Abstract
Chitin is the second most abundant natural polysaccharide after cellulose. But degradation of chitin has never been reported in haloarchaea. In this study, we revealed that Haloferax mediterranei, a metabolically versatile haloarchaeon, could utilize colloidal or powdered chitin for growth and poly(3-hydroxybutyrate-co-3-hydroxyvalerate) (PHBV) accumulation, and the gene cluster (HFX_5025-5039) for the chitin catabolism pathway was experimentally identified. First, reverse transcription polymerase chain reaction results showed that the expression of the genes encoding the four putative chitinases (ChiAHme, ChiBHme, ChiCHme, and ChiDHme, HFX_5036-5039), the LmbE-like deacetylase (DacHme, HFX_5027), and the glycosidase (GlyAHme, HFX_5029) was induced by colloidal or powdered chitin, and chiA Hme, chiB Hme, and chiC Hme were cotranscribed. Knockout of chiABC Hme or chiD Hme had a significant effect on cell growth and PHBV production when chitin was used as the sole carbon source, and the chiABCD Hme knockout mutant lost the capability to utilize chitin. Knockout of dac Hme or glyA Hme also decreased PHBV accumulation on chitin. These results suggested that ChiABCDHme, DacHme, and GlyAHme were indeed involved in chitin degradation in H. mediterranei. Additionally, the chitinase assay showed that each chitinase possessed hydrolytic activity toward colloidal or powdered chitin, and the major product of colloidal chitin hydrolysis by ChiABCDHme was diacetylchitobiose, which was likely further degraded to monosaccharides by DacHme, GlyAHme, and other related enzymes for both cell growth and PHBV biosynthesis. Taken together, this study revealed the genes and enzymes involved in chitin catabolism in haloarchaea for the first time and indicated the potential of H. mediterranei as a whole-cell biocatalyst in chitin bioconversion.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
With the urgent need of transition to a more environmentally sustainable bioeconomy, the biocatalysts that can effectively convert the recalcitrant organic carbon polymers into useful compounds have attracted special interest (Himmel et al. 2007; Vaaje-Kolstad et al. 2010). Chitin is a highly insoluble polymer composed of linear chains of β-1,4-linked N-acetyl-d-glucosamine (GlcNAc) residues, which are tightly held by a large number of intra- and inter-chain hydrogen bonds. Chitin is widespread in nature as a structural component of insect exoskeletons, crustacean shells, and cell walls of most fungi and some algae. Chitin is the second most abundant natural polymer after cellulose and has an estimated biological production of more than 1011 metric tons per year in the aquatic ecosystems alone (Lutz et al. 1994). A large amount of chitin waste is generated from the seafood industry, which has caused severe environmental problems. Thus, bioconversion of chitin waste to value-added products is environmentally and economically beneficial (Songsiriritthigul et al. 2010).
Chitinases catalyze the first step of chitin degradation by cleaving the β-1,4-glycosidic bonds of the chitin chain. According to the amino acid sequence similarity, chitinases fall into either glycoside hydrolase (GH) family 18 or 19 (Tsuji et al. 2010). GH family 18 chitinases are distributed in various organisms, including microbes, plants, and animals, and have a relatively high sequence divergence. Thus, these chitinases are further divided into three subfamilies of A, B, and C (Karlsson and Stenlid 2009). Conversely, GH family 19 chitinases are almost exclusively from plants and possess a high degree of sequence similarity (Tsuji et al. 2010). Additionally, chitinases are classified as endochitinases or exochitinases based on their mode of action. During the degradation of chitin, endochitinases first cleave the β-1,4-glycosidic bonds randomly at the internal sites of the chitin chain to provide a variety of chitooligosaccharides. The exochitinases then remove the GlcNAc2 units from the non-reducing end of the chitooligosaccharides in a progressive manner (Cohen-Kupiec and Chet 1998). Chitinases from bacteria and eukarya have been well studied, but only a limited number of archaeal chitinases have been characterized. To our knowledge, only five archaeal strains, including Thermococcus chitinophagus (Andronopoulou and Vorgias 2004), Thermococcus kodakaraensis KOD1 (Tanaka et al. 2001), Pyrococcus furiosus (Gao et al. 2003; Kreuzer et al. 2013), Sulfolobus tokodaii (Staufenberger et al. 2012), and Halobacterium salinarum strain NRC-1 (Yoshinobu et al. 2006), have been reported to possess chitinases, which all belong to GH18A or GH18C. It is noteworthy that studies regarding these archaeal chitinases have focused primarily on biochemical analysis and structural determination (Nakamura et al. 2008; Oku and Ishikawa 2006; Tsuji et al. 2010).
The chitin catabolism pathway in bacteria and eukarya has also been well characterized; however, the archaeal chitin catabolism pathway has been established only in T. kodakaraensis KOD1 (Tanaka et al. 2004), in which the concerted action of three key enzymes of ChiATk (chitinase, TK1765), DacTk (deacetylase, TK1764), and GlmATk (exo-β-glucosaminidase, TK1754) converts chitin into glucosamine (GlcN). All of the chitin catabolism-related genes cluster together in T. kodakaraensis KOD1. The chitin degradation pathway identified in T. kodakaraensis KOD1 is different from those of bacteria or eukarya. Whether there are alternative chitin degradation pathways in archaea remains to be determined.
