Abstract
Globally, Salmonella infection poses a major public health problem. Here, we report antibiofilm activity and quorum sensing inhibition of aqueous seeds extract of Myristica fragrans (nutmeg) and biosynthesized silver nanoparticles (AgNPs) against multidrug resistant (MDR) Salmonella enterica serovar Typhi (S. Typhi) isolated from typhoid patients and asymptomatic carriers. S. Typhi isolates revealed higher percentage (46%) of biofilm production identified by tissue culture plate (TCP) than Congo red agar (CRA) and tube adherence (TA) methods. The inhibition of biofilm-producing MDR S. Typhi isolates and pigment production of Chromobacterium violaceum (indicator bacteria) demonstrated the quorum sensing potential of nutmeg. The aqueous seed extract of nutmeg exhibited 87% of antibiofilm activity, while the biosynthesized AgNPs showed 99.1% of antibiofilm activity. Molecular docking studies of bioactive compounds of nutmeg against transcriptional regulatory protein RcsB and sensor kinase protein RcsC revealed interaction with the target proteins. It is proposed that biosynthesized AgNPs could be used as one of the effective candidates in treating asymptomatic typhoid carriers or typhoid patients and to control the subsistence of biofilm-producing S. Typhi strains or other pathogenic bacteria in the environment or industrial settings.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Salmonella species are one of the most important pathogens worldwide as they exhibit high morbidity and mortality (Fabrega and Vila 2013). Despite advanced treatment and precautions efforts, every year, millions of cases with new typhoid infections occur worldwide. Following infection, Salmonella enterica serovar Typhi (S. Typhi) colonizes gallbladder and stays there long after subsiding symptoms and also serving as a reservoir for spread of disease further (Gonzalez-Escobedo et al. 2011). However, a significant percentage of typhoid infection can result in asymptomatic carriage of S. Typhi (Reeve 2010; Abida et al. 2011). The clinical observations among typhoid carriers regarding their resistance to antibiotic treatment and extension of infection to the gallbladder are related with biofilm-producing potential of S. Typhi (Swidsinski and Lee 2001). Biofilms are organized communities of microorganisms that adhere to each other and to inert or live substrates and are encased in an extracellular matrix (Monds and O’Toole 2009). Generally, it is considered as a response to stress; biofilms have been concerned in many chronic and acute infections. In many typhoid cases, persistence can lead to establishing biofilm on gallstone both in typhoid acute patients and also in chronic carriers (Adcox et al. 2016). Subsequently, Salmonella species biofilm formation has also been observed in gallbladder infection by in vitro studies (Prouty et al. 2002) and in human typhoid carrier (Crawford et al. 2010; Marshall et al. 2014). However, a significant percentage of typhoid infection can result in asymptomatic carriage of S. Typhi, possibly due to the formation of biofilms that contributes to the development of the carrier state (Reeve 2010; Abida et al. 2011). The duration of S. Typhi clearance from typhoid patients after recovery is directly related to their biofilm-producing ability (Abida et al. 2011).
Typhoidal and non-typhoidal Salmonella biofilms are the most important health problem in industrial, veterinary, and medical settings (Veldman et al. 1995) because they are critical in the failure of the antibiotic treatment in patients and lead to developing and spreading of antibiotic resistance pathogens among healthy people and also responsible for many foodborne outbreaks originated from food industries in the absence of regular sanitation regimens (Shi and Zhu 2009).
Multidrug resistance (MDR) in Salmonella species (Senthilkumar and Prabakaran 2005) can prevent the antibiotics/drugs from accessing their target by a number of mechanisms like mutations in topoisomerase IV (fluoroquinolone resistance) and DNA gyrase (Giraud et al. 2006) or breaking them down to an inactive form (Miro et al. 2004). These mechanisms are mediated by making changes in the cellular envelope, over expressing efflux pump systems, downregulating porins, or by increasing lipopolysaccharide (LPS) component present in the outer membrane (Giraud et al. 2000). Besides these, quorum sensing (QS) and the formation of biofilms also contribute to resistance (Jayaraman and Wood 2008). The QS signals produced by Salmonella enterica is used by some bacterial pathogens to coordinate virulence gene expression (Perrett et al. 2009), activating gene expression responsible for Salmonella pathogenicity island-1 (SPI-1) (Choi et al. 2007; Senthilkumar et al. 2019), and in the motility and colonization (Bearson and Bearson 2008).
Two-component signal transduction systems (2CSTSs) in the bacteria provide with a complex network of regulation that manages motility, pathogenesis, and biofilm development and responding to a numerous environmental signal (Prüß 2017). Bacterial colonization and their survival happen to some extent by means of a phosphotransfer mechanism by 2CSTSs (Zschiedrich et al. 2016). The 2CSTS usually consists of a sensor kinase (histidine kinase) and a response regulator (RR). Mostly, sensor kinases are membrane-bound proteins that autophosphorylate the conserved histidine residue in the presence of ATP, from which the phosphoryl is transferred to a conserved aspartate in the response regulator (Mitrophanov and Groisman 2008). Two-component systems (TCSs) are becoming popular antibacterial drug targets as these are ubiquitous in all bacteria and are essential for their survival, and homology is not detected in humans or other mammals (Thomason and Kay 2000).
We have hitherto stated the occurrence of the MDR S. Typhi isolates (Senthilkumar and Prabakaran 2005; Senthilkumar et al. 2011; Balaji and Senthilkumar 2011) in asymptomatic typhoid carriers (Sasikumar et al. 2005; Senthilkumar et al. 2014a, b, c; Ilakkia et al. 2015, 2016) and antimicrobial activity of nutmeg-mediated biosynthesized AgNPs against the MDR S. Typhi isolates (Senthilkumar et al. 2017) among the food handlers and school children. Therefore, this study was intended to evaluate biosynthesized AgNPs for antiquorum sensing and antibiofilm activity against the MDR S. Typhi isolates and to screen bioactive compounds of M. fragrans (nutmeg) with the transcriptional regulatory protein RcsB and sensor kinase protein RcsC of S. Typhi through molecular docking approach.
Materials and methods
Biosynthesized AgNPs and bacterial pathogens
Nutmeg aqueous seed extract, biosynthesized AgNPs, and MDR S. Typhi isolates MCASMZU1-13 (NCBI accession no. KT037135-38, KT037130-34, KT696507, KT696504-06) from asymptomatic typhoid carriers and typhoid patients illustrated elsewhere (Senthilkumar et al. 2017) were used for the biofilm profiling and antibiofilm assay in this study.
