Abstract
Objective
Solid tumors often establish a procoagulable state that can lead to venous thromboembolism (VTE). Although some of the key genes involved in this process are known, no previous study has compared the “coagulome”, i.e., the expression of coagulation/fibrinolysis genes, across different primary tumor types. It is also unclear whether the coagulome is associated with specific characteristics of the tumor microenvironment (TME). We aimed to address this question.
Methods
We analyzed the expression of the genes F3, PLAU, PLAT, PLAUR, SERPINB2, and SERPINE1 in 32 cancer types using data from The Cancer Genome Atlas (TCGA) and other freely available resources.
Results
We identified specific expression patterns of procoagulant and fibrinolytic genes. The expression of the Tissue Factor (F3) was found to be tumor type dependent, with the highest expression in glioblastoma (GBM), a highly procoagulable tumor type. Conversely, high expression of the fibrinolysis gene cluster PLAU, PLAUR, SERPINE1 was consistently linked to the characteristics of the TME (monocytic infiltration) and high expression of important checkpoints of the immune response, such as PD-L2 and CD276/B7-H3.
Conclusion
These tumor-specific patterns of expression might partially explain the differences in VTE risk among tumor types. We propose that biomarkers of coagulation fibrinolysis might provide valuable information about the TME in cancer patients.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Venous thromboembolism (VTE) frequently occurs in patients with solid tumors and represents a major cause of mortality and morbidity in cancer patients [1, 2]. Depending on the study and their design, cancer patients are reported to have a 4- to sevenfold increase in the relative risk of VTE as compared to the general population or patients without cancer [1, 2]. The pathogenesis of cancer-associated thrombosis (CAT) is complex. General risk factors (typically older age and reduced mobility) and the use of potentially procoagulant anticancer therapies (surgery, chemotherapy, and antiangiogenic drugs) are well-recognized risk factors for VTE [3]. To date, thrombophylaxis is not routinely recommended for all outpatients with cancer, but a regular assessment of the risk of VTE is recommended [4, 5]. Importantly, different types of primary tumors vary greatly in their propensity to cause VTE: glioblastoma multiforme (GBM) and pancreatic adenocarcinoma (PAAD) are typically considered to be the tumor types with the highest risk of VTE [1, 2]. The differences in the risk of VTE across tumor types could partially be accounted for by differences in the prothrombic properties of the tumor tissue, but currently there are no studies that address this possibility across a large set of human tumors.
Tumor cells can directly activate blood clotting by producing and releasing the major procoagulant factor, Tissue Factor (TF), encoded by the gene F3 [3, 6]. TF is a cell-associated receptor that can activate the coagulation factor VII, leading to the activation of the common pathway and activating thrombin, thus promoting coagulation in a large variety of tumor types [6]. TF is typically expressed by cancer cells and the multiple nonmalignant cell types that constitute the tumor microenvironment (TME). It plays a pivotal role in CAT, either at the tumor cell surface or in the form of TF-bearing microparticles that are shed in the TME [7]. GBM, which has been identified as a high-risk tumor type for VTE, express TF at high levels [8]. In GBM, the expression of TF is related to the histological subtype of GBM, the presence of genomic alterations, and possibly also the acquisition of mutations in proto-oncogenes or miRNA [8]. The procoagulant effect of TF is counteracted by fibrinolysis. The Tissue Plasminogen Activator (tPA) and Urokinase Plasminogen Activator (uPA), two serine proteases encoded by the genes PLAT and PLAU, respectively, activate plasminogen, which degrades fibrin. Their activity is inhibited by the serpin inhibitors plasminogen activator inhibitor-1 (PAI-1) and plasminogen activator inhibitor-2 (PAI-2) encoded by the genes SERPINE1 and SERPINB2, respectively. The activity of uPA is increased upon its binding to its glycolipid-anchored receptor, uPAR (encoded by the gene PLAUR) [3]. The qualitative equilibrium achieved between procoagulant and fibrinolytic cascades defines a tumor-specific « coagulome», as was proposed by Rak and colleagues [9, 10].
The tumor microenvironment (TME) consists of a variety of cell types that have a symbiotic relationship and contribute to the tumor ecosystem [11]. Coagulation and fibrinolysis are under complex regulation by inflammation and the local recruitment of leukocytes in the TME [12, 13]. To date however, no study has addressed in depth the link between the coagulome and the cellular nature and properties of the TME. Importantly, the exploitation of genomic data has recently permitted progress toward a better understanding of TME regulation [14]. Genomic data, especially that made available from The Cancer Genome Atlas (TCGA), enable pan-cancer studies covering multiple aspects of cancer biology [15, 16], including the study of the TME [17,18,19]. In the present study, we examined the expression and regulation of the tumor coagulome across the main human tumors. We used RNAseq data to analyze mRNA levels of six key genes of coagulation and fibrinolysis (F3, PLAT, PLAU, PLAUR, SERPINE1 and SERPINB2) to explore their expression in relation to clinical and pathological parameters in 10,071 individual tumor samples and 32 tumor types.