Haloarchaea represent a distinct archaeal group that inhabits hypersaline environments. Knowledge regarding haloarchaeal chitin utilization is very limited; so far, only a chitinase homolog in H. salinarum strain NRC-1 has been heterogeneously expressed and characterized (Yoshinobu et al. 2006). Haloferax mediterranei is a good model for the study of haloarchaeal metabolism (Han et al. 2012), as it can utilize a variety of compounds as the sole carbon source, including carbohydrates, polyalcohols, organic acids, and amino acids (Rodriguez-Valera et al. 1983). In the presence of excessive carbon sources, H. mediterranei is capable of accumulating a large amount of poly(3-hydroxybutyrate-co-3-hydroxyvalerate) (PHBV) (Lu et al. 2008b), a desired bioplastic with a variety of applications such as packaging and medical materials. Interestingly, bioinformatic analysis of the H. mediterranei genome revealed a gene cluster (HFX_5025-5039) that might encode the chitin catabolism pathway. Whether H. mediterranei could use chitin for growth and convert it into PHBV remained to be determined.
This study revealed for the first time that H. mediterranei was able to use colloidal or powdered chitin for cell growth and PHBV production. Notably, four important chitinases involved in chitin degradation by H. mediterranei were identified through bioinformatic prediction, transcription analysis, genetic verification, chitinase assay, and detection of the chitinolytic products. In addition, two other key enzymes, the LmbE-like deacetylase and the glycosidase, were also verified to be involved in chitin utilization by H. mediterranei. Taken together, this study has enriched our understanding of the enzymes in haloarchaeal chitin catabolism and provided the first evidence for the potential of haloarchaea in chitin bioconversion, especially in hypersaline environments.
Materials and methods
Strains and culture conditions
The strains used in this study are listed in Table 1. Colloidal chitin was prepared as described by Tanaka et al. (1999), except that concentrated hydrochloric acid rather than 85 % phosphoric acid was used to dissolve the powdered chitin. Escherichia coli JM109 was cultivated in Luria–Bertani medium at 37 °C (Sambrook et al. 1989). When needed, 100 μg/ml ampicillin was added to the media. H. mediterranei was first cultured at 37 °C in AS-168 medium (Han et al. 2007) for 36 h as seed culture. Subsequently, 2.5 ml of the seed culture was harvested, washed with sterile NaCl solution, and inoculated into 50 ml basal medium (per liter, 200 g NaCl, 20 g MgSO4 · 7H2O, 2 g KCl, 2 g NH4Cl, 37.5 mg KH2PO4, 5 mg FeSO4 · 7H2O, 0.036 mg MnCl2 · 4H2O, 15 g PIPES, pH 7.2) with 5 g/l glucose or colloidal chitin or powdered chitin (Sangon Biotech, Shanghai, China) as the sole carbon source for additional cultivation of 24 h (for reverse transcription polymerase chain reaction, RT-PCR) or until stationary phase (for growth monitoring and PHBV accumulation analysis in H. mediterranei strains). In the experiment of comparing the PHBV production from glucose, colloidal chitin, and powdered chitin, the culture conditions were the same as described above, except that 1 g/l sodium glutamate was added to the basal medium (modified basal medium) for a short lag phase. For H. mediterranei DF50 (a uracil auxotroph of the wild-type strain) and the DF50-based knockout mutants, 50 μg/ml uracil was added into the media (Liu et al. 2011). Cell numbers (expressed as colony-forming units (CFU) per milliliter) were determined by plating an aliquot of the diluted cell culture onto the surface of AS-168 Petri plates, which were then incubated at 37 °C for about 72 h. The colonies were counted and the cell numbers in the original culture were determined.
RT-PCR
Total RNA of H. mediterranei was isolated with the TRIzol reagent (InvitrogenTM-Life Technologies, USA). RT-PCR was performed as described previously (Han et al. 2007), except that the primers for the RT reaction were random hexamer primers (Thermo Scientific) and the specific primers used for the PCR reactions are listed in Table S1.
Knockout mutant construction and verification
Gene knockout was performed as previously described (Lu et al. 2008b). The primers and plasmids used for gene knockout are listed in Table S1 and Table 1, respectively. Briefly, the fragments (approximately 650 bp for each) located immediately upstream or downstream of the target gene were amplified, sequenced, and inserted into the plasmid pHFX (Liu et al. 2011). The resulting plasmids were then transformed into the host strain H. mediterranei DF50 to delete the target gene through homologous recombination. The knockout mutant strains were all verified by PCR using the primers shown in Table S1.
PHBV analysis
The PHBV content and its monomer composition in the cells were determined by gas chromatography as described previously (Han et al. 2007). Benzoic acid was used as the internal standard for quantitative analysis of the PHBV content.
