Abstract
Functional genomics of filamentous fungi has gradually uncovered gene information for constructing ‘cell factories’ and controlling pathogens. Available gene manipulation methods of filamentous fungi include random integration methods, gene targeting technology, gene editing with artificial nucleases and RNA technology. This review describes random gene integration constructed by restriction enzyme-mediated integration (REMI); Agrobacterium-mediated transformation (AMT); transposon-arrayed gene knockout (TAGKO); gene targeting technology, mainly about homologous recombination; and modern gene editing strategies containing transcription activator-like effector nucleases (TALENs) and a clustered regularly interspaced short palindromic repeat/associated protein system (CRISPR/Cas) developed in filamentous fungi and RNA technology including RNA interference (RNAi) and ribozymes. This review describes historical and modern gene manipulation methods in filamentous fungi and presents the molecular tools available to researchers investigating filamentous fungi. The biggest difference of this review from the previous ones is the addition of successful application and details of the promising gene editing tool CRISPR/Cas9 system in filamentous fungi.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Filamentous fungi are of special interest to researchers because of their great capacity to produce diverse valuable metabolites, including antibiotics (Meyer et al. 2011), enzymes (Hoffmeister and Keller 2007), acids (Magnuson and Lasure 2004) and lipids (Tang et al. 2016) (Chen et al. 2015), in addition to detrimental toxins such as aflatoxins (Sørensen et al. 2008) and sterigmatocystins (Keller and Hohn 1997). Molecular tools are needed to manipulate them, such as strengthening biosynthetic pathway to construct microbiological cell factories and disrupting pathogenic genes to control the harmful filamentous fungi. Besides, genome sequences of hundreds of filamentous fungi have been established (http://www.genomesonline.org/; http://genome.jgi.doe.gov/), and effective gene manipulation techniques, especially multiple-gene and target-specific methods, are needed for exploration of genetic information to promote its application in the pharmaceutical field, agricultural field and food industry.
Various transformation methods of filamentous fungi, including protoplast transformation (Case et al. 1979, Turgeon et al. 2010), particle bombardment (Bhairi and Staples 1992), electroporation (Richey et al. 1989) and Agrobacterium -mediated transformation methods (de Groot et al. 1998a) presented in Table 1 (Meyer 2008), have been established. And multiple selection markers summarised in Table 2, (referring to (Fincham 1989)), such as hygromycin B drug resistance (Kück et al. 1989; Punt and van den Hondel 1992) and uracil auxotrophic mutants (Ando et al. 2009; Hao et al. 2014) are available. Those all facilitate the progress of the genome manipulation method in filamentous fungi with addition of the development of corresponding screening systems.
The random integration method is applied for high-throughput mutagenesis in a strain of interest, but its process is tedious. Gene targeting technology based on the homologous recombination method can help us manipulate particular genes but this method is not efficient in filamentous fungi. Gene editing technology appears after the genome sequencing can be used for efficiently uncovering gene function, and one of the most promising tools—CRISPR systems—was successfully used in filamentous fungi. The RNA technology can disturb gene expression in the translational level, but is not applicable in some filamentous fungi. Those molecular tools in filamentous fungi allow researchers to choose proper methods for uncovering gene function and manipulating different genes efficiently.
This review provides an insight into the gene manipulation methods applied in functional genomics in filamentous fungi and details up-to-date gene editing strategies, with a focus on the CRISPR/Cas9 system.
Random integration methods
Gene functions have traditionally been tested with random gene mutation methods using physical mutagens, including ionising radiation, ultraviolet light or radioactive chemicals and chemical mutagens such as alkaloids and benzene (Casselton and Zolan 2002) to determine whether any phenotypic change had occurred. However, the process of isolation and recovery of the mutated gene is tedious. The detection of genetic mutations with molecular tools saves the laborious work involved in mutation isolation and gene identification.
Random gene disruptions can be created through transformation methods (foreign DNA insertion) or by the movement of mobile genetic elements. Other related gene manipulation methods used in filamentous fungi are restriction enzyme-mediated integration (REMI) and transposon-arrayed gene knockout (TAGKO) (Jiang et al. 2013).
Restriction enzyme-mediated integration
REMI was first established in 1991 in Saccharomyces cerevisiae (Schiestl and Petes 1991). The basic process of REMI transformation and recovery of mutated genes is demonstrated in Fig. 1 (Kahmann and Basse 1999). After the enzyme-digested DNA fragment and the enzyme are transformed into the protoplast, the plasmid can be integrated into the genome of the host based on the same sticky ends. The mutated genes can be isolated with their corresponding primers after plasmid excision and ligation after transformation in Escherichia coli. REMI has been successfully used in filamentous fungi, mainly for the identification of pathogenicity genes. One example is the filamentous ascomycete Magnaporthe grisea. Shi et al. screened 10 mutants for sporulation, pathogenicity and auxotrophy among 800 transformants with the REMI method (Shi and Leung 1995). Sweigard et al. also used this method to achieve 27 stable pathogenic mutants among 5538 transformants in M. grisea (Sweigard et al. 1998). Researchers have also found that the use of enzymes in the REMI method improves transformation efficiency (Shi et al. 1995). In addition, this method has already been used in the filamentous fungi Monacrosporium sphaeroides (Xu et al. 2005), Fusarium oxysporum (Imazaki et al. 2007) and Pleurotus eryngii (Noh et al. 2010); approximately 5000 transformants and 2929 transformants were achieved for the first two species, respectively, and 10 to 40 transformants per 106 protoplasts were achieved for the third.
REMI transformation and plasmid rescue. Adapted from REMI (Restriction Enzyme Mediated Integration) and its Impact on the Isolation of Pathogenicity Genes in Fungi Attacking Plants. Doi: https://doi.org/10.1023/A:1008757414036. a A REMI plasmid (RP) often contains a fungal transformation marker for gene transcription and a bacterial transformation marker for plasmid rescue. b The circular or linearised REMI plasmid is transformed into fungal cells together with a restriction enzyme (e.g. BamHI), resulting in two cleavage sites at the transforming plasmid and chromosomal. c Free ends of the two cleaved sites are joined together. d Plasmid rescue can be achieved via plasmid excising from genome in presence of flanking sequences (e.g. by using MluI) and circularising with DNA ligase before transformation in E. coli
Several disadvantages hold back the development of this method. It was initially dependent on protoplasts for the uptake of plasmid DNA and enzymes. The proper choices and concentrations of enzymes must be determined before the transformation. Researchers also found that different restriction enzymes vary significantly in their ability to integrate fragments into the host genome (Manivasakam and Schiestl 1998). If plasmid rescue is needed, essential enzymatic sites must be known.
Agrobacterium-mediated transformation
The Agrobacterium-mediated transformation (AMT) method was traditionally used in plants (Valvekens et al. 1988; Tingay et al. 1997; Komari et al. 1996) and was later applied to yeast (Piers et al. 1996), filamentous fungi (de Groot et al. 1998b) and mammal cells (Kunik et al. 2001). A. tumefacians is a gram-negative plant pathogen that can transfer its T-DNA randomly into genome of the recipient at a random site (Hooykaas and Beijersbergen 1994) upon induction via chemicals, usually acetosyringone. Acetosyringone was used as induction through activating VirA and VirG proteins on the surface of Agrobacterium tumefaciens to send signals. T-DNA is bordered by 25 base pair repeats on each end. And the transfer is initiated at the right border and terminated at the left border in presence of the Vir proteins. The specific AMT process (Michielse et al. 2005) is illustrated in Fig. 2.