Materials and methods
Patient and gene expression data
Basic clinical, pathological, and genomic data (RNA SeqV2 data normalised using RNA-Seq by Expectation Maximization: RSEM) were retrieved using cBioportal at: https://cbioportal.org [20, 21]. The tumor types and the number of samples for each are: acute myeloid leukemia (LAML, n = 173), adrenocortical carcinoma (ACC, n = 78), bladder urothelial carcinoma (BLCA, n = 407), brain lower grade glioma (LGG, n = 514), breast invasive carcinoma (BRCA, n = 1082), cervical squamous cell carcinoma (CESC, n = 294), cholangiocarcinoma (CHOL, n = 36), colorectal adenocarcinoma (COAD, n = 592), diffuse large B-cell lymphoma (DLBC, n = 48), esophageal adenocarcinoma (ESCA, n = 181), glioblastoma multiforme (GBM, n = 160), head and neck squamous cell carcinoma (HNSC, n = 515), kidney chromophobe (KICH, n = 65), kidney renal clear cell carcinoma (KIRC, n = 510), kidney renal papillary cell carcinoma (KIRP, n = 283), liver hepatocellular carcinoma (LIHC, n = 366), lung adenocarcinoma (LUAD, n = 510), lung squamous cell carcinoma (LUSC, n = 484), mesothelioma(MESO, n = 87), ovarian serous cystadenocarcinoma (OV, n = 300), pancreatic adenocarcinoma (PAAD, n = 177), pheochromocytoma and paraganglioma (PCPG, n = 178), prostate adenocarcinoma (PRAD, n = 493), sarcoma (SARC, n = 253), skin cutaneous melanoma (SKCM, n = 443), stomach adenocarcinoma (STAD, n = 412), testicular germ cell tumors (TGCT, n = 149), thymoma (THYM, n = 119), thyroid carcinoma (THCA, n = 498), uterine carcinosarcoma (UCS, n = 57), uterine corpus endometrial carcinoma (UCEC, n = 527), uveal melanoma (UVM, n = 80). The thromboembolic risk for different tumor types was based on the study by Blom et al. [22].
Gene ontology analysis
Gene Set Enrichment Analysis (GSEA) was performed using the Java GSEA desktop application. We followed the standard procedures (https://www.gsea-msigdb.org/gsea/index.jsp). We used curated hallmark gene sets, downloaded from the GSEA website, to compute their overrepresentation in RNAseq tumor samples with high expression of F3 or PLAU (high vs low expression defined by the median). The analyses were done using 1,000 permutations [23].
Tumor microenvironment analysis
The microenvironment cell population counter (MCP counter) method was used to quantify the relative abundance of eight types of immune and stromal cell populations based on the RNA seq data [24]. Panels of immune genes were recovered from the study by Thorsson et al. [17].
Statistics
Comparisons of two groups of numeric data were performed using the unpaired Wilcoxon–Mann–Whitney test. Where appropriate the false-discovery rate (FDR) correction (Bonferroni) was applied to control for multiple testing. p < 0.05 was set as the threshold for significance. Heatmaps were created using the R library gplots—the clustering method used was Ward.D2. The association of F3 and PLAU genes to overall survival (OS) and disease-free survival (DFS) was studied by calculating the hazard ratios (HR) and 95% confidence intervals for each gene (Cox proportional hazards regression model). All statistical analyses were done with R version 3.4.2 (https://www.r-project.org). All correlation analyses were done using R, packages Hmisc, and corrplot, calculating Pearson correlation coefficients r.
Results
Transcriptional regulation of the coagulome in human tumors
Based on the literature, we selected six genes that have been reported to constitute the core coagulome in human tumors: F3, PLAT, PLAU, PLAUR, SERPINE1 and SERPINB2, encoding TF, tPA, uPA, uPAR, PAI-1, and PAI-2, respectively. To examine their patterns of expression in human tumors, we retrieved RNA seq data for the corresponding genes. A pan-cancer comparison revealed great differences among the different types of primary tumors, with up to 250-fold difference in the median expression between tumor types (Fig. 1A). When compared with other tumors, GBM were characterized by the highest F3 mRNA expression levels with an average expression of 3841 RSEM compared to the overall pan-cancer average of 1516 RSEM (p < 2.2e-16) (Fig. 1A). In order to address the existence of different patterns of expression of procoagulant/fibrinolytic proteins among tumors, we performed a pan-cancer analysis, after a normalization step for each gene (Z score), combined with hierarchical clustering by cancer type (Fig. 1B). Using this analysis, we noticed a dissociation between the expression of the procoagulant gene F3 and the pro-fibrinolytic genes PLAU, PLAUR and SERPINE1 in some tumors (Fig. 1B). For example, GBM expressed the highest F3 mRNA levels among the different tumor types, but relatively lower mRNA levels of PLAU (average 1156 RSEM, ranked 18/32 among tumor types), PLAUR (720 RSEM average, ranked 19/32), and SERPINE1 (5714 RSEM average, ranked 9/32) (Fig. 1B). Taking the complete tumor set as a whole, we noticed that the expression of the genes PLAU, PLAUR, and SERPINE1 clustered together (suppl. Figure 1). This analysis was confirmed by measuring the correlation coefficient r for their mutual expression: for the three genes PLAU, PLAUR, and SERPINE1, the correlation coefficients r were higher than 0.3 (Suppl. Figure 1). This initial analysis provided an overview of the tumor coagulome. We concluded that differences exist among the human tumor types and individual tumors in their expression of the coagulome genes.