Biofilm profiling assay
Potent biofilm forming MDR S. Typhi isolates were evaluated by the following three methods.
Congo red agar method
The Congo red agar (CRA) method (Freeman et al. 1989) was performed using brain heart infusion (BHI) agar supplemented with 5% sucrose and Congo red. A darkening of the bacterial colonies with a dry crystalline morphology indicated high biofilm producer, whereas moderate biofilm producer indicated pink with black-centered colonies, and weak biofilm producer indicates pink colonies. The experiment was performed in triplicate.
Tube adherence method
A qualitative assessment of biofilm formation was determined as described by Christensen et al. (1982) with few modifications. Trypticase soy broth (10 mL) with 1% glucose (TSBglu; HiMedia, India) was inoculated with 1% logarithmic phase culture (OD590 at 0.5) and incubated at 37 °C for 24 h. The aliquots were decanted and washed twice with phosphate buffer saline (PBS; pH 7.2). The dried tubes were stained with 0.1% (w/v) crystal violet for 30-min incubation, the excess stain was removed, and the tubes were washed twice with deionized water. The dried tubes were observed for any visible film that lined the wall and the bottom of the tube. The amount of biofilm formation was recorded on the basis of intensity of crystal violet coloration as absent, moderate, and strong. The experiment was performed in triplicate.
Tissue culture plate method
All the isolates screened for biofilm production were confirmed in tissue culture plate (TCP) method by following Christensen et al. (1985) with minor changes. S. Typhi isolates were inoculated in TSBglu broth (HiMedia, India) and brain heart infusion broth with 2% sucrose (BHIsuc) (HiMedia, India) and incubated at 37 °C for 18 h. About 1 mL of broth culture was diluted in 100 mL fresh broth, and 200 μL of this diluted culture was added in individual wells of sterile, polystyrene 96-well flat-bottom tissue culture plate, and broth without culture was served as control to check sterility and non-specific binding of media. After 24-h incubation at 37 °C, the planktonic cells were removed from wells and washed twice with 200 μL of PBS. Biofilm formed by adhered cells in plate was fixed with sodium acetate (2%) and stained by crystal violet (0.1% w/v) for 30 min. Excess stain was rinsed off by thorough washing with deionized water, and the plate was dried. Usually adherent S. Typhi cells formed biofilm on the sides of wells were uniformly stained with crystal violet. Optical density (OD) of the stained adherent bacteria was determined with micro ELISA auto reader (Bio-Rad) at 570 nm. The experiment was performed in triplicate; the data were averaged; and, standard deviation was calculated. The mean OD value obtained from the media control well was deducted from all the test OD values.
Determination of quorum sensing inhibition
The biofilm-inhibiting ability of nutmeg aqueous seed extract and biosynthesized AgNPs was determined using soft agar overlay protocol as described by McLean et al. (2004) with minor changes. Briefly, BHI agar (HiMedia, India) plates were prepared and halved into two parts. The representative MDR S. Typhi MCASMZU1 (typhoid patient) and MCASMZU7 (asymptomatic typhoid carrier) isolates were streaked separately on one-half, and the nutmeg aqueous seed extract was added into the well made on another half. Similarly, the test isolates were streaked on one-half and the biosynthesized AgNPs were added into the well made on another half. After incubation at 30 °C for 24 h, 5 mL of soft agar containing the indicator bacteria Chromobacterium violaceum (ATCC 12472) was overlaid and incubated for growth and pigmentation.
Antibiofilm assay
Biofilm inhibition was carried out in a 96-well plate by adopting a modified method of biofilm inhibition spectrophotometric assay (Gurunathan et al. 2014). About 100 μL of log phase cell suspension (OD590 of 0.5) of each representative MDR S. Typhi MCASMZU1 (typhoid patient) and MCASMZU7 (asymptomatic typhoid carrier) isolates, was added into a 96-well titer plate, and different concentrations of nutmeg aqueous seed extract and biosynthesized AgNPs as 10, 20, 30, 40, and 50 μg mL−1 were added and incubated at 37 °C for 3 days. The wells were stained with 100 μL of 1% (w/v) crystal violet after discarding cell suspension. After 30-min incubation, the dye was removed, thoroughly washed, and added with 95% ethanol for 15 min. This reaction mixture was read spectrophotometrically at 595 nm. The percentage inhibition of biofilm activity was calculated by the following equation: [1-(A 595 of test/A 595 of control)] × 100 (Wei et al. 2006). The experiment was performed in triplicate. The data are expressed as mean ± SD.
Homology modeling and molecular docking
The protein sequences of transcriptional regulatory protein RcsB (accession Q56127) and sensor histidine kinase RcsC (accession Q56128) from S. Typhi were retrieved using ExPasY UniProtKB (Gasteiger et al. 2003; Pundir et al. 2017), and their three-dimensional structures were modeled using the Swiss-Model (Biasini et al. 2014) and Phyre2 (Kelley et al. 2015) web servers. Structure refinement and energy minimization of the predicted models were performed using ModRefiner (Xu and Zhang 2011) and GROMOS96 force field (van Gunsteren et al. 1996), respectively. Energy-refined models were then validated using ProSA (Wiederstein and Sippl 2007) and the Ramachandran plot from PROCHECK (Laskowski et al. 1993).
In order to hypothesize a mechanism for quorum sensing inhibition and antibiofilm activities, homology modeling and molecular docking studies were performed on the Rcs signal transduction system in S. Typhimurium. The molecular interaction between the bioactive compounds in nutmeg aqueous seed extract and transcriptional regulatory protein RcsB and sensor kinase protein RcsC from S. enterica has been performed using AutoDock 4.2 (Morris et al. 2009). The choice of phytochemicals was based on our pervious publication on nutmeg (Senthilkumar et al. 2017), and their three-dimensional structures were retrieved from public database PubChem (Kim et al. 2016). In order to distinguish a drug-like compound from other compounds, Sanjeevini (Jayaram et al. 2012) was used to evaluate the molecular descriptors following Lipinski’s rule of five (Lipinski 2004). Addition of hydrogen atoms, Kollman charges, and polar hydrogens and merging of non-polar hydrogens were performed using the AutoDock Tools (ADT). For RcsB, blind docking was performed by setting a grid box of 60 × 60 × 60, with spacing of 0.7 Å and for RcsC, a grid box of 48 × 46 × 52, with a spacing of 0.7 Å, was positioned using autogrid. The Lamarckian genetic algorithm (LGA) of up to 20 runs was employed with a population size of 150 individuals and maximum number of generations and energy evaluations of 27,000 and 2.5 million, respectively (Yadav et al. 2015). Ligand-binding energy (ΔG) and other parameters for each ligand were calculated and ranked based on the lowest binding energy. The best pose (top ranked) and capacity to form hydrogen bonds (if any) were alone considered for each ligand.
