Abstract
Microbial communities are more effective in degrading natural polymers and xenobiotics than pure cultures. Biodegradation of polyacrylic and polyurethane polymers by bacterial and fungal strains has been addressed, but limited information about their biodegradation by microbial communities exists. The aim of this work was to evaluate the ability of three enriched microbial communities (BP1h, BP3h, and BP7h), selected from deteriorated foam pieces collected in a landfill, to biodegrade the polyacrylic component of the 2K-PU coating Bayhydrol® A2470 and the polyester polyurethane coating NeoRez™ R-9637. Two communities were further selected to quantify extracellular esterase, protease, and urease activities, to identify their taxonomic composition, and to analyze the ability of their isolated members to grow in those polymers. The growth of the three communities was larger in polyester polyurethane than in polyacrylic and their biodegradative activities affected ester, urethane, ether, aromatic, and aliphatic groups of the compounds present in the coatings. From all the communities growing in polyacrylic or in polyester polyurethane, two and five different types of colonies were isolated, respectively. In polyacrylic, extracellular esterase and protease activities were at their maximum level at 7 days of culture, whereas in polyester polyurethane, protease and urease were greatest at 21 days. All the isolated community members were identified as xenobiotics degraders. The complete communities grew better in media with the polymers than the isolated members. This is one of the few studies reporting biodegradation of synthetic polymers by microbial communities and serves as basis for developing synthetic consortia with enhanced degradative abilities.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
In the environment, microorganisms interact by establishing microbial communities with regulated metabolic interdependencies to obtain carbon and energy from complex natural environments and from polluted sites (Vogt and Richnow 2014). Interesting examples include microbial communities able to degrade lignin, a complex natural heteropolymer of hydroxylated and methoxylated phenylpropanoid units, that represents a significant input of organic compounds into soil (Kato et al. 2015); a bacterial consortium, isolated from soil at a fruit-packaging plant, able to degrade thiabendazole, a persistent fungicide used for fruit post-harvest treatment (Perruchon et al. 2017); and salt-tolerant microbial consortia, isolated from an oil field, able to aerobically degrade n-alkanes and polycyclic aromatic hydrocarbons from heavy crude oil (Gurav et al. 2017).
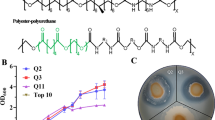
Despite the progress made in studying the role that microbial communities play in the degradation of natural polymers and in the bioremediation of xenobiotic compounds, little is known about microbial communities able to degrade synthetic polymers. Polyurethanes (PU) are synthetic polymers widely used because of its high durability, resistance, and versatility of forms and textures, originated from their chemical composition. PU are synthesized from polyisocyanates and polyols generating the characteristic carbamate group. Different types of polyols are used for PU synthesis, i.e., polyester (PS), polyether (PE), and polyacrylic (PA) polyol, recently included in PU coating formulations. Because of its diverse chemical composition, PU recycling is troublesome, so it is usually incinerated for heat production or disposed in landfills, generating severe pollution problems. PA polyols are frequently used in two components (2K) PU coatings for automotive finish with high chemical resistance and durability. They are obtained from the copolymerization of conventional acrylic monomers, such as ethyl acrylates, butyl acrylates, acrylic acid, methyl methacrylate, or styrene with hydroxylated acrylic monomers such as 2-hydroxyethyl acrylates or 4-hydroxybutyl acrylates, generating complex and highly stable molecules with vinyl functional groups. Acrylic derivatives, such as water-soluble PA acid and polyacrylamide, are broadly used as thickeners, drilling muds, dispersing or flocculation agents, and as soil amendments for improved water management (Mai et al. 2004). Cross-linked polymers of acrylic acid and acrylamide co-monomers are effective superabsorbents largely used in hygiene articles such as sanitary napkins and diapers. The high stability of the carbon-carbon backbone of polyacrylate polymers, their insolubility, and their high molecular weight make their biodegradation challenging (Cameron et al. 2000).
Microbial biodegradation of polyester polyurethane (PS-PU) by pure strains of bacteria and fungi has been extensively documented (reviewed in Howard 2012; Mahajan and Gupta 2015; Wilkes and Aristilde 2017), whereas PA and acrylate oligomers biodegradation have been scarcely analyzed (Cameron et al. 2000; Hayashi et al. 1993; Kawai 1993; Mai et al. 2004; Oprea and Doroftei 2011). On the contrary, at microbial communities’ level, PS-PU biodegradation has been barely investigated. The bacterial strains Bacillus sp. AF8, Pseudomonas sp. AF9, Micrococcus sp. AF10, Arthrobacter sp. AF11, and Corynebacterium sp. AF12, isolated from the surface of PU films after 6 months of soil burial, were used to establish a synthetic consortium with the ability to degrade PU films (Shah et al. 2008). Likewise, by evaluating PU degradation by FTIR and scanning electron microscopy and selecting the best PU-degrading strains based on their extracellular esterase activity, a consortium made of Bacillus subtilis MZA-75 and Pseudomonas aeruginosa MZA-85 was able to utilize PS-PU films more efficiently than individual strains (Shah et al. 2016). Fungal communities involved in biodegradation of PS-PU buried in soil and compost have also been reported (Cosgrove et al. 2007; Zafar et al. 2013). Nevertheless, there are even fewer studies about PA biodegradation by microbial communities. Mixed populations of activated sludge microorganisms, previously exposed to mixtures of low-molecular-weight (Mw < 8000) polyacrylic acids, could extensively metabolize acrylic acid oligomers of up to seven units, although complete mineralization only occurred with monomers and dimers (Larson et al. 1997). The biodegradability of a cross-linked PA acid was characterized in four agricultural microbial-containing soils, measured as the 13CO2 efflux from 13C-labeled PA acid, under different conditions, showing that mineralization occurred at very low rates (0.12–0.24% every 6 months) (Wilske et al. 2014), but no information about the microbial composition of these communities was provided.
In this article, we examine the ability to biodegrade PA and PS-PU of three enriched microbial communities (BP1h, BP3h, BP7h), selected in a mineral medium with a PS-PU coating as the only carbon source, and inoculated with pieces of deteriorated PU foams collected in a municipal landfill. The biodegradative action of these communities on the PA component of a 2K PU coating (Bayhydrol®) and on the PS-PU coating (NeoRez™) is demonstrated by FTIR. In two specific communities, we also measured the extracellular enzymatic activities related to polymer biodegradation and analyzed the bacterial biological diversity. Moreover, we present evidences proving that synergistic interactions between members of the communities exist. This work reports microbial communities that can be used for PA and PS-PU waste treatment, and lays the foundation for the assembly of efficient synthetic consortia for the degradation of these types of materials.