Coagulome gene expression in human tumors. A. Dot plots showing the tumor type ranking according to the mRNA expression levels of six essential components of the tumor coagulome (F3, PLAT, SERPINB2, PLAU, PLAUR and SERPINE1). Data were retrieved from TCGA, with n = 32 tumor types and a total number of n = 10,071 tumors. B. Heatmap comparison of F3, PLAU, PLAUR, PLAT, SERPINB2 and SERPINE1 pattern of expression across different tumor types. For each gene, the expression was normalised by tumor type (z score). Red indicates high expression (positive z score), blue indicates low expression (negative z score)
Correlation between the expression of the coagulation/fibrinolysis genes and the risk of VTE across tumor types
In order to link the pattern of gene expression shown in Fig. 1 with the risk of VTE, we used data published by Blom et al., reporting an analysis of a large Dutch cancer registry covering 66,329 cancer patients, that included a large number of tumor types (including some primary tumors with low incidence) [22]. We carried out a correlation analysis between the mRNA levels of each of our key coagulation/fibrinolysis genes (based on RSEM data from TCGA) and the risk of VTE (measured as a cumulative incidence for 1000 patients) from the study by Blom et al. [22] (Fig. 2). A positive correlation was found between F3 mRNA expression and VTE incidence (Pearson r = 0.53, p = 0.036), suggesting the clinical relevance of the mRNA expression study based on TCGA. To address the possibility that the expression of the F3 and PLAU genes may have a prognostic value, a univariate Cox proportional hazards regression model was used to calculate the hazard ratios for overall survival (OS) and disease-free survival (DFS) across the different types of primary tumors (Supp. Figure 2). A hazard ratio > 1, representing a significantly reduced OS and an unfavorable outcome was observed for F3 in GBM (HR = 1.2, p = 0.036), PLAU in PAAD (1.3, p = 0.000023) and PLAU in HNSCC (HR = 1.2, p = 0.00032) (Suppl. Figure 2).
A correlation between coagulome gene expression and the risk of VTE. The graphs show the correlation between the mRNA gene expression levels (RSEM) of F3, PLAU, PLAUR, SERPINE1, SERPINB2 and PLAT and the risk of VTE (based on data from Blom et al. [22]). The risk of VTE was measured as a cumulative incidence for 1000 patients. A positive correlation was found between F3 mRNA expression and the incidence of VTE (Pearson r = 0.53, p = 0.036)
Gene set enrichment analysis (GSEA)
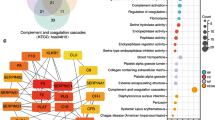
Gene Set Enrichment Analysis (GSEA) revealed that the F3 gene expression profile was the most positively associated with the Hallmark term “TNFA signaling via NFKB” in PAAD with a normalized enrichment score (NES) of 2.06 (p = 0.016 FDR) (Fig. 3A). The second most enriched gene set was “Hypoxia” (NES = 1.97, p = 0.023 FDR). In contrast, PLAU gene expression was most positively associated with the Hallmark term “Epithelial Mesenchymal Transition” (NES = 2.20, p = 0 FDR). The second most enriched term was “Apical junction” (NES = 2.14, p = 0.001 FDR) (Fig. 3B). Interestingly, these observations were confirmed in three additional cancer types: GBM, HNSC and PRAD (Fig. 3C), where F3 expression was positively associated with “TNFA signaling via NFKB”, ranking first for three of the four cancer types. In all four cancer types examined PLAU expression was positively associated with “Epithelial Mesenchymal Transition”. Together, these analyses indicated that F3 and PLAU are associated with different biological processes within the tumor.