Schematic overview of the Agrobacterium tumefaciens T-DNA transfer system. Adapted from Agrobacterium-mediated transformation as a tool for functional genomics in fungi. Doi: https://doi.org/10.1007/s00294-005-0578-0. When acetosyringone or sugars exists, the vir genes encoding the T-DNA transfer machinery of A. tumefaciens are induced. Virulence proteins VirA and VirG are activated first in the presence of acetosyringone, while chromosomally encoded protein, ChvE, interacts with the VirA protein upon recognition of monosaccharides. The VirG protein is activated through accepting the phosphoryl group from the activated VirA proteins. The activated VirG proteins have DNA-binding properties, and acts as a transcriptional activator of itself and other virulence genes located in the virulence region. The VirD2 protein, assisted by VirD1, nick precisely in the bottom strand of both border repeats, leading to T-DNA release. VirC1 can bind the ‘overdrive’ of T-DNA and help T-strand production. After the T-DNA is released, the T-DNA strand will transfer through the bacterial membrane and fungal cell wall via a type IV secretion mechanism. The VirB protein forms a transport pore. The inner membrane VirD assists in the transferring process. The virulence proteins VirF, VirE1/E2 are also exported via the pore. The transferred T-DNA is targeted to the nucleus and randomly integrated into the genome. The precise mechanism of T-DNA integration is not known
Combier et al. transformed the mycelium of Hebeloma cylindrosporum with this method; Southern blot analysis of 83 randomly selected transformants showed a unique plasmid insert pattern, and 60% was composed of single-transferred DNA copies. The left and right borders can achieve 85 and 15% of transformants, respectively, via the thermal asymmetrical interlaced polymerase chain reaction (TAIL-PCR) (Combier et al. 2003). TAIL-PCR was used for recovering unknown (mutated) gene adjacent to known sequences, such as the inserted T-DNA and transposon, through utilising nested known-specific primers with a high melting temperature (Tm > 65 °C) in consecutive reactions together with a short (15–16 nucleotides) arbitrary degenerate (AD) primer with a low Tm (about 45 °C) and 64–256-folds of degeneracy (Liu and Chen 2007). The relative amplification efficiencies of target and nontarget products can be thermally controlled; thus, the mutated gene can be amplified. Gento et al. also used the ATMT method in Colletotrichum lagenarium and obtained 150 to 300 transformants per 106 conidia, and highly efficient gene recovery was achieved via the thermal asymmetrical interlaced polymerase chain reaction (Tsuji et al. 2003).
When the technique is used in filamentous fungi, host cells can include protoplasts (Liu et al. 2015), spores (Hao et al. 2014; Wang and Li 2008), hyphae and sporocarps. This alleviates the need for protoplast isolation as required by REMI. Rogers et al. adopted both REMI and ATMT to identify pathogenic mutants for Coniothyrium minitans and found that 32 transformants (μg DNA−1) were achieved with REMI while 37.8 transformants (5 × 105 germlings−1) were achieved with ATMT. And single-copy DNA integrations occurred in 8% of REMI and 40% of ATMT transformants (Rogers et al. 2004). Although ATMT surpasses REMI in some aspects, it also has some clear disadvantages. ATMT is time consuming, and many of the parameters involved, such as the ratio of host cells to Agrobacterium, the co-cultivation temperature and the co-cultivation time, must be optimised to achieve a high frequency of transformation.
Transposon-arrayed gene knockout
Transposable elements are diverse and ubiquitous in major groups of filamentous fungi, including Ascomycetes, Zygomycetes and Basidiomycetes. They are used for gene mutation due to their ability to transfer among the host cells (Daboussi 1997).
The transposable-arrayed gene knockout technique was first used for high-throughput mutagenesis in M. grisea. The process is illustrated in Fig. 3 (Hamer et al. 2001). The transposon-mediated mutagenesis method has also been established in some other filamentous fungi. Aspergillus niger was found to harbour a non-autonomous transposon named Vader, for which an excision frequency of 1 in 2.2 × 105 was found (Hihlal et al. 2011). A similar method was used in Penicillium griseoroseum to achieve a transformation frequency based on a heterologous transposon, Impala, from F. oxysporum (de Queiroz and Daboussi 2003), and similar methods were established in other filamentous fungi, including Mycosphaerella graminicola (Adachi et al. 2002) and Aspergillus fumigatus (Firon et al. 2003). Although this method can be used interchangeably and requires a high-efficiency transformation approach (Jiang et al. 2013), target manipulation cannot be achieved.
Transposon-arrayed gene knockout. Adapted from Gene discovery and gene function assignment in filamentous fungi. Doi: https://doi.org/10.1073/pnas.091094198. An IVT reaction needs transposons, corresponding transposase and recipient DNA, usually a cosmid vector. Then, transform the IVT products into E. coli. Positive transposon insertion sites can be determined by sequencing. Then, the proper cosmids are digested with the homing endonuclease to release full-length inserts for transformation. Ectopic and TI events are distinguished by PCR analysis, and mutants are subjected to phenotype analysis. IVT in vitro transposition, WT wild type, EI ectopic integration, TI targeted integration
Gene targeting technology
The functions of specific genes can be verified by gene overexpression and downregulation methods through homologous recombination.
Homologous recombination
The homologous recombination (HR) method was conventionally used for target integration for gene knock-in and gene mutation, but it was not as effective in filamentous fungi as in yeast. In S. cerevisiae, 50 bp was sufficient for foreign DNA integration (Hua et al. 1997). In filamentous fungi, larger fragments longer than 1000 bp may not achieve high transformation efficiencies (Kupfer et al. 1997). Although the frequency of homologous recombination of filamentous fungi can be improved by mutating the genes involved in the non-homologous end-joining process, usually ku70 and ku80, the technique still depends upon protoplasts and a proper plasmid with a proper selection marker. Digestion of cell wall of filamentous fungi is usually difficult.
Gene editing technology
“Cre/Loxp” and “FLP/FRT” system
Before appearance of the concept of gene editing, there exists a selection marker recycle method called the “Cre/Loxp” system, allowing multiple genes manipulation in filamentous fungi. The “Cre/Loxp” system was adapted from P1 bacteriophage and was composed with a recombinase named Cre and two corresponding consequent Loxp sites. And the Cre can digest at the two consequent Loxp sites, resulting in a gene, usually a selection marker, within the two consequent Loxp sites being deleted. The Cre/Loxp system was applied in Aspergillus (Krappmann S et al. 2005, Forment et al. 2006, Mizutani et al. 2012, Huang et al. 2016), Trichoderma (Steiger et al. 2011), Neurospora (Honda and Selker 2009), Neotyphodium (Florea et al. 2009) and so on (Zhang et al. 2013, Aguiar et al. 2014) (summarised in Table 3).