Materials and methods
Culture media
For culture enrichment, we used a mineral medium (MM) (in mmol l-1: KH2PO4, 14.7; K2HPO4, 40.2; NH4NO3, 12.5; MgSO4·7H2O, 0.4; ZnSO4·7H2O, 0.0035; CuSO4·7H2O, 0.0004; FeSO4·7H2O, 0.036; MnSO4·6H2O, 0.0077; the final pH was 7.2) (Nakajima-Kambe et al. 1995) with the PS-PU coating Hydroform® (Polyform de México, SA de CV) at 0.3% solids (v/v) as the sole carbon source (MM-Hydroform) (Oceguera-Cervantes et al. 2007). Since Hydroform® is no longer commercially available, characterization of PU degradation was carried out in MM with Bayhydrol® A2470 (MM-Bayhydrol) or with NeoRez™ R-9637 (MM-NeoRez) at 0.3% or 3% solids as the sole carbon source. Bayhydrol® A2470 (Bayer, México) is a hydroxyfunctional polyacrylic (PA) dispersion that is used in combination with aliphatic polyisocyanates as a component in the formulation of waterborne 2K-PU coatings. It is supplied approximately at 45% of solids in water:solvent naphtha® 100 (aromatic hydrocarbons with carbon numbers in the range of C9 through C16):Dowanol® PnB (propylene glycol n-butyl ether (PGBE)):neutralized with dimethylethanolamine (DMEA)/triethanolamine (TEA), in the approximate ratio of 44.3:4:4:2.7. NeoRez™ R-9637 (DSM NeoResins, Inc. México) is a waterborne aliphatic PS-PU dispersion, with 35–37% of solids, which contains 8.4% N-methylpyrrolidone (NMP) and 2% TEA. Luria Bertani (LB) medium was used for isolation of the individual community members. All the chemical reagents used for culture media preparation were RA grade, from J.T. Baker® Chemical. Co.
Isolation of enriched microbial communities able to grow in a polyester polyurethane coating as the only carbon source
Erlenmeyer flasks (125 ml) with 25 ml MM-Hydroform® (0.3% solids) were inoculated with approximately 1 cm3 of different pieces of foams collected at El Bordo Poniente (BP) landfill at Nezahualcóyotl, Estado de México, México. The flasks were incubated at 30 °C and 220 rpm for 7 days. After that time, 2 ml of each culture were inoculated into fresh 25 ml MM-Hydroform and were incubated for 1 week, re-inoculated in the same fresh media and incubated for the same period two more times, giving a total of 28 days of culture in MM-Hydroform. Samples of these communities were preserved in MM-Hydroform/30% glycerol at − 70 °C. One glycerol tube was used to start every culture for the subsequent experiments or to prepare more glycerols following the procedure described before. The original enriched communities were cultivated in MM-Bayhydrol and MM-NeoRez (0.3% in every case) for 2 days. After that time, the communities were maintained in the correspondent media with 30% glycerol at − 70 °C.
Quantification of microbial growth
Because the microbial communities grow forming floccules in the media with coatings as carbon source, growth was quantified by biomass formation, measuring dry weight, and also by measuring the carbon dioxide released by the oxidation of the chemical components present in the coatings as reported previously (Crossno et al. 1996). To this end, pre-cultures of each community were started by inoculating the content of one glycerol tube (0.5 ml) into an Erlenmeyer flask containing 25 ml of MM-Bayhydrol or MM-NeoRez (0.3%) and were incubated overnight at 37 °C and 200 rpm. After that period, cultures were centrifuged at 11,952 g (Rotor SS34 in a Sorvall RC5-C) and 4 °C, for 15 min, and the pellets were washed with 1 ml 50 mmol l-1 phosphate buffer pH 7.2 and weighted. Next, the pellets were disaggregated, resuspended in 1 ml of phosphate buffer, and inoculated in 160 ml of MM-Bayhydrol or MM-NeoRez (3%) at 0.5 mg biomass ml-1. Aliquots (25 ml) of these cell suspensions were distributed into six 125 ml Erlenmeyer flasks to start growth quantification experiments. One flask was harvested every 24 h, centrifuged, washed, and the wet pellets were placed in an oven at 60 °C with vacuum (10 lb. pressure) for 48 h, afterwards the dry weight was measured. These measurements were performed in triplicate. For the measurements of CO2 release, the Erlenmeyer flasks with cell cultures were connected to 14 × 100 mm test tubes containing 5 ml of NaOH (0.5 mol l-1), to capture the CO2 released from microbial respiration, and were incubated at 37 °C and 200 rpm. Every 24 h, a flask and the connected test tube were harvested and placed at 4 °C for 1 h. After that, the NaOH solution of the test tube was titrated with 0.119 mol l-1 HCl, up to pH values of 10.32 and 6.35, the two pKa values of Na2CO3, using a pH meter (Beckman Coulter PHI510). The volumes of HCl needed to titrate Na2CO3 were used to calculate the amounts of CO2 released by the culture, according to the equation cited in Crossno et al. (1996). CO2 values fall within the linear interval of a calibration curve constructed with different amounts of Na2CO3 (from 25 to 250 mg). Similar measurements were performed from non-inoculated cultures and were subtracted from values of the inoculated cultures. Three replicates were performed for each treatment. The growth of the isolated members from the communities BP1h and BP7h cultured in MM-Bayhydrol and MM-NeoRez (3%), respectively, was also measured by CO2 release, in pure cultures as described for complete communities.
Fourier-transform infrared spectroscopy analysis
Each community was cultured in both MM-Bayhydrol and MM-NeoRez (3%) for 21 days at 37 °C and 200 rpm. Cells were pelleted and 0.5 ml aliquots of the supernatant were drawn off, placed in Eppendorf tubes and allowed to evaporate to dryness at 37 °C. Supernatants from uninoculated media incubated in the same conditions and prepared similarly, served as negative controls. FTIR analysis was performed in a 1605 Perkin Elmer spectrometer from 800 to 4000 cm-1. Three biological replicates for each experimental condition were analyzed. Identification of the functional groups in the FTIR spectra for PA, PS-PU, and other chemical components present in the coatings was performed according to McCarthy et al. (1997) and Pardini and Amalvy (2008).