Protein preparation and chitinase assay
For chitinase assay, four plasmids (pWLA, pWLB, pWLC, and pWLD) listed in Table 1 were constructed for the expression of ChiAHme, ChiBHme, ChiCHme, and ChiDHme, respectively. The coding regions under the hsp5 promoter (Lu et al. 2008a), without putative signal peptides, were amplified and inserted into the plasmid pWL502 (Cai et al. 2012) to generate the expression plasmids. These plasmids were then transformed individually into the ΔchiABCD Hme mutant strain. The resulting strains were first cultivated in AS-168 medium without yeast extract and then inoculated as 5 % of the total volume into the basal medium (as described in the “Strains and culture conditions” section) with glucose as the sole carbon source for an additional 36 h of cultivation. These cells were harvested by centrifugation (10,000×g for 10 min at 4 °C) and then suspended with high salt buffer (HSB; 3.4 M KCl in 20 mM Tris–HCl (pH 7.5)) for ultrasonication. Intact cells and debris were removed by centrifugation (10,000×g for 10 min at 4 °C) to obtain crude extracts. The protein concentration of the crude extracts was determined by a bicinchoninic acid protein assay kit (Thermo Scientific). The standard chitinase assay mixture contained 100 μg of crude protein and 3 mg of colloidal or powdered chitin in the HSB in a total volume of 500 μl. After incubation at 37 °C for 15 min, the reaction was terminated by boiling for 3 min, and the reaction mixture was centrifuged to obtain the supernatant. The amount of liberated reducing sugar was measured with a modified Schales procedure (Imoto and Yagishita 1971). One unit of activity was defined as the amount of protein that produces 1 μmol of reducing sugars per minute. The specific activity was expressed as unit per gram crude protein. The chitinase assay with the crude extracts from H. mediterranei DF50 ΔchiABCD Hme harboring the plasmid pWL502 was used as the negative control.
Analysis of the chitinolytic products by high-performance anion exchange chromatography
The chitinase assay was performed as described in the “protein preparation and chitinase assay” section, except that the crude extracts used were the mixture of the four crude extracts containing ChiAHme, ChiBHme, ChiCHme, and ChiDHme, respectively; the substrate was 3 mg of colloidal chitin, and the reaction time was extended to 1 h. The high-performance anion exchange chromatography (HPAEC) analysis of the hydrolysis products was performed as described previously (Lü et al. 2009). The reaction mixture containing the crude extracts from the ΔchiABCD Hme strain with the plasmid pWL502 and 3 mg of colloidal chitin in HSB was used as the negative control. The standards of GlcNAc, GlcNAc2, and GlcNAc3 (Northstar BioProductsTM-Associates of Cape Cod, USA) were used to calibrate the retention time.
Protein sequence analysis and phylogenetic tree construction
Signal peptides were predicted with the SignalP 4.0 server (http://www.cbs.dtu.dk/services/SignalP/). Domain annotation was performed by using the NCBI conserved domain search service (http://www.ncbi.nlm.nih.gov/Structure/cdd/wrpsb.cgi), InterProScan program (http://www.ebi.ac.uk/Tools/pfa/iprscan/), and referring to the CAZy database (http://www.cazy.org/). Protein modeling was performed on the SWISS-MODEL server using the automated mode (http://swissmodel.expasy.org/workspace/index.php?func=modelling_simple1). Sequence homology was analyzed using the BLAST service (http://blast.ncbi.nlm.nih.gov/Blast.cgi) and the GeneDoc program (http://www.nrbsc.org/gfx/genedoc/). The phylogenetic tree was constructed in MEGA4 software using the neighbor-joining method. The topology of the phylogenetic tree was evaluated by bootstrap analysis on the basis of 1,000 replicates.
Results
Bioinformatic analysis of putative chitinases in H. mediterranei
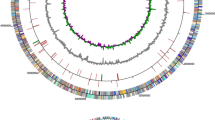
Our recent genome sequencing of H. mediterranei (Han et al. 2012) revealed a gene cluster that might encode the enzymes involved in chitin catabolism. This gene cluster (HFX_5025-5039) includes the genes encoding four potential chitinases (HFX_5036-5039), a putative ABC transport system (HFX_5025, HFX_5030-5032), a glycosidase (HFX_5029), and an LmbE-like deacetylase (HFX_5027), which might be directly involved in chitin degradation. In addition, a putative galactonate dehydratase (HFX_5033) and a glucose-1-dehydrogenase (HFX_5035) were also found in this gene cluster (Fig. 1a and Table S2).
Bioinformatic analysis of the four putative chitinases in H. mediterranei. a The putative chitin catabolism-related gene cluster in H. mediterranei. The locus tag of each gene is indicated. The corresponding gene names are listed in Table S2. b The schematic representation of the domain organization in the four chitinases. The dark line in the catalytic domain represents the conserved “DXDXE” motif in the GH family 18 chitinases
The four putative chitinases, which were designated as ChiAHme, ChiBHme, ChiCHme, and ChiDHme, respectively, may catalyze the first step of chitin degradation by H. mediterranei. Among them, ChiAHme and ChiBHme were more closely related (45 % identity), and ChiCHme and ChiDHme were more similar (41 % identity). The protein sequence analysis revealed that the four chitinases all harbored a signal sequence in the N terminus, followed by a chitin binding domain (ChtBD; accession number IPR003610), a polycystic kidney disease domain (PKD; accession number IPR000601), and a C-terminal catalytic domain (CD; accession number IPR017853) (Fig. 1b). The N-terminal signal sequence may direct the chitinase to the outside of the cell. The ChtBD, a member of the carbohydrate-binding module family 5/12 (CBM5/12), may be responsible for concentrating the CD on the chitin surface and disrupting the hydrogen bonds of the chitin chains (Nakamura et al. 2008). PKD was reported to be involved in the degradation of powdered chitin in a marine bacterium (Orikoshi et al. 2005). The CDs of all the four chitinases adopted the triosephosphate isomerase (β/α)8-barrel fold (Fig. S1a) and had a conserved sequence motif of “DXDXE” (Fig. 1b and Fig. S1b), which are characteristics of GH family 18 chitinases (Tsuji et al. 2010). In addition to these domains, ChiCHme and ChiDHme contained an additional chitinase insertion domain (CID; accession number SSF54556) in the CD (Fig. 1b). The CID was found to occur in the bacterial exochitinases and may enhance the enzyme activity of exochitinases by increasing the depth of the substrate-binding cleft (Li and Greene 2010). Hence, ChiCHme and ChiDHme might be exochitinases, and ChiAHme and ChiBHme might be endochitinases. Phylogenetic tree analysis of the chitinases from archaea and the chitinases with definitive classifications revealed that the archaeal chitinases belonged to either GH18A or GH18C subfamily. ChiCHme and ChiDHme were members of GH18A subfamily, while ChiAHme and ChiBHme belonged to GH18C (Fig. 2 and Table S3). This result was consistent with a previous report that the CID seemed specific to GH18A (Li and Greene 2010).