Gene Ontology GSEA analysis for F3 and PLAU genes across human tumors. A. Ranking of the Hallmark gene sets that were enriched in high-F3 (top 50% in mRNA expression) and high-PLAU (top 50% in mRNA expression) PAAD. B. GSEA analysis revealed an enrichment of the “TNFA signaling via NFKB” gene set in high-F3 PAAD, HNSC, GBM, and PRAD, and an enrichment of the “Epithelial Mesenchymal Transition” gene set in high-PLAU PAAD, HNSC, GBM, PRAD
The coagulome is related to the cellular composition and the immune activity of the TME
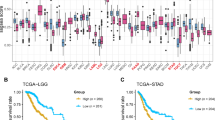
To address the contribution of the heterogeneous cell composition of the tumors and relate it to the gene expression patterns detected previously, we used the algorithm MCP counter, which is based on the detection of cell type-specific mRNA [18, 19, 24]. We calculated a Pearson’s correlation coefficient between each coagulome gene and the tumor infiltration of each cell type analyzed (T cells, CD8 T cells, Cytotoxic T, Natural Killer cells, B cells, monocytic cells, myeloid cells, neutrophils, endothelial cells and fibroblasts) for each cancer type (Suppl. Tables 1–3). A heat map was constructed with the corresponding Pearson coefficients r (Fig. 4). Interestingly, this analysis revealed two predominant patterns, depending on the gene considered: for F3 and PLAT, the correlations between the mRNA levels and the density of the cell populations were relatively stable and tumor type-dependent (Fig. 4). A different pattern was noticed with the genes PLAU, PLAUR, and SERPINE1. The mRNA levels of the corresponding genes were positively correlated with tumor infiltration by cells of the monocytic lineage and fibroblasts, independently of the type of tumor considered (Fig. 4). For the monocytic lineage in a pan-cancer analysis the average correlations were as follows: PLAU r = 0.25, PLAUR r = 0.35, SERPINE1 r = 0.19. These correlations were notably high for example for COAD (PLAU r = 0.42, PLAUR r = 0.40, SERPINE1 r = 0.30) and BLCA (PLAU r = 0.36, PLAUR r = 0.52, SERPINE1 r = 0.28). For fibroblasts in a pan-cancer analysis the average pan-cancer correlations were as follows: PLAU r = 0.35, PLAUR r = 0.34, SERPINE1 r = 0.34. After observing this consistent positive correlation with the monocytic infiltrate, we decided to further examine the possibility of a link with the active status of the immune microenvironment. We carried out a correlation analysis of the expression of the key genes of the coagulome with the 66 immune regulatory genes reported by Thorsson et al. [17], classified into seven categories (co-stimulator, co-inhibitor, ligand, receptor, cell adhesion, antigen presentation, and other) among various cancer types (Fig. 5). This analysis confirmed the existence of a positive correlation between the expression of immune genes and the fibrinolysis gene cluster PLAU, PLAUR and SERPINE1. For PLAU, the strongest association was observed for the genes PDCD1LG2 and CD276, encoding the checkpoints PD-L2 and B7-H3, for which we obtained an average Pearson r coefficient of 0.376 (p < 1.0e-20) and 0.292 (p < 1.0e-20), respectively, in a pan-cancer analysis. Finally, we directly compared the expression levels of these two immune checkpoints in ten of the most frequent primary human tumors. For each tumor type, we selected the tumors with high and low expression of PLAU (upper and lower quartile, respectively), and we compared the mRNA expression levels (RSEM) for the two checkpoints, PD-L2 and B7-H3 (Fig. 6). PDCD1LG2 expression levels were significantly increased in all tumor types. The fold expression of PDCD1LG2 in high PLAU-expressing tumors compared to low PLAU-expressing tumors was as follows: BRCA 2.1-fold increase (p < 2.2e-15, FDR), COAD 6.7-fold increase (p < 2.2e-15, FDR), KIRC 2.1-fold increase (p = 4.61e-9, FDR), LIHC 6.1-fold increase (p < 2.2e-15, FDR), SKCM sevenfold increase (p < 2.2e-15, FDR), PRAD 2.7-fold increase (p < 2.2e-15, FDR), GBM 2.5-fold increase (p = 0.004262, FDR), LUAD threefold increase (p < 2.2e-15, FDR), HNSC 2.5-fold increase (p = 3.314e-13, FDR), and PAAD 2.6-fold increase (p = 1.367e-06, FDR). CD276 expression levels were significantly increased in almost all tumor types, excluding PRAD (p = 0.7281), when comparing high PLAU-expressing tumors and low PLAU-expressing tumors (BRCA 1.6-fold increase (p < 2.2e-15, FDR), COAD 1.6-fold increase (p < 2.2e-15, FDR), KIRC 1.7-fold increase (p < 2.2e-15, FDR), LIHC 1.6-fold increase (p = 5.4e-10, FDR), SKCM 1.2-fold increase (p = 0.038, FDR), GBM 1.7-fold increase (p = 6.5e-07), LUAD 1.5-fold increase (p < 2.2e-15, FDR), HNSC 2.2-fold increase (p < 2.2e-15, FDR), and PAAD 2.1-fold increase (p = 2.98e-14, FDR)). We concluded that the PLAU, PLAUR and SERPINE1 cluster correlated with the presence of a “hot” tumor immune environment across tumor types.