Enzymatic assays
Spectrophotometric quantifications for esterase, protease, and urease activities were performed in the cell-free supernatants of cultures at 0, 5, 12, and 21 days (three replicates for each time), from two selected communities grown in 25 ml MM-Bayhydrol or MM-NeoRez (3%), incubated at 37 °C and 200 rpm. For esterase activity measurement, cell-free supernatants obtained after centrifugation at 11,953 g (Rotor SS34 in a Sorvall RC5-C plus) for 15 min at 4 °C were directly used. For measurements of protease and urease activities, remains of coatings were eliminated from the cell-free supernatant by precipitation with 30% and 10% ammonium sulfate for Bayhydrol and NeoRez, respectively, and subsequently removed by centrifugation as indicated above. Supernatants were dialyzed against 50 mmol l-1 potassium phosphate (pH 7.0) and concentrated by ultrafiltration in Amicon Ultra 10K devices. These samples served as the source of extracellular enzymatic activities in all the cases. Protein concentration was determined according to Bradford’s method (Bradford 1976) by using BSA as standard. Protein aliquots (10 μg) were used for in vitro enzymatic assays. Esterase activity was determined spectrophotometrically at 405 nm by hydrolysis of p-nitrophenyl acetate (p-NPA) (Alfa Aesar, Cat. Num. L00314). The reaction mixture contained 50 mmol l-1 potassium phosphate (pH 7.0), enzyme extract, and 5 mmol l-1 p-NPA in a final volume of 1 ml (Desphande et al. 1984). Lipase from Pseudomonas fluorescens (Sigma Aldrich, Cat. Num. 28602) was used as positive control. Protease activity was determined spectrophotometrically by casein hydrolysis at 280 nm, as previously reported (Lovrien and Matulis 1995), except that the reaction mixture contained 100 mmol l-1 potassium phosphate (pH 7.0), enzyme extract, and 0.5% casein in a final volume of 1 ml and was incubated for 1 h. Proteinase K (Thermo-Scientific. Cat. Num. EO0491) was used as positive control. Urease activity was determined spectrophotometrically by using a phenol hypochlorite assay, quantifying ammonia release at 636 nm. The reaction mixture contained 15 mmol l-1 potassium phosphate (pH 7.0), enzyme extract, and 3.8 mmol l-1 urea in a final volume of 1.3 ml and was incubated for 1 h (Witte and Medina-Escobar 2001). Jack bean urease (Sigma Aldrich, Cat. Num. U-1500) was used as positive control. Negative controls were also performed for all the enzymatic analysis by setting enzymatic assays with mock protein extracts obtained from non-inoculated cultures and assays with reaction mixtures containing protein extracts without substrates.
Isolation and identification of members from selected communities
Aliquots of liquid cultures from BP1h, BP3h, and BP7h grown in MM-Bayhydrol and in MM-NeoRez for 48 h were diluted at 1:200, 1:5000, and 1:10,000 in phosphate buffer; 100 μL from each dilution were spread over solid plates containing MM-Bayhydrol and MM-NeoRez correspondingly, and incubated at 37 °C for 48 h. All the diverse colonies were picked, cultured in liquid LB medium at 37 °C for 24 h, and streaked onto solid LB. This procedure was repeated until pure cultures were obtained. Morphological characterization of the isolated colonies grown for 48 h in LB was carried out under a stereomicroscope (DMW143 Triple Output Zoom). Isolated colonies from each culture were Gram stained and analyzed under a microscope (Motic, Model Ba210 Digital) at × 100. The identification of the isolated bacterial species was carried out by PCR amplification of the 16S rDNA V3 region with the forward primer 341F: 5′-GCCTACGGGAGGCAGCAG-3′ and the reverse primer 518R: 5′-ATTACCGCGGCTGCTGG-3′ (Muyzer et al. 1993), using 200 ng of genomic DNA as template. DNA was extracted from cultures with an OD660 nm of 0.6 at (approximately 3 h) in LB liquid medium, incubated at 37 °C and 200 rpm, according to Ausubel et al. (1997), with some modifications (Solís-González et al. 2018). All the reagents were molecular biology grade from Sigma-Aldrich. Final concentrations in the PCR mix were as follows: 1× Taq buffer, 2.5 mmol l-1 MgCl2, primers 0.4 μmol l-1 each, 250 μmol l-1 dNTPs each, and 1.25 U Taq DNA polymerase (Thermo-Scientific). PCR program was initial denaturation 94 °C/5 min, one cycle; denaturation 94 °C/30 s, annealing 60 °C/30 s, extension 72 °C/30 s, 35 cycles; final extension 72 °C/5 min, one cycle, run in a Veriti thermal cycler Model # 9902 (Applied Biosystem). PCRs were resolved in agarose gels and the products were gel-purified using the GeneJET PCR purification kit (Thermo-Scientific). Sequencing was performed with the same primers used for the PCR amplifications at the Unidad de Síntesis y Secuenciación de ADN (Instituto de Biotecnología, UNAM). Identification was made by sequence comparison to the GenBank database using basic local alignment search tool (BLAST) (Altschul et al. 1990).
Accession numbers
The BP1h and BP7h communities and one representative member of the different types of isolated bacteria were deposited in the Culture Collection at Cepario Facultad de Química, UNAM, World Data Centre for Microorganisms CFQ100, under the accession numbers: BP1h community grown in Bayhydrol, CFQ-B-261; BP7h community grown in NeoRez, CFQ-B-264; BPh1B1, CFQ-B-262; BPh1B4, CFQ-B-263; BP7hN1, CFQ-B-265; BP7hN2, CFQ-B-266; BP7hN5, CFQ-B-267; BP7hN7, CFQ-B-268; BP7hN12, CFQ-B-269. The nucleotide sequences of the 16S rRNAs V3 regions from the isolated members of the BP1h and BP7h communities were deposited in the DNA Data Bank of Japan and have the following accession numbers: BP1hB1, LC425052; BP1hB2, LC425053; BP1hB3, LC425054; BP1hB4, LC425055; BP7hN1, LC425056; BP7hN2, LC425060; BP7hN3, LC425058; BP7hN4, LC425059; BP7hN5, LC425057; BP7hN6, LC425061; BP7hN7, LC425062; BP7hN8, LC425063; BP7hN9, LC425064; BP7hN10, LC425065; BP7hN11, LC425066; BP7hN12, LC425067.
Results
Growth of the enriched microbial communities in polyacrylic and polyester polyurethane coatings as the sole carbon source
Eight communities able to grow in MM-Hydroform (a PS-PU coating) were isolated from eight different PU foam pieces collected at El Bordo Poniente landfill. A preliminary screening identified BP1h, BP3h, and BP7h as the best-grown communities in this medium (data not shown). The ability of these communities to utilize Bayhydrol® A2470, the PA component of a 2K-PU coating, and NeoRez™ R-9637, a PS-PU coating, as the only carbon sources to sustain their growth was quantified by dry weight and by CO2 release. Except for the growth in Bayhydrol measured as dry weight, which displayed inconsistent differences, the three communities showed similar growth in each media during the 5 days of cultivation, by the two techniques used. However, although the differences in biomass are slightly higher in NeoRez than in Bayhydrol, CO2 release was much higher in the former 2.6-fold at 48 h and 3-fold at 120 h, indicating a higher metabolic activity of the communities cultured in NeoRez (Fig. 1).
