Abstract
Ovarian cancer remains one of the most lethal gynecological cancers in the world despite extensive progress in the areas of chemotherapy and surgery. Many studies have postulated that this is because of the profound heterogeneity that underpins response to therapy and prognosis. Standard imaging evaluation using CT or MRI does not take into account this tumoral heterogeneity especially in advanced stages with peritoneal carcinomatosis. As such, newly emergent fields in the assessment of tumor heterogeneity have been proposed using radiomics to evaluate the whole tumor burden heterogeneity as opposed to single biopsy sampling. This review provides an overview of radiomics, radiogenomics, and proteomics and examines the use of these newly emergent fields in assessing tumor heterogeneity and its implications in ovarian cancer.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Ovarian cancer remains one of the most lethal gynecological cancers in the world with the epithelial subtype accounting for more than 90% of all cases and being responsible for the vast majority of ovarian cancer-related deaths [1]. Although progress has been made through refinements in chemotherapeutic and surgical approaches, patient survival and prognosis have only improved slightly in the recent years [2,3,4,5,6,7]. Many studies have postulated that this is because the disease cannot be defined as a single entity, but rather consists of a vast degree of cellular and genomic heterogeneity that dictate response to therapy and prognosis [8,9,10,11,12,13,14,15]. This heterogeneity can be divided into inter- and intra-tumoral heterogeneity [15, 16]. Inter-tumoral heterogeneity is defined as the genotypic and phenotypic variations found between multiple tumor implants in the peritoneal cavity while intra-tumoral heterogeneity is recognized as the coexistence of different cell populations within one single lesion [16]. Two models used to explain this inter- and intra-tumoral heterogeneity have been proposed based on deep spatial and longitudinal DNA sequencing studies of ovarian tumors and matched peritoneal implants [17,18,19]. In linear evolution, distinct populations of cells arise in the primary ovarian tumor, from which cells then break off to form secondary sites. In parallel evolution, both primary ovarian tumors and metastases continue to evolve, with new populations of metastases arising and even reseeding the primary site of metastases (Fig. 1) [18].
Drawing demonstrating the linear and parallel model explaining tumor heterogeneity: In the linear progression model, metastasis is seeded at a late stage of tumor progression, resulting in minimal genetic divergence between the primary tumor and its metastases. In the parallel progression model, metastases are seeded early in tumor progression, so high levels of genetic divergence are expected between the primary tumor and its metastases, even reseeding the primary tumor
Next-generation sequencing has documented the genome of primary ovarian cancer, identifying potential therapeutic targets [20,21,22,23]. Far less analysis has been performed on peritoneal implants due to the difficulty in accessing metastatic tissues, therefore a number of important clinical and biological queries remain unanswered. The large number of peritoneal implants, their accessibility, and their imaging presentation mean that analyzing high-grade serous ovarian carcinoma (HGSOC) by single biopsy sampling and traditional imaging tools is extremely challenging. Computed tomography (CT) and, in selected cases magnetic resonance imaging (MRI), have been used to evaluate the extent of disease and monitor treatment response in patients with HGSOC providing mainly an understaging of the disease burden [24,25,26,27,28,29,30,31]. Tools to capture the heterogeneity of the disease at an imaging level are emerging [32,33,34]. Imaging research is particularly focused on the field of radiomics [33, 35,36,37,38,39]. Radiomics is defined as the extraction of a large level of quantitative data from images which cannot be assessed visually [32, 34]. The quantification of heterogeneity on imaging in ovarian cancer may offer an opportunity to noninvasively evaluate the whole tumor volume, i.e., ovarian tumor and peritoneal implants rather than single biopsy evaluation. This would allow better prediction of outcome, optimal triage, and the ability to offer the best treatment option for every patient [36].
This review provides an overview of radiomics, radiogenomics, and proteomics, and examines the use of these newly emergent fields in assessing tumor heterogeneity and its implications in ovarian cancer.
Ovarian cancer: where are we now in terms of imaging?
Tremendous advances in risk stratification have been made by establishing standardized and evidence-based risk assessment algorithms using both ultrasound and MRI, allowing accurate characterization of the adnexal lesions [40, 41].
For ultrasound, the International Ovarian Tumor Analysis (IOTA) group has developed the simple rules classification system and Assessment of Different Neoplasia in the Adnexa (ADNEX) model to differentiate benign from malignant adnexal masses [42,43,44,45,46,47]. Other ovarian mass characterization and management systems have been proposed, including the Society of Radiologists in Ultrasound consensus statement [48, 49] and the Gynecologic Imaging Reporting and Data System, or GI-RADS [40, 50].
Most recently, the Ovarian-Adnexal Reporting and Data System (O-RADS) published in 2018 has provided a standardized lexicon that includes all pertinent descriptors and definitions of the characteristic ultrasound appearance of normal ovaries and ovarian lesions [40]. Based on this validated reporting system, ultrasound guidelines for lesion management have been proposed in 2020 [51]. These guidelines now include all risk categories with their corresponding management strategies, which had not been presented in any of the previous versions [51].
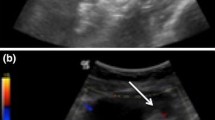
However, approximately 18% to 31% of adnexal lesions detected on ultrasound remain indeterminate, with MRI recognized as the complementary tool to assess indeterminate ovarian lesion on ultrasound [52]. The European Society of Urogenital Radiology (ESUR) recommendation has proposed an algorithmic approach to help differentiate benign from malignant lesions [53]. More recently an O-RADS MRI, similar to O-RADS ultrasound, has been proposed. This 5-category MRI scoring system was initially developed in a retrospective single-center study [41] using indeterminate adnexal masses on ultrasonography and was subsequently validated prospectively on 1340 women. Area under the receiver operating characteristic curve for differentiating benign from malignant lesions was 0.961 (95% CI 0.948–0.971) among experienced readers, with a sensitivity of 0.93 (95% CI 0.89–0.96; 189 of 203 patients) and a specificity of 0.91 (95% CI 0.89–0.93; 848 of 927 patients) [54].
For peritoneal implants, less advances have been made. CT remains the imaging gold standard for preoperative evaluation, with an accuracy ranging between 70 and 90% for implant detection [29, 55,56,57,58]. CT, however, has a limited sensitivity (25% to 50%) for implants less than 1 cm, particularly in locations such as the bowel surface or mesentery [55]. Implant detection is improved by the opacification of the gastrointestinal tract which helps to differentiate bowel from serosal and mesenteric implants. Reformatted images in coronal and sagittal planes are particularly useful for evaluating the subphrenic space [57].
The role of MRI in peritoneal disease staging has recently been evaluated. Promising results and guidelines have been proposed at the 10th Peritoneal Surface Oncology Group International (PSOGI) congress 2016. For now, MRI serves mainly as an adjunct to CT for implant detection. Standard MRI has the same sensitivity as CT in the detection of peritoneal deposits [29, 59]. However, the use of fat suppression, delayed post-contrast imaging, oral contrast agents, and functional imaging such as DWI [60] have allowed the detection sensitivities of MRI to surpass CT [61, 62] with sensitivity and specificity of DWI being 90% and 95.5%, respectively, for implant detection [60].
Response evaluation to chemotherapy by imaging also remains challenging. CT does not provide a quantitative assessment of disease response to cytotoxic therapy and does not reflect molecular events in the tumor (apoptosis, necrosis etc.) as it relies solely on macroscopic changes in tumor size as per Response evaluation criteria in solid tumors (RECIST) criteria [63]. The use of RECIST is often not possible as the carcinomatosis frequently consists of multiple subcentimeter nodules, ill-defined soft tissue masses, confluent disease or disease of a diffuse infiltrative pattern not eligible for RECIST evaluation. Functional imaging using diffusion-weighted imaging (DWI) has been evaluated in the research setting. Using ADC histograms, authors have shown efficacy in treatment monitoring of peritoneal carcinomatosis [64].
However, all evaluation of peritoneal disease by either CT or multiparametric MRI requires the subjective interpretation of a radiologist to inform the clinician on the volume and location of the many implants. This approach inevitably introduces a high degree of variability. Tools for more automated imaging analyses have been tested, not only to reduce this variability, but to provide more objective, clinically relevant information [65]. Following the steps of genomics, in the era of big data, a new field in medical imaging has emerged under the label of ‘’radiomics.’’ Radiomics has been introduced as an emergent tool for postprocessing CT or MR images and developing new quantification metrics [66,67,68,69,70,71,72] linking qualitative and/or qualitative imaging data to clinical endpoints. Compared to Computer Aided Detection (CAD) in which the number of image features involved is usually less than 20, radiomics deals with hundreds to thousands of imaging features extracted from large-scale radiological images. These quantitative metrics have been shown to provide important insights into tumor biology, linking quantitative imaging data to meaningful clinical endpoints. Of particular interest is the potential ability of image-derived metrics to reliably identify important tumor subregions noninvasively and derive predictive imaging biomarkers.
Radiomics: how does it work? (Fig. 2)
The process of radiomics analysis encompasses the following steps:1/image acquisition and segmentation, 2/feature extraction, 3/feature selection, and 4/ model construction [73, 74].