Similarly, a “FLP/FRT” system, composed of FLP recombinase with corresponding FRT sites, is also used for marker recycling in filamentous fungi, but is derived from the yeast S. cerevisiae. Kopke et al. firstly demonstrated successful application of the “FLP/FRT” system in Penicillium chrysogenum and Sordaria macrospora in 2010 (Kopke et al. 2010). Later, this method was used in Ustilago maydis (Khrunyk et al. 2013) and Acremonium chrysogenum (Bloemendal et al. 2014).
The above two methods offer possibilities for multigene manipulation but need two steps to achieve gene recycle. Take the “Cre/Loxp” system for example, the Loxp sites as well as a marker gene need to be integrated into the host genome first, then the Cre needs expression to finish the gene deletion process. Thus, a precise selection process of the second step is necessary for successful application of the two methods.
Zinc-finger nucleases (ZFNs), transcription activator-like effector nucleases (TALENs) and clustered regulatory interspaced short palindromic repeat/associated systems (CRISPR/Cas) are gene manipulation systems that use nucleases. CRISPR/Cas9 is the most widely used. Although the ZFN method was established earlier than TALENs and CRISPR/Cas9 and has been applied to gene editing in plants and animals, we were unable to find any examples of its use in filamentous fungi. Therefore, we focus on TALENs and CRISPR/Cas9.
Transcription activator-like effector nucleases
Transcription activator-like effectors were first found in 2009 in the plant pathogen Xanthomonas and contain a central region of tandem direct repeats with 34 amino acids (Boch et al. 2009). Figure 4 is a simple illustration of the transcription activator-like effector nuclease (TALEN) method. Two specific amino acids, often the 12th and 13th and known as repeat-variable di-residues, are two hypervariable amino acids that determine the single nucleotide in the target DNA sequence (Christian et al. 2010). The artificial TALENs are fused with a DNA catalytic part FokI to produce a specific double-strand break. Compared with the three bases recognised by one ZFN monomer, one TALEN monomer recognises one base and therefore offers a larger, more flexible target range (Sun and Zhao 2013).
Transcription activator-like effector nuclease (TALEN) in complex with target DNA. TALEN target sites contain two TALE binding sites with a spacer sequence separation of varying lengths (12–20 bp). Each TALE repeats recognise the specific base pair through two hypervariable residues and then linked FokI cleavage at target site. RVD repeat-variable di-residue
This technology was first used in filamentous fungi in the rice blast fungus Pyricularia oryzae in 2015 (Arazoe et al. 2015), and the established platinum-fungal TALEN system raised the targeted gene replacement efficiency through homologous recombination by as much as 100%. This technique has not yet been reported in any other genus of filamentous fungi.
The technical challenge for this technology is cloning extensive identical repeat sequences (Gaj et al. 2013). Several methods have been explored to overcome this problem, such as the ‘Golden Gate’ assembly method (Cermak et al. 2011), high-throughput solid-phase assembly (Briggs et al. 2012) and ligation-independent cloning techniques (Schmid-Burgk et al. 2013).
Clustered regularly interspaced palindromic repeat/associated systems
The best-known gene editing method in the field of artificial nuclease-based gene manipulation is CRISPR/Cas9.
Mechanism of CRISPR/Cas systems
CRISPR/Cas systems in bacteria and archaebacteria play a biological role in adaptive immunity systems against foreign DNA (Ran et al. 2013a; Doudna and Charpentier 2014). After a virus or phage injects DNA into the bacterium, parts of its DNA can be integrated into the CRISPR array in the genome (Wang et al. 2014). When the foreign DNA invades the bacterium again, the mature CRISPR RNA (crRNA) combined with a trans-activating RAN (tracrRNA) can combine and lead the Cas9 protein to degrade the DNA based on complementary sequencing (Ran et al. 2013b; Hsu et al. 2014). The process is summarised in Fig. 5.
Schematic overview of a CRISPR/Cas system. a Acquisition of the type II CRISPR/Cas system. The invaded DNA is processed as new spacer into the CRISPR array after the first attack. When the same DNA starts a second attack, the spacer expresses into the crRNA-tracr RNA complex and leads the Cas9 protein to the invading DNA and cleavage on the corresponding sequence. b Mechanisms of the type II CRISPR/Cas system. There is only Cas9 in a type II CRISPR/Cas system. Cas9 has two function domains: HNH, cleaving the responding sequence attached with sgRNA, and RuvC, cleaving the remaining complementary sequence. The PAM sites are usually upstream of the cleavage sites
Of the three CRISPR/Cas systems, type II CRISPR/Cas has been extensively explored for its simple mono-protein composition (Bassett et al. 2013; Friedland et al. 2013; Gratz et al. 2013). After a linker loop is added between the crRNA and tracrRNA, more convenient single-strand RNA is generated for gene editing in various cells and organisms (Bassett et al. 2013; Jinek et al. 2012). The first specific 20-nucleotide at the 5′ end of the gRNA corresponding to the tracrRNA decides the target position, and the remaining sequence at the 3′ end forms a stem-loop structure that is necessary for the activity of Cas9 protein. Due to the low content of sgRNA, a plasmid can be designed that will encode multiple sgRNAs. Multigene manipulation can thus be easily achieved, which makes unnecessary the complex procedure used in the above two nucleases guided by proteins (Shalem et al. 2014).
The commonly used CRISPR/Cas9 systems have two characteristics: they are protospacer adjacent motif (PAM) dependent and act up to six base pairs off-target. The mechanism can be understood in terms of its biological role in defending against foreign DNA. Kleinstiver et al. modified PAM sites in human cells and broadened the targeting range of Staphylococcus aureus Cas9, although off-target effect remains the same (Kleinstiver et al. 2015).
Many studies have been performed to improve Cas9 specificity and reduce off-target effects, including reducing the active Cas9 amount (Davis et al. 2015; Hsu et al. 2013), using inactivated Cas9 such as Cas9 nickase combined with FokI nuclease domain (Guilinger et al. 2014; Tsai et al. 2014) and using a truncated guide sequence (Kleinstiver et al. 2015), among others (Slaymaker et al. 2016).
Application of CRISPR/Cas systems
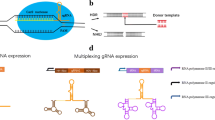
Many biotechnological companies, such as Addgene, offer an online service for PAM sequence searching and sgRNA construction and provide synthesised products. Researchers have already constructed specific plasmids used to establish the CRISPR/Cas9 system in filamentous fungi. The four plasmids AFUM_pyrG, AN_argB, bleR and hygR each carry a common selection marker to allow convenient operation in different filamentous fungi (Nødvig et al. 2015).
CRISPR/Cas9 systems have been established in several filamentous fungi since 2015, and the list continues to grow. The first was Trichoderma reesei. A controllable and conditional CRISPR/Cas9 system was established and used to disrupt the ura5 gene, and a 200-bp homologous length can induce site-specific mutations with 93% recombination frequency with the help of the CRISPR/Cas9 system (Liu et al. 2015). This system has been established in Trichoderma, Aspergillus, Penicillium and the model organism Neurospora crassa (Table 4).
