Abstract
A fungal strain, Penicillium chrysogenum A096, was isolated from an Arctic sediment sample. Its culture supernatant inhibited mycelial growth of some plant pathogenic fungi. After saturation of P. chrysogenum A096 culture supernatant with ammonium sulfate and ion exchange chromatography, a novel antifungal protein (Pc-Arctin) was purified and identified by matrix assisted laser desorption ionization-time of flight-time of flight-mass spectrometry (MALDI-TOF-TOF-MS). The gene encoding for Pc-Arctin consisting of 195 nucleotides was cloned from P. chrysogenum A096 to confirm the mass spectrometry result. Pc-Arctin displays antifungal activity against Paecilomyces variotii, Alternaria longipes, and Trichoderma viride at minimum inhibitory concentrations (MIC) of 24, 48, and 192 ng/disc, respectively. Pc-Arctin was most sensitive to proteinase K and then to trypsin but insensitive to papain. Pc-Arctin possesses high thermostability and cannot be antagonized by common surfactants, except for sodium dodecyl sulfate (SDS). Divalent ions, such as Mn2+, Mg2+, and Zn2+, inhibited the antifungal activity of Pc-Arctin. Hemagglutination assays showed that Pc-Arctin had no hemagglutinating or hemolytic activity against red blood cells (RBC) from rabbits, rats, and guinea pigs. Therefore, Pc-Arctin from Arctic P. chrysogenum may represent a novel antifungal protein with potential for application in controlling plant pathogenic fungal infection.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
To date, about 300 species of fungi have been reported to display pathogenic or allergenic potentials (Gupte et al. 2002). Fungal infections of humans and animals have become a severe problem around the world. Until now, the number of drugs for efficient treatment of fungal invasion has been very limited, and there is an increasing need for new antifungal agents (Chen and Sorrell 2007). The side effects of the currently available antifungal drugs, such as nephrotoxicity, fever, and taste disturbance, have caused widespread concern (Chen and Sorrell 2007; Denning 2003; Lemke et al. 2005). Therefore, it is crucial for the research community to find new sources of antifungal drugs. Natural products from microbial sources from less-exploited environments are emerging as attractive options.
A tremendous challenge in agricultural industry is to control plant diseases caused by pathogenic fungi (Meyer 2008). Currently, chemical pesticides are the preferred options for suppressing growth of fungi in the agricultural industry. However, broad use of chemical pesticides can cause harmful effects on the environment and human health. Therefore, new substitutes for these chemical fungicides that are less harmful to the environment and to humans are urgently needed.
Antimicrobial proteins that are ubiquitous in nature are suitable fungicide candidates because of their natural origin and reduced side effects (Lay and Anderson 2005; Woo et al. 2002; Zasloff 2002; Zhang et al. 2008). Antimicrobial proteins function as the first defensive system against microbial invasion and display strong growth-inhibitory activities against many pathogens, including bacteria, fungi, and viruses (Lehrer and Ganz 1999; Marx et al. 2008). Several proteins derived from the imperfect filamentous fungi Aspergillus and Penicillium have broad-spectrum antifungal properties (Gun Lee et al. 1999; Marx et al. 1995; Skouri-Gargouri and Gargouri 2008; Wnendt et al. 1994). Most of these proteins are small molecules rich in lysine and cysteine residues that form disulfide bridges that are postulated to contribute to their high stability against high temperatures and protease degradation (Marx 2004; Rodriguez-Martin et al. 2010; Skouri-Gargouri and Gargouri 2008). Recent data revealed that such an antifungal protein Penicillium antifungal protein (PAF) could interfere with calcium homeostasis in sensitive fungi and certain types of chitin synthases are critical for fungi sensitive to antifungal protein (AFP) (Binder et al. 2010; Ouedraogo et al. 2011). Indeed, expression of some antifungal proteins in transgenic plants has enhanced resistance against pathogens, indicating a promising means for plant protection (Coca et al. 2004; Mitsuhara et al. 2000; Moreno et al. 2005; Sharma et al. 2000).
The Arctic represents an extremely cold and uncharted environment, where large numbers of diverse biological resources remain to be exploited. Here, we report purification and characterization of a novel antifungal protein, Pc-Arctin, from the fungal strain Penicillium chrysogenum A096 that was isolated from a sediment sample of Arctic origin. Antifungal activity and physicochemical properties of Pc-Arctin were studied, and its hemagglutinating and hemolytic activities against red blood cells (RBC) from several animals were also analyzed. These results suggest that Pc-Arctin may have a good potential for application in controlling plant diseases caused by pathogenic fungi.