Results and discussion
Biofilm profiling of MDR S. Typhi isolates
Biofilm profiling of MDR S. Typhi isolates were found to be 38.5% strong biofilm producer, 30.8% moderate biofilm producer, and 30.8% weak biofilm producer by the CRA method. In the TCP method, 46% of MDR S. Typhi isolates showed mean OD value > 0.240, indicating high biofilm producers, 38.5% of MDR S. Typhi isolates were moderate biofilm producers (mean OD value 0.120–0.240), and 15% were found to be weak biofilm producers (mean OD value of < 0.120). In the tube adherence (TA) method, 15% of MDR S. Typhi isolates were strongly positive, 46% were moderately positive, and 38.5% were weak or non-biofilm producers (Table 1). These results indicate that high percentage of biofilm producers were profiled in the TCP method than the other methods including CRA and TA method. Similarly, the TCP method is a more reliable method for the detection of biofilm from clinical bacterial isolates (Afreenish et al. 2011). Agarwal et al. (2011) reported that majority of Salmonella strains (87, 57.61%) were found to be moderate biofilm producers, while 22.52% were weak producers, and 19.21% of strains were strong biofilm producers. Sereno et al. (2017) also found that the rates of 13.6, 59.1, and 27.3% of isolates were moderate, weak, and non-biofilm producers among 100 MDR Salmonella strains from poultry carcasses. The profiling of biofilm production by S. Typhi strains may identify that the high-risk patients would be the key for effective therapeutic management and gallbladder infection prevention (Enea et al. 2017). Because, bacterial cells growing in biofilm are physiologically distinct from planktonic cells of the same bacteria and are embedded within a self-produced matrix of extra cellular polymeric substance (EPS) (Monroe 2007; Hall-Stoodley et al. 2006), which can increase antibiotic resistance by up to 1000-fold (Stewart and Costerton 2001).
Quorum sensing inhibition
Biofilm assists cells to live close to each other in order to facilitate the exchange of plasmids and free DNA that enable them to overcome different environmental stresses. The bacteria in biofilm use chemical communication that helps them to coordinate their metabolism and other complex processes (Ankita et al. 2017). Quorum sensing of bacteria is mediated by Cyclic di-GMP (Sharma et al. 2014) and uses it as a checkpoint to proceed through the distinct stages of biofilm development until they fully commit to the biofilm lifestyle including Salmonella enterica (Valentini and Filloux 2016).
The inhibition of the growth of each representative MDR S. Typhi MCASMZU1 (typhoid patient) and MCASMZU7 (asymptomatic typhoid carrier) isolates and inhibition of pigment production by the indicator bacteria, C. violaceum, around the well containing nutmeg aqueous seed extract and AgNPs revealed the quorum sensing inhibition potential of nutmeg.
We previously reported that nutmeg aqueous seed extract consists of methane, oxybis [dichloro - and 1,4-benzenediol, 2-bromo, 1H-cyclopenta [c] furan- 3-(3aH)-one, 6,6a-dihydro-1-(1,3-dioxolan-2-yl)-, and (3aR, 1-t, and these compounds are effective in killing S. Typhi isolates (Senthilkumar et al. 2017) and in inhibiting biofilm formation (Seghal et al. 2010). This study results also presumed that the compounds of nutmeg aqueous seed extract might be responsible for the quorum sensing inhibition of S. Typhi and hence decided to perform a molecular interaction study with AutoDock 4.2.
Antibiofilm effects of nutmeg aqueous seed extract and biosynthesized AgNPs
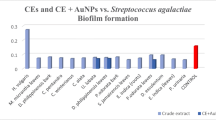
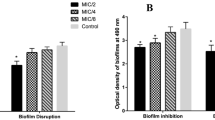
Biofilm inhibition studies carried out using nutmeg aqueous seed extract and biosynthesized silver nanoparticles at all the tested concentrations have successfully inhibited biofilm formation by the representative MDR S. Typhi isolates from typhoid patients and asymptomatic carriers. This study also revealed that the nutmeg aqueous seed extract showed 87% antibiofilm activity while the biosynthesized AgNPs showed 99.1% of antibiofilm activity at the concentration of 50 μg mL−1. This result concludes that AgNPs significantly (p > 0.05) inhibited biofilm formation than the nutmeg aqueous seed extract but in dose-dependent manner (Fig. 1a, b). Similarly, Gurunathan et al. (2014) reported that the efficiency of biosynthesized AgNPs exhibited enhanced biofilm inhibition than nutmeg aqueous seed extract. The high concentration of nanoparticles leads to diffusion through the medium by which they inhibit bacterial growth by more than 90% (Murugan et al. 2013).
Senthilkumar et al. (2017) reported antimicrobial potential of nutmeg aqueous seed extract and its AgNPs against the MDR S. Typhi isolates and proved that nutmeg aqueous seed extract could be used to treat bacterial diseases. Biosynthesis of AgNPs using nutmeg aqueous seed extract is an excellent, cost-effective, non-toxic, eco-friendly approach, and highly effective for treating human pathogens. Among natural plant extracts, stem bark (Murugan et al. 2014) and seed extract (Senthilkumar et al. 2017) have many advantages during sample collection process, as they are not destroying the parent plant.