Growth of microbial communities in Bayhydrol (PA) and NeoRez (PS-PU) coatings as sole carbon source. Growth was estimated by biomass production measured by a, b dry weight and c, d CO2 release in communities BP1h (☐), BP3h (○), and BP7h (△) cultured in a, c MM-Bayhydrol 3% and b, d MM-NeoRez 3% at 37 °C and 200 rpm. n = 3. Bars represent standard deviations
Biodegradation of polyacrylic and polyester polyurethane coatings
FTIR spectra of Bayhydrol and NeoRez cell-free media supernatants, each of them cultured with the three communities, were compared to the spectra of supernatants from non-inoculated media, incubated at the same conditions. For each coating, FTIR spectra generated by the three communities were very similar (Fig. 2). The changes in Bayhydrol were an increase in the 3264 cm-1 signal that corresponds to the OH group and in the 1724 cm-1 signal associated with the C=O group from the hydroxyfunctional PA; a strong decrease in the 1574 cm-1 signal of amine C–N stretch from DMEA and TEA; considerable increments in the signals at 1452 and 1146 cm-1, related to aliphatic CH2 bending and CH2 twisting, respectively, from the aliphatic carbons of PA, PGBE, DMEA, TEA, and naphtha components, and in the 1388 cm-1 signal of the C–C stretch from the benzene ring of the aromatic carbons present in naphtha components; and considerable decrements at the 1084, 1037, and 980 cm-1 signals that correspond to the aliphatic symmetric C–O–C stretch from PGBE and PA (Fig. 2a). The FTIR spectra analysis generated by the biodegradative activity of the three communities on the PS-PU NeoRez showed an increment in the 3314 cm-1 signal that correspond to N–H stretch; an overlapping of the 1727 cm-1 and 1672 cm-1 signals corresponding to the C=O stretch from the urethane and the polyester polyols, probably as a result of ester polyols degradation; a slight increase in the 1524 cm-1, a decrease in the 1226 cm-1 and an increase in the 1040 cm-1 signals that correspond to N–H bending plus C–N stretch, C–N stretching, and C–O–C stretch, all from the urethane group respectively; and a slight decrease in the 1450 cm-1 that correspond to aliphatic CH2 bending (Fig. 2b).
Microbial diversity and identification of members from selected communities grown in polyacrylic and polyester polyurethane coatings
The morphological diversity of microorganisms present in the communities grown in MM-Bayhydrol or in MM-NeoRez was analyzed by spreading diluted aliquots onto plates of the same solid media, correspondingly. Colonies with different morphologies grew, but when they were re-streaked in the same solid media with the coatings as carbon sources in order to obtain pure cultures, many colonies did not grow. Thus, colonies’ isolation was performed on LB plates, where colonies’ growth was evident. Re-streaking on LB plates was repeated until pure cultures were obtained. All the colonies obtained from the initial plates with the coatings as carbon source grew on LB plates. From each of the three enriched microbial communities grown in Bayhydrol, only one type of colony morphology, with two different sizes, were observed, thus we isolated two representative colonies from each size; colonies B1 and B2 were slightly larger (5 mm) than colonies B3 and B4 (2 mm) (Table 1). A representative photograph of the colony morphology (BP1) observed in BP1h grown in Bayhydrol is shown (Fig. 3a). In contrast, from each of the three communities grown in NeoRez, five different types of colonies were observed. We selected 12 colonies (N1-N12) representing the five morphologies. Description of colonies’ morphology and Gram stain properties is presented in Table 1. Representative photographs of colonies from communities grown in NeoRez are shown (Fig. 3b-f).
Morphological diversity of bacterial colonies isolated from BP1h grown in Bayhydrol and BP7h grown in NeoRez. Diluted aliquots of liquid cultures of BP1h grown in Bayhydrol and BP7h grown in NeoRez were spread onto agar plates with the correspondent media. Colonies were picked and repeatedly streaked on LB plates up to obtaining pure cultures. Four (B1-B4) and 12 (N1-N12) isolates were obtained from BP1h and BP7h respectively, cultured on LB plates for 48 h at 37 °C and photographed. Bacterial strains were identified by sequencing 16S rDNA V3 region. Representative photographs show the isolates obtained from BP1h aAcinetobacter sp. and BP7h bAcinetobacter sp., cParacoccus sp., dBacillus sp., eHydrogenophaga sp., and fMicrobacterium sp. Macroscopic images were obtained at ×5 magnification
As the three communities grew similarly in each coating, they affected each polymer in similar way, and presented the same type of colonies’ morphology, we selected BP1h grown in Bayhydrol and BP7h grown in NeoRez to identify their members by 16S rDNA V3 region sequencing. All the identified members showed 98–100% identity to genera previously reported as xenobiotic degraders. The four members of BP1h grown in Bayhydrol (B1-B4) were identified as Acinetobacter sp. (Fig. 3a). In BP7h grown in NeoRez, N1, N4, N8, and N9 were identified as members of the genus Acinetobacter (Fig. 3b); N2, N3, N6, and N11 belonged to the genus Paracoccus (Fig. 3c); N5 and N10 were Bacillus sp. (Fig. 3d); N7 was Hydrogenophaga sp. (Fig. 3e); and N12 was identified as Microbacterium sp. (Fig. 3f). These results show a clear correlation between the types of colony morphology presented by the different members isolated from the communities and their specific identity at genera level.
Extracellular enzymatic activities related to biodegradation of coatings
Extracellular enzymatic activities associated with hydrolysis of the chemical components of the coatings (esterase, protease, and urease) were measured in cell-free supernatants of BP1h grown in MM-Bayhydrol and in BP7h grown in MM-Neorez, at 0, 5, 12, and 21 days. Esterase was the most prominent activity by far in both communities. In BP1h, esterase and protease activities were higher at the beginning of the culture (13 mmol h-1 mg protein-1 and 8.5 μmol h-1 mg protein-1, respectively), afterwards, the activities decreased but rose slightly at the end of the cultivation period (Fig. 4a, b). Urease activity was very low at the beginning and showed a discrete increase to 2 μmol h-1 mg protein-1 at the end of the analysis (Fig. 4c). In BP7h, extracellular esterase, protease, and urease activities increased throughout the culture period, reaching maximal values of 2.7 mmol h-1 mg protein-1, 9 μmol h-1 mg protein-1 and 6.7 μmol h-1 mg protein-1, respectively, at the end of the analysis (Fig. 4a–c).