Materials and methods
Tested strains
Antifungal activity was assayed against different species of fungi, including Paecilomyces variotii (CGMCC 3.776, China General Microbiological Culture Collection Center), Colletotrichum gloeosporioides (ACCC 31200, Agricultural Cultural Collection of China), Fusarium oxysporum (ACCC 31352), Aspergillus niger (ACCC 31856), Penicillium expansum (ACCC 30898), Trichoderma viride (ACCC 30902), Rhizoctonia solani (ACCC 36316), Alternaria longipes (ACCC 30002), Sclerotinia sclerotiorum (ACCC 36081), and Pythium aphanidermatum (ACCC 31412). All fungi were purchased from the Agricultural Cultural Collection of China, except for P. variotii, which was obtained from CGMCC, Institute of Microbiology Chinese Academy of Sciences.
Isolation and identification of an antifungal strain
The specimen used for strain isolation was collected from an Arctic sediment (N 78°57′09.2″, E 12°00′02.3″). The specimen was diluted with sterilized seawater, and approximately 50 μL of the diluted sample was sprayed on plates containing different types of medium such as SGY (1 % soybean meal, 1 % glucose, 0.5 % yeast extract, and 0.25 % NaCl), YEM (1 % yeast extract, 1 % malt extract, and 0.4 % glucose), and YTM (0.5 % yeast extract, 0.3 % tryptone, and 2.5 % mannitol). Plates were incubated at 28 °C for growth. A total of 18 strains were purified based on their morphological feature and inoculated into corresponding liquid media for further growth to evaluate their antifungal potential. Supernatant of the strain A096 cultured in SGY liquid medium showed remarkable inhibitory activity against several pathogenic fungi. Spores of this strain, grown on SGY agar plates, were washed with 40 % glycerol and preserved at −80 °C. This strain has been deposited at the China Center for Type Culture Collection with preservation number M2010270 (Wuhan, China).
To further identify the strain A096 in terms of the ribosomal internal transcribed spacer (ITS) gene sequence, its ITS gene was amplified using the primers ITS5 (GGAAGTAAAAGTCGTAACAAGG) and ITS4 (TCCTCCGCTTATTGATATGC) (White et al. 1990). The polymerase chain reaction (PCR) program included an initial denaturation at 95 °C for 4 min, followed by 36 cycles of 95 °C for 1.5 min, 52 °C for 1.5 min, and 72 °C for 1.5 min, with a final extension at 72 °C for 10 min. The amplified ITS products were sequenced at Sangon Biotech (Shanghai, China). A BLASTn search on the National Center for Biotechnology Information (NCBI) web was performed to analyze the obtained sequences. The ITS sequence has been submitted to the NCBI GenBank with accession number JQ015265.
Antifungal activity detection
The bioactive strain A096 was cultured in shaking incubators at 28 °C, 180 rpm for 7 days. Culture supernatant was collected by vacuum filtration and further sterilized by filtering through an aseptic 0.22-μμm filter membrane. Antifungal activity was estimated by a paper disc diffusion assay. Fungi were grown on potato dextrose agar (PDA) plates. One 0.65-cm diameter piece of cylinder agar with fungal growth was placed on the center of a PDA plate until the mycelium developed at 28 °C. Sterile paper discs of 0.65-cm diameter were placed 0.8 cm from the rim of the growing mycelium. Approximately 40 μμL of culture supernatant from strains being tested for antifungal activity were was loaded onto each paper disc, with culture medium as the negative control. Plates were sealed with Parafilm (Sangon Biotech, Shanghai, China) and incubated at 28 °C until the mycelium enveloped the blank control discs and formed a crescent shape around discs containing culture supernatant (Wong and Ng 2003) .
Purification of antifungal protein
For large-scale culture, strain A096 was primarily inoculated on SGY plates. Spores were precultured in 200-mL SGY liquid medium in 1-L Erlenmeyer flasks at 28 °C in a shaker for 1 day. Seed cultures were transferred to 4.5-L SGY liquid medium in a BIOSTAT B plus fermenter (B. Braun, Germany) and cultured at 28 °C with agitation at 200 rpm for 3 days. Supernatant was collected by vacuum filtration, fully saturated with ammonium sulfate, stored at 4 °C overnight and centrifuged at 14,000 g at 4 °C. Precipitates were harvested, dissolved in distilled water, and dialyzed against distilled water. The solution was centrifuged at 10,000 g for 10 min at 4 °C to remove the insoluble components. After concentration by lyophilization, the remaining components were redissolved in a small volume of distilled water to prepare a crude protein extract.