-
(1)
Image acquisition represents the first step. Radiomics analysis can be applied to any type of medical images, including X-ray, US, CT, MRI, and PET-CT. Before radiomic features are calculated, a region of interest or volume of interest encompassing the dedicated area has to be defined. Currently, there are a number of reliable algorithms capable of automatic or semi-automatic lesion segmentation, limiting the need for human input, helping overcome subjectivity and time cost associated with manual segmentation [75, 76]. However, in some instances, performance of these algorithms is jeopardized by the complexity of the task mainly due to the complex appearance of the lesion (ill-defined borders, locations adjacent and not being easily distinguishable from adjacent anatomical structures). Under these circumstances, expert manual segmentation and delineation of the ROI on one chosen slice (2D) or on the entire volume (3D) remain the only valid option [77,78,79,80].
-
(2)
Feature extraction, requiring the use of dedicated software or open-source programming packages, offers many options, from pre-processing tools including normalization, resampling, and discretization, to customizing the extraction and enabling the application of additional filters on the original images [81,82,83].
The process of extraction returns a large amount of quantitative imaging metrics, which can be categorized in 6 different groups: a/First-Order Statistics are features related to the histogram, including energy, entropy, kurtosis, skewness … b/Shape-based features, such as volume, surface area, sphericity… The following groups include textural features, and assess the spatial distribution of pixel intensities: c/Gray-Level Co-occurence Matrix, d/Gray-Level Run Length Matrix, e/Gray-Level Size Zone Matrix, and f/Gray-Level Dependence Matrix.
A large number of features (between 50 and 5000) are usually extracted. These are then decreased through feature qualification in order to select only the features that are informative and reproducible and to prevent the risk of overfitting [84]. Overfitting is defined as the construction of an analytical process that corresponds too closely to a specific dataset, with the model performing poorly in the validation phase in contrast to its learning phase, and consequently failing to generalize.
-
(3)
Feature selection The process of feature selection usually starts with removing redundant features [85]. Secondly, inter-observer and intra-observer feature variability are assessed. Highly variable features are not considered in model construction as they may yield too hazardous results [86]. Thirdly, only important features with high correlation to the studied endpoint are selected. The number of optimal selected features might vary depending on the dataset, but no consensus on the exact number has been achieved. Yet, Parmar et al. suggest that number should be equal to the square root of the number of observations in a correlated set of features. Selected features constitute what we—in the literature—refer to as the ‘’Radiomics signature’ [87]’.
-
(4)
Model construction and performance assessment The goal is to develop a mathematical or a statistical model to be associated with a diagnosis, tumor response, or patient outcome. In other words, the model is equivalent to an algorithm that analyzes training data and infers a hypothesis to predict a variable when given quantitative imaging features. Different algorithms have been proposed. The most popular include the following: random forest (RF) [88], least absolute shrinkage and selection operator (LASSO) [89], artificial neural networks (ANN) [90], support vector machine (SVM) [91], and minimum redundancy maximum relevance (mRMR) [92]. No consensus on the pre-eminent algorithm has been achieved, as one algorithm might outperform another depending on the dataset type and variables [93]. To quantify the discrimination performance, the AUC or the Harrell concordance index is used. Validation of the model could be assessed either internally or externally. In internal validation, the most common procedure is the ‘’leave-one out’’ cross-validation (LOOCV) where all the data are used for training except for one data point, which is left out for testing and validation [94]. Another common method is the bootstrap, which consists of generating a large series of data (bootstrap sample) [95]. Each bootstrap sample represents a patient chosen randomly, combined with its corresponding features and outcome, and this process is reiterated over the entire cohort of patients.
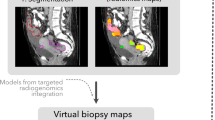
Radiomics and radiogenomics in ovarian cancer (Fig. 3) (Table 1)
Work on radiomics and radiogenomics has been mainly linked with genomic advances thanks to The Cancer Genome Atlas (TCGA) research network [96, 97]. Based on the TCGA data, a prognostic algorithm for HGSOC known as Classification of Ovarian Cancer (CLOVAR) has been defined with four subtypes: differentiated, immunoreactive, mesenchymal, and proliferative [98, 99]. Patients with mesenchymal subtype have been shown to have a higher rate of platinum resistance (63%) compared with patients with other subtypes (23%), as well as shorter median survival (23 vs 46 months for patients). Initially, Vargas et al. explored the relationships between subjective qualitative CT features and CLOVAR subtypes of HGSOC [100]. The authors found that CLOVAR mesenchymal subtype was significantly associated with higher risk of peritoneal involvement and the presence of mesenteric infiltration on CT. These results may explain the reported poorer prognosis of patients with the CLOVAR mesenchymal subtype of HGSOC [99]. However, in a subsequent study, the same research team showed the limit of subjective qualitative assessment of CT with poor inter-observer agreement in the assessment of this feature (mesenteric infiltration) (α = 0.23) when reviewed by eight readers. As such, the limitations of subjective evaluation support the use of for more automated or semiautomated analysis [101].
In another study including 38 patients, Vargas et al. developed 12 quantitative metrics to capture spatial inter-site imaging heterogeneity in high-grade serous ovarian cancer [102]. The authors demonstrated that metrics capturing the differences in texture similarities across sites were associated with shorter overall survival (inter-site similarity entropy, similarity-level cluster shade, and inter-site similarity-level cluster prominence; p ≤ 0.05) and incomplete surgical resection (similarity -level cluster shade, inter-site similarity-level cluster prominence, and inter-site cluster variance) [102].
Rizzo et al. evaluated whether CT radiomics features extracted from the primary tumor alone or combined with clinical data were associated with residual tumor at surgery in 101 patients with ovarian cancer and were able to predict the risk of disease progression within 12 months [33]. The authors found that features related to mass size, randomness and homogeneity were associated with residual tumor at surgery. At multivariate analysis, the risk of progression at 12 months was associated with one feature, F2 shape/Max3DDiameter, related to tumor size [33]. Adding this radiomic feature to a clinically based model significantly increased prediction of progression at 12 months by 14% (AUC = 0.73 for clinical model vs 0.87 for clinical radiomic model) [33].
More recently, Meier et al. assessed the associations between inter-site texture heterogeneity parameters derived from CT, survival, and BRCA mutation status in women with high-grade serous ovarian cancer [103]. The authors demonstrated that high inter-site cluster variance was associated with lower progression-free survival (PFS) (p = 0.006) and overall survival (OS) (p = 0.003). Higher inter-site cluster prominence was associated with lower PFS (p = 0.02) and higher inter-site cluster entropy correlated with lower OS (p = 0.01). High values of all three metrics were significantly associated with lower complete surgical resection status in BRCA-negative patients (SE p = 0.039, SCV p = 0.006, SCP p = 0.02), but not in BRCA-positive patients (SE p = 0.7, SCV p = 0.91, SCP p = 0.67). However, none of the metrics were able to distinguish between BRCA mutation carrier and nonmutation carrier [103].
With the aim of developing a novel radiomics-determined mathematical descriptor of epithelial ovarian cancer tumor risk phenotype with a predictive value, Lu et al. extracted 657 quantitative mathematical descriptors from the preoperative CT scans of 364 ovarian cancer patients at their initial presentation [104]. The cohorts were divided into discovery dataset (n = 136), validation dataset (n = 77), and external dataset evaluated with the TCGA (n = 70). Forty-two radiomic features were significantly associated with OS; these 42 radiomic features were further reduced to 4 weighted features using least absolute shrinkage and selection operator method. The weighted sum of these four radiomic features gave a “radiomic prognostic vector” score for each tumor that reliably identified the 5% of patients with median overall survival of less than 2 years [104].
Textural features are gaining interest not only from CT studies, but also from MR studies and ex vivo studies. For example, Song et al. evaluated the efficiency of 2- and 3-class classification predictive tasks constructed from radiomics features extracted from Dynamic contrast enhancement (DCE)-MRI in discriminating among benign, borderline, and malignant ovarian tumors in 104 ovarian lesions [105]. The 2-class classification task was divided into three subtasks: benign vs. borderline (task A), benign vs. malignant (task B), and borderline vs. malignant (task C). For the 3-class classification task, 104 lesions were randomly divided into training (72 lesions) and validation (32 lesions) cohorts. For the 2-class classification task, the radiomics model showed a good diagnostic ability with the highest AUCs of 0.899, 0.865, and 0.893 for tasks A, B, and C, respectively. The 3-class classification task demonstrated a good discrimination performance with AUCs of 0.893, 0.944, and 0.891 for the benign, borderline, and malignant groups, respectively [105]. More recently, Zhang et al. evaluated again the diagnostic performance of a radiomics model based on MR images to discriminate benign ovarian tumors from malignancies using features from multiple sequences (T1, T2WI, T2WI-FS, DWI, and multiple phase T1 contrast). MR radiomics including features from all the sequences acquired showed higher accuracy than that of the radiologists (90.6% and 83.5%, respectively) [106]. Interestingly, in this study, the most common diagnostic error among the radiologists was the classification of borderline ovarian tumors into the benign group. Therefore, when borderline ovarian tumors were excluded, the radiologist’s diagnostic performance in differentiating benign from malignant tumors was comparable to the computer’s performance [106]. In the same paper, the authors aimed to assess whether MR radiomics could differentiate between type I and type II epithelial ovarian cancer, showing a model accuracy for this endpoint of 83% and an AUC of 85%. In this analysis, the authors also showed that the diagnostic performance was higher when features extracted from all the sequences were included [106]. Finally, the authors demonstrate that after scoring the features by the least absolute shrinkage and selection operator method (LASSO), the model divided patients into good and poor prognosis groups with an AUC of 0.899 and that T1-weighted imaging (WI) radiomics features were more likely associated with the clinical outcome than the other sequences [106]. Similarly, Jian et al. developed an MR combined radiomic model extracted from the combination of the four following sequences (T2WI-FS, DWI, ADC, and CE-T1WI) which were able to differentiate type I and type II epithelial ovarian cancer. This model achieved an AUC of 0.834 and 0.847 in internal and external validation cohorts, respectively [107].
