Kramer M, Mohr B, Aulitzky W, Bodenstein H, Tischler HJ, Stuhlmann R, Schuler U, Stölzel F, von Bonin M, Wandt H, Schäfer-Eckart K, Schaich M, Ehninger G (2011) Long-term prognosis of acute myeloid leukemia according to the new genetic risk classification of the European LeukemiaNet recommendations: evaluation of the proposed reporting system. J Clin Oncol. doi:10.1200/JCO.2010.32.8500
Bhatt VR, Akhtari M, Bociek RG, Sanmann JN, Yuan J, Dave BJ, Sanger WG, Kessinger A, Armitage JO (2014) Allogeneic stem cell transplantation for Philadelphia chromosome-positive acute myeloid leukemia. J Natl Compr Cancer Netw 12(7):963–968
Ueda K, Horiike S, Zen K, Misawa S, Taniwaki M (2006) Complete cytogenetic and molecular response to treatment with imatinib mesylate for Philadelphia chromosome positive acute myeloid leukemia with multilineage dysplasia. Leuk Lymphoma 47(9):1967–1969
Hehlmann R (2012) How I treat CML blast crisis. Blood. doi:10.1182/blood-2012-03-380147
Baccarani M, Deininger MW, Rosti G, Hochhaus A, Soverini S, Apperley JF, Cervantes F, Clark RE, Cortes JE, Guilhot F, Hjorth-Hansen H, Hughes TP, Kantarjian HM, Kim DW, Larson RA, Lipton JH, Mahon FX, Martinelli G, Mayer J, Müller MC, Niederwieser D, Pane F, Radich JP, Rousselot P, Saglio G, Saußele S, Schiffer C, Silver R, Simonsson B, Steegmann JL, Goldman JM, Hehlmann R (2013) European LeukemiaNet recommendations for the management of chronic myeloid leukemia: 2013. Blood. doi:10.1182/blood-2013-05-501569
Wassmann B, Pfeifer H, Goekbuget N, Beelen DW, Beck J, Stelljes M, Bornhäuser M, Reichle A, Perz J, Haas R, Ganser A, Schmid M, Kanz L, Lenz G, Kaufmann M, Binckebanck A, Brück P, Reutzel R, Gschaidmeier H, Schwartz S, Hoelzer D, Ottmann OG (2006) Alternating versus concurrent schedules of imatinib and chemotherapy as front-line therapy for Philadelphia-positive acute lymphoblastic leukemia (Ph + ALL). Blood 108(5):1469–1477
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
This work was not supported by any grant.
Conflict of interest
J.W. receives research support and/or honoraria from Novartis, BMS, and Celgene; T.B. received research support from Novartis; N.R.N. receives a scholarship from Medac. Bernd Dörken declares that he has no conflicts of interest.
Electronic supplementary material
ESM 1
(PDF 611 kb)
Rights and permissions
About this article
Cite this article
Neuendorff, N.R., Burmeister, T., Dörken, B. et al. BCR-ABL-positive acute myeloid leukemia: a new entity? Analysis of clinical and molecular features. Ann Hematol 95, 1211–1221 (2016). https://doi.org/10.1007/s00277-016-2721-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00277-016-2721-z