The crude proteins and fractions separated by ion exchange chromatography were tested for antifungal activity against P. variotii. Ion exchange chromatography was performed with an AKTA FPLC system (GE Healthcare, USA). The crude protein solution was loaded onto a DEAE Sepharose Fast Flow column (GE Healthcare) that was preequilibrated with starting buffer A (pH 8.1, 20 mM Tris–HCl) for the primary purification step. The unabsorbed fraction A was washed with starting buffer A and subsequently with elution buffer B (pH 8.1, 20 mM Tris–HCl, 1 M NaCl) to desorb the absorbed fraction B. The bioactive fraction A was concentrated by ultrafiltration in a Vivaspin 15R (molecular weight cutoff 5000, Sartorius, Germany). Subsequently, fraction A was separated on a CM Sepharose Fast Flow column (GE Healthcare) preequilibrated with starting buffer A. The unabsorbed fraction C (washed with starting buffer A) and absorbed fraction D (eluted with a 0–100 % linear gradient of elution buffer B) were pooled, respectively. The results indicated that the absorbed component D was the active fraction. Purity of the active component was determined by 15 % sodium dodecyl sulfate (SDS)-polyacrylamide gel electrophoresis (PAGE) and silver nitrate staining.
Mass spectrometry identification of the active protein
The active protein band was extracted from the stained SDS-PAGE gel; washed three times with sterile, distilled water; and treated with trypsin before matrix assisted laser desorption ionization-time of flight-time of flight-mass spectrometry (MALDI-TOF-TOF-MS) identification of the purified protein at Shanghai Institutes for Biological Sciences, Chinese Academy of Sciences. The protein was identified by database search on Mascot server (http://www.matrixscience.com/search_form_select.html) with MALDI-TOF–TOF data.
RNA extraction and Pc-Arctin gene cloning
The spores of P. chrysogenum A096 were inoculated into SGY liquid medium and cultured at 28 °C, 180 rpm for 3 days. Then, fungi mycelium was collected by vacuum filtration. The mycelium was grinded into powder and total RNA was extracted with TRIzol Reagent (Invitrogen, USA) according to the manufacturer’s instruction. Total RNA was then dissolved in diethypyrocarbonate (DEPC)-treated water and preserved at −80 °C.
First-strand cDNA was synthesized from 0.5 μg of total RNA above using oligo-dT-adaptor primer (TaKaRa, Dalian, China) and used as the template for PCR amplification of Pc-Arctin gene. Primers were designed based on the nucleotide sequence encoding the predicted mature peptide of the hypothetical protein Pc21g12970 (CAP96194): forward primer: 5′-GACACCTGCGGAGGCGGCTAT-3′; reverse primer: 5′-CTAGCATCTGGCTCCCCCCTGG-3′. PCR was performed with an initial denaturation step of 3 min at 95 °C and then 30 cycles were run as follows: 30-s denaturation at 95 °C, 30-s annealing at 56 °C, and 30-s extension at 72 °C, followed by a cycle of 72 °C for 5 min. The PCR product was purified and cloned into pMD18-T vector (TaKaRa, Dalian, China). The obtained positive clone was sequenced by M13+/− sequencing primers. Sequence similarity analysis was performed using BLAST program (http://blast.ncbi.nlm.nih.gov/Blast.cgi). Signal peptide prediction was made using SignalP 4.1 Server (http://www.cbs.dtu.dk/services/SignalP/). Multiple sequence alignment was carried out using the DNAMAN program (Lynnon Biosoft).
MIC determination of Pc-Arctin
Minimum inhibitory concentration (MIC) of Pc-Arctin against different pathogenic fungi was determined by paper disc dilution method (Woo et al. 2002). Twofold serial dilutions of the active protein solution ranging from 76.8 to 0.6 ng/μL were prepared, and 40 μL of each diluted solution was added onto paper discs placed 0.8 cm from the growing mold on a PDA plate. The plates were placed at 28 °C for several days, depending on the tested pathogen. The MIC was determined as the lowest concentration of active protein that could inhibit visible mold growth and calculated as the total protein added on each paper disc (nanogram per disc). Starting buffer A without active compound was used as blank control.
Effect of proteases on the antifungal activity of Pc-Arctin
To understand sensitivity of Pc-Arctin to proteases, the purified protein was subjected to treatment with 1 mg/mL proteinase K (BBI, USA), papain (Worthington, USA), and trypsin (Worthington, USA) at 55 °C for 2 h, at 37 °C for 24 h, and at 25 °C for 24 h and 48 h, respectively. The treatment effect was analyzed by observing the residual antifungal activity of Pc-Arctin, where Pc-Arctin without treatment with the proteases was used as positive control. The proteases and starting buffer A were used as blank controls.
Effect of surfactant on the antifungal activity of Pc-Arctin
For surfactant-resistant test, several surfactants including 0.05 % SDS, 0.05 % Triton X-100, 8 % urea, and 1 % Tween 20, were dissolved in starting buffer A. These surfactants were all purchased from Sangon Biotech (Shanghai, China). Pc-Arctin was mixed with each surfactant solution at room temperature for 1 h and tested against P. variotii to determine whether the surfactants negatively affected its antifungal function. The active protein without added surfactant was used as the positive control, and starting buffer A and the surfactant solutions were the negative controls.