Extracellular enzymatic activities of selected communities grown in Bayhydrol and NeoRez. a Esterase, b protease, and c urease activities from cell-free supernatants of BP1h cultured in MM-Bayhydrol (3%) (white columns), and BP7h cultured in MM-NeoRez (3%) (gray columns) were measured at 0 (no activity detected), 5, 12, and 21 days. n = 3. Bars represent standard deviation
Ability of the members from selected communities to grow as pure cultures in polyacrylic and polyester polyurethane coatings
The growth of members isolated from BP1h grown in MM-Bayhydrol and BP7h grown in MM-Neorez was analyzed at 72 and 120 h of culture, and the results were compared to the growth of the complete community. The growth of each of the four members isolated from BP1h was around 65% of the growth observed in the complete community: 230 μg CO2 ml-1 vs 368 μg CO2 ml-1 and 332 μg CO2 ml-1 vs 504 μg CO2 ml-1 at 72 and 120 h cultivation, respectively (Fig. 5a). In contrast, the growth of each of the 12 members of BP7h was from 2.3 to 20.6% of the growth observed in the complete community: 80–248 μg CO2 ml-1 vs 1200 μg CO2 ml-1 and 40–328 mg CO2 ml-1 vs 1724 μg CO2 ml-1 culture, at 72 and 120 h cultivation, respectively (Fig. 5b).
Growth of the isolated members of the communities in Bayhydrol (PA) and NeoRez (PS-PU) coatings as sole carbon sources. CO2 release was measured at 0 h (negligible, black columns), 72 h (white columns), and 120 h (dotted columns) from 25 ml cultures. a Isolated members (B1-B4) and complete BP1h community incubated in MM-Bayhydrol (3%). b Isolated members (N1-N12) and complete BP7h community incubated in MM-NeoRez (3%) are shown. n = 3. Bars represent standard deviation
Discussion
Three microbial communities, BP1h, BP3h, and BP7h were selected by culture enrichment in a mineral medium with a PS-PU coating as the sole carbon source, inoculated with pieces of deteriorated PU foams, and their biodegradative activities were analyzed on two different coatings, Bayhydrol® A2470, a PA type and NeoRez™ R-9637, a PS-PU type. The higher growth, the larger number of colony types, and the broader taxonomic diversity observed in the three communities cultured in NeoRez, compared to what they presented when were cultured in Bayhydrol could be because of the polyester nature of NeoRez, compared to the polyacrylic nature of Bayhydrol. It is well documented that PS-PU is more susceptible to microbial biodegradation than polyether-PU for example (reviewed in Howard 2012; Mahajan and Gupta 2015; Wilkes and Aristilde 2017), whereas works analyzing biodegradation of PA report that polyacrylic is more recalcitrant than acrylic monomers, dimers, or trimers (Cameron et al. 2000; Hayashi et al. 1993; Mai et al. 2004). For instance, complete mineralization of acrylic acid oligomers by mixed populations of activated sludge microorganisms only occurred with monomers and dimers, but not with larger polyacrylic molecules (Kawai 1993; Larson et al. 1997). The biodegradability analysis of a cross-linked polyacrylic acid by agricultural microbial-containing soils showed that mineralization occurred at rates as low as 0.12–0.24% every 6 months (Wilske et al. 2014). Also, in the biodegradation of films of PU acrylate–acrylated epoxidized soybean oil (AESO)-based cross-linked blends, by the fungus Chaetomium globosum, the blends with more AESO were more biodegradable than the mere PU acrylate (Oprea and Doroftei 2011). As a matter of fact, the high stability of the carbon-carbon backbone of polyacrylate polymers, their insolubility, and their high molecular weight make their biodegradation quite difficult (Cameron et al. 2000).
We observed that changes in some specific functional groups of the coatings’ compounds (Fig. 2) generated by the communities were correlated to the biodegradative extracellular enzymatic activities detected in the supernatant. In Bayhydrol incubated with the BP1h community, the increase in the carbonyl signal of PA can be generated by the high esterase activity that was constantly present during the cultivation period (Fig. 4). Similarly, the decrease in the amine groups (1574 cm-1 signal) of DMEA and TEA can be the result of the constant protease and urease activities acting on the C–N bonds, observed during the whole culture period. On the other hand, the increments in the methyl signals (1452, 1146, and 1388 cm-1), that probably are the result of aliphatic and aromatic C–C bond cleavages of the coating components, and the strong reduction in the ether group signals (1084, 1037, and 980 cm-1) of PA and PGBE suggest that enzymes involved in alkane and ether degradation such as mono and dioxygenases, dehydrogenases, and other type of ether attacking enzymes must be produced by the microbial communities in order to use the Bayhydrol components as carbon source. Studies reporting the biodegradation of polyethylene glycols, compounds related to PGBE, DMEA, and TEA have been published. The utilization of DMEA by Methanococcoides vulcani sp. nov. strain SLH33T, an anaerobic, marine methanogen (L'Haridon et al. 2014), and the degradation of TEA by Pseudomonas pseudoalcaligenes and Alcaligenes faecalis in a cooling-lubrication circuit have been reported (Klammsteiner et al. 2018). In neither case, the mechanisms underlying these abilities were envisaged and although detailed works describing the mechanism for polyethylene glycol degradation in bacteria by successive oxidations with alcohol and aldehyde dehydrogenases and oxidases have been reported (Kawai 2010), no information for degradation of PGBE has been published. In NeoRez, changes in the signals of the PS-PU functional groups were observed, indicating the activity of the microbial communities over the polymer (Fig. 2). The activity of enzymes related to PS-PU biodegradation, esterase, protease, and urease was mainly present after the middle of the incubation period in BP7h (Fig. 4). Similarly, enzymes involved in the break of ether bonds must also be active. These results show that the members of the community secrete extracellular enzymes involved in the biodegradation of the different components of the coatings, as pure bacterial cultures with this ability do (Howard 2002; Oceguera-Cervantes et al. 2007; Rowe and Howard 2002). Further research is needed to identify the specific enzymes involved in the degradation of Bayhydrol and NeoRez components by these communities. In particular, discovering the enzymes that attack the urethane and the ether bonds would be of great relevance because the highly recalcitrant nature of these bonds.