Other applications of the CRISPR/Cas9 systems used in bacteria, such as CRISPR interference (Cleto et al. 2016; Qi et al. 2013) and activation (Cheng et al. 2013), may also be used for filamentous fungi because the inactive form of Cas9 nucleases also exhibits versatile abilities. With inactivation of either of the functional parts of the HNH and RuvC domain, the Cas9 nucleases become nickases that induce single-strand breaks. With both inactive domains, Cas9 nucleases become repressor-like proteins that can block the transcription progress, which is called CRISPR interference (CRISPRi) (Larson et al. 2013). If a regulatory part, a repressor or an enhancer, is combined with a Cas9 protein with DNA targeting ability, a single gene or a series of genes can be regulated. Researchers have compared the effect of CRISPRi and shRNA in the identification of essential genes in RT-112 cells and found that a CRISPR-based screening method performed the best (Evers et al. 2016).
Integration of Cas9 protein and sgRNA
The key challenge to the establishment of this system in filamentous fungi is the method of integrating the Cas9 protein and sgRNA into the host. Methods include DNA and mRNA techniques. Both components can be co-transformed by one plasmid or solo transformed by being transformed twice. In most filamentous fungi, the Cas9 DNA is integrated into the host’s genome, and some filamentous fungi such as Ustilago maydis use an auto-replicating plasmid with Cas9 DNA in the cytoplasm. The auto-replicating plasmid allows transient expression of Cas9 to avoid the disadvantage of permanent Cas9 (Schuster et al. 2016), which was found to aggravate the off-target effect.
In addition, a RNA-guided endonuclease technique using a preassembled duplex combined with purified Cas9 proteins has been successfully used in the filamentous fungus P. chrysogenum (Pohl et al. 2016) and in human cells, zebrafish embryos, bacteria (Kim et al. 2014) and plants (Woo et al. 2015). This preassembled ribonucleoproteins method is able to improve on-target efficiency and reduce off-target inefficiency compared with the traditional plasmid transformation method, with which it is difficult to successfully integrate the Cas9 protein into the genome and in which continuous expression of the Cas9 protein can aggravate the off-target effects.
SgRNA
RNA polymerase III promoters, such as U6 and T7 promoters, drive the expression of sgRNA. The endogenous RNA polymerase III promoter may not exist in filamentous fungi, so a heterogenous RNA polymerase III promoter, usually U6, is often used. Alternative methods, such as using sgRNA derived from the RNA polymerase II promoter, and RNA processing strategies have also shown the capability of sgRNA synthesis.
The sgRNA derived from the RNA polymerase II promoter in P. oryzae has the ability to guide the Cas9 protein but shows less activity than sgRNA derived from the RNA polymerase III promoter (Arazoe et al. 2015). Researchers have also used ribozymes to design the artificial gene RGR (ribozyme-RNA-ribozyme), whose transcripts can undergo self-catalysed cleavage and release the designed gRNA using the RNA polymerase II promoter in vitro and in yeast. The results showed efficient Cas9-mediated target DNA cleavage (Gao and Zhao 2014). Other integrated RNA methods, including RNA-triple-helix structures, introns and microRNAs, have also been attempted in human cells (Nissim et al. 2014), and those methods may also be capable of operating well in filamentous fungi.
RNA technology
Besides the molecular tools used for DNA manipulation, some RNA technology which can verify gene function through disturbing mRNA translation was also applied in filamentous fungi. Those technologies include RNA interference and ribozymes.
RNA interference
RNA interference (RNAi) is an available method for the verification of gene function when the necessary gene knock methods are lacking. In filamentous fungi, RNA quelling was first discovered in N. crassa; six pathways produced various small RNAs (Li et al. 2010). The mechanisms of two RNAi-related phenomena including quelling and meiotic silencing of N. crassa were demonstrated by Romano et al. (Romano and Macino 1992). In summary, the aberrant RNA can trigger cells to inactivate the connate mRNA, thus quelling gene expression in the cell.
The RNAi technology has been successfully used in more than 30 species of filamentous fungi and fungus-like organisms such as Mucor and Aspergillus. Kuck et al. gave the list of filamentous fungi with successful application of the RANi technology in 2010 (Kück and Hoff 2010). Some filamentous fungi, such as Mortierella alpina (Chen et al. 2015), are added into the list in the following years.
The results of RNA interference cannot be predicted and varies among experiments and laboratories. This method is not applicable to some filamentous fungi, such as U. maydis, because the necessary component in the RNAi silencing pathway is absent (Jiang et al. 2013).
Ribozymes
Ribozymes are special RNAs capable of catalysing RNA. Among the different kinds of natural ribozymes, hammerhead ribozymes and hairpin ribozymes are extensively used due to their small size and high cleavage activity. Up to now, only the hammerhead ribozymes were successfully used in filamentous fungi. The first reported case was in Aspergillus giganteus with seven different hammerhead ribozymes designed targeting the mRNA of the beta-glucuronidase transcript (uidA). (Mueller et al. 2006) The result showed that the ribozymes could reduce uidA expression up to 100%.
Although various forms of products for small RNAs are available, such as short-hairpin RNA (shRNA) (Hannon 2002), double-strand RNA transcribed in both directions under dual promoters, antisense single-strand RNA (Hamilton and Baulcombe 1999) and directly synthetic double-strand RNA, different kinds of filamentous fungi have different levels of ability to take up foreign RNAs, thus leading to different silencing effects. Moreover, the heritability of this molecular tool is not stable. Those disadvantages need to be taken into consideration before choosing the right gene manipulation tools.
Summary and outlook
All gene manipulation systems require choices about DNA vectors, transformation methods and gene editing strategies. Gene manipulation plays a key role in this gene function exploration generation. The gene manipulation methods for prokaryotes have developed more quickly than those for filamentous fungi. Because of the complexities of ploidy, propagation and the mycelial structure of filamentous fungi, the search for a versatile, effective and stable genetic tool for use in filamentous fungi faces many challenges. The special composition of fungal cell wall makes it difficult for recombinant DNA to enter the host. These factors may account for the slow progress in the development of molecular genetic tools for use in filamentous fungi. The further development of these molecular tools still faces these difficulties.
As for gene editing strategies, the commonly used homologous recombination method and RNA interference are mainly used to uncover the role of a single gene among multiple pathway genes due to the technological barriers. With the development of gene editing technologies, new tools such as TALENs and the CRISPR/Cas system, which simplify the gene manipulation process and improve gene targeting efficiencies, have arisen to meet the needs of a new generation. Gene manipulation with the CRISPR/Cas9 system in filamentous fungi mainly involves gene mutation, but other types of gene manipulation such as gene motivation and gene interference are likely to be applied in the future. The RNA editing tool C2c2, which is already used in bacteria (Abudayyeh et al. 2016) and was derived from the CRISPR/Cas system, may also be useful in filamentous fungi. The novel CRISPR/Cas systems CRISPR/CasX and CRISPR/CasY (Burstein et al. 2016) offer new choices for the future, and gene editing tools that use DNA-guided artificial nucleases (Qi et al. 2016) may open up new techniques for the gene manipulation field.
