Abstract
Background
The molecular bases for parathyroid carcinomas present in conjunction with sporadic primary hyperparathyroidism are not fully elucidated. Gene copy number variations (CNVs) play an important role in tumorigenesis. The aim of the current study was to explore whether the CNVs of specific tumor-associated genes are involved in parathyroid carcinogenesis.
Methods
A multiplex ligation-dependent probe amplification method was used to compare differences in copy number in 39 common tumor-associated genes among 7 patients with parathyroid carcinoma and 14 age- and sex-matched subjects with parathyroid adenoma.
Results
It was shown that amplification of CCND1, a gene encoding cyclin D1, was more prevalent in parathyroid carcinomas than in adenomas (71 vs. 21 %, p = 0.056). This result was confirmed quantitatively by real-time polymerase chain reaction. Expression of CCND1 mRNA level was significantly higher in carcinomas than in adenomas (p = 0.003). Western blot and immunohistochemical analysis also demonstrated higher expression of CCND1 in carcinoma specimens than in adenoma samples.
Conclusions
It is thus inferred that gain in copy number of CCND1 is implicated in the molecular pathogenesis of parathyroid carcinoma.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Sporadic primary hyperparathyroidism (PHPT) is a common endocrine disease caused by over-production of parathyroid hormone (PTH) by functioning benign or malignant parathyroid tumor(s) [1]. Pathologically, sporadic PHPT can be distinguished as adenoma (85 %), hyperplasia (10–15 %), or carcinoma (<1 %). Although parathyroid carcinoma is a rare cause of PHPT in most Western countries, in Asian populations such as Japan, Korea, India, and China (including our own cohort) it accounts for 4–6 % of PHPT cases [2–5].
The clinical manifestations, biochemical abnormalities, and prognosis of parathyroid carcinoma are much severer and poorer than benign cases [5, 6]. The initial en bloc resection represents the best chance for cure of parathyroid carcinoma [6, 7]. The etiology of parathyroid tumor is still largely unknown, although several oncogenes—cyclin D1/parathyroid adenoma 1 gene (PRAD1)—and tumor suppressor genes—e.g., retinoblastoma (Rb), p53, and hyperparathyroidism type 2/cell division cycle 73 (HRPT2/CDC73) gene—are involved in the development of parathyroid adenoma and carcinoma [1, 6, 8].
In recent years, the importance of gene copy number variations (CNVs) in the initiation and progression of tumors [9] and in other human diseases [10–12] has been increasingly recognized. Previous comparative studies using genomic hybridization (CGH) have found that loss of chromosomes 22, 11, and 17 might be associated with the development of parathyroid adenoma, and the loss of chromosome 1q and the acquisition of chromosome 5 segments might be associated with parathyroid carcinoma [13, 14]. However, such a technique cannot map the disease to a specific gene.
Multiplex ligation-dependent probe amplification (MLPA) is a more sensitive and high throughput approach to identify gene CNVs in the chromosomes, including small intragenic rearrangements by hybridizing sequence-specific probe to genomic DNA [15–17]. This technique allows us to test as many as 40–50 targets in one reaction to quantify the relative copy number of each DNA sequence [18]. MLPA has become a widely used research and diagnostic tool for many human diseases, such as breast cancer [19–21], gastric cancer [22], leukemia [23, 24], and insulinoma [25] among others. Although our knowledge of the genetics of parathyroid tumorigenesis has improved in recent years [26, 27], it is still unclear which genes exhibit differences in copy numbers in parathyroid adenoma versus parathyroid carcinoma.
In the present study, we employed MLPA to compare differences in copy numbers in 39 common tumor-associated genes between patients with parathyroid carcinoma and those with adenoma, with the aim of exploring whether the CNV of a specific gene is involved in carcinogenesis in the parathyroid.
Materials and methods
Patients and samples
From 2000 to 2010, a total of 235 PHPT patients were operated on in our center. Overall, 14 of them proved to be parathyroid carcinoma [5]. However, only 7 malignant parathyroid specimens were available for current study. The other tumor samples had insufficient material to investigate after pathological analysis. Another 14 sex- and age-matched benign PHPT cases were selected as controls for a 1:2 ratio.
The diagnosis of parathyroid carcinoma was established pathologically based on surgically dissected tumor specimens: the presence of prominent trabecular growth, the presence of thick fibrous bands within the tumor, increased mitotic activity, capsular penetration, vascular invasion, extraparathyroid spread, or distant metastases [28]. Genomic DNA was extracted from fresh-frozen tumor specimens by standard procedures using the Qiagen DNA extraction Kit (Qiagen, Hilden, Germany) .