Homology modeling and energy minimization of modeled structures
A protein’s ability to establish interaction with other molecules or to have varied functions depends upon their tertiary or 3D structure (Hasan et al. 2011; Alshatwi et al. 2011). As there are no crystal structures available in the protein data bank (PDB), the tertiary structures of RcsC and RcsB have been modeled using the web-based servers like Swiss-Model and Phyre2. Template search was done using BLAST (Altschul et al. 1997) and HHBlits (Remmert et al. 2012) against the SWISS-MODEL template library (SMTL version September 21, 2017; PDB release September 15, 2017). For transcriptional regulatory protein RcsB, PDB ID: 5i4c with 99.32 and sensor kinase protein RcsC, PDB ID: 2ayx with 80.97, sequence identity was used as templates for building models by Swiss-Model. Based on the target-template alignment, models were built using ProMod3. Final geometry of the resulting models was then optimized by using a force field. Phyre2 used template PDB ID: 5o8y for RcsB and for RcsC 7 templates (PDB ID: 5idj, 3d36, 2c2a, 4gcz, 2ayx, 4ew8, 4i5s) and was selected to model the protein based on heuristics to increase the confidence, percentage of identity, and coverage of the alignment. Phyre2 uses the hidden Markov method to generate alignments (Kelley and Sternberg 2009). Besides that, Phyre2 uses Poing, an ab initio folding simulation algorithm, to model the regions of the query where no detectable similarities to the known structures are noted (Jefferys et al. 2010). Out of 948 amino acids, 343 residues were modeled using the ab initio method. Poing algorithm uses information from multiple templates with known structures in order to develop the final model. The “intensive” mode in Phyre2 was used to build the 3D structure models for RcsB and RcsC. Structure refinement and energy minimization were performed to correct distorted geometries of predicted models. Figure 2 shows the refined protein structures from Swiss-Model and Phyre2. For RcsB and RcsC, Swiss-Model generated the models with lengths 131 (aa 2–132) and 247 (aa 701–947), respectively, whereas Phyre2 generated the models with lengths 216 (aa 1–216) and 948 (aa 1–948), respectively.
Model validation for RcsB and RcsC
Validation of the model is very critical for the modeled proteins as the structure is being used further downstream simulations for better understanding the protein’s biological function. Figure 3 shows the phi/psi Ramachandran plot of energy-minimized structures generated by PROCHECK. The plot shows phi/psi dihedral angles between N-Cα and Cα-C planar peptide bonds in the protein’s backbone. Though theoretically, any combination of these angles is possible; biologically limited combinations are seen due to steric clashes that occur in the proteins’ backbone structure (Goswami 2015). The numbers of residues in the most favored regions, additionally allowed, generously allowed, and disallowed regions are as presented in Table 2. The energy-minimized models were further evaluated using the ProSA web server. ProSA predicts the overall model quality and represent as Z-score by measuring the total energy deviation of the structure using random conformations and molecular mechanics force field (Wiederstein and Sippl 2007). This score is then plotted against the experimentally determined protein structures in the PDB. This helps us to visualize if the refined structure falls within the ranges of the native proteins (Doss et al. 2012). Z-score for energy-minimized PDB structure from Swiss-Model for RcsB and RcsC were − 7.82 and − 6.96, respectively. On the other hand, Z-score for energy-minimized PDB structure from Phyre2 were − 8.72 and − 7.31, respectively. These results indicated that the model generated from Phyre2 was better. Thus, based on model length and Z-score, the model generated by Phyre2 was chosen for further docking studies.
Molecular docking of bioactive compounds in nutmeg
Based on the promising in vitro results, it was thought worthy to screen the bioactive compounds in nutmeg aqueous seed extract, inculcating both in silico and in vitro results by performing molecular docking studies. Considering transcriptional regulatory protein RcsB and sensor kinase protein RcsC from S. enterica as the target receptor, comparative and automated docking studies with the bioactive compounds in nutmeg were performed using the LGA in the docking program AutoDock 4.2. Additionally, as positive control, ciprofloxacin was docked with each protein structure in order to compare docking energies with the phytochemicals. For protein targets, standard protonation states on neutral pH were used as a rigid model structure; no relaxation of the protein was performed during the docking studies. All the ligand molecule structures were energy minimized and screened for Lipinski’s rule of five before docking. The molecular descriptors of the ligands used in the study are presented in Table 3. Figures 4 and 5 show the docked images of the candidate ligands including the control drug, i.e., ciprofloxacin. Table 4 shows the lowest energy (strongest docking) scores for each of the candidate ligands including the standard in each protein target. In silico studies revealed that all the bioactive compounds showed good binding energy towards both the target proteins and established bonds with one or more amino acids.
Docked poses of bioactive compounds in nutmeg with the transcriptional regulatory protein RcsB. a Methane, oxybis [dichloro-, b 1H-cyclopenta [c] furan-3-(3aH)-one, 6,6a-dihydro-1-(1,3-dioxolan-2-yl)-, (3aR, 1-t, c octadecane, 6-methyl-, d heptadecane, 2,6,10,14-tetramethyl-, e BIS (2-ethylhextl) phthalate, f 4H-pyran-4-one,2,3-dihydro-3,5-dihydroxy- 6-methyl-, g 3,4-dichlorophenethylamine, h 1,4-benzenediol, 2-bromo-, i ciprofloxacin
Docked poses of bioactive compounds in nutmeg with the sensor histidine kinase RcsC. a Methane, oxybis [dichloro-, b 1H-cyclopenta [c] furan-3-(3aH)-one, 6,6a-dihydro-1-(1,3-dioxolan-2-yl)-, (3aR, 1-t, c Octadecane, 6-methyl-, d heptadecane, 2,6,10,14-tetramethyl-, e BIS (2-ethylhextl) phthalate, f 4H-pyran-4-one, 2, 3-dihydro-3,5-dihydroxy- 6-methyl-, g 3,4-dichlorophenethylamine, h 1,4-benzenediol, 2-bromo-, i ciprofloxacin
The Rcs signaling system follows the two-component signaling mechanism, though it consists of more than two proteins. The RcsCDB system has a numerous of functions, not just activating colanic acid synthesis (Sharma et al. 2017; Oropeza et al. 2015) but also downregulates genes like regulator for curli, csgD (Ferrieres and Clarke 2003), and flagellar gene, flhD (Francez-Charlot et al. 2003).