The coagulome gene expression pattern correlates with immune cell infiltration. A Pearson correlation analysis between the expression of the indicated genes and the tumor infiltration with different cell types was performed. The infiltration was calculated for eight types of immune cells and two stromal cell types using the MCP counter algorithm. The corresponding heatmaps show the Pearson r for each of the six genes analyzed. Red corresponds to a positive correlation, blue to a negative correlation. For each of the genes a pan-cancer correlation was also performed
A correlation between the expression of the coagulome and genes encoding immune regulatory molecules. For each gene of the coagulome, we show the Pearson coefficient r with the expression levels of the genes involved in local immune regulation (n = 66), as previously identified by Thorsson et al. [17]. These analyses were done by cancer type
PDCD1LG2 and CD276 expression in the most frequent primary tumors, stratified according to PLAU expression. For each of the 10 most common primary tumors, we compared the mRNA levels of PDCD1LG2 and CD276 in tumors with high PLAU (top 25% in PLAU mRNA expression) vs low PLAU (bottom 25%). The expression levels are given as RSEM (RNA-Seq by Expectation–Maximization). **p < 0.01; ***p < 0.001, ns = not significant using the Wilcoxon–Mann–Whitney test
Discussion
In the present study, we used TCGA, the only large database with RNAseq data for most human tumor types, in order to establish the landscape of the cancer coagulome. We found great differences between primary human tumor types, and observed that pro-coagulant and pro-fibrinolytic genes are regulated separately: the pattern of expression of F3 did not match that of the fibrinolytic genes (PLAT or the cluster PLAU/PLAUR/SERPINE1). Each type of primary tumor is characterized by a specific balance between the pro-coagulant and fibrinolytic genes. In support of the clinical relevance of our findings, we found that the most thrombogenic tumors, such as GBM and PAAD, express high levels of F3 mRNA [1, 2]. A positive correlation was seen between F3 expression and the VTE risk. This correlation was; however, limited, with a Pearson’s coefficient of r = 0.53, probably reflecting the contribution of multiple genes and the complex regulation of coagulation/fibrinolysis in solid tumors. An optimal prediction of the risk of VTE based on tumor gene expression would likely necessitate machine learning and more refined mathematical modeling, but this was not the aim of the present study. Importantly however, we noted that some tumor types, such as HNSC, express both high levels of F3 and high levels of the fibrinolytic genes. In this respect, we provide support to the recently formulated hypothesis that a high fibrinolytic activity might counterbalance the pro-coagulant effect of TF and explain the paradoxical low risk of VTE in these tumors [25]. While our study might partially explain why different primary tumor types have a different risk of developing VTE, it has a number of important limitations. The first is the lack of validation of our conclusions in an independent cohort, an obstacle that we could not address because of the lack of a study that would match TCGA in terms of primary tumor coverage. Another limitation resides in the recruitment of patients differing in the stages of cancer and the treatment received, variables that we could not address in the present study because of lack of precise clinical data. Compared to previous studies, including those published by Rak and colleagues [8, 9], our study nevertheless represents the first attempt to chart the landscape of the human tumor coagulome. Clearly, this analysis is open for further studies that may include novel actors of coagulation as their contribution to VTE unfolds.
Interestingly, we found a correlation between the tumor coagulome and some of the characteristics of the TME. The expression of F3 correlated with the hallmark gene set “TNFα-signaling via NFκB” and there was also a weak, yet consistent positive correlation with tumor infiltration by neutrophils. These results are in agreement with previous studies showing that the TNFα has a direct pro-coagulant effect in CAT [12, 13]. Upon exposure to TNFα, cancer cells increase their expression of TF and produce TF-bearing microparticles with potent local pro-coagulant effects [12, 13]. The presence of neutrophils that release their chromatin as NET (Neutrophils–Extracellular Traps) is also an inflammatory feature that has been reported in CAT [12, 13]. Our study therefore highlights the coordinated interplay between local inflammation and coagulation in the TME. Interestingly, the expression of the PLAU/PLAUR/SERPINE1 gene cluster was related to the hallmark gene set “Epithelial–Mesenchymal Transition”. This cluster was also consistently correlated with tumor infiltration with cells of the fibroblastic and monocytic lineages across different tumor types. These findings are in agreement with previous studies that reported that PAI-1, encoded by SERPINE1 is expressed by cancer cells undergoing EMT [26]. EMT is a process of phenotypic plasticity that has multiple roles in organ development, wound healing, tumor progression and response to therapeutics [27]. The existence of possible reciprocal regulation between EMT and the TME is a matter of discussion [27,28,29]. This withstanding, our findings support the notion that the procoagulant and fibrinolytic systems, besides their antagonistic action on VTE, may have subtly different and nonoverlapping effects on the TME.