The Ethics Committee of Rui-jin Hospital, Shanghai Jiao-Tong University School of Medicine approved the study.
Multiplex ligation-dependent probe amplification
MLPA analysis was carried out using SALSA P006 Chromosomal Aberration MLPA Kits (MRC-Holland, Amsterdam, The Netherlands). The P006 kit contains probes for 39 loci. Among them are many genes frequently involved in the pathogenesis of various tumors and spanning almost all chromosomes (e.g., CCND1, RB, P53). All of the reactions were carried out in PTC-225 DNA Engine Tetrad (MJ Research, San Francisco, CA, USA). PCR products were analyzed using a Beckman Coulter CEQ 8800 sequencer (Beckman Coulter, Fullerton, CA, USA).
Data analysis was performed with a fragment analysis module with the detailed method introduced in the literature [25].
Real-time polymerase chain reaction
Total RNA was isolated with Trizol reagent (Invitrogen, Carlsbad, CA, USA) and reverse transcribed from a Random Primers (Promega, Madison, WI, USA) according to the manufacturer’s instructions. The real-time polymerase chain reaction (RT-PCR) was performed with a Roche Light Cycler 480 system using SYBR-Premix Ex TaqTM (Takara, Otsu, Japan). PCRs were performed in triplicate, and the glyceradehyde-3-phosphate dehydrogenase (GAPDH) gene was amplified as an internal control. The primers sequences used to detect mRNA expression levels were as follows: GAPDH, forward primer 5′-ATGGGGAAGGTGAAGGTCG-3′ and reverse primer 5′-GGGGTCATTGATGGCAACAATA-3′; CCND1; forward primer 5′-GTGCTGCGAAGTGGAAACC-3′ and reverse primer 5′-ATCCAGGTGGCGACGATCT-3′.
Real-time PCR was also used to quantitate the CNVs with the following primers: albumin (ALB) forward primer 5′-ACACGCCTTTGGCACAATGA-3′ and reverse primer 5′-CCCTGGAATAAGCCGAGCTA-3′; CCND1 forward primer 5′-AGGACGTAATTGGTGGCAGG-3′ and reverse primer 5′-GCCAGATACTGGGCTCATCC-3′.
Immunohistochemistry
Immunohistochemistry (IHC) analysis was performed to confirm the gene changes revealed by MLPA. The 4- to 5-μm sections were cut from paraffin blocks and heated at 60 °C overnight. Slides were deparaffinized and heated in citric acid 0.01 mol/L (pH 6.0) for 15 min at 95 °C followed by slow cooling for antigen retrieval. Slides were treated with 0.3 % hydrogen peroxide to block endogenous peroxidase activity. To reduce nonspecific background staining, slides were incubated with 2 % goat serum for 10 min at room temperature. They were then incubated for 60 min at 37 °C with rabbit monoclonal anti-cyclin D1 (CCND1) diluted 1:100 (Thermo Fisher Scientific, Loughborough, Leicestershire, UK), stained using Envision-Plus reagents (Dako, Carpinteria, CA, USA) and diaminobenzidine as chromogen, then counterstained with hematoxylin. Positive CCND1 immunoreactivity showed nuclear staining. The cutoff parameters used for classification of normal and overexpression are <10 % nuclear positive and >30 % nuclear positive, respectively [29].
Western blot
Western blot analysis was performed on 6 parathyroid carcinoma and 12 adenoma tissue samples. Parathyroid tissues were lysed in radioimmunoprecipitation (RIPA) buffer containing 50 mM Tris–HCl (pH 8), 150 mM NaCl, 5 mM MgCl2, 2 mM EDTA, 1 Mm NaF, 1 % NP40, and 0.1 % sodium dodecyl sulfate (SDS). The cell lysates were loaded onto 10 % SDS-polyacrylamide gel electrophoresis and transferred to polyvinylidenedifluoride membranes (Millipore, Bedford, MA, USA). The membranes were blocked with 10 % nonfat milk and then incubated with CCND1 antibodies (Thermo Fisher Scientific) followed by incubation with horseradish peroxidase-conjugated secondary antibodies. The proteins were visualized with enhanced chemiluminescence (ECL) reagents (Amersham Pharmacia, Little Chalfont, Buckinghamshire, UK) according to the manufacturer’s protocol.
Statistical analysis
Data are expressed as the mean ± SD or median (interquartile range). Group comparisons were performed using Student’s t test. Differences in frequency of variables were tested by the χ 2 test or Fisher’s exact test. A value of p < 0.05 denoted the presence of a significant difference.