RcsB is basically a response regulator and forms a component of the Rcs signaling system that controls the transcription of several other genes. RcsB can function both in a RcsA-dependent or RcsA-independent manner by binding to the regulatory DNA regions. The protein has two domains: amino acids 5 to 124 form the response regulatory domain and the amino acids 144 to 209 HTH luxR-type domain. More specifically, it is the amino acids 168 to 187 that contribute to the helix-turn-helix (H-T-H) motif. The HTH motifs are common in all known DNA-binding proteins which regulates gene expression. The first helix of the HTH motif helps in stabilizing the structure while the second helix binds to DNA through a number of hydrogen bonds and hydrophobic interactions (Sauer et al. 1982; Brennan and Matthews 1989). In this study, compound 4H-pyran-4-one, 2,3-dihydro-3, 5-dihydroxy- 6-methyl- established three hydrogen bonds with the amino acids VAL168 and THR169 with a docking score of − 3.43. Compounds 3,4-dichlorophenethylamine (Fig. 4g) and 1,4-benzenediol, 2-bromo- (Fig. 4h) on the other hand established two hydrogen bonds with amino acids LYS173, ASN176, and ASN176, ARG177, respectively. Though compound 1H-cyclopenta [c] furan-3-(3aH)-one, 6,6a-dihydro-1-(1,3-dioxolan-2-yl)-, (3aR, 1-t exhibited the highest docking score of − 4.08 among the phytocompounds, it managed to establish only one hydrogen bond with the amino acid ASN176 (Fig. 4; Table 4).
Bem et al. (2015) have reviewed excellently on how novel antibiotics can target histidine kinases (HKs), especially those HKs that are part of the bacterial two-component systems (TCSs). TCS signaling pathway involves autophosphorylation of a histidine kinase (HK), phosphotransfer of the phosphoryl group to a cognate response regulator (RR), which in turn, modulates the expression of target genes (Casino et al. 2010). A HK usually contains four main domains, namely a periplasmic sensor domain (Sp), cytoplasmic sensor domains (Sc), dimerization and phosphotransfer domain (DHp), and catalytic and ATP-binding domain (CA) (Bem et al. 2015). HK autophosphorylation is mainly mediated by the CA domain, which binds ATP and phosphorylates the HK at a conserved histidine residue (HIS479) in the DHp domain. In most HKs, the CA and DHp domains are conserved whereas the remaining sensor domains like PAS, GAF, HAMP, and periplasmic domains are quiet variable and not found in all HKs (Casino et al. 2010). TCS signaling is blocked by dephosphorylation of the RR, which can be auto induced or mediated by the cognate HK or by auxiliary proteins (Stock et al. 2000; Parashar et al. 2011). Several studies have shown that TCSs are essential for antibiotic resistance, coordinated expression of virulence factors, and, in some cases, for bacterial growth and viability. Because of this, TCSs have been considered potent targets for antibacterial drug design (Thomason and Kay 2000).
In our study, RcsC, a membrane-associated histidine kinase was chosen as the second target. Here, 1,4-benzenediol, 2-bromo- with a docking score of − 3.47 established three hydrogen bonds with the amino acids PHE473, ALA475, and HIS479 (Fig. 5h). Compounds 4H-pyran-4-one, 2, 3-dihydro-3,5-dihydroxy- 6-methyl- (docking score of − 3.0) (Fig. 5f) and 3,4-dichlorophenethylamine (docking score of − 5.2) (Fig. 5g) formed two hydrogen bonds with amino acids HIS479, THR483, ALA475, and GLU480, respectively. From the results, it is clearly evident that compounds 1,4-benzenediol, 2-bromo- and 4H-pyran-4-one, 2,3-dihydro-3,5-dihydroxy- 6-methyl- interact with HIS479 responsible for the phosphorylation (Fig. 5). Moreover, compound 3,4-dichlorophenethylamine exhibited higher docking score compared with the control ciprofloxacin (Table 4).
Because of the sequence similarities of HKs’ CA domain from different organisms, it can be suggested that these putative HK inhibitors from nutmeg would exhibit polypharmacology (inhibiting multiple targets) which could slow down antimicrobial resistance development. But, it has to be noted that the docking method has its own limitations (Yuriev et al. 2011; Yuriev and Ramsland 2013). Moreover, this docking study has only examined natural ligand docking and does not take into account in vivo breakdown or other metabolites derived as a result of metabolism. Also, binding energies do not tell whether the ligands that bind strongly function as agonists or antagonists (Powers and Setzer 2015).
Conclusion
Treating MDR and biofilm-producing pathogenic bacterial diseases poses serious challenge in the clinical settings and hence, exploring new agents that can interfere with QS signal molecules and/or inhibition of the biofilm development are of useful future goals that should be considered. This study revealed the inhibition of the growth of MDR S. Typhi isolates and inhibition of pigment production by the indicator bacteria by nutmeg aqueous seed extract and further, AgNPs disclosed the quorum sensing inhibition potential of nutmeg. This is the first report of the antibiofilm activity of biosynthesized nutmeg AgNPs. Our in silico study showed how the phytochemicals interact with the target Rcs system. Consequently, the present study is evidenced to boost the biomedicine value of the nutmeg-based biosynthesized AgNPs which could be used to control the asymptomatic carriers, development of asymptomatic carriers, and existence of biofilm-producing S. Typhi strains and other pathogenic bacterial strains in the host, environment, or industrial settings.