Through these gene editing strategies, further eukaryotic regulatory mechanisms of anabolism, catabolism, growth and phenotype can be elucidated and many more filamentous species may become available as ‘cell factories’ for use in the medical, agricultural and food industries.
References
Abudayyeh OO, Gootenberg JS, Konermann S, Joung J, Slaymaker IM, Cox DB, Shmakov S, Makarova KS, Semenova E, Minakhin L, Severinov K, Regev A, Lander ES, Koonin EV, Zhang F (2016) C2c2 is a single-component programmable RNA-guided RNA-targeting CRISPR effector. Science 353(6299):557–567. https://doi.org/10.1126/science.aaf5573
Adachi K, Nelson GH, Peoples KA, Frank SA, Montenegro-Chamorro MV, DeZwaan TM, Ramamurthy L, Shuster JR, Hamer L, Tanzer MM (2002) Efficient gene identification and targeted gene disruption in the wheat blotch fungus Mycosphaerella graminicola using TAGKO. Curr Genet 42(2):123–127. https://doi.org/10.1007/s00294-002-0339-2
Ando A, Sumida Y, Negoro H, Suroto DA, Ogawa J, Sakuradani E, Shimizu S (2009) Establishment of Agrobacterium tumefaciens-mediated transformation of an oleaginous fungus, Mortierella alpina 1S-4, and its application for eicosapentaenoic acid producer breeding. Appl Environ Microbiol 75(17):5529–5536. https://doi.org/10.1128/AEM.00648-09
Arazoe T, Miyoshi K, Yamato T, Ogawa T, Ohsato S, Arie T, Kuwata S (2015) Tailor-made CRISPR/Cas system for highly efficient targeted gene replacement in the rice blast fungus. Biotechnol Bioeng 112(12):2543–2549. https://doi.org/10.1002/bit.25662
Bassett AR, Tibbit C, Ponting CP, Liu JL (2013) Highly efficient targeted mutagenesis of Drosophila with the CRISPR/Cas9 system. Science 4(1):220–228. https://doi.org/10.1016/j.celrep.2013.06.020
Bhairi SM, Staples RC (1992) Transient expression of the ß-glucuronidase gene introduced into Uromyces appendiculatus uredospores by particle bombardment. Phytopathology 82:986-989 https://www.Apsnet.Org/publications/phytopathology/backissues/documents/1992Articles/Phyto82n09_986.PDF accessed 27 July 2017
Boch J, Scholze H, Schornack S, Landgraf A, Hahn S, Kay S, Lahaye T, Nickstadt A, Bonas U (2009) Breaking the code of DNA binding specificity of TAL-type III effectors. Science 326(5959):1509–1512. https://doi.org/10.1126/science.1178811
Briggs AW, Rios X, Chari R, Yang LH, Zhang F, Mali P, Church GM (2012) Iterative capped assembly: rapid and scalable synthesis of repeat-module DNA such as TAL effectors from individual monomers. Nucleic Acids Res 40(15):e117–e126. https://doi.org/10.1093/nar/gks624
Burstein D, Harrington LB, Strutt SC, Probst AJ, Anantharaman K, Thomas BC, Doudna JA, Banfield JF (2016) New CRISPR–Cas systems from uncultivated microbes. Nature 542:237–241. https://doi.org/10.1038/nature21059
Case ME, Schweizer M, Kushner SR, Giles HN (1979) Efficient transformation of Neurospora crassa by utilizing hybrid plasmid DNA. Proc Natl Acad Sci U S A 76(10):5259–5263. http://www.pnas.org/content/76/10/5259
Casselton L, Zolan M (2002) The art and design of genetic screens: filamentous fungi. Nat Rev Genet 3(9):683–697. https://doi.org/10.1038/nrg889
Cermak T, Doyle EL, Christian M, Wang L, Zhang Y, Schmidt C, Baller JA, Somia NV, Bogdanove AJ, Voytas DF (2011) Efficient design and assembly of custom TALEN and other TAL effector-based constructs for DNA targeting. Nucleic Acids Res 39(12):e82–e92. https://doi.org/10.1093/nar/gkr218
Chen HQ, Hao GF, Wang L, Wang HC, Gu ZN, Liu LM, Zhang H, Chen W, Chen YQ (2015) Identification of a critical determinant that enables efficient fatty acid synthesis in oleaginous fungi. Sci Rep 5:11247–11256. https://doi.org/10.1038/srep11247
Cheng AW, Wang H, Yang H, Shi L, Katz Y, Theunissen TW, Rangarajan S, Shivalila CS, Dadon DB, Jaenisch R (2013) Multiplexed activation of endogenous genes by CRISPR-on, an RNA-guided transcriptional activator system. Cell Res 23(10):1163–1171. https://doi.org/10.1038/cr.2013.122
Christian M, Cermak T, Doyle EL, Schmidt C, Zhang F, Hummel A, Bogdanove AJ, Voytas DF (2010) Targeting DNA double-strand breaks with TAL effector nucleases. Genetics 186(2):757–761. https://doi.org/10.1534/genetics.110.120717
Cleto S, Jensen JV, Wendisch VF, Lu TK (2016) Corynebacterium glutamicum metabolic engineering with CRISPR interference (CRISPRi). ACS Synth Biol 5(5):375–385. https://doi.org/10.1021/acssynbio.5b00216
Combier JP, Melayah D, Raffier C, Gay G, Marmeisse R (2003) Agrobacterium tumefaciens-mediated transformation as a tool for insertional mutagenesis in the symbiotic ectomycorrhizal fungus Hebeloma cylindrosporum. FEMS Microbiol Lett 220(1):141–148. https://doi.org/10.1016/s0378-1097(03)00089-2
Daboussi MJ (1997) Fungal transposable elements and genome evolution contemporary issues. In Pierre C (ed) Contemporary issues in genetics and evolution, 1st edn. Springer Netherlands, pp 253–260. https://doi.org/10.1007/978-94-011-4898-6_25
Davis KM, Pattanayak V, Thompson DB, Zuris JA, Liu DR (2015) Small molecule–triggered Cas9 protein with improved genome-editing specificity. Nat Chem Biol 11:316–318. https://doi.org/10.1038/nchembio.1793
Doudna JA, Charpentier E (2014) The new frontier of genome engineering with CRISPR-Cas9. Science 346(6213):1077–1087. https://doi.org/10.1126/science.1258096
de Groot MJ, Bundock P, Hooykaas PJ, Beijersbergen AG (1998) Agrobacterium tumefaciens-mediated transformation of filamentous fungi. Nat Biotechnol 16:839–842. https://doi.org/10.1038/nbt0998-839
de Queiroz MV, Daboussi M-J (2003) Impala, a transposon from Fusarium oxysporum, is active in the genome of Penicillium griseoroseum. FEME Microbiol Lett 218:317–321. https://doi.org/10.1111/j.1574-6968.2003.tb11535.x
Evers B, Jastrzebski K, Heijmans JP, Grernrum W, Beijersbergen RL, Bernards R (2016) CRISPR knockout screening outperforms shRNA and CRISPRi in identifying essential genes. Nat Biotechnol 34(6):631–633. https://doi.org/10.1038/nbt.3536