Significantly, all the genera identified in BP1h and BP7h have been reported as xenobiotic degraders. The fact that only Acinetobacter was identified in BP1h grown in Bayhydrol reveals the ability of this genus to degrade not only the aliphatic chains of the PA, naphtha components, PGBE, DMEA, and TEA, but also to attack the ester, amide, and ether groups, as we observed in the FTIR spectra. Acinetobacter sp. has great versatility for using xenobiotics such as phenol (Briganti et al. 1997), toluene (Allen et al. 1999), di-n-butyl phthalate (Fang et al. 2017), or even chemically complex compounds such as lignin (Buchan et al. 2001) as carbon sources. It has the ability to degrade oil (Rusansky et al. 1987) and long-chain alkanes (Asperger et al. 1981). Specifically, Acinetobacter oleivorans PF1, isolated from diesel-contaminated soil, showed alkane degradation in a minimal medium with diesel as the sole carbon source (Balseiro-Romero et al. 2017) and Acinetobacter genospecies 11 W2 was able to consume acrylic monomers, dimers, and trimers, but not polymers (PA) (Kawai 1993). A different scenario is observed in BP7h cultured in NeoRez, in which five different genera, Acinetobacter, Paracoccus, Bacillus, Hydrogenophaga, and Microbacterium, were identified. A. gerneri P7 and A. calcoaceticus are able to degrade the PS-PU coating Impranil (Howard et al. 2010, 2012) and the PU-painting of military aircrafts (El-Sayed et al. 1996), respectively. Acinetobacter and Paracoccus are able to mineralize NMP (Jong et al. 2010), and Paracoccus sp. is also able to hydrolyze amide bonds in pesticides (Zhang et al. 2012) and has been used to enhance bioremediation of soil contaminated with polycyclic aromatic hydrocarbons (PAH) (Teng et al. 2010). Thus, it is feasible that these two bacteria degrade TEA, NMP, and alkanes and that they also attack the carbamate group in the polymer. Also, several Bacillus species have been reported as able to grow in Impranil (Blake et al. 1998), to degrade PU films (Shah et al. 2013), and poly(ether-urethane-urea) (Rafiemanzelat et al. 2015). Hydrogenophaga sp. has been reported as able to biodegrade 4-aminobenzenesulfonate (Gan et al. 2012) and PAHs of high molecular weight (Zaisheng et al. 2017), whereas Microbacterium sp. has the ability to degrade nylon oligomers (Esikova et al. 2014) and acrylic monomers, dimers, and trimers, but not polymers (PA) (Kawai 1993). Since Paracoccus, Hydrogenophaga, and Microbacterium have not been reported as PU degraders, it will be interesting to determine if they are able to attack PU, or if their function in the community is to degrade the additives present in NeoRez, NMP, and TEA.
An interesting result observed in this work was that the individual members of the two analyzed communities, BP1h grown in Bayhydrol and BP7h grown in NeoRez, were not able to independently grow in the media with the coatings as the sole carbon source, at the level the complete communities did (Fig. 5). Even in BP1h, where only two different bacteria from the same genus were isolated, the growth of the isolated members did not reach the growth of the whole community. Moreover, a stronger effect was observed in the BP7h community, in which five different genera were identified, as the growth of each of the 12 isolated populations was less than 20% of the growth of the complete community. The scarce growth that the isolated members of the BP7h community showed in NeoRez indicates that the activity of more than one member of the community is needed for degrading the components of the coating. Although some species such as Acinetobacter have a strong degradative activity over different xenobiotic compounds, a complex network of interactions where some members might have specialized functions seems to operate. Nevertheless, the existence of other bacterial species, not detected by the methods used in this study that may contribute to the performance of the whole community cannot be ruled out. Further analyses will be of great relevance to define which species are the most degradative, and to understand the level of interaction each community member plays during xenobiotics biodegradation.
The results reported in this article set the basis for future works in which robust synthetic consortia, with enhanced degrading abilities towards different xenobiotics may be assembled.
References
Allen A, Hilliard N, Howard GT (1999) Purification and characterization of a soluble polyurethane degrading enzyme from Comamonas acidovorans. Int Biodeterior Biodegradation 43(1–2):37–41. https://doi.org/10.1016/S0964-8305(98)00066-3
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215(3):403–410. https://doi.org/10.1016/S0022-2836(05)80360-2
Asperger O, Naumann A, Kleber HP (1981) Occurrence of cytochrome P-450 in Acinetobacter strains after growth on N-hexadecane. FEMS Microbiol Lett 11(4):309–312. https://doi.org/10.1111/j.1574-6968.1981.tb06986.x
Ausubel FM, Brent R, Kingston RE, Moore DD, Seidman JG, Smith JA, Struhl K (1997) Current protocols in molecular biology. John Wiley & Sons, Inc, New York
Balseiro-Romero M, Gkorezis P, Kidd PS, Van Hamme J, Weyens N, Monterroso C, Vangronsveld J (2017) Characterization and degradation potential of diesel-degrading bacterial strains for application in bioremediation. Int J Phytoremediation 19(10):955–963. https://doi.org/10.1080/15226514.2017.1337065
Blake RC, Norton WN, Howard GT (1998) Adherence and growth of a Bacillus species on an insoluble polyester polyurethane. Int Biodeterior Biodegradation 42(1):63–73. https://doi.org/10.1016/S0964-8305(98)00048-1
Bradford MM (1976) Rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72(1–2):248–254. https://doi.org/10.1016/0003-2697(76)90527-3
Briganti F, Pessione E, Giunta C, Scozzafava A (1997) Purification, biochemical properties and substrate specificity of a catechol 1,2-dioxygenase from a phenol degrading Acinetobacter radioresistans. FEBS Lett 416(1):61–64. https://doi.org/10.1016/S0014-5793(97)01167-8