Effect of metal ions on the antifungal activity of Pc-Arctin
To examine the effect of metal ions on Pc-Arctin activity, selected metal ions such as K+, Ca2+, Co2+, Ni2+, Cu2+, Zn2+, Mg2+, and Mn2+ were dissolved in starting buffer A at each different concentration. The Pc-Arctin protein was treated with different ion solutions at room temperature for 1 h before testing for antifungal activity. The Pc-Arctin protein without ion solution treatment was used as the positive control, and starting buffer A and ion solutions were used as blank negative controls.
Effect of heat treatment on the antifungal activity of Pc-Arctin
To evaluate thermostability of the antifungal protein, the Pc-Arctin protein was heated at 50°C, 60 °C, 70 °C, 80 °C, 90 °C, and 100 °C for 30 min, 100 °C for 1 h, and autoclaved (121 °C for 20 min). After cooling the protein samples, the residual antifungal activity of the pure protein was tested against sensitive fungi. Pc-Arctin without heat treatment and starting buffer A were used as the positive control and blank control, respectively.
Detection of hemagglutination activity
To evaluate the hemagglutinating activity of the Pc-Arctin, twofold serial dilutions of the protein solution (25 μL) in buffer C (pH 8.10, 20 mM Tris, 0.1 M NaCl) were mixed with equal volumes of 2 % RBC suspensions from rabbits, rats, and guinea pigs on an agglutinating board. The results were evaluated when RBCs in the negative control (buffer C without active protein) were fully sedimented. Then, whether RBCs of different origins respond to Pc-Arctin is to be judged.
Detection of hemolytic activity
The procedure used here was the same as for hemagglutination activity detection, except that serial dilutions of the protein were mixed with equal volumes of 4 % RBC suspensions originating from different animals species. The mixtures were incubated at room temperature until all RBCs were lysed by 0.05 % Triton X-100 (dissolved in buffer C and used as the positive control). The effect of the Pc-Arctin protein on erythrocyte membranes was estimated by calculating the amount of hemoglobin released from disrupted erythrocytes, which was determined spectrophotometrically at λ540 nm (Wong et al. 2008).
Results
Strain identification and detection of antifungal activity
Aerial mycelium of the fungal strain A096 did not develop well, but many green spores were produced that finally turned gray-brown. A light yellow substance was produced on PDA plates. Based on the morphological and molecular information, this fungal strain was assigned to P. chrysogenum A096 and has been deposited at the China Center for Type Culture Collection in Wuhan, China. The preservation number of this strain is M2010270.
The supernatant of P. chrysogenum A096 was added on paper discs at a distance of 0.8 cm from the rim of each growing mycelium of P. variotii, T. viride, and A. longipes. The appearance of crescent mycelium rim around paper discs indicates the mycelium inhibitory activity of the supernatant of P. chrysogenum A096. Preliminary analysis of its antifungal components revealed that the precipitate obtained by saturating the culture supernatant with ammonium sulfate exerted antifungal activity. The crude extract was then used for further purification.
Purification of Pc-Arctin
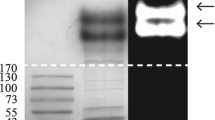
Further purification of the bioactive protein was performed using ion exchange chromatography and was monitored with the antifungal assay. The precipitate with antifungal activity was first loaded onto DEAE Sepharose Fast Flow column, and only fractions A and B were isolated (Fig. 1a), of which only fraction A possessed antifungal activity. Subsequently, the partially purified fraction A was loaded onto a CM Sepharose Fast Flow column, and two major peaks, C and D, were generated (Fig. 1b). The results of antifungal assay indicated that only fraction D contained a bioactive component (Fig. 1c). The active component showed only a single band on a SDS-PAGE gel (Fig. 1d). This protein band was excised from the silver-stained gel and treated with trypsin for mass spectrometry analysis. Three-peptide sequences of this protein were identified by MALDI-TOF–TOF and an online search (Table 1). This bioactive protein was designated Pc-Arctin. The sequences of these three peptides of Pc-Arctin were all contained in a hypothetical protein Pc21g12970 (CAP96194), which function has not been known (van den Berg et al. 2008).