Fang YF, Tyler BM (2016) Efficient disruption and replacement of an effector gene in the oomycete Phytophthora sojae using CRISPR/Cas9. Mol Plant Pathol 17(1):127–139. https://doi.org/10.1111/mpp.12318
Fincham JR (1989) Transformation in fungi. Microbiol Rev 53(1):148–170
Firon A, Villalba F, Beffa R, d'Enfert C (2003) Identification of essential genes in the human fungal pathogen Aspergillus fumigatus by transposon mutagenesis. Eukaryot Cell 2(2):245–253. https://doi.org/10.1128/ec.2.2.247-255.2003
Florea S, Andreeva K, Machado C, Mirabito PM, Schardl CL (2009) Elimination of marker genes from transformed filamentous fungi by unselected transient transfection with a Cre-expressing plasmid. Fungal Genet Biol 46(10):721–730. https://doi.org/10.1016/j.fgb.2009.06.010
Forment JV, Ramon D, MacCabe AP (2006) Consecutive gene deletions in Aspergillus nidulans: application of the Cre/loxP system. Curr Genet 50(3):217–224. https://doi.org/10.1007/s00294-006-0081-2
Friedland AE, Tzur YB, Esvelt KM, Colaiácovo MP, Church GM, Calarco JA (2013) Heritable genome editing in C. elegans via a CRISPR-Cas9 system. Nat Methods 10:741–742. https://doi.org/10.1038/nmeth.2532
Gaj T, Gersbach CA, Barbas CF (2013) ZFN, TALEN, and CRISPR/Cas-based methods for genome engineering. Trends Biotechnol 31(7):397–405. https://doi.org/10.1016/j.tibtech.2013.04.004
Gao Y, Zhao Y (2014) Self-processing of ribozyme-flanked RNAs into guide RNAs in vitro and in vivo for CRISPR-mediated genome editing. J Integr Plant Biol 56(4):343–349. https://doi.org/10.1111/jipb.12152
Gratz SJ, Cumming AM, Nguyen JN, Hamm DC, Donohue LK, Harrison MM, Wildonger J, O’Connor-Giles KM (2013) Genome engineering of Drosophila with the CRISPR RNA-guided Cas9 nuclease. Genetics 194(4):1029–1035. https://doi.org/10.1534/genetics.113.152710
Guilinger JP, Thompson DB, Liu DR (2014) Fusion of catalytically inactive Cas9 to FokI nuclease improves the specificity of genome modification. Nat Biotechnol 32(6):577–583. https://doi.org/10.1038/nbt.2909
Hamer L, Adachi K, Montenegro-Chamorro MV, Tanzer MM, Mahanty SK, Lo C, Tarpey RW, Skalchunes AR, Heiniger RW, Frank SA, Darveaux BA, Lampe DJ, Slater TM, Ramamurthy L, DeZwaan TD, Nelson GH, Shuster JR, Woessner J, Hamer JE (2001) Gene discovery and gene function assignment in filamentous fungi. Proc Natl Acad Sci U S A 98(9):5110–5115. https://doi.org/10.1073/pnas.091094198
Hamilton AJ, Baulcombe DC (1999) A species of small antisense RNA in posttranscriptional gene silencing in plants. Science 286(5441):950–952. https://doi.org/10.1126/science.286.5441.950
Hannon GJ (2002) RNA interference. Nature 418:244–251. https://doi.org/10.1038/418244a
Hao GF, Chen HQ, Wang L, Gu ZN, Song YD, Zhang H, Chen W, Chen YQ (2014) Role of malic enzyme during fatty acid synthesis in the oleaginous fungus Mortierella alpina. Appl Environ Microbiol 80(9):2762–2768. https://doi.org/10.1128/AEM.00140-14
Hihlal E, Braumann I, van den Berg M, Kempken F (2011) Suitability of Vader for transposon-mediated mutagenesis in Aspergillus niger. Appl and Environ Microbiol 77(7):2332–2335. https://doi.org/10.1128/AEM.02688-10
Hoffmeister D, Keller NP (2007) Natural products of filamentous fungi: enzymes, genes, and their regulation. Nat Prod Rep 24(2):393–416. https://doi.org/10.1039/b603084j
Hooykaas PJ, Beijersbergen AG (1994) The virulence system of Agrobacterium tumefaciens. Annu Rev Phytopathol 32:157–181. https://doi.org/10.1146/annurev.py.32.090194.001105
Hsu PD, Lander ES, Zhang F (2014) Development and applications of CRISPR-Cas9 for genome engineering. Cell 157(6):1262–1278. https://doi.org/10.1016/j.cell.2014.05.010
Hsu PD, Scott DA, Weinstein JA, Ran FA, Konermann S, Agarwala V, Li Y, Fine EJ, Wu X, Shalem O, Cradick TJ, Marraffini LA, Bao G, Zhang F (2013) DNA targeting specificity of RNA-guided Cas9 nucleases. Nat Biotechnol 31:827–832. https://doi.org/10.1038/nbt.2647
Hua SB, Qiu M, Chan E, Zhu L, Luo Y (1997) Minimum length of sequence homology required for in vivo cloning by homologous recombination in yeast. Plasmid 38(2):91–96. https://doi.org/10.1006/plas.1997.1305
Imazaki I, Kurahashi M, Iida Y, Tsuge T (2007) Fow2, a Zn(II)2Cys6-type transcription regulator, controls plant infection of the vascular wilt fungus Fusarium oxysporum. Mol Microbiol 63(3):737–753. https://doi.org/10.1111/j.1365-2958.2006.05554.x
Jiang DW, Zhu W, Wang YC, Sun C, Zhang K-Q, Yang JK (2013) Molecular tools for functional genomics in filamentous fungi: recent advances and new strategies. Biotechnol Adv 31(8):1562–1574. https://doi.org/10.1016/j.biotechadv.2013.08.005
Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E (2012) A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337(6096):816–821 doi:https://doi.org/10.1126/science.1225829
Kahmann R, Basse C (1999) REMI (restriction enzyme mediated integration) and its impact on the isolation of pathogenicity genes in fungi attacking plants. Eur J Plant Pathol 105(3):221–229. https://doi.org/10.1023/A:1008757414036
Katayama T, Tanaka Y, Okabe T, Nakamura H, Fujii W, Kitamoto K, Maruyama J (2016) Development of a genome editing technique using the CRISPR/Cas9 system in the industrial filamentous fungus Aspergillus oryzae. Biotechnol Lett 38(4):637–642. https://doi.org/10.1007/s10529-015-2015-x
Keller NP, Hohn TM (1997) Metabolic pathway gene clusters in filamentous fungi. Fungal Genet and Biol 21:17–29. https://doi.org/10.1006/fgbi.1997.0970
Kim S, Kim D, Cho SW, Kim J, Kim JS (2014) Highly efficient RNA-guided genome editing in human cells via delivery of purified Cas9 ribonucleoproteins. Genome Res 24(6):1012–1019. https://doi.org/10.1101/gr.171322.113