Importantly, our study suggests that tumors with high expression of the PLAU/PLAUR/SERPINE1 gene cluster are characterized by a “hot” immune microenvironment. Our data are in complete agreement with the recent study by Kubaka et al. (2018), showing that PAI-1 plays an active role in the regulation of the recruitment and functional polarization of CD163+ve Tumor-associated macrophages [30]. The recent introduction of immune checkpoint blockers (ICB) represents a major advancement for various types of tumors [31]. There is currently a great need for biomarkers that could anticipate their efficacy in individual patients. The observation that the expression of PLAU correlates with the mRNA levels of two important checkpoints of the immune response, PD-L2 and CD276/B7-H3 [32, 33], is interesting in this regard. We propose that the biomarkers of fibrinolysis might be useful for the assessment of the presence of immune checkpoints or the active immune status of the TME, a possibility that has to the best of our knowledge not yet been addressed [34]. The tumor coagulome is also a direct target of several therapeutics that are approved for use in humans, and it is an actionable component of the TME [35]. We propose that the landscape of human tumor coagulome that we report in this study will be a valuable resource for future research exploring the contribution of vascular biology to the TME and to the outcome of cancer immunotherapy [36, 37].
Abbreviations
- LAML:
-
Acute myeloid leukemia
- ACC:
-
Adrenocortical carcinoma
- BLCA:
-
Bladder urothelial carcinoma
- LGG:
-
Brain lower grade glioma
- BRCA:
-
Breast invasive carcinoma
- CAT:
-
Cancer associated thrombosis
- CESC:
-
Cervical squamous cell carcinoma
- CHOL:
-
Cholangiocarcinoma
- COAD:
-
Colorectal adenocarcinoma
- DFS:
-
Disease-free survival
- DLBC:
-
Diffuse large B-cell lymphoma
- EMT:
-
Epithelial mesenchymal transition
- ESCA:
-
Esophageal adenocarcinoma
- FDR:
-
False-discovery rate
- GBM:
-
Glioblastoma multiforme
- GSEA:
-
Gene set enrichment analysis
- HNSC:
-
Head and neck squamous cell carcinoma
- ICB:
-
Immune checkpoint blockers
- KICH:
-
Kidney chromophobe
- KIRCH:
-
Kidney renal clear cell carcinoma
- KIRP:
-
Kidney renal papillary cell carcinoma
- LIHC:
-
Liver hepatocellular carcinoma
- LUAD:
-
Lung adenocarcinoma
- LUSC:
-
Lung squamous cell carcinoma
- MESO:
-
Mesothelioma
- NES:
-
Normalized enrichment score
- OS:
-
Overall survival
- OV:
-
Ovarian serous cystadenocarcinoma
- PAAD:
-
Pancreatic adenocarcinoma
- PAI-1/2:
-
Plasminogen activator inhibitor-1/2
- PCPG:
-
Pheochromocytoma and paraganglioma
- PRAD:
-
Prostate adenocarcinoma
- RSEM:
-
RNA-seq by expectation maximization
- SARC:
-
Sarcoma
- SKCM:
-
Skin cutaneous melanoma
- STAD:
-
Stomach adenocarcinoma
- TCGA:
-
The Cancer Genome Atlas
- TGCT:
-
Testicular germ cell tumors
- THYM:
-
Thymoma
- THCA:
-
Thyroid carcinoma
- TF:
-
Tissue factor
- tPA:
-
Plasminogen activator
- TME:
-
Tumor microenvironment
- UCS:
-
Uterine Carcinosarcoma
- UCEC:
-
Uterine corpus endometrial carcinoma
- uPA:
-
Urokinase plasminogen activator
- UVM:
-
Uveal melanoma
- VTE:
-
Venous thromboembolism
References
Chew HK, Wun T, Harvey D, Zhou H, White RH (2006) Incidence of venous thromboembolism and its effect on survival among patients with common cancers. Arch Intern Med 166:458–464. https://doi.org/10.1001/archinte.166.4.458
Timp JF, Braekkan SK, Versteeg HH, Cannegieter SC (2013) Epidemiology of cancer-associated venous thrombosis. Blood 122:1712–1723. https://doi.org/10.1182/blood-2013-04-460121
Falanga A, Schieppati F, Russo L (2019) Mechanisms of Thrombosis in Cancer Patients. Cancer Treat Res 179:11–36. https://doi.org/10.1007/978-3-030-20315-3_2
Farge D, Frere C, Connors JM, Ay C, Khorana AA, Munoz A et al (2019) international clinical practice guidelines for the treatment and prophylaxis of venous thromboembolism in patients with cancer. Lancet Oncol 20:e566–e581. https://doi.org/10.1016/S1470-2045(19)30336-5
Key NS, Khorana AA, Kuderer NM, Bohlke K, Lee AYY, Arcelus JI et al (2020) Venous Thromboembolism Prophylaxis and Treatment in Patients With Cancer: ASCO Clinical Practice Guideline Update. J Clin Oncol 38:496–520. https://doi.org/10.1200/JCO.19.01461