Results
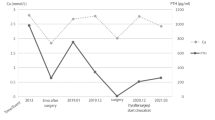
The clinical features of benign and malignant PHPT patients in this study as a group or individually are described in Tables 1 and 2, respectively. Vascular, capsular, or extraparathyroid invasion or distant metastasis was present in malignant parathyroid tumors based on histologic examination (Table 2). Patients with parathyroid carcinoma had significantly higher serum calcium, albumin-corrected serum calcium and PTH levels as well as a lager tumor size than those with benign disease (Table 1). In our series, three of the seven patients with a malignancy died because of recurrent hypercalcemia within 2 years after surgery. The PHPT carcinoma patients were followed for 5 years (0.08–7.0 years). The adenoma patients were also followed for 5 years (4–8 years) (Table 2).
It was revealed that all of the 21 tested specimens had multiple copy number losses or gains (Fig. 1). The average total genetic copy number aberrations in malignant and benign disease were 7.86 ± 1.95 and 5.79 ± 3.39, respectively. There was no significant difference between two groups (p = 0.168).
The most obvious difference between parathyroid carcinoma and adenoma was amplification of the CCND1 gene locus, which is located on chromosome 11q13. In parathyroid carcinoma, amplification of the CCND1 gene locus was detected in 5 of the 7 (71.4 %) samples, whereas only 3 of the 14 (21.4 %) benign samples showed gain of a CCND1 gene locus (p = 0.056) (Fig. 1; Table 3). Although 71.4 % carcinoma samples demonstrating loss of the TP53 gene locus on 17p13.1, 4 of 14 (28.6 %) benign samples also had such a change. The difference between the two groups was not significant (p = 0.159).
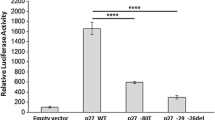
It was also revealed that CCND1 mRNA level was significantly higher in carcinoma than in adenoma (p = 0.003) (Fig. 2). To further quantitate CCND1 gene copy number in the malignant and benign tumor specimens, the RT-PCR method was applied. The results derived from this method were consistent with those found with the MLPA approach (Table 3). In addition, IHC analysis revealed that 6 of the 7 malignant tumors (85.7 %) showed positive CCND1 immunoreactivity, whereas it was seen in only 4 of the 14 benign tumors (28.6 %) (p = 0.024) (Table 3; Fig. 3). Western blot analysis showed that 5 of 6 (83.33 %) parathyroid carcinoma showed positive CCND1 immunoreactivity, and only 2 of 12 (16.67 %) benign tumors did so (p = 0.013) (Fig. 4).
Discussion
The present investigation revealed that gain in copy number of CCND1 is implicated in the molecular pathogenesis of sporadic parathyroid carcinoma.
CCND1 encodes the protein cyclin D1. As early as during the 1990s, its role in the pathogenesis of parathyroid tumors as an oncogene and later as a major regulatory protein in the cell cycle was established [30]. It was demonstrated that cyclin D1 gene expression is significantly higher in parathyroid adenomas and carcinomas than in normal parathyroid gland [31, 32]. Mice overexpressing cyclin D1 exhibited enlarged parathyroid glands and elevated serum calcium and serum PTH concentrations [33]. In the present study, we noted that nearly 70 % of parathyroid carcinoma samples had increased CCND1 copy numbers, whereas an increase was present in only 20 % of parathyroid adenomas, which was further confirmed quantitatively by RT-PCR. In line with this observation, both the mRNA and protein expression level of this gene were significantly higher in the parathyroid carcinomas than in the adenomas. It is thus fair to hypothesize that gene amplification is the principal mechanism of CCND1 overexpression in parathyroid carcinomas.
Indeed, previous studies have suggested a greater role of cyclin D1 overexpression in parathyroid carcinoma relative to adenoma [32, 34]. In one series, it was shown that up to 90 % of parathyroid carcinoma specimens had overexpressed cyclin D1, whereas only 40–60 % of benign parathyroid specimens displayed cyclin D1 overexpression [35]. However, these findings, including ours, still could not establish the causality between cyclin D1 and parathyroid carcinoma. More in-depth molecular studies are needed to address this question.
The underling mechanism responsible for the increased CCND1 copy number in parathyroid carcinomas was not clear. It has been noted that parafibromin, a protein encoded by HRPT2/CDC73 gene can block cyclin D1 expression and thus inhibit cell proliferation. Also, loss of parafibromin causes overexpression of cyclin D1 [32]. We compared the HRPT2 mRNA expression levels between parathyroid carcinomas and adenomas but failed to find any difference between the two groups (data not shown). However, because of the limited number of cases in the current study, no conclusion can be drawn as to the relation between HRPT2/CDC73 mutation and CCND1 CNVs. Our data should be interpreted with caution. Because germline DNA was not collected from all of the patients, we could not exclude the possibility that some areas of DNA gain and loss in the CCND1 gene shown by MLPA in parathyroid tumors could be germline variants rather than somatic mutations. On the other hand, the genes included in this MLPA panel are not necessarily the pathogenically relevant gene in an amplicon or area of loss of heterozygosity. They may only be markers for a region of DNA gain or loss, with the true pathogenic target being elsewhere in the same region.