The phylogenetic tree of the chitinases from archaea and the chitinases with certain subfamily classifications of GH18A, GH18B, GH18C, and GH19. Numbers at nodes represent the percentage bootstrap values based on 1,000 replicates. Only values above 70 % are considered significant and are shown here. The scale bar indicates a difference of 0.2 substitutions per site. The accession numbers of these proteins are shown in Table S3
Bioconversion of chitin to PHBV by H. mediterranei
To test the capability of chitin utilization by H. mediterranei, the commercially available powdered chitin or the hydrochloric acid-treated colloidal chitin was used as the carbon source. During the colloidal chitin preparation, some hydrogen bonds and glycosidic bonds are disrupted; thus, colloidal chitin has a lower degree of crystallinity and polymerization than powdered chitin (Kurita 2006). Such structural changes in the colloidal chitin make it more susceptible to degradation. H. mediterranei was found to be able to grow on colloidal or powdered chitin, and it could also produce PHBV from colloidal or powdered chitin. The final PHBV concentration from colloidal chitin was nearly as high as that from glucose in shake flask culture (1 g/l sodium glutamate added only in this experiment, see “Materials and methods”), although the accumulation rate was slower from colloidal chitin than that from glucose during the first 48 h (Fig. 3). This relatively slow accumulation rate is possibly due to the need of a degradation process, which converts the polymeric chitin into usable monomers. A lower PHBV concentration was obtained from powdered chitin than from colloidal chitin (Fig. 3), possibly because the highly crystalline structure of powdered chitin is not readily accessible to the chitinases. These results show for the first time that H. mediterranei can efficiently utilize colloidal or powdered chitin as a carbon source for PHBV production.
The time course of PHBV accumulation from glucose (filled square), colloidal chitin (empty circle), or powdered chitin (empty triangle) in H. mediterranei wild-type strain. H. mediterranei cells were cultivated in modified basal medium with 5 g/l glucose, or colloidal chitin, or powdered chitin, respectively (see “Materials and methods”)
Induced expression of the four chitinase genes
To investigate the involvement of the four putative chitinases (ChiAHme, ChiBHme, ChiCHme, and ChiDHme) in chitin degradation, RT-PCR was performed to examine their transcripts under the growth conditions with different carbon sources, including glucose, colloidal chitin, and powdered chitin. The results showed that all of the four chitinase genes were transcribed when colloidal or powdered chitin was provided as the sole carbon source, but not when glucose was used as the carbon source (Fig. 4a). These results indicated that these four chitinase genes were induced by colloidal or powdered chitin and might participate in chitin catabolism. Moreover, the RT-PCR results suggested that chiA Hme, chiB Hme, and chiC Hme were cotranscribed, while chiD Hme was transcribed alone (Fig. 4b). This is a very interesting transcriptional profile. But the transcriptional regulation mechanism of the four chitinase genes remains to be clarified.
RT-PCR analysis of the transcripts of the four chitinase genes. a Expression of the four chitinase genes under different culture conditions. b Transcriptional pattern of the four chitinase genes. The primer positions (F1/R1 and F2/R2) are indicated in the gene cluster of chiABCD Hme. Lanes M marker; lanes Glu, Col, and Pow the reverse transcripts of total RNA from H. mediterranei grown with glucose (Glu), colloidal chitin (Col), and powdered chitin (Pow), respectively, as the template; lanes + genome as the template, positive control; lanes − negative control. The primers used here are listed in Table S1
Genetic determination of the involvement of ChiABCDHme in chitin utilization
Based on the transcriptional pattern of the four chitinase genes, three knockout mutants were constructed using H. mediterranei DF50 (Liu et al. 2011) as the host strain. These mutants included the knockout of cotranscribed chiA Hme, chiB Hme, and chiC Hme in combination, the knockout of chiD Hme, and the knockout of the gene cluster of chiABCD Hme, which were named ΔchiABC Hme, ΔchiD Hme, and ΔchiABCD Hme, respectively. A growth curve was constructed by plotting the CFU per milliliter versus time. As shown in the growth curves, H. mediterranei DF50 was able to grow well with colloidal or powdered chitin as the sole carbon source, while the two knockout mutants, ΔchiABC Hme and ΔchiD Hme, could hardly grow on the powdered chitin and ΔchiABCD Hme could grow on neither powdered nor colloidal chitin (Fig. 5a). These results suggested that the four chitinases were indeed involved in the degradation of colloidal and powdered chitin by H. mediterranei, and both ChiABCHme and ChiDHme were indispensible for the utilization of powdered chitin.