References
Abida R, Yasra S, Aamir A, Amer J, Asma H, Abdul H (2011) Effect of biofilm formation on the excretion of Salmonella enterica serovar Typhi in feces. Int J Infect Dis 15:e747–e752
Adcox HE, Vasicek EM, Dwivedi V, Hoang KV, Turner J, Gunn JS (2016) Salmonella colonization. Infect Immun 84(11):3243–3251
Afreenish H, Javaid U, Fathima K, Maria O, Ali K, Muhammad I (2011) Evaluation of different detection methods of biofilm formation in the clinical isolates. Braz J Infect Dis 15(4):305–311
Agarwal RK, Singh S, Bhilegaonkar KN, Singh VP (2011) Optimization of microtitre plate assay for the testing of biofilm formation ability in different Salmonella serotypes. Int Food Res J 18:1493–1498
Alshatwi AA, Hasan TN, Syed NA, Shafi G (2011) Predicting the possibility of two newly isolated phenetheren ring containing compounds from Aristolochia manshuriensis as CDK2 inhibitors. Bioinform 7:334–338
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Ankita S, Jaya G, Sunil K, Awanish K (2017) Gut biofilm forming bacteria in inflammatory bowel disease. Microb Pathog. https://doi.org/10.1016/j.micpath.2017.09.041
Balaji C, Senthilkumar B (2011) Screening, phylogenetic analysis and antibiotic sensitivity pattern of Salmonella enterica serovar Typhi isolates from typhoid asymptomatic carriers. Asian Pac J Trop Med 4(10):769–772
Bearson BL, Bearson SM (2008) The role of the QseC quorum-sensing sensor kinase in colonization and norepinephrine-enhanced motility of Salmonella enterica serovar Typhimurium. Microb Pathog 44(4):271–278
Bem AE, Velikova N, Pellicer MT, Pv B, Marina A, Wells JM (2015) Bacterial histidine kinases as novel antibacterial drug targets. ACS Chem Biol 10:213–224
Biasini M, Bienert S, Waterhouse A, Arnold K, Studer G, Schmidt T, Kiefer F, Cassarino TG, Bertoni M, Bordoli L, Schwede T (2014) SWISS-MODEL: modelling protein tertiary and quaternary structure using evolutionary information. Nucleic Acids Res 42:W252–W258
Brennan RG, Matthews BW (1989) The helix-turn-helix DNA binding motif. J Biol Chem 264(4):1903–1906
Casino P, Rubio V, Marina A (2010) The mechanism of signal transduction by two-component systems. Curr Opin Struct Biol 20:763–771
Choi J, Shin D, Ryu S (2007) Implication of quorum sensing in Salmonella enterica serovar Typhimurium virulence: the luxS gene is necessary for expression of genes in pathogenicity island 1. Infect Immun 75(10):4885–4890
Christensen GD, Simpson WA, Bisno AL, Beachey EH (1982) Adherence of slime producing strains of Staphylococcus epidermidis to smooth surfaces. Infect Immun 37:318–326
Christensen GD, Simpson WA, Younger JJ, Baddour LM, Barrett FF, Melton DM, Beachey EH (1985) Adherence of coagulase-negative staphylococci to plastic tissue culture plates: a quantitative model for the adherence of staphylococci to medical devices. J Clin Microbiol 22(6):996–1006
Crawford RW, Reyesb RR, Ramirez-Aguilarb ML, Azuelac OC, Arandad CA, Gunna JS (2010) Gallstones play a significant role in Salmonella spp. gallbladder colonization and carriage. PNAS 107(9):4353–4358
Doss CG, Rajith B, Garwasis N, Mathew PR, Raju AS, Apoorva K, William D, Sadhana NR, Himani T, Dike IP (2012) Screening of mutations affecting protein stability and dynamics of FGFR1—a simulation analysis. Appl Transl Genom 1:37–43
Enea GDD, Ilaria C, Martina P, Luigi T, Fabrizio E (2017) Biofilm producing Salmonella Typhi: chronic colonization and development of gallbladder cancer. Int J Mol Sci 18:1887
Fàbrega A, Vila J (2013) Salmonella enterica serovar Typhimurium skills to succeed in the host: virulence and regulation. Clin Microbiol Rev 26:308–341
Ferrieres L, Clarke DJ (2003) The RcsC sensor kinase is required for normal biofilm formation in Escherichia coli K-12 and controls the expression of a regulon in response to growth on a solid surface. Mol Microbiol 50:1665–1682
Francez-Charlot A, Laugel B, Van Gemert A, Dubarry N, Wiorowski F, Castanie-Cornet MP, Gutierrez C, Cam K (2003) RcsCDB His-Asp phosphorelay system negatively regulates the flhDC operon in Escherichia coli. Mol Microbiol 49:823–832
Freeman DJ, Falkiner FR, Keane CT (1989) New method for detecting slime production by coagulase negative staphylococci. J Clin Pathol 42:872–874
Gasteiger E, Gattiker A, Hoogland C, Ivanyi I, Appel RD, Bairoch A (2003) ExPASy: the proteomics server for in-depth protein knowledge and analysis. Nucleic Acids Res 31:3784–3788
Giraud E, Cloeckaert A, Kerboeuf D, Chaslus-Dancla E (2000) Evidence for active efflux as the primary mechanism of resistance to ciprofloxacin in Salmonella enterica serovar Typhimurium. Antimicrob Agents Chemother 44(5):1223–1228
Giraud E, Baucheron S, Cloeckaert A (2006) Resistance to fluoroquinolones in Salmonella: emerging mechanisms and resistance prevention strategies. Microbes Infect 8(7):1937–1944
Gonzalez-Escobedo G, Marshall JM, Gunn JS (2011) Chronic and acute infection of the gall bladder by Salmonella Typhi: understanding the carrier state. Nat Rev Microbiol 9(1):9–14
Goswami AM (2015) Structural modeling and in silico analysis of non-synonymous single nucleotide polymorphisms of human 3β-hydroxysteroid dehydrogenase type 2. Meta Gene 8:162–172
Gurunathan S, Han JW, Kwon DN, Kim JH (2014) Enhanced antibacterial and anti-biofilm activities of silver nanoparticles against Gram-negative and Gram-positive bacteria. Nanoscale Res Lett 31:373
Hall-Stoodley L, Hu FZ, Gieseke A, Nistico L, Nguyen D, Hayes J, Forbes M, Greenberg DP, Dice B, Burrows A, Wackym PA, Stoodley P, Post JC, Ehrlich GD, Kerschner E (2006) Direct detection of bacterial biofilms on the middle-ear mucosa of children with chronic otitis media. JAMA 296:202–211
Hasan TN, Grace BL, Masoodi TA, Shafi G, Alshatwi AA, Sivashanmugam P (2011) Affinity of estrogens for human progesterone receptor A and B monomers and risk of breast cancer: a comparative molecular modeling study. Adv Appl Bioinforma Chem 4:29–36