This study has several other limitations. First, the sample size is very small. Second, this study was performed without normal controls because of the difficulty of obtaining normal parathyroid tissues. It was reported that the frequency of cyclin D1 expression in normal parathyroid tissue is rarely low (<6 %) [35]. Third, only common genes associated with cancers were selected for MLPA analysis. Some other genes, such as HRPT2 and multiple endocrine neoplasia type 1 (MEN1), which are also involved in the molecular pathogenesis of parathyroid tumor [27], were not tested in this study. It was known that 11q13 allelic loss occurs frequently in 25–40 % of parathyroid adenomas. The loss of this region in parathyroid adenomas is closely linked to the MEN1 gene [36, 37], but this target sequence was not included in the MLPA kit that we used. Thus, all of the limitations of our study may bias its results.
Conclusions
Our study provided some clues that the copy number increase in the CCND1 gene may be one of the underlying mechanisms responsible for sporadic parathyroid carcinogenesis.
References
Fraser WD (2009) Hyperparathyroidism. Lancet 374:145–158
Obara T, Okamoto T, Kanbe M et al (1997) Functioning parathyroid carcinoma: clinicopathologic features and rational treatment. Semin Surg Oncol 13:134–141
Lee YS, Hong SW, Jeong JJ et al (2010) Parathyroid carcinoma: a 16-year experience in a single institution. Endocr J 57:493–497
Pradeep PV, Jayashree B, Mishra A et al (2011) Systematic review of primary hyperparathyroidism in India: the past, present, and the future trends. Int J Endocrinol 2011:921814
Zhao L, Liu JM, He XY et al (2013) The changing clinical patterns of primary hyperparathyroidism in Chinese patients: data from 2000 to 2010 in a single clinical center. J Clin Endocrinol Metab 98:721–728
Shane E (2001) Clinical review 122: parathyroid carcinoma. J Clin Endocrinol Metab 86:485–493
Ricci G, Assenza M, Barreca M et al (2012) Parathyroid carcinoma: the importance of high clinical suspicion for a correct management. Int J Surg Oncol 2012:649148
Bricaire L, Odou MF, Cardot-Bauters C et al (2013) Frequent large germline HRPT2 deletions in a French National cohort of patients with primary hyperparathyroidism. J Clin Endocrinol Metab 98:E403–E408
Lee C, Iafrate AJ, Brothman AR (2007) Copy number variations and clinical cytogenetic diagnosis of constitutional disorders. Nat Genet 39:S48–S54
Rogers AJ, Chu JH, Darvishi K et al (2013) Copy number variation prevalence in known asthma genes and their impact on asthma susceptibility. Clin Exp Allergy 43:455–462
Zahnleiter D, Uebe S, Ekici AB et al (2013) Rare copy number variants are a common cause of short stature. PLoS Genet 9:e1003365
Wheeler E, Huang N, Bochukova EG et al (2013) Genome-wide SNP and CNV analysis identifies common and low-frequency variants associated with severe early-onset obesity. Nat Genet 45:513–517
Agarwal SK, Schrock E, Kester MB et al (1998) Comparative genomic hybridization analysis of human parathyroid tumors. Cancer Genet Cytogenet 106:30–36
Kytola S, Farnebo F, Obara T et al (2000) Patterns of chromosomal imbalances in parathyroid carcinomas. Am J Pathol 157:579–586
Schouten JP, McElgunn CJ, Waaijer R et al (2002) Relative quantification of 40 nucleic acid sequences by multiplex ligation-dependent probe amplification. Nucleic Acids Res 3:e57
Hogervorst FB, Nederlof PM, Gille JJ et al (2003) Large genomic deletions and duplications in the BRCA1 gene identified by a novel quantitative method. Cancer Res 63:1449–1453
Eldering E, Spek CA, Aberson HL et al (2003) Expression profiling via novel multiplex assay allows rapid assessment of gene regulation in defined signalling pathways. Nucleic Acids Res 31:e153
Stuppia L, Antonucci I, Palka G et al (2012) Use of the MLPA assay in the molecular diagnosis of gene copy number alterations in human genetic diseases. Int J Mol Sci 13:3245–3276