With the aim of comparing the ability of clinical features, conventional MR image features, ADC value, T2WI, DWI, DCE- MR radiomics, and a combined multiple features model to predict the type of epithelial ovarian cancer, Qian et al. retrospectively evaluated 61 patients with epithelial ovarian cancer. In this study, the model included T2WI, DWI, and DCE-MRI, and a multisequence model. 1070 radiomics features of each sequence were extracted. Then, univariate analysis and LASSO were used to select important features. The authors demonstrated that the traditional model achieved the best diagnostic performance, while the diagnostic efficacy of the mixed model was slightly increased, thus concluding that the traditional model represents a noninvasive and reliable tool that can better distinguish EOC type I from EOC type II preoperatively [108].
Wang et al. proposed a classification model to identify ex vivo benign, malignant, and normal ovarian tissue [109]. The authors extracted 75 three-dimensional texture features from second harmonic generation images of human ovarian tissue from 31 patients, using radiomics-based feature extraction methods. Combining the minimum redundancy maximum relevancy feature selection method and a tree-based pipeline optimization tool, they classified benign, malignant, and normal fresh ex vivo ovarian tissue, with performance values at AUC-ROC of 0.98, 0.96, and 0.94, respectively [109].
Proteomics
Proteomics is the analysis of the entire protein complement of a cell, tissue, or organism under a specific, defined set of conditions [110]. Proteomics can complement genomic analysis by uncovering pathways and processes of cancer biology to match with particular clinical phenotypes [111]. Pairwise analysis of proteomic and genomic data from the same tumor has shown that proteomics give added information beyond that elicited only through genomic analysis [112].
In ovarian cancer, an analysis of proteomics utility in predicting recurrence and survival time was performed by the PRotein-driven index of OVARian cancer (PROVAR) study team in 2013 [113]. PROVAR analyzed 412 ovarian cancer cases from TCGA [113]. They found that 5 proteins were associated with longer progression-free survival (PFS) and greater overall survival (OS), namely: androgen receptor (AR); phosphorylated tafazzin (pTAZ); BH3-interacting domain death agonist (BID); phosphorylated epidermal growth factor receptor (pEGFR); and 70 kilodalton heat shock protein (HSP70) [113]. Conversely, 4 proteins were highly expressed in patients with shorter PFS and shorter OS: signal transducer and activator of transcription 5 α (STAT5 α); phosphorylated protein kinase C-α (pPKCα); phosphorylated dual-specificity mitogen-activated protein kinase 1 (pMEK1); and eukaryotic translation elongation factor 2 (EEF2) [113].
Of the markers discovered in the PROVAR study, AR proved the most robust [113]. No change in its abundance was noted between the primary and metastatic tumors, which suggested that the downregulation of AR happens early in carcinogenesis [113].
The clinical proteomic tumor analysis consortium (CPTAC), in their 2016 study, used mass spectroscopy to categorize 174 ovarian tumors from TCGA [111]. They integrated previously sequenced genomic data from the TCGA study [114] with proteomic measurements to gain insight into the effect genomic copy number alterations (CNAs) had on the proteome. They found there was a convergence of CNA targets on a set of biological functions associated with motility/invasion and immune regulation, both classical of cancer cells [111]. They compared the presence of phosphorylated peptides in tumors of patients who survived < 3 years and those who survived > 5 years and found that PDGFR-beta (associated with angiogenesis), rhoA-regulatory, and integrin-like kinase pathways (both associated with cell mobility and invasion) were most abundant in short survivors [111]. The proliferation-associated transcription factor serum response factor (SRF) and the target proteins it regulates were also found to be more abundant in these short-surviving patients [111]. Their study found that combining analysis of protein quantity and phosphorylation status gave greater statistical significance when correlated with overall survival, emphasizing the utility of assessing protein phosphorylation in understanding pathway activity thus linking proteotype with phenotype [111].
Recent work has integrated proteomic analysis with CT-based qualitative and radiomic features [115]. In a retrospective hypothesis-generating study of 20 patients with HGSOC awaiting primary cytoreductive surgery, the abundance of certain proteins was found to be associated with CT-based qualitative traits, namely cysteine-rich protein 2 (CRIP2 – a tumor suppressor which regulates cell proliferation) was negatively correlated with mesenteric disease; and Melanoma Antigen Gene A4 (MAGEA4a—of the MAGE family, which are broadly expressed in many tumor types [116]) was positively correlated with supradiaphragmatic lymphadenopathy [115] (overexpression of MAGE is a predictor of poor PFS and OS in ovarian cancer [116,117,118]).
Texture feature extraction of intra-and inter-site heterogeneity (cluster-site entropy, cluster standard deviation, and cluster dissimilarity) was calculated and integrated with the proteomic data for those tumors. It showed an association between low levels of argininosuccinate synthase 1 (ASS1) and more heterogenous tumors [115]. This finding is in agreement with our knowledge of ASS1, low levels of which correlate with platinum therapy resistance and poor prognosis [119], and more heterogenous tumors, which are also predictive of worse survival [120].
Proteomics has the potential to facilitate the paradigm shift from treatment based on trial and error guided by anatomy and genetic profiling of tumors, to tailored, individualized treatment-based specifically on a tumors molecular profile [121]. To this end, large-scale prospective trials such as those planned by the APOLLO group (Applied Proteogenomics OrganizationaL Learning and Outcomes) will give greater insight into pathways of ovarian cancer and potential therapeutic liabilities and allow discovery and full validation of markers of disease heterogeneity, treatment resistance, and predictors of outcome [121]. Further large prospective studies integrating proteomic, genomic, and imaging tools are also necessary to potentiate the development of radiomic and radiogenomic tools towards true precision medicine [122].
Critics and challenges
Radiomics is in the early stages of development, with improvements to each step of the entire analytical process still required [123].
Firstly, one of its main limitations is the lack of standard protocol in terms of feature selection or model construction. A large number of feature selection algorithms and classifiers are available for use. An algorithm appropriate for a particular dataset might be inappropriate for another [124]. There is also great variety in the methods employed in most radiomic studies in ovarian cancer, with the frequent use of in house software raising issues for generalizability and reproducibility.
Secondly, feature reproducibility might be compromised when images are obtained from different vendors. CT features vary from one scanner to another depending on the parameters being used and from one reconstruction algorithm to another. In PET imaging, textural features are most sensitive to reconstruction algorithms and post-filtering level. In MRI, textural features depend on the field of view, magnetic field strength, slice thickness, and the contrast agent. Although image pre-processing is critical to achieving a level of homogeneity within data, thus overcoming the heterogeneity that can taint the quality of the samples being studied, there is no consensus regarding the protocol to be followed for image pre-processing [125,126,127].
Thirdly, most radiomics studies involve small cohorts, in disproportion to the dimensionality of the features, thus greatly increasing the risk of overfitting. Moreover, most studies do not test their models on an independent validation dataset, which has been reported to inflate the risk alpha. This is the case in the studies published on ovarian cancer, with the number of patients included ranging from 20 to 364. Most of these studies did not have a validation cohort [33, 102, 103, 105, 107, 109]. Institutions should work together in order to improve data sharing as this might be a useful way to conduct robust studies and increased the speed of radiomics adoption into routine clinical practice [84].
The emergence of artificial intelligence (AI) and its progressively wider impact on radiology may overcome some limitations of radiomics such as the need for manual “human” segmentation. AI algorithms are not only able to analyze the predefined segmentation but also directly analyze the images in order to automatically design its own radiomic features without underlying human evaluation. Moreover, AI offers a better capability of handling a massive amount of data compared with the traditional statistical methods.
Conclusion
Although many issues need to be addressed, radiomics is a potential game changer shifting radiology from the traditional visual analysis to a more objective and automated analysis. By integrating imaging biomarkers automatically derived from imaging data to clinical, genomics, and/or proteomics data, radiomics offers huge opportunities to better capture tumor behavior. Radiomics raises particular hope in ovarian cancer to better capture the whole disease heterogeneity and offer a new tool to predict tumor aggressiveness and response to therapy.