Ilakkia S, Senbagam D, Senthilkumar B, Sivakumar P (2015) A prevalence study of typhoid fever and convalescent phase asymptomatic typhoid carriers among the schoolchildren in the northern part of Tamil Nadu. J Public Health 23(6):373–378. https://doi.org/10.1007/s10389-015-0655-x
Ilakkia S, Senbagam D, Senthilkumar B (2016) Analysis of TLR polymorphisms in typhoid patients and asymptomatic typhoid carriers among the schoolchildren. Egy J Med Hum Genet 17:353–357. https://doi.org/10.1016/j.ejmhg.2015.12.2010
Jayaram B, Singh T, Mukherjee G, Mathur A, Shekhar S, Shekhar V (2012) Sanjeevini: a freely accessible web-server for target directed lead molecule discovery. BMC Bioinform 13:S7
Jayaraman A, Wood TK (2008) Bacterial quorum sensing: signals, circuits, and implications for biofilms and disease. Annu Rev Biomed Eng 10:145–167
Jefferys BR, Kelley LA, Sternberg MJ (2010) Protein folding requires crowd control in a simulated cell. J Mol Biol 397:1329–1338
Kelley LA, Sternberg MJ (2009) Protein structure prediction on the Web: a case study using the Phyre server. Nat Protoc 4(3):363–371
Kelley LA, Mezulis S, Yates CM, Wass MN, Sternberg MJ (2015) The Phyre2 web portal for protein modeling, prediction and analysis. Nat Protoc 10(6):845–858
Kim S, Thiessen PA, Bolton EE, Chen J, Fu G, Gindulyte A, Han L, He J, He S, Shoemaker BA, Wang J, Yu B, Zhang J, Bryant SH (2016) PubChem substance and compound databases. Nucleic Acids Res 44(D1):D1202–D1213
Laskowski RA, MacArthur MW, Moss DS, Thornton JM (1993) PROCHECK-a program to check the stereo chemical quality of protein structures. J Appl Crystallogr 26:283–291
Lipinski CA (2004) Lead- and drug-like compounds: the rule-of-five revolution. Drug Discov Today Technol 1(4):337–341
Marshall JM, Flechtner AD, La Perle KM, Gunn JS (2014) Visualization of extracellular matrix components within sectioned Salmonella biofilms on the surface of human gallstones. PLoS One 9:e9–e89243
McLean RJC, Pierson LS, Fuqua C (2004) A simple screening protocol for the identification of quorum signal antagonists. J Microbiol Methods 58:351–360
Miro E, Verges C, Garcia I, Mirelis B, Navarro F, Coll P, Prats G, Martinez-Martinez L (2004) Resistance to quinolones and beta-lactams in Salmonella enterica due to mutations in topoisomerase-encoding genes, altered cell permeability and expression of an active efflux system. Enferm Infect Microbiol Clin 22(4):204–211
Mitrophanov AY, Groisman EA (2008) Signal integration in bacterial two-component regulatory systems. Genes Dev 22:2601–2611
Monds RD, O’Toole GA (2009) The developmental model of microbial biofilms: ten years of a paradigm up for review. Trends Microbiol 17:73–87
Monroe D (2007) Looking for chinks in the armor of bacterial biofilms. PLoS Biol 5(11):e307
Morris GM, Huey R, Lindstrom W, Sanner MF, Belew RK, Goodsell DS, Olson AJ (2009) Autodock4 and AutoDockTools4: automated docking with selective receptor flexiblity. J Comput Chem 16:2785–2791
Murugan K, Selvanayaki K, Kalyanasundaram VB, Saleh A-S (2013) Nanotechnological approach for exploring the antibiofilm a potential of an ethanomedicinal herb Andrographis paniculata for controlling lung infection causing Pseudomonas aeruginosa. Dig J Nanomat Biostru 8(1):117–126
Murugan K, Senthilkumar B, Senbagam D, Saleh Al S (2014) Biosynthesis of silver nanoparticles using Acacia leucophloea extract and their antibacterial activity. Int J Nanomedicine 9(1):2431–2438
Oropeza R, Salgado-Bravo R, Calva E (2015) Deletion analysis of RcsC reveals a novel signalling pathway controlling poly-N-acetyl glucosamine synthesis and biofilm formation in Escherichia coli. Microbiol 161:903–913
Parashar V, Mirouze N, Dubnau DA, Neiditch MB (2011) Structural basis of response regulator dephosphorylation by Rap phosphatases. PLoS Biol 9:e1000589
Perrett CA, Karavolos MH, Humphrey S, Mastroeni P, Martinez-Argudo I, Spencer H, Bulmer D, Winzer K, McGhie E, Koronakis V, Williams P, Khan CM, Jepson MA (2009) LuxS-based quorum sensing does not affect the ability of Salmonella enterica serovar Typhimurium to express the SPI-1 type 3 secretion system, induce membrane ruffles, or invade epithelial cells. J Bacteriol 191(23):7253–7259
Powers CN, Setzer WN (2015) A molecular docking study of phytochemical estrogen mimics from dietary herbal supplements. In Silico Pharmoclo 3:4. https://doi.org/10.1186/s40203-015-0008-z
Prouty AM, Schwesinger WH, Gunn JS (2002) Biofilm formation and interaction with the surfaces of gallstones by Salmonella spp. Infect Immun 70:2640–2649
Prüß BM (2017) Involvement of two-component signaling on bacterial motility and biofilm development. J Bacteriol 199:e00259–e00217
Pundir S, Martin MJ, O’Donovan C (2017) UniProt protein knowledgebase. Methods Mol Biol 1558:41–55
Reeve KE (2010) Salmonella binding to and biofilm formation on cholesterol/gallstone surfaces in the chronic carrier state. Undergraduate Honors Thesis. School of Allied Medical Professions, The Ohio State University
Remmert M, Biegert A, Hauser A, Soding J (2012) HHblits: lightning-fast iterative protein sequence searching by HMM-HMM alignment. Nat Methods 9:173–175
Sasikumar C, Kanan V, Senthilkumar B (2005) Asymptomatic typhoid carriers in Namakkal District Tamil Nadu. J Environ Biol 26:113–115
Sauer RT, Yocum RR, Doolittle RF, Lewis M, Pabo CO (1982) Homology among DNA-binding proteins suggests use of a conserved super-secondary structure. Nature 298(5873):447–451
Seghal KG, Sabarathnam B, Selvin J (2010) Biofilm disruption potential of a glycolipid biosurfactant from marine Brevibacterium casei. FEMS Immunol Med Microbiol 59(3):432–438
Senthilkumar B, Prabakaran G (2005) Multidrug resistant Salmonella typhi in asymptomatic typhoid carriers among food handlers in Namakkal District Tamil Nadu. Indian J Med Microbiol 23(2):92–94