Purification of active protein by ion exchange chromatography and detection of its purity and antifungal activity using P. variotii as the sensitive fungal species. a Chromatographic profile of P. chrysogenum A096 crude protein extract on a DEAE Sepharose Fast Flow column using an AKTA FPLC system. A unabsorbed fraction, B absorbed fraction. b Chromatographic profile of unabsorbed fraction A on a CM Sepharose Fast Flow column using an AKTA FPLC system. C unabsorbed fraction, D absorbed fraction. c Detection of inhibitory activity of peak D (disc 1) against P. variotii with buffer C (disc 2) as blank control. d 15 % SDS-PAGE profile of Pc-Arctin. Lane 1 standard protein markers, lane 2 purified antifungal protein Pc-Arctin
Pc-Arctin gene cloning
To clone the gene encoding Pc-Arctin, RT-PCR was performed with primers designed based on the predicted mature peptide of the hypothetical protein Pc21g12970. The sequencing result showed that the obtained Pc-Arctin gene is 195 nucleotides long, encoding a peptide of 64 amino acids (Fig. 2a). The deduced Pc-Arctin contained eight cysteine residues, which are required for the formation of disulfide bond. The three peptides identified by mass spectrometry were all included in the deduced Pc-Arctin (Fig. 2a), indicating that the gene cloned here encodes Pc-Arctin. Homology comparison revealed that Pc-Arctin shares 100 % sequence identity to the mature peptide of hypothetical protein Pc21g12970 (CAP96194), while it shares only 10.77 % and 18.46 % sequence identities to PAF from P. chrysogenum (Batta et al. 2009) and AFP from Aspergillus giganteus (Wnendt et al. 1994), two known typical basic antifungal proteins from filamentous fungi. By searching the Antimicrobial Peptide Database (http://aps.unmc.edu/AP/main.php) with the deduced Pc-Arctin, only five antimicrobial peptides Ee-CBP (Antimicrobial Peptide Database ID AP00915), SIalpha1 (AP01528), Divercin V41 (AP00842), Halocin C8 (AP01193), and Cathelicidin-AL (AP01898) were found to have low sequence similarities to Pc-Arctin, ranging from 20.3 to 27.7 % (Fig. 2b).
a Nucleotide and deduced amino acid sequences of Pc-Arctin gene. Three-peptide sequences of Pc-Arctin identified by MALDI-TOF–TOF are shown with red background; filled triangle indicates cysteine residues in the deduced Pc-Arctin. b Alignment of the deduced Pc-Arctin amino acid sequence with those of two typical basic antifungal proteins PAF and AFP from filamentous fungi and five other antifungal proteins. Five antifungal proteins Ee-CBP (AP00915), SIalpha1 (AP01528), Divercin V41 (AP00842), Halocin C8 (AP01193), and Cathelicidin-AL (AP01898) were obtained by searching the Antimicrobial Peptide Database (http://aps.unmc.edu/AP/main.php) with the deduced Pc-Arctin
MIC determination of Pc-Arctin
The purified Pc-Arctin protein was tested against different pathogenic fungi in a plate diffusion assay. Pc-Arctin showed the strongest antifungal activity against P. variotii, followed by A. longipes and T. viride with MIC of 24, 48, and 192 ng/disc, respectively, indicating that different pathogenic fungi exhibited different sensitivity against Pc-Arctin.
Physiochemical properties of Pc-Arctin
To estimate the application potential of Pc-Arctin, the purified protein was treated with proteases, surfactants, ions, and high temperatures. Pc-Arctin was sensitive to proteinase K, since it exhibited no antifungal activity against P. variotti when treated with 1 mg/mL proteinase K at 55 °C for 2 h, at 37 °C for 24 h, or at 25 °C for 48 h, but it can retain remarkable residual activity when treated with proteinase K at 25 °C for 24 h (Fig. 3a1–a4). When treated with trypsin at a concentration of 1 mg/mL at 37 °C for 24 h or at 25 °C for 48 h, Pc-Arctin could not keep all of its antifungal activity (Fig. 3b2–b4). However, it was able to stand stable no matter if it was treated with 1 mg/mL trypsin at 55 °C for 2 h or with 1 mg/mL papain at different temperatures for 48 h (Fig. 3b1, c1–c4).