Buchan A, Neidle EL, Moran MA (2001) Diversity of the ring-cleaving dioxygenase gene pcaH in a salt marsh bacterial community. Appl Environ Microbiol 67(12):5801–5809. https://doi.org/10.1128/AEM.67.12.5801-5809.2001
Cameron MD, Post ZD, Stahll JD, Haselbach J, Aust SD (2000) Cellobiose dehydrogenase-dependent biodegradation of polyacrylate polymers by Phanerochaete chrysosporium. Environ Sci Pollut R 7(3):130–134. https://doi.org/10.1065/espr2000.04.022.1
Cosgrove L, McGeechan PL, Robson GD, Handley PS (2007) Fungal communities associated with degradation of polyester polyurethane in soil. Appl Environ Microbiol 73(18):5817–5824. https://doi.org/10.1128/AEM.01083-07
Crossno SK, Kalbus LH, Kalbus GE (1996) Determination of carbon dioxide by titration: new experiments for general, physical, and quantitative analysis course. J Chem Educ 73(2):175–176. https://doi.org/10.1021/ed073p175
Desphande MV, Eriksson KE, Pettersson LG (1984) An assay for selective determination of exo-1,4-β-glucanases in a mixture of cellulolytic enzymes. Anal Biochem 138(2):481–487. https://doi.org/10.1016/0003-2697(84)90843-1
El-Sayed AHMM, Mahmoud WM, Davis EM, Coughlin RW (1996) Biodegradation of polyurethane coatings by hydrocarbon-degrading bacteria. Int Biodeterior Biodegradation 37(1–2):69–79. https://doi.org/10.1016/0964-8305(95)00091-7
Esikova TZ, Akatova EV, Taran SA (2014) Bacteria that degrade low-molecular linear epsilon-caprolactam oligomers. Appl Biochem Microbiol 50(5):463–470. https://doi.org/10.1134/S0003683814050044
Fang Y, Zhang LS, Wang J, Zhou Y, Ye BC (2017) Identification of the di-n-butyl phthalate-biodegrading strains and the biodegradation pathway in strain LMB-1. Appl Biochem Microbiol 53(3):310–317. https://doi.org/10.1134/S000368381703005X
Gan HM, Chew TH, Tay YL, Lye FS, Yahya A (2012) Genome sequence of Hydrogenophaga sp. strain PBC, a 4-aminobenzenesulfonate-degrading bacterium. J Bacteriol 194(17):4759–4760. https://doi.org/10.1128/JB.00990-12
Gurav R, Lyu H, Ma J, Tang J, Liu Q, Zhang H (2017) Degradation of n-alkanes and PAHs from the heavy crude oil using salt-tolerant bacterial consortia and analysis of their catabolic genes. Environ Sci Pollut R 24(12):11392–11403. https://doi.org/10.1007/s11356-017-8446-2
Hayashi T, Mokouyama M, Sakano K, Tani Y (1993) Degradation of a sodium acrylate oligomer by an Arthrobacter sp. Appl Environ Microbiol 59(5):1555–1559
Howard GT (2002) Biodegradation of polyurethane: a review. Int Biodeterior Biodegradation 49(4):245–252. https://doi.org/10.1016/S0964-8305(02)00051-3
Howard GT (2012) Polyurethane biodegradation. In: Singh SN (ed) Microbial degradation of xenobiotics. Springer-Verlag, Heidelberg, pp 189–211
Howard GT, Duos B, Watson E (2010) Characterization of the soil microbial community associated with the decomposition of a swine carcass. Int Biodeterior Biodegradation 64(4):300–304. https://doi.org/10.1016/j.ibiod.2010.02.006
Howard GT, Norton WN, Burks T (2012) Growth of Acinetobacter gerneri P7 on polyurethane and the purification and characterization of a polyurethanase enzyme. Biodegradation 23(4):561–573. https://doi.org/10.1007/s10532-011-9533-6
Jong KL, Woo JL, Yong-Ju C, Doo HF, Yong-Woo L, Jinwook C (2010) Variation of bacterial community immobilized in polyethylene glycol carrier during mineralization of xenobiotics analyzed by TGGE technique. Korean J Chem Eng 27(6):1816–1821. https://doi.org/10.1007/s11814-010-0291-7
Kato S, Chino K, Kamimura N, Masai E, Yumoto I, Kamagata Y (2015) Methanogenic degradation of lignin-derived monoaromatic compounds by microbial enrichments from rice paddy field soil. Sci Rep 5:14295. https://doi.org/10.1038/srep14295
Kawai F (1993) Bacterial degradation of acrylic oligomers and polymers. Appl Microbiol Biotechnol 39:382–385. https://doi.org/10.1007/BF00192097
Kawai F (2010) The biochemistry and molecular biology of xenobiotic polymer degradation by microorganisms. Biosci Biotechnol Biochem 74(9):1743–1759. https://doi.org/10.1271/bbb.100394
Klammsteiner T, Insam H, Probst M (2018) Microbiota in a cooling-lubrication circuit and an option for controlling triethanolamine biodegradation. Biofouling 34(5):519–531. https://doi.org/10.1080/08927014.2018.1468887
Larson RJ, Bookland EA, Williams RT, Yocom KM, Saucy DA, Freeman MB, Swift G (1997) Biodegradation of acrylic acid polymers and oligomers by mixed microbial communities in activated sludge. J Environ Polym Degrad 5(1):41–48. https://doi.org/10.1007/BF02763567
L'Haridon S, Chalopin M, Colombo D, Toffin L (2014) Methanococcoides vulcani sp. nov., a marine methylotrophic methanogen that uses betaine, choline and N,N-dimethylethanolamine for methanogenesis, isolated from a mud volcano, and emended description of the genus Methanococcoides. Int J Syst Evol Microbiol 64(Pt6):1978–1983. https://doi.org/10.1099/ijs.0.058289-0
Lovrien R, Matulis D (1995) Assays for total protein. In: Coligan JE, Dunn BM, Ploegh HL, Speicher DW, Wingfield PT (eds) Current protocols in protein science. John Wiley & Sons, Inc, New York, pp 3.4.4–3.4.24
Mahajan N, Gupta P (2015) New insights into the microbial degradation of polyurethanes. RSC Adv 5:41839–41854. https://doi.org/10.1039/c5ra04589d
Mai C, Schormann W, Majcherczyk A, Hüttermann A (2004) Degradation of acrylic copolymers by white-rot fungi. Appl Microbiol Biotechnol 65(4):479–487. https://doi.org/10.1007/s00253-004-1668-5
McCarthy SJ, Meijs GF, Mitchell N, Gunatillake PA, Heath G, Brandwood A, Schindhelm K (1997) In-vivo degradation of polyurethanes: transmission-FTIR microscopic characterization of polyurethanes sectioned by cryomicrotomy. Biomaterials 18(21):1387–1409. https://doi.org/10.1016/S0142-9612(97)00083-5
Muyzer G, de Waal EC, Uitterlinden AG (1993) Profiling of complex microbial populations by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes coding for 16S rRNA. Appl Environ Microbiol 59(3):695–700
Nakajima-Kambe T, Onuma F, Kimpara N, Nakahara T (1995) Isolation and characterization of a bacterium which utilizes polyester polyurethane as a sole carbon and nitrogen source. FEMS Microbiol Lett 129(1):39–42. https://doi.org/10.1111/j.1574-6968.1995.tb07554.x