Utilization of colloidal or powdered chitin for growth and PHBV accumulation by H. mediterranei DF50 and the recombinant strains. a The growth curves of the H. mediterranei strains (DF50, ΔchiABC Hme, ΔchiD Hme, and ΔchiABCD Hme) grown with colloidal (Col) or powdered (Pow) chitin as the sole carbon source were constructed by plotting the CFU per milliliter versus time. DF50 with Pow, filled square; DF50 with Col, empty circle; ΔchiABC Hme with Pow, empty inverted triangle; ΔchiD Hme with Pow, filled triangle; ΔchiABCD Hme with Col, empty diamond; ΔchiABCD Hme with Pow, cross. b The final PHBV concentration of H. mediterranei strains (DF50, ΔchiABC Hme, ΔchiD Hme, and ΔchiABCD Hme) at the stationary phase (maximum PHBV production) from colloidal (dark gray columns) or powdered (light gray columns) chitin. The data in b represent mean values ± SD of three independent experiments
As PHBV is deposited as a kind of intracellular carbon and energy storage compound in the presence of excessive carbon source, the PHBV accumulation capabilities could to some degree reflect the carbon source utilization capabilities of the strains. Thus, the final PHBV accumulation of H. mediterranei DF50 and the chitinase knockouts at the stationary phase (maximum PHBV production) were investigated when chitin was used as the sole carbon source. It was found that when colloidal chitin was used as the sole carbon source, ΔchiABC Hme and ΔchiD Hme strains accumulated less PHBV than DF50. Specifically, knockout of chiD Hme resulted in a 28 % decrease in the final PHBV concentration from colloidal chitin compared with DF50, and the final PHBV concentration in ΔchiABC Hme was reduced by 86 % (Fig. 5b). As for the ΔchiABCD Hme strain, no PHBV was detected, which was consistent with no observed growth of this strain on colloidal chitin (Fig. 5a, b). When powdered chitin was used as the sole carbon source, PHBV was only detected in the DF50 strain but not in the three chitinase knockouts, which was in agreement with the result that the three chitinase knockouts could not grow on powdered chitin at all (Fig. 5a, b). These results showed that ChiDHme and the combination of ChiAHme, ChiBHme, and ChiCHme could degrade colloidal chitin individually, but the chitin degradation efficiency of ChiDHme was lower than the combination of ChiAHme, ChiBHme, and ChiCHme. Furthermore, the presence of all the four chitinases was more efficient in degrading colloidal chitin than the combination of ChiAHme, ChiBHme, and ChiCHme, suggesting a combined action of these four chitinases in colloidal chitin degradation. Additionally, only the combined action of the four chitinases could produce enough usable carbon from powdered chitin for growth and PHBV accumulation. Therefore, the growth curves and PHBV accumulation analysis both confirmed the significance of the four chitinases in chitin degradation.
Enzyme activity of the four chitinases and detection of the colloidal chitin hydrolysis products
To biochemically demonstrate the functions of the four chitinases in chitin degradation in H. mediterranei, the chitinase activity was assayed with colloidal or powdered chitin as the substrate. The four chitinase genes, depleted of the signal sequence for intracellular expression, were introduced individually under the hsp5 promoter (Lu et al. 2008a) into the host strain ΔchiABCD Hme. The resulting strains were cultivated with glucose as the sole carbon source, and under this condition, the chitin catabolism-related genes except the introduced chiA Hme, chiB Hme, chiC Hme, or chiD Hme were not transcribed. The crude extracts were subject to the chitinase assay. All of the four crude extracts with chitinases showed hydrolytic activities toward both colloidal and powdered chitin (Fig. 6a). The crude extracts with ChiDHme showed a specific activity of 253.2 U/g crude protein toward colloidal chitin and 140.1 U/g crude protein toward powdered chitin, and ChiCHme showed a specific activity of 135.6 U/g crude protein toward colloidal chitin and 49.7 U/g crude protein toward powdered chitin. The specific activity of the crude extracts with ChiBHme was 90.4 U/g crude protein with colloidal chitin and 13.6 U/g crude protein with powdered chitin, respectively. As for the crude extracts containing ChiAHme, it exhibited a specific activity of 31.6 U/g crude protein with colloidal chitin and 4.5 U/g crude protein with powdered chitin (Fig. 6a). In addition, the reaction products from hydrolysis of colloidal chitin were also detected using HPAEC. Hydrolysis of colloidal chitin by the four chitinases produced a mixture of GlcNAc, GlcNAc2, and GlcNAc3, with the diacetylchitobiose GlcNAc2 being the dominant product, whereas no hydrolysis product was generated in the chitinase-negative control (Fig. 6b). Therefore, the role of the four chitinases in chitin degradation was further confirmed by these in vitro results.