Moelans CB, Monsuur HN, de Pinth JH et al (2011) ESR1 amplification is rare in breast cancer and is associated with high grade and high proliferation: a multiplex ligation-dependent probe amplification study. Cell Oncol (Dordr) 34:489–494
Berge EO, Knappskog S, Geisler S et al (2010) Identification and characterization of retinoblastoma gene mutations disturbing apoptosis in human breast cancers. Mol Cancer 9:173
Reintjes N, Li Y, Becker A et al (2013) Activating somatic FGFR2 mutations in breast cancer. PLoS ONE 8:e60264
Milne AN, Leguit R, Corver WE et al (2010) Loss of CDC4/FBXW7 in gastric carcinoma. Cell Oncol 32:347–359
Hess CJ, Denkers F, Ossenkoppele GJ et al (2004) Gene expression profiling of minimal residual disease in acute myeloid leukaemia by novel multiplex-PCR-based method. Leukemia 18:1981–1988
Veronese L, Tournilhac O, Combes P et al (2013) Contribution of MLPA to routine diagnostic testing of recurrent genomic aberrations in chronic lymphocytic leukemia. Cancer Genet 206:19–25
Jia H, Jiang X, Zhao Z et al (2009) High frequency of down-regulation of E-cadherin detected in benign sporadic insulinomas by multiplex ligation-dependent probe amplification. Hum Pathol 40:1336–1341
Newey PJ, Nesbit MA, Rimmer AJ et al (2012) Whole-exome sequencing studies of nonhereditary (sporadic) parathyroid adenomas. J Clin Endocrinol Metab 97:E1995–E2005
Cromer MK, Starker LF, Choi M et al (2012) Identification of somatic mutations in parathyroid tumors using whole-exome sequencing. J Clin Endocrinol Metab 97:E1774–E1781
Bergero N, De Pompa R, Sacerdote C et al (2005) Galectin-3 expression in parathyroid carcinoma: immunohistochemical study of 26 cases. Hum Pathol 36:908–914
Haven CJ, Howell VM, Eilers PH et al (2004) Gene expression of parathyroid tumors: molecular subclassification and identification of the potential malignant phenotype. Cancer Res 64:7405–7411
Mohebati A, Shaha A, Shah J (2012) Parathyroid carcinoma: challenges in diagnosis and treatment. Hematol Oncol Clin North Am 26:1221–1238
Forsberg L, Bjorck E, Hashemi J et al (2005) Distinction in gene expression profiles demonstrated in parathyroid adenomas by high-density oligoarray technology. Eur J Endocrinol 152:459–470
Woodard GE, Lin L, Zhang JH et al (2005) Parafibromin, product of the hyperparathyroidism-jaw tumor syndrome gene HRPT2, regulates cyclin D1/PRAD1 expression. Oncogene 24:1272–1276
Imanishi Y, Hosokawa Y, Yoshimoto K et al (2001) Primary hyperparathyroidism caused by parathyroid-targeted overexpression of cyclin D1 in transgenic mice. J Clin Invest 107:1093–1102
Hsi ED, Zukerberg LR, Yang WI et al (1996) Cyclin D1/PRAD1 expression in parathyroid adenomas: an immunohistochemical study. J Clin Endocrinol Metab 81:1736–1739
Vasef MA, Brynes RK, Sturm M et al (1999) Expression of cyclin D1 in parathyroid carcinomas, adenomas, and hyperplasias: a paraffin immunohistochemical study. Mod Pathol 12:412–416
Tahara H, Smith AP, Gas RD et al (1996) Genomic localization of novel candidate tumor suppressor gene loci in human parathyroid adenomas. Cancer Res 56:599–605
Friedman E, De Marco L, Gejman PV et al (1992) Allelic loss from chromosome 11 in parathyroid tumors. Cancer Res 52:6804–6809
Acknowledgments
The National Natural Science Foundation of China supported this study (81200647, 81370977, 81170804, 81000359) along with the Shanghai Municipal Health Bureau (XBR2011013 and 2012-235) and Shanghai young teachers of universities support program from Shanghai education committee (2013).
Conflict of interest
None.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Lin Zhao, Li-hao Sun and Dong-mei Liu have contributed equally to this work.
Rights and permissions
About this article
Cite this article
Zhao, L., Sun, Lh., Liu, Dm. et al. Copy Number Variation in CCND1 Gene Is Implicated in the Pathogenesis of Sporadic Parathyroid Carcinoma. World J Surg 38, 1730–1737 (2014). https://doi.org/10.1007/s00268-014-2455-9
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00268-014-2455-9