Oceguera-Cervantes A, Carrillo-García A, López N, Bolaños-Nuñez S, Cruz-Gómez MJ, Wacher C, Loza-Tavera H (2007) Characterization of the polyurethanolytic activity of two Alicycliphilus sp. strains able to degrade polyurethane and N-methylpyrrolidone. Appl Environ Microbiol 73(19):6214–6223. https://doi.org/10.1128/AEM.01230-07
Oprea S, Doroftei F (2011) Biodegradation of polyurethane acrylate with acrylated epoxidized soybean oil blend elastomers by Chaetomium globosum. Int Biodeterior Biodegradation 65(3):533–538. https://doi.org/10.1016/j.ibiod.2010.09.011
Pardini OR, Amalvy JI (2008) FTIR, 1H-NMR spectra, and thermal characterization of water-based polyurethane/acrylic hybrids. J Appl Polym Sci 107:1207–1214. https://doi.org/10.1002/app.27188
Perruchon C, Pantoleon A, Veroutis D, Gallego-Blanco S, Martin-Laurent F, Liadaki K, Karpouzas DG (2017) Characterization of the biodegradation, bioremediation and detoxification capacity of a bacterial consortium able to degrade the fungicide thiabendazole. Biodegradation 28(5-6):383–394. https://doi.org/10.1007/s10532-017-9803-z
Rafiemanzelat F, Jafari M, Emtiazi G (2015) Study of biological degradation of new poly(ether-urethane-urea)s containing cyclopeptide moiety and PEG by Bacillus amyloliquefaciens isolated from soil. Appl Biochem Biotechnol 177(4):842–860. https://doi.org/10.1007/s12010-015-1782-0
Rowe L, Howard GT (2002) Growth of Bacillus subtilis on polyurethane and the purification and characterization of a polyurethanase-lipase enzyme. Int Biodetrior Biodegradation 50(1):33–40. https://doi.org/10.1016/S0964-8305(02)00047-1
Rusansky S, Avigad R, Michaeli S, Gutnick DL (1987) Involvement of a plasmid in growth on and dispersion of crude oil by Acinetobacter calcoaceticus RA57. Appl Environ Microbiol 53(8):1918–1923
Shah AA, Hasan F, Akhter JI, Hameed A, Ahmed S (2008) Degradation of polyurethane by novel bacterial consortium isolated from soil. Ann Microbiol 58:381–386. https://doi.org/10.1007/BF03175532
Shah Z, Krumholz L, Aktas DF, Hasan F, Khattak M, Shah AA (2013) Degradation of polyester polyurethane by a newly isolated soil bacterium, Bacillus subtilis strain MZA-75. Biodegradation 24(6):865–877. https://doi.org/10.1007/s10532-013-9634-5
Shah Z, Gulzar M, Hasan F, Shah AA (2016) Degradation of polyester polyurethane by an indigenously developed consortium of Pseudomonas and Bacillus species isolated from soil. Polym Degrad Stab 134:349–356. https://doi.org/10.1016/j.polymdegradstab.2016.11.003
Solís-González CJ, Domínguez-Malfavón L, Vargas-Suárez M, Gaytán I, Cevallos MA, Lozano L, Cruz-Gómez MJ, Loza-Tavera H (2018) Novel metabolic pathway for N-Methylpyrrolidone degradation in Alicycliphilus sp. BQ1 Appl Environ Microbiol 84(1): pii: e02136-17). https://doi.org/10.1128/AEM.02136-17
Teng Y, Luo Y, Sun M, Liu Z, Li Z, Christie P (2010) Effect of bioaugmentation by Paracoccus sp. strain HPD-2 on the solid microbial community and removal of polycyclic aromatic hydrocarbons from and aged contaminated soil. Bioresour Technol 101(10):3437–3443. https://doi.org/10.1016/j.biortech.2009.12.088
Vogt C, Richnow HH (2014) Bioremediation via in situ microbial degradation of organic pollutants. Adv Biochem Eng Biotechnol 142:123–146. https://doi.org/10.1007/10_2013_266
Wilkes RA, Aristilde L (2017) Degradation and metabolism of synthetic plastics and associated products by Pseudomonas sp.: capabilities and challenges. J Appl Microbiol 123(3):582–593. https://doi.org/10.1111/jam.13472
Wilske B, Bai M, Lindenstruth B, Bach M, Rezaie Z, Frede HG, Breuer L (2014) Biodegradability of a polyacrylate superabsorbent in agricultural soil. Environ Sci Pollut Res 21(16):9453–9460. https://doi.org/10.1007/s11356-013-2103-1
Witte CP, Medina-Escobar N (2001) In-gel detection of urease with nitroblue tetrazolium and quantification of the enzyme from different crop plants using the indophenols reaction. Anal Biochem 290(1):102–107. https://doi.org/10.1006/abio.2000.4933
Zafar U, Houlden A, Robson GD (2013) Fungal communities associated with the biodegradation of polyester polyurethane buried under compost at different temperatures. Appl Environ Microbiol 79(23):7313–7324. https://doi.org/10.1128/AEM.02536-13
Zaisheng Y, Yu Z, Huifang W, Mingzhong Y, Haichen Z, Zheng H, Helong J (2017) Isolation and characterization of a bacterial strain Hydrogenophaga sp. PYR1 for anaerobic pyrene and benzo[a]pyrene biodegradation. RSC Adv 7(74):46690–46698. https://doi.org/10.1039/c7ra09274a
Zhang J, Yin JG, Hang BJ, Cai S, He J, Zhou SG, Li SP (2012) Cloning of novel arylamidase gene from Paracoccus sp. strain FLN-7 that hydrolyzes amide pesticide. Appl Environ Microb 78(14):4848–4855. https://doi.org/10.1128/AEM.00320-12
Acknowledgements
V.F.C. acknowledges Programa 121, Formación Básica en Investigación, Facultad de Química, Universidad Nacional Autónoma de México for her scholarship. We thank Chem. Maricela Gutiérrez Franco for FTIR analysis carried out at Unidad de Servicios de Apoyo a la Investigación y a la Industria (USAII), Facultad de Química, UNAM. We also thank José Galván and Carlos Galván from Pinturas y Solventes de México, S.A. de C.V. for providing the coatings used in this study.
Funding
This study was funded by Programa de Apoyo a Proyectos de Investigación e Innovación Tecnológica, Dirección General de Asuntos del Personal Académico, Universidad Nacional Autónoma de México grants IN217114 and IN223317, and Programa de Apoyo a la Investigación y el Posgrado, Facultad de Química, Universidad Nacional Autónoma de México, grant 5000-9117.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Vargas-Suárez, M., Fernández-Cruz, V. & Loza-Tavera, H. Biodegradation of polyacrylic and polyester polyurethane coatings by enriched microbial communities. Appl Microbiol Biotechnol 103, 3225–3236 (2019). https://doi.org/10.1007/s00253-019-09660-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-019-09660-y