References
Torre LA, Trabert B, DeSantis CE, Miller KD, Samimi G, Runowicz CD, Gaudet MM, Jemal A, Siegel RL (2018) Ovarian cancer statistics, 2018. CA Cancer J Clin 68 (4):284-296. https://doi.org/10.3322/caac.21456
Arend R, Martinez A, Szul T, Birrer MJ (2019) Biomarkers in ovarian cancer: To be or not to be. Cancer 125 Suppl 24:4563-4572. https://doi.org/10.1002/cncr.32595
Fujiwara K, Hasegawa K, Nagao S (2019) Landscape of systemic therapy for ovarian cancer in 2019: Primary therapy. Cancer 125 Suppl 24:4582-4586. https://doi.org/10.1002/cncr.32475
Lee JM, Minasian L, Kohn EC (2019) New strategies in ovarian cancer treatment. Cancer 125 Suppl 24:4623-4629. https://doi.org/10.1002/cncr.32544
Bogani G, Lopez S, Mantiero M, Ducceschi M, Bosio S, Ruisi S, Sarpietro G, Guerrisi R, Brusadelli C, Dell'Acqua A, Di Donato V, Raspagliesi F (2020) Immunotherapy for platinum-resistant ovarian cancer. Gynecol Oncol. https://doi.org/10.1016/j.ygyno.2020.05.681
Bowtell DD (2010) The genesis and evolution of high-grade serous ovarian cancer. Nat Rev Cancer 10 (11):803-808. https://doi.org/10.1038/nrc2946
Patch AM, Christie EL, Etemadmoghadam D, Garsed DW, George J, Fereday S, Nones K, Cowin P, Alsop K, Bailey PJ, Kassahn KS, Newell F, Quinn MC, Kazakoff S, Quek K, Wilhelm-Benartzi C, Curry E, Leong HS, Australian Ovarian Cancer Study G, Hamilton A, Mileshkin L, Au-Yeung G, Kennedy C, Hung J, Chiew YE, Harnett P, Friedlander M, Quinn M, Pyman J, Cordner S, O'Brien P, Leditschke J, Young G, Strachan K, Waring P, Azar W, Mitchell C, Traficante N, Hendley J, Thorne H, Shackleton M, Miller DK, Arnau GM, Tothill RW, Holloway TP, Semple T, Harliwong I, Nourse C, Nourbakhsh E, Manning S, Idrisoglu S, Bruxner TJ, Christ AN, Poudel B, Holmes O, Anderson M, Leonard C, Lonie A, Hall N, Wood S, Taylor DF, Xu Q, Fink JL, Waddell N, Drapkin R, Stronach E, Gabra H, Brown R, Jewell A, Nagaraj SH, Markham E, Wilson PJ, Ellul J, McNally O, Doyle MA, Vedururu R, Stewart C, Lengyel E, Pearson JV, Waddell N, deFazio A, Grimmond SM, Bowtell DD (2015) Whole-genome characterization of chemoresistant ovarian cancer. Nature 521 (7553):489-494. https://doi.org/10.1038/nature14410
Takaya H, Nakai H, Sakai K, Nishio K, Murakami K, Mandai M, Matsumura N (2020) Intratumor heterogeneity and homologous recombination deficiency of high-grade serous ovarian cancer are associated with prognosis and molecular subtype and change in treatment course. Gynecol Oncol 156 (2):415-422. https://doi.org/10.1016/j.ygyno.2019.11.013
Nelson L, Tighe A, Golder A, Littler S, Bakker B, Moralli D, Murtuza Baker S, Donaldson IJ, Spierings DCJ, Wardenaar R, Neale B, Burghel GJ, Winter-Roach B, Edmondson R, Clamp AR, Jayson GC, Desai S, Green CM, Hayes A, Foijer F, Morgan RD, Taylor SS (2020) A living biobank of ovarian cancer ex vivo models reveals profound mitotic heterogeneity. Nat Commun 11 (1):822. https://doi.org/10.1038/s41467-020-14551-2
Jimenez-Sanchez A, Cybulska P, Mager KL, Koplev S, Cast O, Couturier DL, Memon D, Selenica P, Nikolovski I, Mazaheri Y, Bykov Y, Geyer FC, Macintyre G, Gavarro LM, Drews RM, Gill MB, Papanastasiou AD, Sosa RE, Soslow RA, Walther T, Shen R, Chi DS, Park KJ, Hollmann T, Reis-Filho JS, Markowetz F, Beltrao P, Vargas HA, Zamarin D, Brenton JD, Snyder A, Weigelt B, Sala E, Miller ML (2020) Unraveling tumor-immune heterogeneity in advanced ovarian cancer uncovers immunogenic effect of chemotherapy. Nat Genet 52 (6):582-593. https://doi.org/10.1038/s41588-020-0630-5
Gao Y, Chen L, Cai G, Xiong X, Wu Y, Ma D, Li SC, Gao Q (2020) Heterogeneity of immune microenvironment in ovarian cancer and its clinical significance: a retrospective study. Oncoimmunology 9 (1):1760067. https://doi.org/10.1080/2162402X.2020.1760067
Tan TZ, Heong V, Ye J, Lim D, Low J, Choolani M, Scott C, Tan DSP, Huang RY (2019) Decoding transcriptomic intra-tumour heterogeneity to guide personalised medicine in ovarian cancer. J Pathol 247 (3):305-319. https://doi.org/10.1002/path.5191
Roberts CM, Cardenas C, Tedja R (2019) The Role of Intra-Tumoral Heterogeneity and Its Clinical Relevance in Epithelial Ovarian Cancer Recurrence and Metastasis. Cancers (Basel) 11 (8). https://doi.org/10.3390/cancers11081083
Garziera M, Roncato R, Montico M, De Mattia E, Gagno S, Poletto E, Scalone S, Canzonieri V, Giorda G, Sorio R, Cecchin E, Toffoli G (2019) New Challenges in Tumor Mutation Heterogeneity in Advanced Ovarian Cancer by a Targeted Next-Generation Sequencing (NGS) Approach. Cells 8 (6). https://doi.org/10.3390/cells8060584
Lee JY, Yoon JK, Kim B, Kim S, Kim MA, Lim H, Bang D, Song YS (2015) Tumor evolution and intratumor heterogeneity of an epithelial ovarian cancer investigated using next-generation sequencing. BMC Cancer 15:85. https://doi.org/10.1186/s12885-015-1077-4
Tong JG, Valdes YR, Sivapragasam M, Barrett JW, Bell JC, Stojdl D, DiMattia GE, Shepherd TG (2017) Spatial and temporal epithelial ovarian cancer cell heterogeneity impacts Maraba virus oncolytic potential. BMC Cancer 17 (1):594. https://doi.org/10.1186/s12885-017-3600-2
Subramanian DN, Zethoven M, McInerny S, Morgan JA, Rowley SM, Lee JEA, Li N, Gorringe KL, James PA, Campbell IG (2020) Exome sequencing of familial high-grade serous ovarian carcinoma reveals heterogeneity for rare candidate susceptibility genes. Nat Commun 11 (1):1640. https://doi.org/10.1038/s41467-020-15461-z
Masoodi T, Siraj S, Siraj AK, Azam S, Qadri Z, Parvathareddy SK, Tulbah A, Al-Dayel F, AlHusaini H, AlOmar O, Al-Badawi IA, Alkuraya FS, Al-Kuraya KS (2020) Genetic heterogeneity and evolutionary history of high-grade ovarian carcinoma and matched distant metastases. Br J Cancer 122 (8):1219-1230. https://doi.org/10.1038/s41416-020-0763-4
Cooke SL, Ng CK, Melnyk N, Garcia MJ, Hardcastle T, Temple J, Langdon S, Huntsman D, Brenton JD (2010) Genomic analysis of genetic heterogeneity and evolution in high-grade serous ovarian carcinoma. Oncogene 29 (35):4905-4913. https://doi.org/10.1038/onc.2010.245
Shah RH, Scott SN, Brannon AR, Levine DA, Lin O, Berger MF (2015) Comprehensive mutation profiling by next-generation sequencing of effusion fluids from patients with high-grade serous ovarian carcinoma. Cancer Cytopathol 123 (5):289-297. https://doi.org/10.1002/cncy.21522
Fumagalli C, Rappa A, Casadio C, Betella I, Colombo N, Barberis M, Guerini-Rocco E (2020) Next-generation sequencing-based BRCA testing on cytological specimens from ovarian cancer ascites reveals high concordance with tumour tissue analysis. J Clin Pathol 73 (3):168-171. https://doi.org/10.1136/jclinpath-2019-206127
Mandilaras V, Garg S, Cabanero M, Tan Q, Pastrello C, Burnier J, Karakasis K, Wang L, Dhani NC, Butler MO, Bedard PL, Siu LL, Clarke B, Shaw PA, Stockley T, Jurisica I, Oza AM, Lheureux S (2019) TP53 mutations in high grade serous ovarian cancer and impact on clinical outcomes: a comparison of next generation sequencing and bioinformatics analyses. Int J Gynecol Cancer. https://doi.org/10.1136/ijgc-2018-000087
Ross JS, Ali SM, Wang K, Palmer G, Yelensky R, Lipson D, Miller VA, Zajchowski D, Shawver LK, Stephens PJ (2013) Comprehensive genomic profiling of epithelial ovarian cancer by next generation sequencing-based diagnostic assay reveals new routes to targeted therapies. Gynecol Oncol 130 (3):554-559. https://doi.org/10.1016/j.ygyno.2013.06.019
Rizzo S, De Piano F, Buscarino V, Pagan E, Bagnardi V, Zanagnolo V, Colombo N, Maggioni A, Del Grande M, Del Grande F, Bellomi M, Aletti G (2020) Pre-operative evaluation of epithelial ovarian cancer patients: Role of whole body diffusion weighted imaging MR and CT scans in the selection of patients suitable for primary debulking surgery. A single-centre study. Eur J Radiol 123:108786. https://doi.org/10.1016/j.ejrad.2019.108786
Li M, Tan J, Zhang Y, Ai C, Wang H, Zhang H, Jin Y, Chen Y (2020) Assessing CT imaging features combined with CEA and CA125 levels to identify endometriosis-associated ovarian cancer. Abdom Radiol (NY). https://doi.org/10.1007/s00261-020-02571-x
Silverman PM, Osborne M, Dunnick NR, Bandy LC (1988) CT prior to second-look operation in ovarian cancer. AJR Am J Roentgenol 150 (4):829-832. https://doi.org/10.2214/ajr.150.4.829