Antifungal activity of 0.2 μg Pc-Arctin treated with 1 mg/mL proteinase K, trypsin, and papain at various temperatures for different time points. a1–a4 Antifungal activity of 0.2 μg Pc-Arctin treated with 1 mg/mL proteinase K at 55 °C for 2 h, at 37 °C for 24 h, and at 25 °C for 24 h and 48 h, respectively. b1–b4 Antifungal activity of 0.2 μg Pc-Arctin treated with 1 mg/mL trypsin at 55 °C for 2 h, at 37 °C for 24 h, and at 25 °C for 24 h and 48 h, respectively. c1–c4 Antifungal activity of 0.2 μg Pc-Arctin treated with 1 mg/mL papain at 55 °C for 2 h, at 37 °C for 24 h, and at 25 °C for 24 h and 48 h, respectively. Numbers in each picture refer to samples loaded on each corresponding paper disc. 1 refers to proteinase K, trypsin, and papain as control, respectively; 2 refers to Pc-Arctin treated with 1 mg/mL proteinase K, trypsin, and papain, respectively; 3 refers to Pc-Arctin without treatment with 1 mg/mL proteinase K, trypsin, and papain but incubated at various corresponding temperatures, respectively; 4 refers to Pc-Arctin stored at 4 °C as positive control; 5 refers to starting buffer A as blank control
Pc-Arctin remained active when treated with surfactants, such as 0.05 % Triton X-100, 8 % urea, and 1 % Tween 20. However, 0.05 % SDS reduced the antifungal activity of Pc-Arctin (Fig. 4). The antifungal activity of Pc-Arctin remained intact in the presence of different ions (0.5 M K+, 0.5 M Ca2+, and 0.01 M Cu2+, Co2+, and Ni2+) but was impaired when incubated with 0.5 M Mg2+, 0.5 M Mn2+, and 0.03 M Zn2+ (Table 2). Pc-Arctin is thermostable and maintains undamaged activity when heated at 100 °C up to 1 h or autoclaved at 121 °C for 20 min (Fig. 5).
Analysis of Pc-Arctin stability following exposure to high temperatures. a 1 Pc-Arctin without heat treatment, 2–7 Pc-Arctin treated at 50, 60, 70, 80, 90, and 100 °C, respectively, for 30 min, 8 Pc-Arctin treated by autoclaving (121 °C for 20 min), 9 starting buffer A as negative control. b 1 Pc-Arctin without heat treatment, 2–5 Pc-Arctin treated at 100 °C for 30, 40, 50, and 60 min, respectively, 6 starting buffer A as negative control
Finally, cytotoxicity of Pc-Arctin was assessed by hemagglutination and hemolysis assays. Neither hemagglutination nor hemolytic activities were observed when different concentrations of Pc-Arctin (0.3 to 20 μg/mL) were incubated with RBCs from rabbits, rats, and guinea pigs (data not shown).
Discussion
In the process of pursuing novel antifungal compounds from strains isolated from Arctic sediment samples, P. chrysogenum A096 was identified to have distinct antifungal activity. Further analysis revealed that a protein, named Pc-Arctin, was secreted into the culture supernatant and exhibited antifungal activity. Pc-Arctin was identified as a homologue of the hypothetical protein Pc21g12970 (CAP96194) by MALDI-TOF–TOF analysis. The gene encoding for Pc-Arctin was then cloned from P. chrysogenum A096 to confirm the mass spectrometry result. Although Pc-Arctin shares 100 % sequence identity to the mature peptide of hypothetical protein Pc21g12970 with an unknown function (van den Berg et al. 2008), it exhibits rather low sequence identities to two known typical basic antifungal proteins from filamentous fungi, PAF from P. chrysogenum and AFP from A. giganteus, and to five antimicrobial peptides Ee-CBP (AP00915), SIalpha1 (AP01528), Divercin V41 (AP00842), Halocin C8 (AP01193), and Cathelicidin-AL (AP01898). Based on the results from antifungal activity assays and amino acid sequence alignment, we suggest that Pc-Arctin may represent a novel antifungal protein from species P. chrysogenum.
The MICs for Pc-Arctin were as low as 24, 48, and 192 ng/disc against P. variotii, A. longipes, and T. viride, respectively. The antifungal activity of Pc-Arctin remained after treatment with several surfactants, except for 0.05 % SDS that slightly decreased Pc-Arctin activity. Moreover, Pc-Arctin was tolerant to several ions, but 0.5 M Mn2+, 0.5 M Mg2+, and 0.03 M Zn2+ decreased the antifungal activity. One possible reason is that certain types of ions may interact with specific receptors that are targeted by antifungal proteins, and the saturation of these receptors with ions could reduce the interaction between the receptors and the proteins (Kaiserer et al. 2003; Skouri-Gargouri et al. 2010; Theis et al. 2003). The investigation of stability of Pc-Arctin against proteases is of great interest, which may be helpful for evaluating its application potential in plant protection. Our results showed that Pc-Arctin was most sensitive to proteinase K followed by trypsin, but insensitive to papain (Fig. 3). The different sensitivity of Pc-Arctin to the proteases may be due to the specific digestion character of these proteases.
Studies have revealed that AFP from A. giganteus and PAF from P. chrysogenum contain eight and six cysteine residues, respectively, which are involved in the formation of disulfide bridges contributing to the heat stability of each antifungal protein (Marx et al. 2008; Meyer 2008). Moreover, the disulfide bridges in PAF have been considered to be indispensable for its antifungal action (Batta et al. 2009). Here, the purified Pc-Arctin is also thermostable and remains antifungal activity after high-temperature treatment (Fig. 5). The deduced Pc-Arctin contains eight cysteine residues, which may form four pairs of disulfide bridges (Fig. 2a). It is, therefore, supposed that the high thermostability of Pc-Arctin might be attributed to the formation of disulfide bridges. However, further studies are needed to determine whether these eight cysteine residues in Pc-Arctin are involved in its thermostability and antifungal action.