Grover SP, Mackman N (2018) Tissue Factor: An Essential Mediator of Hemostasis and Trigger of Thrombosis. Arterioscler Thromb Vasc Biol 38:709–725. https://doi.org/10.1161/ATVBAHA.117.309846
Rondon AMR, Kroone C, Kapteijn MY, Versteeg HH, Buijs JT (2019) Role of Tissue Factor in Tumor Progression and Cancer-Associated Thrombosis. Semin Thromb Hemost 45:396–412
Magnus N, Meehan B, Garnier D, Hashemi M, Montermini L, Lee TH et al (2014) The contribution of tumor and host tissue factor expression to oncogene-driven gliomagenesis. Biochem Biophys Res Commun 454:262–268. https://doi.org/10.1016/j.bbrc.2014.10.041
Tawil N, Chennakrishnaiah S, Bassawon R, Johnson R, D’Asti E, Rak J (2018) Single cell coagulomes as constituents of the oncogene-driven coagulant phenotype in brain tumours. Thromb Res 164:S136–S142. https://doi.org/10.1016/j.thromres.2018.01.021
Tawil N, Bassawon R, Rak J (2019) Oncogenes and Clotting Factors: The Emerging Role of Tumor Cell Genome and Epigenome in Cancer-Associated Thrombosis. Semin Thromb Hemost 45:373–384. https://doi.org/10.1055/s-0039-1687891
Binnewies M, Roberts EW, Kersten K, Chan V, Fearon DF, Merad M et al (2018) Understanding the tumor immune microenvironment (TIME) for effective therapy. Nat Med 24:541–550. https://doi.org/10.1038/s41591-018-0014-x
Foley JH, Conway EM (2016) Cross Talk Pathways Between Coagulation and Inflammation. Circ Res 118:1392–1408. https://doi.org/10.1161/CIRCRESAHA.116.306853
Date K, Ettelaie C, Maraveyas A (2017) Tissue factor-bearing microparticles and inflammation: a potential mechanism for the development of venous thromboembolism in cancer. J Thromb Haemost 15:2289–2299. https://doi.org/10.1111/jth.13871
Lau D, Bobe AM, Khan AA (2019) RNA Sequencing of the Tumor Microenvironment in Precision Cancer Immunotherapy. Trends Cancer 5:149–156. https://doi.org/10.1016/j.trecan.2019.02.006
Hutter C, Zenklusen JC (2018) The Cancer Genome Atlas: Creating Lasting Value beyond Its Data. Cell 173:283–285. https://doi.org/10.1016/j.cell.2018.03.042
Blum A, Wang P, Zenklusen JC (2018) TCGA-Analyzed Tumors. Cell 173:530. https://doi.org/10.1016/j.cell.2018.03.059
Thorsson V, Gibbs DL, Brown SD, Wolf D, Bortone DS, Ou Yang TH et al (2018) The Immune Landscape of Cancer. Immunity 48:812–830. https://doi.org/10.1016/j.immuni.2018.03.023
Job S, Rapoud D, Dos Santos A, Gonzalez P, Desterke C, Pascal G et al (2019) Identification of four immune subtypes characterized by distinct composition and functions of tumor microenvironment in intrahepatic cholangiocarcinoma. Hepatology. https://doi.org/10.1002/hep.31092
Helmink BA, Reddy SM, Gao J, Zhang S, Basar R, Thakur R et al (2020) B cells and tertiary lymphoid structures promote immunotherapy response. Nature 577:549–555. https://doi.org/10.1038/s41586-019-1922-8
Cerami E, Gao J, Dogrusoz U, Gross BE, Sumer SO, Aksoy BA et al (2012) The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov 2:401–404. https://doi.org/10.1158/2159-8290.CD-12-0095
Gao J, Aksoy BA, Dogrusoz U, Dresdner G, Gross B, Sumer SO et al. (2013) Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal 6, pl1. doi: https://doi.org/10.1126/scisignal.2004088.
Blom JW, Vanderschoot JP, Oostindiër MJ, Osanto S, van der Meer FJ, Rosendaal FR (2006) Incidence of venous thrombosis in a large cohort of 66,329 cancer patients: results of a record linkage study. J Thromb Haemost 4:529–535. https://doi.org/10.1111/j.1538-7836.2006.01804.x
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA et al (2005) Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA 102:15545–15550. https://doi.org/10.1073/pnas.0506580102
Becht E, Giraldo NA, Lacroix L, Buttard B, Elarouci N, Petitprez F et al (2016) Estimating the population abundance of tissue-infiltrating immune and stromal cell populations using gene expression. Genome Biol 17:218. https://doi.org/10.1186/s13059-016-1070-5
Haen P, Mege D, Crescence L, Dignat-George F, Dubois C, Panicot-Dubois L (2019) Thrombosis Risk Associated with Head and Neck Cancer: A Review. Int J Mol Sci. 20, pii:E2838. doi: https://doi.org/10.3390/ijms20112838.