Biochemical determination of the four chitinases. a The specific activities of the four chitinases toward colloidal (dark gray columns) or powdered (light gray columns) chitin. The experiment was performed in triplicate and the representative data are shown here. b HPAEC detection of the colloidal chitin hydrolysis products. The product was the hydrolysis product of colloidal chitin catalyzed by the mixture of crude extracts with ChiAHme, ChiBHme, ChiCHme, and ChiDHme. The spectrum of standards of GlcNAc, GlcNAc2, and GlcNAc3 is shown. The product from the assay with colloidal chitin and the crude extracts of ΔchiABCD Hme harboring pWL502 was set as negative control
The role of DacHme and GlyAHme in chitin utilization
In addition to the four chitinases, the uncharacterized LmbE-like deacetylase (DacHme, HFX_5027) and the glycosidase (GlyAHme, HFX_5029) might also play important roles in chitin utilization for H. mediterranei. To determine the involvement of dac Hme and glyA Hme in chitin utilization by H. mediterranei, RT-PCR analysis was performed at first. The results showed that the expression of dac Hme and glyA Hme was induced by colloidal or powdered chitin, but not by glucose (Fig. 7a), indicating that dac Hme and glyA Hme might be involved in chitin utilization. Additionally, the knockout mutants of dac Hme and glyA Hme were constructed to examine their capability to utilize chitin, which could be reflected by the PHBV accumulation capability from chitin, as mentioned above. Hence, the PHBV concentration of the strains of DF50, Δdac Hme, and ΔglyA Hme at the stationary phase (maximum PHBV production) was obtained when chitin was used as the sole carbon source. The results showed that deletion of dac Hme led to dramatic decreases in the final PHBV concentration from colloidal chitin (a reduction of 81 %) and powdered chitin (a reduction of 91 %) compared with DF50 (Fig. 7b). A small amount of PHBV was still produced from colloidal or powdered chitin by the Δdac Hme strain, which may be due to the presence of a small number of deacetylated units of GlcN in the chitin chain (Mathur and Narang 1990) or the presence of other proteins with the same function as DacHme. Additionally, the ΔglyA Hme strain could only produce a very small amount of PHBV from colloidal chitin (approximately 0.03 g/l) and none from powdered chitin (Fig. 7b). These results indicated the important roles of DacHme and GlyAHme in chitin utilization by H. mediterranei. But the function of DacHme and GlyAHme in the degradation of chitin remains to be determined. Identification of these key enzymes in chitin utilization would advance our understanding of the chitin catabolism in haloarchaea.
Demonstration of the involvement of dac Hme and glyA Hme in chitin utilization by H. mediterranei. a RT-PCR analysis of the expression of dac Hme and glyA Hme under different culture conditions. Lane M marker; lanes Glu, Col, and Pow the reverse transcripts of the total RNA from H. mediterranei grown with glucose (Glu), colloidal chitin (Col), and powdered chitin (Pow), respectively, as the template; lanes + genome as the template, positive control; lanes − negative control. The primers used here are listed in Table S1. b The final PHBV concentration of H. mediterranei strains (DF50, Δdac Hme, and ΔglyA Hme) at the stationary phase (maximum PHBV production) from colloidal (the columns in dark gray) or powdered chitin (the columns in light gray)
Discussion
In this study, we report that H. mediterranei is able to efficiently degrade colloidal and powdered chitin for growth and PHBV accumulation and reveal the key enzymes involved in the chitin catabolism pathway. This is the first study of chitin catabolism and bioconversion in haloarchaea.
The characterized or annotated archaeal chitinases so far are listed in Table S4, except the three identified chitinases from T. chitonophagus with no sequences in NCBI GenBank (Andronopoulou and Vorgias 2004). These chitinases were mainly from Euryarchaeota, including eight species within the class Halobacteria, four Thermococci (T. chitonophagus included), and three Methanomicrobia. Besides, one species of Crenarchaeota (S. tokodaii) also possessed a chitinase, though it is not the classical GH18 family or GH19 family chitinase (Staufenberger et al. 2012). This indicated the potential of archaea for chitin bioconversion. The capability of H. mediterranei to degrade powdered chitin has ecological and economical significance because powdered chitin does not have to be pretreated with acid. In some microorganisms, a type of oxidative enzyme classified as CBMs is responsible for disrupting the highly crystalline structure of powdered chitin (Vaaje-Kolstad et al. 2010). Bioinformatic analysis revealed that this type of enzyme was present in the haloarchaeon of Halomicrobium mukohataei (accession number YP_003177200) but was not present in other archaea with available genome sequences, including H. mediterranei. ChtBD is able to disrupt the hydrogen bonds of the chitin chains (Nakamura et al. 2008), and the role of PKD in powdered chitin degradation has also been reported in bacterial chitinases (Orikoshi et al. 2005). Therefore, the ChtBD and PKD of the chitinases from H. mediterranei may act together to disrupt the hydrogen bonds of powdered chitin.