Pc-Arctin did not show hemagglutinating or hemolytic activity against RBCs from several animal species. Testing these activities is a fundamental strategy for analyzing safety and selectivity of proteins showing application potential (Theis and Stahl 2004). A protein with no cytotoxicity against RBCs can be preliminarily regarded as safe (van’t Hof et al. 2001).
In summary, a novel antifungal protein, Pc-Arctin, was identified from a fungal strain, P. chrysogenum A096, isolated from a sediment sample of Arctic origin. Pc-Arctin displays antifungal activity well against P. variotii, A. longipes, and T. viride, and its antifungal activity is not affected by some metal ions and common surfactants (except for SDS). It was able to keep its antifungal activity by treatment with trypsin and papain, more or less, except for proteinase K. Pc-Arctin also possesses high thermostability and displays no cytolytic activity against RBCs with the concentrations tested. The good stability and antifungal activity of Pc-Arctin indicates that the supernatant of the culture of P. chrysogenum could be directly applied for control of pathogenic fungi in field. Therefore, Pc-Arctin characterized here may represent a novel antifungal protein with potential for application in controlling plant pathogenic fungal infection.
References
Batta G, Barna T, Gáspári Z, Sándor S, Kövér KE, Binder U, Sarg B, Kaiserer L, Chhillar AK, Eigentler A, Leiter E, Hegedüs N, Pócsi I, Lindner H, Marx F (2009) Functional aspects of the solution structure and dynamics of PAF—a highly-stable antifungal protein from Penicillium chrysogenum. FEBS J 276(10):2875–2890
Binder U, Chu M, Read ND, Marx F (2010) The antifungal activity of the Penicillium chrysogenum protein PAF disrupts calcium homeostasis in Neurospora crassa. Eukaryot Cell 9(9):1374–1382
Chen SC, Sorrell TC (2007) Antifungal agents. Med J Aust 187(7):404–409
Coca M, Bortolotti C, Rufat M, Penas G, Eritja R, Tharreau D, del Pozo AM, Messeguer J, San Segundo B (2004) Transgenic rice plants expressing the antifungal AFP protein from Aspergillus giganteus show enhanced resistance to the rice blast fungus Magnaporthe grisea. Plant Mol Biol 54(2):245–259
Denning DW (2003) Echinocandin antifungal drugs. Lancet 362(9390):1142–1151
Gun Lee D, Shin SY, Maeng CY, Jin ZZ, Kim KL, Hahm KS (1999) Isolation and characterization of a novel antifungal peptide from Aspergillus niger. Biochem Biophys Res Commun 263(3):646–651
Gupte M, Kulkarni P, Ganguli BN (2002) Antifungal antibiotics. Appl Microbiol Biotechnol 58(1):46–57
Kaiserer L, Oberparleiter C, Weiler-Gorz R, Burgstaller W, Leiter E, Marx F (2003) Characterization of the Penicillium chrysogenum antifungal protein PAF. Arch Microbiol 180(3):204–210
Lay FT, Anderson MA (2005) Defensins—components of the innate immune system in plants. Curr Protein Pept Sci 6(1):85–101
Lehrer RI, Ganz T (1999) Antimicrobial peptides in mammalian and insect host defence. Curr Opin Immunol 11(1):23–27
Lemke A, Kiderlen AF, Kayser O (2005) Amphotericin B. Appl Microbiol Biotechnol 68(2):151–162
Marx F (2004) Small, basic antifungal proteins secreted from filamentous ascomycetes: a comparative study regarding expression, structure, function and potential application. Appl Microbiol Biotechnol 65(2):133–142
Marx F, Haas H, Reindl M, Stoffler G, Lottspeich F, Redl B (1995) Cloning, structural organization and regulation of expression of the Penicillium chrysogenum paf gene encoding an abundantly secreted protein with antifungal activity. Gene 167(1–2):167–171
Marx F, Binder U, Leiter E, Pocsi I (2008) The Penicillium chrysogenum antifungal protein PAF, a promising tool for the development of new antifungal therapies and fungal cell biology studies. Cell Mol Life Sci 65(3):445–454
Meyer V (2008) A small protein that fights fungi: AFP as a new promising antifungal agent of biotechnological value. Appl Microbiol Biotechnol 78(1):17–28
Mitsuhara I, Matsufuru H, Ohshima M, Kaku H, Nakajima Y, Murai N, Natori S, Ohashi Y (2000) Induced expression of sarcotoxin IA enhanced host resistance against both bacterial and fungal pathogens in transgenic tobacco. Mol Plant Microbe Interact 13(8):860–868