Togashi K (2003) Ovarian cancer: the clinical role of US, CT, and MRI. Eur Radiol 13 Suppl 4:L87-104. https://doi.org/10.1007/s00330-003-1964-y
Burger IA, Goldman DA, Vargas HA, Kattan MW, Yu C, Kou L, Andikyan V, Chi DS, Hricak H, Sala E (2015) Incorporation of postoperative CT data into clinical models to predict 5-year overall and recurrence free survival after primary cytoreductive surgery for advanced ovarian cancer. Gynecol Oncol 138 (3):554-559. https://doi.org/10.1016/j.ygyno.2015.06.010
Forstner R, Hricak H, Occhipinti KA, Powell CB, Frankel SD, Stern JL (1995) Ovarian cancer: staging with CT and MR imaging. Radiology 197 (3):619-626. https://doi.org/10.1148/radiology.197.3.7480729
Fultz PJ, Jacobs CV, Hall WJ, Gottlieb R, Rubens D, Totterman SM, Meyers S, Angel C, Del Priore G, Warshal DP, Zou KH, Shapiro DE (1999) Ovarian cancer: comparison of observer performance for four methods of interpreting CT scans. Radiology 212 (2):401-410. https://doi.org/10.1148/radiology.212.2.r99au19401
Kurtz AB, Tsimikas JV, Tempany CM, Hamper UM, Arger PH, Bree RL, Wechsler RJ, Francis IR, Kuhlman JE, Siegelman ES, Mitchell DG, Silverman SG, Brown DL, Sheth S, Coleman BG, Ellis JH, Kurman RJ, Caudry DJ, McNeil BJ (1999) Diagnosis and staging of ovarian cancer: comparative values of Doppler and conventional US, CT, and MR imaging correlated with surgery and histopathologic analysis--report of the Radiology Diagnostic Oncology Group. Radiology 212 (1):19-27. https://doi.org/10.1148/radiology.212.1.r99jl3619
Lambin P, Rios-Velazquez E, Leijenaar R, Carvalho S, van Stiphout RG, Granton P, Zegers CM, Gillies R, Boellard R, Dekker A, Aerts HJ (2012) Radiomics: extracting more information from medical images using advanced feature analysis. Eur J Cancer 48 (4):441-446. https://doi.org/10.1016/j.ejca.2011.11.036
Rizzo S, Botta F, Raimondi S, Origgi D, Buscarino V, Colarieti A, Tomao F, Aletti G, Zanagnolo V, Del Grande M, Colombo N, Bellomi M (2018) Radiomics of high-grade serous ovarian cancer: association between quantitative CT features, residual tumour and disease progression within 12 months. Eur Radiol 28 (11):4849-4859. https://doi.org/10.1007/s00330-018-5389-z
Kumar V, Gu Y, Basu S, Berglund A, Eschrich SA, Schabath MB, Forster K, Aerts HJ, Dekker A, Fenstermacher D, Goldgof DB, Hall LO, Lambin P, Balagurunathan Y, Gatenby RA, Gillies RJ (2012) Radiomics: the process and the challenges. Magn Reson Imaging 30 (9):1234-1248. https://doi.org/10.1016/j.mri.2012.06.010
Wei W, Rong Y, Liu Z, Zhou B, Tang Z, Wang S, Dong D, Zang Y, Guo Y, Tian J (2018) Radiomics: a Novel CT-Based Method of Predicting Postoperative Recurrence in Ovarian Cancer. Conf Proc IEEE Eng Med Biol Soc 2018:4130-4133. https://doi.org/10.1109/EMBC.2018.8513351
Nougaret S, Tardieu M, Vargas HA, Reinhold C, Vande Perre S, Bonanno N, Sala E, Thomassin-Naggara I (2019) Ovarian cancer: An update on imaging in the era of radiomics. Diagn Interv Imaging 100 (10):647-655. https://doi.org/10.1016/j.diii.2018.11.007
Lubner MG (2019) Reflections on radiogenomics and oncologic radiomics. Abdom Radiol (NY) 44 (6):1959. https://doi.org/10.1007/s00261-019-02047-7
Taghavi M, Trebeschi S, Simoes R, Meek DB, Beckers RCJ, Lambregts DMJ, Verhoef C, Houwers JB, van der Heide UA, Beets-Tan RGH, Maas M (2020) Machine learning-based analysis of CT radiomics model for prediction of colorectal metachronous liver metastases. Abdom Radiol (NY). https://doi.org/10.1007/s00261-020-02624-1
van Griethuysen JJM, Lambregts DMJ, Trebeschi S, Lahaye MJ, Bakers FCH, Vliegen RFA, Beets GL, Aerts H, Beets-Tan RGH (2020) Radiomics performs comparable to morphologic assessment by expert radiologists for prediction of response to neoadjuvant chemoradiotherapy on baseline staging MRI in rectal cancer. Abdom Radiol (NY) 45 (3):632-643. https://doi.org/10.1007/s00261-019-02321-8
Andreotti RF, Timmerman D, Benacerraf BR, Bennett GL, Bourne T, Brown DL, Coleman BG, Frates MC, Froyman W, Goldstein SR, Hamper UM, Horrow MM, Hernanz-Schulman M, Reinhold C, Strachowski LM, Glanc P (2018) Ovarian-Adnexal Reporting Lexicon for Ultrasound: A White Paper of the ACR Ovarian-Adnexal Reporting and Data System Committee. J Am Coll Radiol 15 (10):1415-1429. https://doi.org/10.1016/j.jacr.2018.07.004
Thomassin-Naggara I, Aubert E, Rockall A, Jalaguier-Coudray A, Rouzier R, Darai E, Bazot M (2013) Adnexal masses: development and preliminary validation of an MR imaging scoring system. Radiology 267 (2):432-443. https://doi.org/10.1148/radiol.13121161
Patel-Lippmann KK, Sadowski EA, Robbins JB, Paroder V, Barroilhet L, Maddox E, McMahon T, Sampene E, Wasnik AP, Blaty AD, Maturen KE (2020) Comparison of International Ovarian Tumor Analysis Simple Rules to Society of Radiologists in Ultrasound Guidelines for Detection of Malignancy in Adnexal Cysts. AJR Am J Roentgenol 214 (3):694-700. https://doi.org/10.2214/AJR.18.20630
Abramowicz JS, Timmerman D (2017) Ovarian mass-differentiating benign from malignant: the value of the International Ovarian Tumor Analysis ultrasound rules. Am J Obstet Gynecol 217 (6):652-660. https://doi.org/10.1016/j.ajog.2017.07.019
Timmerman D, Van Calster B, Testa A, Savelli L, Fischerova D, Froyman W, Wynants L, Van Holsbeke C, Epstein E, Franchi D, Kaijser J, Czekierdowski A, Guerriero S, Fruscio R, Leone FPG, Rossi A, Landolfo C, Vergote I, Bourne T, Valentin L (2016) Predicting the risk of malignancy in adnexal masses based on the Simple Rules from the International Ovarian Tumor Analysis group. Am J Obstet Gynecol 214 (4):424-437. https://doi.org/10.1016/j.ajog.2016.01.007
Dakhly DMR, Gaafar HM, Sediek MM, Ibrahim MF, Momtaz M (2019) Diagnostic value of the International Ovarian Tumor Analysis (IOTA) simple rules versus pattern recognition to differentiate between malignant and benign ovarian masses. Int J Gynaecol Obstet 147 (3):344-349. https://doi.org/10.1002/ijgo.12970
Timmerman D, Testa AC, Bourne T, Ferrazzi E, Ameye L, Konstantinovic ML, Van Calster B, Collins WP, Vergote I, Van Huffel S, Valentin L, International Ovarian Tumor Analysis G (2005) Logistic regression model to distinguish between the benign and malignant adnexal mass before surgery: a multicenter study by the International Ovarian Tumor Analysis Group. J Clin Oncol 23 (34):8794-8801. https://doi.org/10.1200/JCO.2005.01.7632
Sladkevicius P, Valentin L (2013) Intra- and interobserver agreement when describing adnexal masses using the International Ovarian Tumor Analysis terms and definitions: a study on three-dimensional ultrasound volumes. Ultrasound Obstet Gynecol 41 (3):318-327. https://doi.org/10.1002/uog.12289
Levine D, Brown DL, Andreotti RF, Benacerraf B, Benson CB, Brewster WR, Coleman B, Depriest P, Doubilet PM, Goldstein SR, Hamper UM, Hecht JL, Horrow M, Hur HC, Marnach M, Patel MD, Platt LD, Puscheck E, Smith-Bindman R (2010) Management of asymptomatic ovarian and other adnexal cysts imaged at US: Society of Radiologists in Ultrasound Consensus Conference Statement. Radiology 256 (3):943-954. https://doi.org/10.1148/radiol.10100213
Levine D, Brown DL, Andreotti RF, Benacerraf B, Benson CB, Brewster WR, Coleman B, DePriest P, Doubilet PM, Goldstein SR, Hamper UM, Hecht JL, Horrow M, Hur HC, Marnach M, Patel MD, Platt LD, Puscheck E, Smith-Bindman R, Society of Radiologists in U (2010) Management of asymptomatic ovarian and other adnexal cysts imaged at US Society of Radiologists in Ultrasound consensus conference statement. Ultrasound Q 26 (3):121-131. https://doi.org/10.1097/RUQ.0b013e3181f09099
Amor F, Vaccaro H, Alcazar JL, Leon M, Craig JM, Martinez J (2009) Gynecologic imaging reporting and data system: a new proposal for classifying adnexal masses on the basis of sonographic findings. J Ultrasound Med 28 (3):285-291. https://doi.org/10.7863/jum.2009.28.3.285
Andreotti RF, Timmerman D, Strachowski LM, Froyman W, Benacerraf BR, Bennett GL, Bourne T, Brown DL, Coleman BG, Frates MC, Goldstein SR, Hamper UM, Horrow MM, Hernanz-Schulman M, Reinhold C, Rose SL, Whitcomb BP, Wolfman WL, Glanc P (2020) O-RADS US Risk Stratification and Management System: A Consensus Guideline from the ACR Ovarian-Adnexal Reporting and Data System Committee. Radiology 294 (1):168-185. https://doi.org/10.1148/radiol.2019191150