Pavón MA, Arroyo-Solera I, Céspedes MV, Casanova I, León X, Mangues R (2016) uPA/uPAR and SERPINE1 in head and neck cancer: role in tumor resistance, metastasis, prognosis and therapy. Oncotarget 7:57351–57366. https://doi.org/10.18632/oncotarget.10344
Dongre A, Weinberg RA (2019) New insights into the mechanisms of epithelial-mesenchymal transition and implications for cancer. Nat Rev Mol Cell Biol 20:69–84. https://doi.org/10.1038/s41580-018-0080-4
Lou Y, Diao L, Cuentas ER, Denning WL, Chen L, Fan YH et al (2016) Epithelial-Mesenchymal Transition Is Associated with a Distinct Tumor Microenvironment Including Elevation of Inflammatory Signals and Multiple Immune Checkpoints in Lung Adenocarcinoma. Clin Cancer Res 22:3630–3642. https://doi.org/10.1158/1078-0432.CCR-15-1434
Mak MP, Tong P, Diao L, Cardnell RJ, Gibbons DL, William WN et al (2016) A Patient-Derived, Pan-Cancer EMT Signature Identifies Global Molecular Alterations and Immune Target Enrichment Following Epithelial-to-Mesenchymal Transition. Clin Cancer Res 22:609–620. https://doi.org/10.1158/1078-0432.CCR-15-0876
Kubala MH, Punj V, Placencio-Hickok VR, Fang H, Fernandez GE, Sposto R, DeClerck YA (2018) Plasminogen Activator Inhibitor-1 Promotes the Recruitment and Polarization of Macrophages in Cancer. Cell Rep 25:2177–2191. https://doi.org/10.1016/j.celrep.2018.10.082
Wei SC, Duffy CR, Allison JP (2018) Fundamental Mechanisms of Immune Checkpoint Blockade Therapy. Cancer Discov 8:1069–1086. https://doi.org/10.1158/2159-8290.CD-18-0367
Okadome K, Baba Y, Nomoto D, Yagi T, Kalikawe R, Harada K et al (2020) Prognostic and clinical impact of PD-L2 and PD-L1 expression in a cohort of 437 oesophageal cancers. Br J Cancer 122:1535–1543. https://doi.org/10.1038/s41416-020-0811-0
Picarda E, Ohaegbulam KC, Zang X (2016) Molecular Pathways: Targeting B7–H3 (CD276) for Human Cancer Immunotherapy. Clin Cancer Res 22:3425–3431. https://doi.org/10.1158/1078-0432.CCR-15-2428
Faille D, Bourrienne MC, de Raucourt E, de Chaisemartin L, Granger V, Lacroix R et al (2018) Biomarkers for the risk of thrombosis in pancreatic adenocarcinoma are related to cancer process. Oncotarget 9:26453–26465. https://doi.org/10.18632/oncotarget.25458
Zhang B, Pang Z, Hu Y (2020) Targeting hemostasis-related moieties for tumor treatment. Thromb Res 187:186–196. https://doi.org/10.1016/j.thromres.2020.01.019
Munn LL, Jain RK (2019) Vascular regulation of antitumor immunity. Science 365:544–545. https://doi.org/10.1126/science.aaw7875
Mpekris F, Voutouri C, Baish JW, Duda DG, Munn LL, Stylianopoulos T, Jain RK (2020) Combining microenvironment normalization strategies to improve cancer immunotherapy. Proc Natl Acad Sci USA 117:3728–3737. https://doi.org/10.1073/pnas.1919764117
Acknowledgements
We would like to thank Dr Aurélien de Reynies (Paris, France) for help with MCP counter. We are grateful to the patients, physicians and scientists involved in TCGA. We are also grateful to Ligue contre le Cancer and CHU Amiens Picardie for supporting research in our laboratory.
Funding
This project was supported by grants obtained from Ligue contre le Cancer, comité de la Somme attributed to Z.S. and A.G.
Author information
Authors and Affiliations
Contributions
Z.S. and A.G. conducted the analyses and wrote the manuscript. S.S., F.C., M.L. V.S. and M.A.S. critically analyzed the study and corrected the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethics approval and data accessibility
Not applicable. All data used in this study are freely available through the TCGA portal (https://portal.gdc.cancer.gov/) and previously published study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Saidak, Z., Soudet, S., Lottin, M. et al. A pan-cancer analysis of the human tumor coagulome and its link to the tumor immune microenvironment. Cancer Immunol Immunother 70, 923–933 (2021). https://doi.org/10.1007/s00262-020-02739-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00262-020-02739-w