Kleinstiver BP, Prew MS, Tsai SQ, Nguyen NT, Topkar VV, Zheng ZL, Joung JK (2015) Broadening the targeting range of Staphylococcus aureus CRISPR-Cas9 by modifying PAM recognition. Nat Biotechnol 33:1293–1298. https://doi.org/10.1038/nbt.3404
Komari T, Hiei Y, Saito Y, Murai N, Kumashiro T (1996) Vectors carrying two separate T-DNAs for co-transformation of higher plants mediated by Agrobacterium tumefaciens and segregation of transformants free from selection markers. Plant J 10(1):165–174. https://doi.org/10.1046/j.1365-313X.1996.10010165.x
Kunik T, Tzfira T, Kapulnik Y, Gafni Y, Dingwall C, Citovsky V (2001) Genetic transformation of HeLa cells by Agrobacterium. Proc Natl Acad Sci U S A 98(4):1871–1876. https://doi.org/10.1073/pnas.041327598
Kupfer DM, Reece CA, Sandra WC. Clifton, Roe BA, Prade RA (1997) Multicellular ascomycetous fungal genomes contain more than 8000 genes. Fungal Genet Biol 21(3):364–372 doi:https://doi.org/10.1006/fgbi.1997.0982
Kück U, Hoff B (2010) New tools for the genetic manipulation of filamentous fungi. Appl Microbiol Biotechnol 86(1):51–62. https://doi.org/10.1007/s00253-009-2416-7
Kück U, Walz M, Mohr G, Mracek M (1989) The 5′-sequence of the isopenicillin N-synthetase gene (pcbC) from Cephalosporium acremonium directs the expression of the prokaryotic hygromycin B phosphotransferase gene (hph) in Aspergillus niger. Appl Microbiol Biotechnol 31(4):358–365. https://doi.org/10.1007/BF00257605
Larson MH, Gilbert LA, Wang X, Lim WA, Weissman JS, Qi LS (2013) CRISPR interference (CRISPRi) for sequence-specific control of gene expression. Nat Protoc 8:2180–2196. https://doi.org/10.1038/nprot.2013.132
Li L, Chang SS, Liu Y (2010) RNA interference pathways in filamentous fungi. Cell Mol Life Sci 67(22):3849–3862. https://doi.org/10.1007/s00018-010-0471-y
Liu R, Chen L, Jiang YP, Zhou ZH, Zou G (2015) Efficient genome editing in filamentous fungus Trichoderma reesei using the CRISPR/Cas9 system. Cell Discov 1:15007. https://doi.org/10.1038/celldisc.2015.7
Liu YG, Chen Y (2007) High-efficiency thermal asymmetric interlaced PCR for amplification of unknown flanking sequences. Bio Techniques 43:649–656. https://doi.org/10.2144/000112601
Magnuson JK, Lasure LL (2004) Organic acid production by filamentous fungi. In: Tkacz JS, Lange L (eds) Advances in fungal biotechnology for industry, agriculture, and medicine. Springer, Boston. https://doi.org/10.1007/978-1-4419-8859-1_12
Manivasakam P, Schiestl RH (1998) Nonhomologous end joining during restriction enzyme-mediated DNA integration in Saccharyomyces cerevisiae. Mol Cell Biol 18(3):1736–1745. https://doi.org/10.1128/MCB.18.3.1736
Matsu-ura T, Baek M, Kwon J, Hong C (2015) Efficient gene editing in Neurospora crassa with CRISPR technology. Fungal Biol Biotechnol 2(4):1–7. https://doi.org/10.1186/s40694-015-0015-1
Meyer V (2008) Genetic engineering of filamentous fungi—progress, obstacles and future trends. Biotechnol Adv 26(2):177–185. https://doi.org/10.1016/j.biotechadv.2007.12.001
Meyer V, Wu B, Ram AF (2011) Aspergillus as a multi-purpose cell factory: current status and perspectives. Biotechnol Lett 33(3):469–476. https://doi.org/10.1007/s10529-010-0473-8
Michielse CB, Hooykaas PJ, van den Hondel CA, Ram AF (2005) Agrobacterium-mediated transformation as a tool for functional genomics in fungi. Curr Genet 48(1):1–17. https://doi.org/10.1007/s00294-005-0578-0
Mueller D, Stahl U, Meyer V (2006) Application of hammerhead ribozymes in filamentous fungi. J Microbiol Meth 65(3):585–595. https://doi.org/10.1016/j.mimet.2005.10.003
Nissim L, Perli SD, Fridkin A, Perez-Pinera P, Lu TK (2014) Multiplexed and programmable regulation of gene networks with an integrated RNA and CRISPR/Cas toolkit in human cells. Mol Cell 54(4):698–710. https://doi.org/10.1016/j.molcel.2014.04.022
Noh W, Kim S-W, Dong-Won B, Kim JY, Ro HS (2010) Genetic introduction of foreign genes to Pleurotus eryngii by restriction enzyme-mediated integration. J Microbiol 48(2):253–256. https://doi.org/10.1007/s12275-010-9278-7
Nødvig CS, Nielsen JB, Kogle ME, Mortensen UH (2015) A CRISPR-Cas9 system for genetic engineering of filamentous fungi. PLoS One 10(7):18. https://doi.org/10.1371/journal.pone.0133085
Piers KL, Heath JD, Liang X, Stephens KM, Nester EW (1996) Agrobacterium tumefaciens-mediated transformation of yeast. Proc Natl Acad Sci U S A 93(4):1613–1618 http://www.pnas.org/content/93/4/1613.short
Pohl C, Kiel JA, Driessen AJ, Bovenberg RA, Nygard Y (2016) CRISPR/Cas9 based genome editing of Penicillium chrysogenum. ACS Synth Biol 5(7):754–764. https://doi.org/10.1021/acssynbio.6b00082
Punt PJ, van den Hondel CA (1992) Transformation of filamentous fungi based on hygromycin B and phlenomycin resistance markers. Method Enzymol 216:447–457. https://doi.org/10.1016/0076-6879(92)16041-H
Qi JL, Dong ZG, Shi YW, Wang X, Qin YY, Wang YM, Liu D (2016) NgAgo-based fabp11a gene knockdown causes eye developmental defects in zebrafish. Cell Res 26:1349–1352. https://doi.org/10.1038/cr.2016.134
Qi LS, Larson MH, Gilbert LA, Doudna JA, Weissman JS, Arkin AP, Lim WA (2013) Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression. Cell 152(5):1173–1183. https://doi.org/10.1016/j.cell.2013.02.022
Ran FA, Hsu PD, Wright J, Agarwala V, Scott DA, Zhang F (2013a) Genome engineering using the CRISPR-Cas9 system. Nat Protoc 8:2281–2308. https://doi.org/10.1038/nprot.2013.143
Ran FA, Hsu PD, Lin CY, Gootenberg JS, Konermann S, Trevino AE, Scott DA, Inoue A, Matoba S, Zhang Y, Zhang F (2013b) Double nicking by RNA-guided CRISPR Cas9 for enhanced genome editing specificity. Cell 154(6):1380–1389. https://doi.org/10.1016/j.cell.2013.08.021
Richey MG, Marek ET, Schardl CL, Smith DA (1989) Transformation of filamentous fungi with plasmid DNA by electroporation. Phytopathology 79:844–847 https://wwwapsnetorg/publications/phytopathology/backissues/Documents/1989Articles/Phyto79n08_844pdf Acessed (27 July 2017)