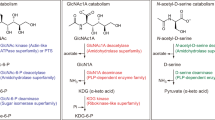
So far, archaeal chitin catabolism pathway is established only in T. kodakaraensis KOD1. Chitin is first broken down into GlcNAc2 by ChiATk. GlcNAc2 is deacetylated by DacTk to generate GlcN-GlcNAc and then cleaved into GlcN and GlcNAc by GlmATk. The generated GlcNAc is also deacetylated by DacTk to produce GlcN (Tanaka et al. 2004). In this study, we identified the key enzymes for chitin catabolism in H. mediterranei including four chitinases (ChiABCDHme), LmbE-like deacetylase (DacHme), and glycosidase (GlyAHme). When compared with the chitin degradation pathway in T. kodakaraensis KOD1, the LmbE-like deacetylase exhibited 31 % identity to DacTk and was assumed to be the deacetylase of H. mediterranei. Nevertheless, the homolog of GlyAHme was not found in T. kodakaraensis KOD1, and the homolog of the key enzyme for chitin degradation in T. kodakaraensis KOD1, GlmATk, was also not found in H. mediterranei. Further analysis revealed that GlmATk was a member of GH family 42, while GlyAHme belonged to GH family 3. The enzymes of GH family 3 have broad substrate specificity and exhibit diverse catalytic properties, including β-glucosidases, β-xylosidases, β-N-acetylhexosaminidases, and even the combination of them (Faure 2002). Additionally, there have been some reports about the enzymes of GH family 3 being involved in chitin catabolism, such as the β-N-acetylglucosaminidase of the marine bacterium Vibrio furnissii CIP 102972 (accession number ZP_05880052) (Keyhani and Roseman 1996). GlyAHme showed 34 % identity to the β-N-acetylglucosaminidase from V. furnissii CIP 102972 and might have the catalytic activity of β-N-acetylglucosaminidase. These results indicate that DacHme and GlyAHme might be involved in the further catabolism of GlcNAc2 for both cell growth and PHBV biosynthesis, although their exact function remains to be demonstrated. It is noteworthy that horizontal gene transfer might occur in the archaeal chitin catabolism pathway because archaea represents an ancient form of life, while chitin is mainly present in highly evolved organisms (Tanaka et al. 1999). Hence, it will be intriguing to discover more novel chitin degradation pathways in archaea and discuss the evolutionary process of different chitin degradation pathways operating in bacteria, eukarya, and archaea.
References
Andronopoulou E, Vorgias CE (2004) Multiple components and induction mechanism of the chitinolytic system of the hyperthermophilic archaeon Thermococcus chitonophagus. Appl Microbiol Biotechnol 65:694–702. doi:10.1007/s00253-004-1640-4
Cai S, Cai L, Liu H, Liu X, Han J, Zhou J, Xiang H (2012) Identification of the haloarchaeal phasin (PhaP) that functions in polyhydroxyalkanoate accumulation and granule formation in Haloferax mediterranei. Appl Environ Microbiol 78:1946–1952. doi:10.1128/AEM.07114-11
Cohen-Kupiec R, Chet I (1998) The molecular biology of chitin digestion. Curr Opin Biotechnol 9:270–277. doi:10.1016/S0958-1669(98)80058-X
Faure D (2002) The family-3 glycoside hydrolases: from housekeeping functions to host–microbe interactions. Appl Environ Microbiol 68:1485–1490. doi:10.1128/AEM.68.4.1485-1490.2002
Gao J, Bauer MW, Shockley KR, Pysz MA, Kelly RM (2003) Growth of hyperthermophilic archaeon Pyrococcus futiosus on chitin involves two family 18 chitinases. Appl Environ Microbiol 69:3119–3128. doi:10.1128/AEM.69.6.3119-3128.2003
Han J, Lu Q, Zhou L, Zhou J, Xiang H (2007) Molecular characterization of the phaEC Hm genes, required for biosynthesis of poly(3-hydroxybutyrate) in the extremely halophilic archaeon Haloarcula marismortui. Appl Environ Microbiol 73:6058–6065. doi:10.1128/AEM.00953-07
Han J, Zhang F, Hou J, Liu XQ, Li M, Liu HL, Cai L, Zhang B, Chen YP, Zhou J, Hu SN, Xiang H (2012) Complete genome sequence of the metabolically versatile halophilic archaeon Haloferax mediterranei, a poly(3-hydroxybutyrate-co-3-hydroxyvalerate) producer. J Bacteriol 194:4463–4464. doi:10.1128/JB.00880-12
Himmel ME, Ding SY, Johnson DK, Adney WS, Nimlos MR, Brady JW, Foust TD (2007) Biomass recalcitrance: engineering plants and enzymes for biofuels production. Science 315:804–807. doi:10.1126/science.1137016
Imoto T, Yagishita K (1971) A simple activity measurement of lysozyme. Agric Biol Chem 35:1154–1156
Karlsson M, Stenlid J (2009) Evolution of family 18 glycoside hydrolases: diversity, domain structures and phylogenetic relationships. J Mol Microbiol Biotechnol 16:208–223. doi:10.1159/000151220
Keyhani NO, Roseman S (1996) The chitin catabolic cascade in the marine bacterium Vibrio furnissii. Molecular cloning, isolation, and characterization of a periplasmic β-N-acetylglucosaminidase. J Biol Chem 271:33425–33432. doi:10.1074/jbc.271.52.33425
Kreuzer M, Schmutzler K, Waege I, Thomm M, Hausner W (2013) Genetic engineering of Pyrococcus furiosus to use chitin as a carbon source. BMC Biotechnol 13:9. doi:10.1186/1472-6750-13-9
Kurita K (2006) Chitin and chitosan: functional biopolymers from marine crustaceans. Mar Biotechnol (NY) 8:203–226. doi:10.1007/s10126-005-0097-5