Mohaghegh P, Rockall AG (2012) Imaging strategy for early ovarian cancer: characterization of adnexal masses with conventional and advanced imaging techniques. Radiographics 32 (6):1751-1773. https://doi.org/10.1148/rg.326125520
Forstner R, Thomassin-Naggara I, Cunha TM, Kinkel K, Masselli G, Kubik-Huch R, Spencer JA, Rockall A (2017) ESUR recommendations for MR imaging of the sonographically indeterminate adnexal mass: an update. Eur Radiol 27 (6):2248-2257. https://doi.org/10.1007/s00330-016-4600-3
Thomassin-Naggara I, Poncelet E, Jalaguier-Coudray A, Guerra A, Fournier LS, Stojanovic S, Millet I, Bharwani N, Juhan V, Cunha TM, Masselli G, Balleyguier C, Malhaire C, Perrot NF, Sadowski EA, Bazot M, Taourel P, Porcher R, Darai E, Reinhold C, Rockall AG (2020) Ovarian-Adnexal Reporting Data System Magnetic Resonance Imaging (O-RADS MRI) Score for Risk Stratification of Sonographically Indeterminate Adnexal Masses. JAMA Netw Open 3 (1):e1919896. https://doi.org/10.1001/jamanetworkopen.2019.19896
Coakley FV, Choi PH, Gougoutas CA, Pothuri B, Venkatraman E, Chi D, Bergman A, Hricak H (2002) Peritoneal metastases: detection with spiral CT in patients with ovarian cancer. Radiology 223 (2):495-499
Tempany CM, Zou KH, Silverman SG, Brown DL, Kurtz AB, McNeil BJ (2000) Staging of advanced ovarian cancer: comparison of imaging modalities--report from the Radiological Diagnostic Oncology Group. Radiology 215 (3):761-767
Forstner R, Sala E, Kinkel K, Spencer JA ESUR guidelines: ovarian cancer staging and follow-up. Eur Radiol 20 (12):2773–2780
Javitt MC (2007) ACR Appropriateness Criteria on staging and follow-up of ovarian cancer. J Am Coll Radiol 4 (9):586-589
Qayyum A, Coakley FV, Westphalen AC, Hricak H, Okuno WT, Powell B (2005) Role of CT and MR imaging in predicting optimal cytoreduction of newly diagnosed primary epithelial ovarian cancer. Gynecol Oncol 96 (2):301-306
Fujii S, Matsusue E, Kanasaki Y, Kanamori Y, Nakanishi J, Sugihara S, Kigawa J, Terakawa N, Ogawa T (2008) Detection of peritoneal dissemination in gynecological malignancy: evaluation by diffusion-weighted MR imaging. Eur Radiol 18 (1):18-23
Low RN, Barone RM, Lacey C, Sigeti JS, Alzate GD, Sebrechts CP (1997) Peritoneal tumor: MR imaging with dilute oral barium and intravenous gadolinium-containing contrast agents compared with unenhanced MR imaging and CT. Radiology 204 (2):513-520
Low RN, Semelka RC, Worawattanakul S, Alzate GD, Sigeti JS (1999) Extrahepatic abdominal imaging in patients with malignancy: comparison of MR imaging and helical CT, with subsequent surgical correlation. Radiology 210 (3):625-632
Eisenhauer EA, Therasse P, Bogaerts J, Schwartz LH, Sargent D, Ford R, Dancey J, Arbuck S, Gwyther S, Mooney M, Rubinstein L, Shankar L, Dodd L, Kaplan R, Lacombe D, Verweij J (2009) New response evaluation criteria in solid tumours: revised RECIST guideline (version 1.1). Eur J Cancer 45 (2):228–247
Kyriazi S, Collins DJ, Messiou C, Pennert K, Davidson RL, Giles SL, Kaye SB, Desouza NM Metastatic ovarian and primary peritoneal cancer: assessing chemotherapy response with diffusion-weighted MR imaging--value of histogram analysis of apparent diffusion coefficients. Radiology 261 (1):182–192
Gillies RJ, Kinahan PE, Hricak H (2016) Radiomics: Images Are More than Pictures, They Are Data. Radiology 278 (2):563-577. https://doi.org/10.1148/radiol.2015151169
Hu T, Wang S, Huang L, Wang J, Shi D, Li Y, Tong T, Peng W (2018) A clinical-radiomics nomogram for the preoperative prediction of lung metastasis in colorectal cancer patients with indeterminate pulmonary nodules. Eur Radiol. https://doi.org/10.1007/s00330-018-5539-3
Ortiz-Ramon R, Larroza A, Ruiz-Espana S, Arana E, Moratal D (2018) Classifying brain metastases by their primary site of origin using a radiomics approach based on texture analysis: a feasibility study. Eur Radiol. https://doi.org/10.1007/s00330-018-5463-6
She Y, Zhang L, Zhu H, Dai C, Xie D, Xie H, Zhang W, Zhao L, Zou L, Fei K, Sun X, Chen C (2018) The predictive value of CT-based radiomics in differentiating indolent from invasive lung adenocarcinoma in patients with pulmonary nodules. Eur Radiol. https://doi.org/10.1007/s00330-018-5509-9
Tan X, Ma Z, Yan L, Ye W, Liu Z, Liang C (2018) Radiomics nomogram outperforms size criteria in discriminating lymph node metastasis in resectable esophageal squamous cell carcinoma. Eur Radiol. https://doi.org/10.1007/s00330-018-5581-1
Wang J, Wu CJ, Bao ML, Zhang J, Wang XN, Zhang YD (2017) Machine learning-based analysis of MR radiomics can help to improve the diagnostic performance of PI-RADS v2 in clinically relevant prostate cancer. Eur Radiol 27 (10):4082-4090. https://doi.org/10.1007/s00330-017-4800-5
Matzner-Lober E, Suehs CM, Dohan A, Molinari N (2018) Thoughts on entering correlated imaging variables into a multivariable model: Application to radiomics and texture analysis. Diagn Interv Imaging 99 (5):269-270. https://doi.org/10.1016/j.diii.2018.04.011
Soyer P (2018) Agreement and observer variability. Diagn Interv Imaging 99 (2):53-54. https://doi.org/10.1016/j.diii.2018.01.009
Lubner MG, Smith AD, Sandrasegaran K, Sahani DV, Pickhardt PJ (2017) CT Texture Analysis: Definitions, Applications, Biologic Correlates, and Challenges. Radiographics 37 (5):1483-1503. https://doi.org/10.1148/rg.2017170056
Nougaret S, Tibermacine H, Tardieu M, Sala E (2019) Radiomics: an Introductory Guide to What It May Foretell. Curr Oncol Rep 21 (8):70. https://doi.org/10.1007/s11912-019-0815-1
Veeraraghavan H, Dashevsky BZ, Onishi N, Sadinski M, Morris E, Deasy JO, Sutton EJ (2018) Appearance Constrained Semi-Automatic Segmentation from DCE-MRI is Reproducible and Feasible for Breast Cancer Radiomics: A Feasibility Study. Sci Rep 8 (1):4838. https://doi.org/10.1038/s41598-018-22980-9
[76] Choi Y, Nam Y, Lee YS, Kim J, Ahn KJ, Jang J, Shin NY, Kim BS, Jeon SS (2020) IDH1 mutation prediction using MR-based radiomics in glioblastoma: comparison between manual and fully automated deep learning-based approach of tumor segmentation. Eur J Radiol 128:109031. https://doi.org/10.1016/j.ejrad.2020.109031
van Heeswijk MM, Lambregts DM, van Griethuysen JJ, Oei S, Rao SX, de Graaff CA, Vliegen RF, Beets GL, Papanikolaou N, Beets-Tan RG (2016) Automated and Semiautomated Segmentation of Rectal Tumor Volumes on Diffusion-Weighted MRI: Can It Replace Manual Volumetry? Int J Radiat Oncol Biol Phys 94 (4):824-831. https://doi.org/10.1016/j.ijrobp.2015.12.017
Fung YL, Ng KET, Vogrin SJ, Meade C, Ngo M, Collins SJ, Bowden SC (2019) Comparative Utility of Manual versus Automated Segmentation of Hippocampus and Entorhinal Cortex Volumes in a Memory Clinic Sample. J Alzheimers Dis 68 (1):159-171. https://doi.org/10.3233/JAD-181172
de Sitter A, Verhoeven T, Burggraaff J, Liu Y, Simoes J, Ruggieri S, Palotai M, Brouwer I, Versteeg A, Wottschel V, Ropele S, Rocca MA, Gasperini C, Gallo A, Yiannakas MC, Rovira A, Enzinger C, Filippi M, De Stefano N, Kappos L, Frederiksen JL, Uitdehaag BMJ, Barkhof F, Guttmann CRG, Vrenken H, Group MS (2020) Reduced accuracy of MRI deep grey matter segmentation in multiple sclerosis: an evaluation of four automated methods against manual reference segmentations in a multi-center cohort. J Neurol. https://doi.org/10.1007/s00415-020-10023-1
Smits LP, van Wijk DF, Duivenvoorden R, Xu D, Yuan C, Stroes ES, Nederveen AJ (2016) Manual versus Automated Carotid Artery Plaque Component Segmentation in High and Lower Quality 3.0 Tesla MRI Scans. PLoS One 11 (12):e0164267. https://doi.org/10.1371/journal.pone.0164267
Elhalawani H, Lin TA, Volpe S, Mohamed ASR, White AL, Zafereo J, Wong AJ, Berends JE, AboHashem S, Williams B, Aymard JM, Kanwar A, Perni S, Rock CD, Cooksey L, Campbell S, Yang P, Nguyen K, Ger RB, Cardenas CE, Fave XJ, Sansone C, Piantadosi G, Marrone S, Liu R, Huang C, Yu K, Li T, Yu Y, Zhang Y, Zhu H, Morris JS, Baladandayuthapani V, Shumway JW, Ghosh A, Pohlmann A, Phoulady HA, Goyal V, Canahuate G, Marai GE, Vock D, Lai SY, Mackin DS, Court LE, Freymann J, Farahani K, Kaplathy-Cramer J, Fuller CD (2018) Machine Learning Applications in Head and Neck Radiation Oncology: Lessons From Open-Source Radiomics Challenges. Front Oncol 8:294. https://doi.org/10.3389/fonc.2018.00294