Moreno AB, Peñas G, Rufat M, Bravo JM, Estopà M, Messeguer J, San Segundo B (2005) Pathogen-induced production of the antifungal AFP protein from Aspergillus giganteus confers resistance to the blast fungus Magnaporthe grisea in transgenic rice. Mol Plant Microbe Interact 18(9):960–972
Ouedraogo JP, Hagen S, Spielvogel A, Engelhardt S, Meyer V (2011) Survival strategies of yeast and filamentous fungi against the antifungal protein AFP. J Biol Chem 286(16):13859–13868
Rodriguez-Martin A, Acosta R, Liddell S, Nunez F, Benito MJ, Asensio MA (2010) Characterization of the novel antifungal protein PgAFP and the encoding gene of Penicillium chrysogenum. Peptides 31(4):541–547
Sharma A, Sharma R, Imamura M, Yamakawa M, Machii H (2000) Transgenic expression of cecropin B, an antibacterial peptide from Bombyx mori, confers enhanced resistance to bacterial leaf blight in rice. FEBS Lett 484(1):7–11
Skouri-Gargouri H, Gargouri A (2008) First isolation of a novel thermostable antifungal peptide secreted by Aspergillus clavatus. Peptides 29(11):1871–1877
Skouri-Gargouri H, Jellouli-Chaker N, Gargouri A (2010) Factors affecting production and stability of the AcAFP antifungal peptide secreted by Aspergillus clavatus. Appl Microbiol Biotechnol 86(2):535–543
Theis T, Stahl U (2004) Antifungal proteins: targets, mechanisms and prospective applications. Cell Mol Life Sci 61(4):437–455
Theis T, Wedde M, Meyer V, Stahl U (2003) The antifungal protein from Aspergillus giganteus causes membrane permeabilization. Antimicrob Agents Chemother 47(2):588–593
van den Berg MA, Albang R, Albermann K, Badger JH, Daran JM, Driessen AJ, Garcia-Estrada C, Fedorova ND, Harris DM, Heijne WH, Joardar V, Kiel JA, Kovalchuk A, Martin JF, Nierman WC, Nijland JG, Pronk JT, Roubos JA, van der Klei IJ, van Peij NN, Veenhuis M, von Dohren H, Wagner C, Wortman J, Bovenberg RA (2008) Genome sequencing and analysis of the filamentous fungus Penicillium chrysogenum. Nat Biotechnol 26(10):1161–1168
van’t Hof W, Veerman EC, Helmerhorst EJ, Amerongen AV (2001) Antimicrobial peptides: properties and applicability. Biol Chem 382(4):597–619
White TF, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky FS, White TT (eds) PCR protocol: a guide to methods and applications. Academic, San Diego, pp 315–322
Wnendt S, Ulbrich N, Stahl U (1994) Molecular cloning, sequence analysis and expression of the gene encoding an antifungal-protein from Aspergillus giganteus. Curr Genet 25(6):519–523
Wong JH, Ng TB (2003) Gymnin, a potent defensin-like antifungal peptide from the Yunnan bean (Gymnocladus chinensis Baill). Peptides 24(7):963–968
Wong JH, Hao J, Cao Z, Qiao M, Xu H, Bai Y, Ng TB (2008) An antifungal protein from Bacillus amyloliquefaciens. J Appl Microbiol 105(6):1888–1898
Woo JH, Kitamura E, Myouga H, Kamei Y (2002) An antifungal protein from the marine bacterium Streptomyces sp. strain AP77 is specific for Pythium porphyrae, a causative agent of red rot disease in Porphyra spp. Appl Environ Microbiol 68(6):2666–2675
Zasloff M (2002) Antimicrobial peptides of multicellular organisms. Nature 415(6870):389–395
Zhang B, Xie C, Yang X (2008) A novel small antifungal peptide from Bacillus strain B-TL2 isolated from tobacco stems. Peptides 29(3):350–355
Acknowledgments
We thank Xingzhong Qiao and Tianhua Zhong for their assistance with the experiments. The work was supported by grants from the Scientific Research Project of the Marine Public Welfare Industry of China (201005032, 201205020), the Nature Science Foundation of China (40930847), and China Polar Environment Comprehensive investigation and Assessment Program (CHINARE2013-01-06 and CHINARE2013-04-03).
Conflict of interest
None.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Chen, Z., Ao, J., Yang, W. et al. Purification and characterization of a novel antifungal protein secreted by Penicillium chrysogenum from an Arctic sediment. Appl Microbiol Biotechnol 97, 10381–10390 (2013). https://doi.org/10.1007/s00253-013-4800-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-013-4800-6