Senthilkumar B, Venkatachiranjeevi P, Rajasekarapandian M (2011) Antibacterial effects of viable antibiotics and pomegranate (Punica granatum) bark extracts on Salmonella typhi and Salmonella paratyphi A isolates from asymptomatic typhoid carriers in Ongole, Andhrapradesh, India. J Pure Appl Microbiol 5(1):337–342
Senthilkumar B, Anbarasu K, Senbagam D, Rajasekarapandian M (2014a) Induction of deletion mutation on ompR gene of Salmonella enterica serovar Typhi isolates from asymptomatic typhoid carriers to evolve attenuated strains for vaccine development. Asian Pac J Trop Med 7(12):933–939
Senthilkumar B, Senbagam D, Rajasekarapandian M (2014b) An epidemiological surveillance of asymptomatic typhoid carriers associated in respect to socioeconomic status in India. J Public Health 22:297–301. https://doi.org/10.1007/s10389-012-0545-4
Senthilkumar B, Sivakumar P, Madhanraj R, Senbagam D, Ilakkia S (2014c) A comparative analysis of TLR5 polymorphism and clinical parameters in typhoid patients and asymptomatic typhoid carriers. J Public Health 22:131–137. https://doi.org/10.1007/s10389-013-0604-5
Senthilkumar B, Ilakkia S, Selvam K, Senbagam D, Nachimuthu S, Guruswami G (2017) Biosynthesis of silver nanoparticles using Myristica fragrans seed (nutmeg) extract and its antibacterial activity against multidrug resistant (MDR) Salmonella enterica serovar Typhi isolates. Environ Sci Pollut Res 24:14758–14769. https://doi.org/10.1007/s11356-017-9065-7
Senthilkumar B, Senbagam D, Prahalathan C, Anbarasu K (2019) Gateways of pathogenic bacterial entry into host cells-Salmonella. In: Balamurugan K, Prithika U (eds) Pocket guide to biomedical Sciences,1st edn. CRC Press, Taylor & Francis Group, USA, pp 59–78. https://doi.org/10.1201/b22196
Sereno MJ, Ziech RE, Druziani JT, Pereira JG, Bersot LS (2017) Antimicrobial susceptibility and biofilm production by Salmonella sp. strains isolated from frozen poultry carcasses. Braz J Poultry Sci. https://doi.org/10.1590/1806-9061-2016-0268
Sharma IM, Petchiappan A, Chatterji D (2014) Quorum sensing and biofilm formation in mycobacteria: role of c-di-GMP and methods to study this second messenger. IUBMB Life 66(2014):823e834
Sharma VK, Bayles DO, Alt DP, Looft T, Brunelle BW, Stasko JA (2017) Disruption of rcsB by a duplicated sequence in a curli-producing Escherichia coli O157:H7 results in differential gene expression in relation to biofilm formation, stress responses and metabolism. BMC Microbiol 17:56
Shi X, Zhu X (2009) Biofilm formation and food safety in food industries. Trends Food Sci Technol 20:407–413. https://doi.org/10.1016/j.tifs.2009.01.054
Stewart PS, Costerton JW (2001) Antibiotic resistance of bacteria in biofilms. Lancet 358(9276):135–138
Stock AM, Robinson VL, Goudrea PN (2000) Two-component signal transduction. Annu Rev Biochem 69:183–215
Swidsinski A, Lee SP (2001) The role of bacteria in gallstone pathogenesis. Front Biosci 6:E93–E103
Thomason P, Kay R (2000) Eukaryotic signal transduction via histidine-aspartate phosphorelay. J Cell Sci 113:3141–3150
Valentini M, Filloux A (2016) Biofilms and cyclic di-GMP (c-di-GMP) signaling: lessons from Pseudomonas aeruginosa and other bacteria. J Biol Chem 291:12547–12555
van Gunsteren WF, Billeter SR, Eising AA, Hünenberger PH, Krüger P, Mark AE, Scott WRP, Tironi IG (1996) Biomolecular simulation: the GROMOS96 manual and user guide. Verlagder Fachvereine Hochschulverlag AG an der ETH Zürich: 1–1042
Veldman A, Vahl HA, Borggreve GJ, Fuller DC (1995) A survey of the incidence of Salmonella species and Enterobacteriaceae in poultry feeds and feed components. Vet Rec 136(7):169–172
Wei GX, Campagna AN, Bobek LA (2006) Effect of MUC7 peptides on the growth of bacteria and on Streptococcus mutans biofilm. J Antimicrob Chemother 57:1100–1109
Wiederstein M, Sippl MJ (2007) ProSA-web: interactive web service for the recognition of errors in three-dimensional structures of proteins. Nucleic Acids Res 35(Web Server issue):W407–W410. https://doi.org/10.1093/nar/gkm290
Xu D, Zhang Y (2011) Improving the physical realism and structural accuracy of protein models by a two-step atomic-level energy minimization. Biophys J 101:2525–2534
Yadav RP, Ibrahim KS, Gurusubramanian G, Kumar NS (2015) In silico docking studies of non-azadirachtinlimonoids against ecdysone receptor of Helicover paarmigera (Hubner) (Lepidoptera: Noctuidae). Med Chem Res 24(6):2621–2631
Yuriev E, Ramsland PA (2013) Latest developments in molecular docking: 2010–2011 in review. J Mol Recognit 26:215–239
Yuriev E, Agostino M, Ramsland PA (2011) Challenges and advances in computational docking: 2009 in review. J Mol Recognit 24:149–164
Zschiedrich CP, Keidel V, Szurmant H (2016) Molecular mechanisms of two-component signal transduction. J Mol Biol 428:3752–3775
Acknowledgments
SB is thankful to the Department of Biotechnology (DBT), New Delhi, Govt. of India for the financial assistance in the form of Advanced State Biotech Hub (BT/04/NE/2009) to Mizoram University. KSI acknowledges the Department of Science and Technology – Science and Engineering Research Board (DST-SERB) for the financial assistance through Young Scientist award (YSS/2014/000657).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interests
The authors declare that they have no competing interests.
Ethics approval and consent to participate
Not applicable.
Additional information
Responsible editor: Philippe Garrigues
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Balakrishnan, S., Ibrahim, K.S., Duraisamy, S. et al. Antiquorum sensing and antibiofilm potential of biosynthesized silver nanoparticles of Myristica fragrans seed extract against MDR Salmonella enterica serovar Typhi isolates from asymptomatic typhoid carriers and typhoid patients. Environ Sci Pollut Res 27, 2844–2856 (2020). https://doi.org/10.1007/s11356-019-07169-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-019-07169-5