Pfaehler E, Zwanenburg A, de Jong JR, Boellaard R (2019) RaCaT: An open source and easy to use radiomics calculator tool. PLoS One 14 (2):e0212223. https://doi.org/10.1371/journal.pone.0212223
Gotz M, Nolden M, Maier-Hein K (2019) MITK Phenotyping: An open-source toolchain for image-based personalized medicine with radiomics. Radiother Oncol 131:108-111. https://doi.org/10.1016/j.radonc.2018.11.021
Vallieres M, Zwanenburg A, Badic B, Cheze Le Rest C, Visvikis D, Hatt M (2018) Responsible Radiomics Research for Faster Clinical Translation. J Nucl Med 59 (2):189-193. https://doi.org/10.2967/jnumed.117.200501
Berenguer R, Pastor-Juan MDR, Canales-Vazquez J, Castro-Garcia M, Villas MV, Mansilla Legorburo F, Sabater S (2018) Radiomics of CT Features May Be Nonreproducible and Redundant: Influence of CT Acquisition Parameters. Radiology 288 (2):407-415. https://doi.org/10.1148/radiol.2018172361
Kuo MD, Jamshidi N (2014) Behind the numbers: Decoding molecular phenotypes with radiogenomics--guiding principles and technical considerations. Radiology 270 (2):320-325. https://doi.org/10.1148/radiol.13132195
Parmar C, Grossmann P, Bussink J, Lambin P, Aerts H (2015) Machine Learning methods for Quantitative Radiomic Biomarkers. Sci Rep 5:13087. https://doi.org/10.1038/srep13087
Kakino R, Nakamura M, Mitsuyoshi T, Shintani T, Kokubo M, Negoro Y, Fushiki M, Ogura M, Itasaka S, Yamauchi C, Otsu S, Sakamoto T, Sakamoto M, Araki N, Hirashima H, Adachi T, Matsuo Y, Mizowaki T (2020) Application and limitation of radiomics approach to prognostic prediction for lung stereotactic body radiotherapy using breath-hold CT images with random survival forest: A multi-institutional study. Med Phys. https://doi.org/10.1002/mp.14380
Antonacci Y, Toppi J, Mattia D, Pietrabissa A, Astolfi L (2019) Single-trial Connectivity Estimation through the Least Absolute Shrinkage and Selection Operator. Conf Proc IEEE Eng Med Biol Soc 2019:6422-6425. https://doi.org/10.1109/EMBC.2019.8857909
Currie G, Iqbal B, Kiat H (2019) Intelligent Imaging: Radiomics and Artificial Neural Networks in Heart Failure. J Med Imaging Radiat Sci 50 (4):571-574. https://doi.org/10.1016/j.jmir.2019.08.006
Xu L, Yang P, Liang W, Liu W, Wang W, Luo C, Wang J, Peng Z, Xing L, Huang M, Zheng S, Niu T (2019) A radiomics approach based on support vector machine using MR images for preoperative lymph node status evaluation in intrahepatic cholangiocarcinoma. Theranostics 9 (18):5374-5385. https://doi.org/10.7150/thno.34149
Radovic M, Ghalwash M, Filipovic N, Obradovic Z (2017) Minimum redundancy maximum relevance feature selection approach for temporal gene expression data. BMC Bioinformatics 18 (1):9. https://doi.org/10.1186/s12859-016-1423-9
Delzell DAP, Magnuson S, Peter T, Smith M, Smith BJ (2019) Machine Learning and Feature Selection Methods for Disease Classification With Application to Lung Cancer Screening Image Data. Front Oncol 9:1393. https://doi.org/10.3389/fonc.2019.01393
Kearns M, Ron D (1999) Algorithmic stability and sanity-check bounds for leave-one-out cross-validation. Neural Comput 11 (6):1427-1453. https://doi.org/10.1162/089976699300016304
Conrad M, Rizki MM (1980) Computational illustration of the bootstrap effect. Biosystems 13 (1-2):57-64. https://doi.org/10.1016/0303-2647(80)90005-2
Blum A, Wang P, Zenklusen JC (2018) SnapShot: TCGA-Analyzed Tumors. Cell 173 (2):530. https://doi.org/10.1016/j.cell.2018.03.059
Liu J, Lichtenberg T, Hoadley KA, Poisson LM, Lazar AJ, Cherniack AD, Kovatich AJ, Benz CC, Levine DA, Lee AV, Omberg L, Wolf DM, Shriver CD, Thorsson V, Cancer Genome Atlas Research N, Hu H (2018) An Integrated TCGA Pan-Cancer Clinical Data Resource to Drive High-Quality Survival Outcome Analytics. Cell 173 (2):400–416 e411. https://doi.org/10.1016/j.cell.2018.02.052
Tothill RW, Tinker AV, George J, Brown R, Fox SB, Lade S, Johnson DS, Trivett MK, Etemadmoghadam D, Locandro B, Traficante N, Fereday S, Hung JA, Chiew YE, Haviv I, Australian Ovarian Cancer Study G, Gertig D, DeFazio A, Bowtell DD (2008) Novel molecular subtypes of serous and endometrioid ovarian cancer linked to clinical outcome. Clin Cancer Res 14 (16):5198-5208. https://doi.org/10.1158/1078-0432.CCR-08-0196
Verhaak RG, Tamayo P, Yang JY, Hubbard D, Zhang H, Creighton CJ, Fereday S, Lawrence M, Carter SL, Mermel CH, Kostic AD, Etemadmoghadam D, Saksena G, Cibulskis K, Duraisamy S, Levanon K, Sougnez C, Tsherniak A, Gomez S, Onofrio R, Gabriel S, Chin L, Zhang N, Spellman PT, Zhang Y, Akbani R, Hoadley KA, Kahn A, Kobel M, Huntsman D, Soslow RA, Defazio A, Birrer MJ, Gray JW, Weinstein JN, Bowtell DD, Drapkin R, Mesirov JP, Getz G, Levine DA, Meyerson M, Cancer Genome Atlas Research N (2013) Prognostically relevant gene signatures of high-grade serous ovarian carcinoma. J Clin Invest 123 (1):517-525. https://doi.org/10.1172/JCI65833
Vargas HA, Wassberg C, Fox JJ, Wibmer A, Goldman DA, Kuk D, Gonen M, Larson SM, Morris MJ, Scher HI, Hricak H (2015) Response. Radiology 274 (2):625
Vargas HA, Huang EP, Lakhman Y, Ippolito JE, Bhosale P, Mellnick V, Shinagare AB, Anello M, Kirby J, Fevrier-Sullivan B, Freymann J, Jaffe CC, Sala E (2017) Radiogenomics of High-Grade Serous Ovarian Cancer: Multireader Multi-Institutional Study from the Cancer Genome Atlas Ovarian Cancer Imaging Research Group. Radiology 285 (2):482-492. https://doi.org/10.1148/radiol.2017161870
Vargas HA, Veeraraghavan H, Micco M, Nougaret S, Lakhman Y, Meier AA, Sosa R, Soslow RA, Levine DA, Weigelt B, Aghajanian C, Hricak H, Deasy J, Snyder A, Sala E (2017) A novel representation of inter-site tumour heterogeneity from pre-treatment computed tomography textures classifies ovarian cancers by clinical outcome. Eur Radiol 27 (9):3991-4001. https://doi.org/10.1007/s00330-017-4779-y
Meier A, Veeraraghavan H, Nougaret S, Lakhman Y, Sosa R, Soslow RA, Sutton EJ, Hricak H, Sala E, Vargas HA (2019) Association between CT-texture-derived tumor heterogeneity, outcomes, and BRCA mutation status in patients with high-grade serous ovarian cancer. Abdom Radiol (NY) 44 (6):2040-2047. https://doi.org/10.1007/s00261-018-1840-5
Lu H, Arshad M, Thornton A, Avesani G, Cunnea P, Curry E, Kanavati F, Liang J, Nixon K, Williams ST, Hassan MA, Bowtell DDL, Gabra H, Fotopoulou C, Rockall A, Aboagye EO (2019) A mathematical-descriptor of tumor-mesoscopic-structure from computed-tomography images annotates prognostic- and molecular-phenotypes of epithelial ovarian cancer. Nat Commun 10 (1):764. https://doi.org/10.1038/s41467-019-08718-9
Song XL, Ren JL, Zhao D, Wang L, Ren H, Niu J (2020) Radiomics derived from dynamic contrast-enhanced MRI pharmacokinetic protocol features: the value of precision diagnosis ovarian neoplasms. Eur Radiol. https://doi.org/10.1007/s00330-020-07112-0
Zhang H, Mao Y, Chen X, Wu G, Liu X, Zhang P, Bai Y, Lu P, Yao W, Wang Y, Yu J, Zhang G (2019) Magnetic resonance imaging radiomics in categorizing ovarian masses and predicting clinical outcome: a preliminary study. Eur Radiol 29 (7):3358-3371. https://doi.org/10.1007/s00330-019-06124-9
Jian J, Li Y, Pickhardt PJ, Xia W, He Z, Zhang R, Zhao S, Zhao X, Cai S, Zhang J, Zhang G, Jiang J, Zhang Y, Wang K, Lin G, Feng F, Wu X, Gao X, Qiang J (2020) MR image-based radiomics to differentiate type Iota and type IotaIota epithelial ovarian cancers. Eur Radiol. https://doi.org/10.1007/s00330-020-07091-2
Qian L, Ren J, Liu A, Gao Y, Hao F, Zhao L, Wu H, Niu G (2020) MR imaging of epithelial ovarian cancer: a combined model to predict histologic subtypes. Eur Radiol. https://doi.org/10.1007/s00330-020-06993-5
Wang G, Sun Y, Chen Y, Gao Q, Peng D, Lin H, Zhan Z, Liu Z, Zhuo S (2020) Rapid identification of human ovarian cancer in second harmonic generation images using radiomics feature analyses and tree-based pipeline optimization tool. J Biophotonics:e202000050. https://doi.org/10.1002/jbio.202000050
Medicine NLo (2020) Proteomics MeSH Descriptor Data https://meshb.nlm.nih.gov/record/ui?ui=D040901.