Rogers CW, Challen MP, Green JR, Whipps JM (2004) Use of REMI and Agrobacterium-mediated transformation to identify pathogenicity mutants for the biocontrol fungus, Coniothyrium minitans. FEMS Microbiol Lett 241:207–214. https://doi.org/10.1016/j.femsle.2004.10.022
Romano N, Macino G (1992) Quelling: transient inactivation of gene expression in Neurospora crassa by transformation with homologous sequences. Mol Microbiol 6(22):3343–3353. https://doi.org/10.1111/j.1365-2958.1992.tb02202.x
Schiestl RH, Petes TD (1991) Integration of DNA fragments by illegitimate recombination in Saccharomyces cerevisiae. Proc Natl Acad Sci U S A 88(17):7585–7589 http://www.pnas.org/content/88/17/7585.full.pdf
Schmid-Burgk JL, Schmidt T, Vera K, Höning K, Hornung V (2013) A ligation-independent cloning technique for high-throughput assembly of transcription activator-like effector genes. Nat Biotechnol 31:76–81. https://doi.org/10.1038/nbt.2460
Schuster M, Schweizer G, Reissmann S, Kahmann R (2016) Genome editing in Ustilago maydis using the CRISPR-Cas system. Fungal Genet Biol 89:3–9. https://doi.org/10.1016/j.fgb.2015.09.001
Shi Z, Christian D, Leung H (1995) Enhanced transformation in Magnaporthe grisea by restriction enzyme mediated integration of plasmid DNA. Phytopathology 85(3):329-333 https://www.apsnet.org/publications/phytopathology/backissues/Documents/1995Articles/Phyto85n03_329.Ppdf Accessed 27 July 2017
Shi Z, Leung H (1995) Genetic analysis of sporulation in Magnaporthe grisea sporulation by chemical and insertional mutagenesis. Mol Plant-Microbe Inter 8(6):949–958. https://doi.org/10.1094/MPMI-8-0949. Accessed 27 July 2017
Shalem O, Sanjana NE, Hartenian E, Shi X, Scott DA, Mikkelsen TS, Heckl D, Ebert BL, Root DE, Doench JG, Zhang F (2014) Genome-scale CRISPR-Cas9 knockout screening in human cells. Science 343(6166):84–87. https://doi.org/10.1126/science.1247005
Slaymaker IM, Gao LY, Zetsche B, Scott DA, Yan WX, Zhang F (2016) Rationally engineered Cas9 nucleases with improved specificity. Science 351(6268):84–88. https://doi.org/10.1126/science.aad5227
Sun N, Zhao H (2013) Transcription activator-like effector nucleases (TALENs): a highly efficient and versatile tool for genome editing. Biotechnol Bioeng 110(7):1811–1821. https://doi.org/10.1002/bit.24890
Sweigard JA, Carroll AM, Farrall L, Chumley FG, Valent B (1998) Magnaporthe grisea pathogenicity genes obtained through insertional mutagenesis. Mol Plant Microbe In 11(5):404–412. https://doi.org/10.1094/MPMI.1998.11.5.404
Sørensen LM, Jacobsen T, Nielsen PV, Frisvad JC, Koch AG (2008) Mycobiota in the processing areas of two different meat products. Int J Food Microbiol 124(1):58–64. https://doi.org/10.1016/j.ijfoodmicro.2008.02.019
Tang X, Zan XY, Zhao LN, Chen HQ, Chen YQ, Chen W, Song YD, Ratledge C (2016) Proteomics analysis of high lipid-producing strain Mucor circinelloides WJ11: an explanation for the mechanism of lipid accumulation at the proteomic level. Microb Cell Factories 15:35. https://doi.org/10.1186/s12934-016-0428-4
Tingay S, McElroy D, Kalla R, Fieg S, Wang M, Thornton S, Brettell R (1997) Agrobacterium tumefaciens-mediated barley transformation. Plant J 11(6):1369–1376. https://doi.org/10.1046/j.1365-313X.1997.11061369.x
Tsai SQ, Wyvekens N, Khayter C, Foden JA, Thapar V, Reyon D, Goodwin MJ, Aryee MJ, Joung JK (2014) Dimeric CRISPR RNA-guided FokI nucleases for highly specific genome editing. Nat Biotechnol 32:569–576. https://doi.org/10.1038/nbt.2908
Tsuji G, Fujii S, Fujihara N, Hirose C, Tsuge S, Shiraishi T, Kubo Y (2003) Agrobacterium tumefaciens-mediated transformation for random insertional mutagenesis in Colletotrichum lagenarium. J Gen Plant Pathol 69(4):230–239. https://doi.org/10.1007/s10327-003-0040-4
Turgeon BG, Condon B, Liu J, Zhang N (2010) Protoplast transformation of filamentous fungi. Methods Mol Biol 638:3–19. https://doi.org/10.1007/978-1-60761-611-5_1
Valvekens D, Montagu MV, Lijsebettens MV (1988) Agrobacterium tumefaciens-mediated transformation of Arabidopsis thaliana root explants by using kanamycin selection. Proc Natl Acad Sci U S A 85(15):5536–5540 http://www.pnas.org/content/85/15/5536.full.pdf
Wang JY, Li HY (2008) Agrobacterium tumefaciens-mediated genetic transformation of the phytopathogenic fungus Penicillium digitatum. J Zhejiang Univ Sci B 9(10):823–828. https://doi.org/10.1631/jzus.B0860006
Wang T, Wei JJ, Sabatini DM, Lander ES (2014) Genetic screens in human cells using the CRISPR-Cas9 system. Science 343(6166):80–84. https://doi.org/10.1126/science.1246981
Woo JW, Kim J, Kwon S, Corvalán C, Cho SW, Kim H, Kim SG, Kim ST, Choe S, Kim JS (2015) DNA-free genome editing in plants with preassembled CRISPR-Cas9 ribonucleoproteins. Nat Biotechnol 33:1162–1164 doi:https://doi.org/10.1038/nbt.3389
Xu J, He MM, Mo MH, Huang XW, Zhang KQ (2005) Transformation and mutagenesis of the nematode-trapping fungus Monacrosporium sphaeroides by restriction enzyme mediated integration (REMI). J Microbiol 43(5):417–423
Funding
This study was supported by the National Science Foundation of China (NSFC) (31722041, 31530056), the Fundamental Research Funds for the Central Universities (JUSRP51702A) and the program of ‘Collaborative innovation center of food safety and quality control in Jiangsu Province’.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with animals performed by any of the authors.
Rights and permissions
About this article
Cite this article
Wang, S., Chen, H., Tang, X. et al. Molecular tools for gene manipulation in filamentous fungi. Appl Microbiol Biotechnol 101, 8063–8075 (2017). https://doi.org/10.1007/s00253-017-8486-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-017-8486-z