Li H, Greene LH (2010) Sequence and structural analysis of the chitinase insertion domain reveals two conserved motifs involved in chitin-binding. PLoS One 5:e8654. doi:10.1371/journal.pone.0008654
Liu HL, Han J, Liu XQ, Zhou J, Xiang H (2011) Development of pyrF-based gene knockout systems for genome-wide manipulation of the archaea Haloferax mediterranei and Haloarcula hispanica. J Genet Genomics 38:261–269. doi:10.1016/j.jgg.2011.05.003
Lu Q, Han J, Zhou L, Coker JA, DasSarma P, DasSarma S, Xiang H (2008a) Dissection of the regulatory mechanism of a heat-shock responsive promoter in Haloarchaea: a new paradigm for general transcription factor directed archaeal gene regulation. Nucleic Acids Res 36:3031–3042. doi:10.1093/nar/gkn152
Lu QH, Han J, Zhou LG, Zhou J, Xiang H (2008b) Genetic and biochemical characterization of the poly(3-hydroxybutyrate-co-3-hydroxyvalerate) synthase in Haloferax mediterranei. J Bacteriol 190:4173–4180. doi:10.1128/JB.00134-08
Lü Y, Yang H, Hu H, Wang Y, Rao Z, Jin C (2009) Mutation of Trp137 to glutamate completely removes transglycosyl activity associated with the Aspergillus fumigatus AfChiB1. Glycoconj J 26:525–534. doi:10.1007/s10719-008-9203-z
Lutz RA, Shank TM, Fornari DJ, Haymon RM, Lilley MD, Vondamm KL, Desbruyeres D (1994) Rapid growth at deep-sea vents. Nature 371:663–664. doi:10.1038/371663a0
Mathur NK, Narang CK (1990) Chitin and chitosan, versatile polysaccharides from marine animals. J Chem Educ 67:938–942. doi:10.1021/ed067p938
Nakamura T, Mine S, Hagihara Y, Ishikawa K, Ikegami T, Uegaki K (2008) Tertiary structure and carbohydrate recognition by the chitin-binding domain of a hyperthermophilic chitinase from Pyrococcus furiosus. J Mol Biol 381:670–680. doi:10.1016/j.jmb.2008.06.006
Oku T, Ishikawa K (2006) Analysis of the hyperthermophilic chitinase from Pyrococcus furiosus: activity toward crystalline chitin. Biosci Biotechnol Biochem 70:1696–1701. doi:10.1271/bbb.60031
Orikoshi H, Nakayama S, Hanato C, Miyamoto K, Tsujibo H (2005) Role of the N-terminal polycystic kidney disease domain in chitin degradation by chitinase A from a marine bacterium, Alteromonas sp. strain O-7. J Appl Microbiol 99:551–557. doi:10.1111/j.1365-2672.2005.02630.x
Rodriguez-Valera F, Juez G, Kushner DJ (1983) Halobacterium mediterranei spec, nov., a new carbohydrate-utilizing extreme halophile. Syst Appl Microbiol 4:369–381. doi:10.1016/S0723-2020(83)80021-6
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual, 2nd edn. Cold Spring Harbor Laboratory, Cold Spring Harbor
Songsiriritthigul C, Lapboonrueng S, Pechsrichuang P, Pesatcha P, Yamabhai M (2010) Expression and characterization of Bacillus licheniformis chitinase (ChiA), suitable for bioconversion of chitin waste. Bioresour Technol 101:4096–4103. doi:10.1016/j.biortech.2010.01.036
Staufenberger T, Imhoff JF, Labes A (2012) First crenarchaeal chitinase found in Sulfolobus tokodaii. Microbiol Res 167:262–269. doi:10.1016/j.micres.2011.11.001
Tanaka T, Fujiwara S, Nishikori S, Fukui T, Takagi M, Imanaka T (1999) A unique chitinase with dual active sites and triple substrate binding sites from the hyperthermophilic archaeon Pyrococcus kodakaraensis KOD1. Appl Environ Microbiol 65:5338–5344
Tanaka T, Fukui T, Fujiwara S, Atomi H, Imanaka T (2004) Concerted action of diacetylchitobiose deacetylase and exo-β-D-glucosaminidase in a novel chitinolytic pathway in the hyperthermophilic archaeon Thermococcus kodakaraensis KOD1. J Biol Chem 279:30021–30027. doi:10.1074/jbc.M314187200
Tanaka T, Fukui T, Imanaka T (2001) Different cleavage specificities of the dual catalytic domains in chitinase from the hyperthermophilic archaeon Thermococcus kodakaraensis KOD1. J Biol Chem 276:35629–35635. doi:10.1074/jbc.M105919200
Tsuji H, Nishimura S, Inui T, Kado Y, Ishikawa K, Nakamura T, Uegaki K (2010) Kinetic and crystallographic analyses of the catalytic domain of chitinase from Pyrococcus furiosus—the role of conserved residues in the active site. FEBS J 277:2683–2695. doi:10.1111/j.1742-464X.2010.07685.x
Vaaje-Kolstad G, Westereng B, Horn SJ, Liu Z, Zhai H, Sørlie M, Eijsink VG (2010) An oxidative enzyme boosting the enzymatic conversion of recalcitrant polysaccharides. Science 330:219–222. doi:10.1126/science.1192231
Yoshinobu H, Motosuke S, Keita O, Rie Y, Kimiko E, Toshiaki F, Satoshi N (2006) Characterization of recombinant family 18 chitinase from extremely halophilic archaeon Halobacterium salinarum strain NRC-1. Chitin Chitosan Res 12:201
Acknowledgments
This work was financially supported by grants from the National Natural Science Foundation of China (grant nos. 30830004, 30925001, and 31000023) and the Chinese Academy of Sciences (KSCX2-EW-G-2-4).
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 343 kb)
Rights and permissions
About this article
Cite this article
Hou, J., Han, J., Cai, L. et al. Characterization of genes for chitin catabolism in Haloferax mediterranei . Appl Microbiol Biotechnol 98, 1185–1194 (2014). https://doi.org/10.1007/s00253-013-4969-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-013-4969-8