Zhang H, Liu T, Zhang Z, Payne SH, Zhang B, McDermott JE, Zhou J-Y, Petyuk VA, Chen L, Ray D, Sun S, Yang F, Chen L, Wang J, Shah P, Cha SW, Aiyetan P, Woo S, Tian Y, Gritsenko MA, Clauss TR, Choi C, Monroe ME, Thomas S, Nie S, Wu C, Moore RJ, Yu K-H, Tabb DL, Fenyö D, Bafna V, Wang Y, Rodriguez H, Boja ES, Hiltke T, Rivers RC, Sokoll L, Zhu H, Shih I-M, Cope L, Pandey A, Zhang B, Snyder MP, Levine DA, Smith RD, Chan DW, Rodland KD, Carr SA, Gillette MA, Klauser KR, Kuhn E, Mani DR, Mertins P, Ketchum KA, Thangudu R, Cai S, Oberti M, Paulovich AG, Whiteaker JR, Edwards NJ, McGarvey PB, Madhavan S, Wang P, Chan DW, Pandey A, Shih I-M, Zhang H, Zhang Z, Zhu H, Cope L, Whiteley GA, Skates SJ, White FM, Levine DA, Boja ES, Kinsinger CR, Hiltke T, Mesri M, Rivers RC, Rodriguez H, Shaw KM, Stein SE, Fenyo D, Liu T, McDermott JE, Payne SH, Rodland KD, Smith RD, Rudnick P, Snyder M, Zhao Y, Chen X, Ransohoff DF, Hoofnagle AN, Liebler DC, Sanders ME, Shi Z, Slebos RJC, Tabb DL, Zhang B, Zimmerman LJ, Wang Y, Davies SR, Ding L, Ellis MJC, Townsend RR (2016) Integrated Proteogenomic Characterization of Human High-Grade Serous Ovarian Cancer. Cell 166 (3):755-765. https://doi.org/10.1016/j.cell.2016.05.069
Zhang B, Whiteaker JR, Hoofnagle AN, Baird GS, Rodland KD, Paulovich AG (2019) Clinical potential of mass spectrometry-based proteogenomics. Nat Rev Clin Oncol 16 (4):256-268. https://doi.org/10.1038/s41571-018-0135-7
Yang J-Y, Yoshihara K, Tanaka K, Hatae M, Masuzaki H, Itamochi H, Cancer Genome Atlas Research N, Takano M, Ushijima K, Tanyi JL, Coukos G, Lu Y, Mills GB, Verhaak RGW (2013) Predicting time to ovarian carcinoma recurrence using protein markers. J Clin Invest 123 (9):3740-3750. https://doi.org/10.1172/JCI68509
Integrated genomic analyses of ovarian carcinoma (2011). Nature 474 (7353):609–615. https://doi.org/10.1038/nature10166
Beer L, Sahin H, Bateman NW, Blazic I, Vargas HA, Veeraraghavan H, Kirby J, Fevrier-Sullivan B, Freymann JB, Jaffe CC, Brenton J, Miccó M, Nougaret S, Darcy KM, Maxwell GL, Conrads TP, Huang E, Sala E (2020) Integration of proteomics with CT-based qualitative and radiomic features in high-grade serous ovarian cancer patients: an exploratory analysis. Eur Radiol 30 (8):4306-4316. https://doi.org/10.1007/s00330-020-06755-3
Weon JL, Potts PR (2015) The MAGE protein family and cancer. Curr Opin Cell Biol 37:1-8. https://doi.org/10.1016/j.ceb.2015.08.002
Yakirevich E, Sabo E, Lavie O, Mazareb S, Spagnoli G, Resnick M (2003) Expression of the MAGE-A4 and NY-ESO-1 cancer-testis antigens in serous ovarian neoplasms. Clinical cancer research : an official journal of the American Association for Cancer Research 9:6453-6460
Xu Y, Wang C, Zhang Y, Jia L, Huang J (2015) Overexpression of MAGE-A9 Is Predictive of Poor Prognosis in Epithelial Ovarian Cancer. Sci Rep 5:12104. https://doi.org/10.1038/srep12104
Nicholson LJ, Smith PR, Hiller L, Szlosarek PW, Kimberley C, Sehouli J, Koensgen D, Mustea A, Schmid P, Crook T (2009) Epigenetic silencing of argininosuccinate synthetase confers resistance to platinum-induced cell death but collateral sensitivity to arginine auxotrophy in ovarian cancer. International Journal of Cancer 125 (6):1454-1463. https://doi.org/10.1002/ijc.24546
Vargas HA, Veeraraghavan H, Micco M, Nougaret S, Lakhman Y, Meier AA, Sosa R, Soslow RA, Levine DA, Weigelt B, Aghajanian C, Hricak H, Deasy J, Snyder A, Sala E (2017) A novel representation of inter-site tumour heterogeneity from pre-treatment computed tomography textures classifies ovarian cancers by clinical outcome. European Radiology 27 (9):3991-4001. https://doi.org/10.1007/s00330-017-4779-y
Fiore L, Rodriguez H, Shriver C (2017) Collaboration to Accelerate Proteogenomics Cancer Care: The Department of Veterans Affairs, Department of Defense, and the National Cancer Institute's Applied Proteogenomics OrganizationaL Learning and Outcomes (APOLLO) Network. Clinical Pharmacology & Therapeutics 101 (5):619-621. https://doi.org/10.1002/cpt.658
Pinker K, Chin J, Melsaether AN, Morris EA, Moy L (2018) Precision Medicine and Radiogenomics in Breast Cancer: New Approaches toward Diagnosis and Treatment. Radiology 287 (3):732-747. https://doi.org/10.1148/radiol.2018172171
Zwanenburg A, Leger S, Vallières M, Löck S (2018) Initiative for the IBS. Image biomarker standardisation initiative. https://www.arxivorg/abs/161207003
Vokes TJ, Pham A, Wilkie J, Kocherginsky M, Ma SL, Chinander M, Karrison T, Bris O, Giger ML (2008) Reproducibility and sources of variability in radiographic texture analysis of densitometric calcaneal images. J Clin Densitom 11 (2):211-220. https://doi.org/10.1016/j.jocd.2007.10.004
Mwangi B, Tian TS, Soares JC (2014) A review of feature reduction techniques in neuroimaging. Neuroinformatics 12 (2):229-244. https://doi.org/10.1007/s12021-013-9204-3
Vallieres M, Laberge S, Diamant A, El Naqa I (2017) Enhancement of multimodality texture-based prediction models via optimization of PET and MR image acquisition protocols: a proof of concept. Phys Med Biol 62 (22):8536-8565. https://doi.org/10.1088/1361-6560/aa8a49
Parmar C, Rios Velazquez E, Leijenaar R, Jermoumi M, Carvalho S, Mak RH, Mitra S, Shankar BU, Kikinis R, Haibe-Kains B, Lambin P, Aerts HJ (2014) Robust Radiomics feature quantification using semiautomatic volumetric segmentation. PLoS One 9 (7):e102107. https://doi.org/10.1371/journal.pone.0102107
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declared that they have no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Nougaret, S., McCague, C., Tibermacine, H. et al. Radiomics and radiogenomics in ovarian cancer: a literature review. Abdom Radiol 46, 2308–2322 (2021). https://doi.org/10.1007/s00261-020-02820-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00261-020-02820-z