Abstract
Corynebacterium glutamicum, the industrial microbe traditionally used for the production of amino acids, proved its value for the fermentative production of diverse products through genetic/metabolic engineering. A successful demonstration of the heterologous expression of arabinose and xylose utilization genes made them interesting biocatalysts for pentose fermentation, which are the main components in lignocellulosic hydrolysates. Its ability to withstand substantial amount of general growth inhibitors like furfurals, hydroxyl methyl furfurals and organic acids generated from the acid/alkali hydrolysis of lignocellulosics in growth arrested conditions and its ability to produce amino acids like glutamate and lysine in acid hydrolysates of rice straw and wheat bran, indicate the future prospective of this bacterium as a potent biocatalyst in fermentation biotechnology. However, the efforts so far on these lines have not yet been reviewed, and hence an attempt is made to look into the efficacy and prospects of C. glutamicum to utilize the normally non-fermentable pentose sugars from lignocellulosic biomass for the production of commodity chemicals.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Industrial applications of Corynebacterium glutamicum have been well known since 1957, when Kinoshita and co-workers isolated it from soil sample collected from the Ueno Zoo in Tokyo. To date, it remains as the most widely used industrial strain for the amino acid production mainly l-glutamate and l-lysine, (Leuchtenberger et al. 2005; Liebl 1992; Takors et al. 2007). Extensive studies on the metabolic pathways (Ikeda et al. 2006; Marx et al. 1996, 1997; Sonntag et al. 1995; Wendisch 2007) resulted in genetically engineered C. glutamicum with enhanced product range including various amino acids such as l-phenylalanine, l-aspartate, l-tryptophan, l-arginine, l-valine, nucleic acids like purines, vitamins such as riboflavin and pantothenic acid (Burkovski 2008; Eggeling and Bott 2005), and significant amounts of organic acids like acetic, lactic and succinic acid (Inui et al. 2004b; Kinoshita 1985; Terasawa and Yukawa 1993). In addition to these, the efficacy of C. glutamicum to express the recombinant proteins such as the transglutaminase of Streptomyces mobaraensis, the subtilisin-like serine protease of Streptomyces albogriseolus and human epidermal growth factor hEGF were also demonstrated (Date et al. 2003, 2006; Kikuchi et al. 2003). Yet another product of industrial importance is poly(3-hydroxybutyrate), produced by manipulating the biosynthetic pathway of C. glutamicum by the introduction of the phbCAB operon derived from Ralstonia eutropha (Jo et al. 2006). Putrescine and cadaverine are also products of corynebacterial metabolism through metabolic engineering. Cadaverine production was realized by replacing l-homoserine dehydrogenase gene (hom) of a lysine producing strain with l-lysine decarboxylase gene (cadA) of Escherichia coli (Mimitsuka et al. 2007). On the other hand, heterologous expression of genes encoding enzymes of the arginine- and ornithine decarboxylase pathways from E. coli enabled the production of putrescine (Schneider and Wendisch 2010). Recent advances in metabolic engineering strongly recommend C. glutamicum for its ability to transform renewable biomasses to value-added products.
Pentose sugar-rich lignocellulosic biomass, an abundant renewable source of energy, holds a lion’s share of the world’s total agricultural, industrial and forest residual biomass and has been underutilized for a long time (Dien et al. 2003). It generally consists of 10–20% lignin, 40–50% cellulose and 20–30% hemicellulose (Wyman 1999). Bioconversion of lignocellulosic biomass to industrially important products is possible only after its conversion to monomeric sugars, mostly by enzymatic hydrolysis. A pretreatment or prehydrolysis by acid or alkali at higher temperature before enzyme hydrolysis can increase the accessibility of enzyme for its substrate there by increasing the efficiency of hydrolysis (Anuj et al. 2011).
Pretreatment of lignocellulosics can give rise to both cellulose and hemicellulose. A direct conversion of cellulose and hemicellulose in to value-added products by fermentation using C. glutamicum is almost impossible unless, it is converted to its monomeric forms: pentoses and hexoses. Hexoses can be readily taken up by almost all organisms. But pentose sugars cannot be utilized by most of the industrial strains including wild-type C. glutamicum under normal conditions (Collins and Cummins 1984; Lee 1997). In order to attain a maximum of 60–80% process efficiency, the organism has to utilize both pentoses and hexoses. Even though some of the industrial strains are capable of pentose fermentation (Jeffries et al. 2000) as shown in Table 1, most of them are sensitive to inhibitory by-product of lignocellulosic biomass pretreatment. But C. glutamicum showed remarkable resistance towards inhibitors like furfural, 5-hydroxymethylfurfural, 4-hydroxybenzaldehyde (Sakai et al. 2007) under growth arrested conditions. In our previous work (Gopinath et al. 2011), we demonstrated the ability of C. glutamicum to grow under normal fermentation conditions in acid hydrolysates of rice straw and wheat bran to produce amino acids and hence a direct fermentation of pretreated biomass is possible without any detoxification. But all isolates so far identified as C. glutamicum lacked the genes for pentose metabolism (except ATCC31831, which possess arabinose metabolising genes). However, different strategies have been adopted in order to overcome this limitation, and this mini-review is mainly focused on summarizing these developments in C. glutamicum for its ability to convert pentose sugars to value-added commodities.
Pentose utilization by wild-type C. glutamicum
The pentose utilization capability and the associated process performance differ extensively among C. glutamicum strains even though they are reported to utilize hexoses and mixed sugars efficiently.
Out of the different strains of C. glutamicum isolated so far (Liebl 2005), the genome sequences of two isolates, C. glutamicum ATCC13032 (Kalinowski et al. 2003; Ikeda and Nakagawa 2003) and C. glutamicum R (Yukawa et al. 2007), have been determined and annotated independently. Both of these sequences confirmed the absence of arabinose metabolising genes in these strains. Interestingly, another strain, C. glutamicum ATCC31831 possesses a functional cluster of genes for l-arabinose utilization (Kawaguchi et al. 2009). It encodes the l-arabinose transporter araE (probably a low affinity H+ symporter), the l-arabinose isomerase araA (catalyzes isomerization of l-arabinose to l-ribulose), the ribulokinase araB (catalyzes the phosphorylation of l-ribulose to l-ribulose-5-phosphate) and the l-ribulose-5- phosphate 4-epimerase, araD, which catalyzes the formation of the PPP intermediate d-xylulose-5-phosphate from l-ribulose-5-phosphate (Sprenger 1995). According to Kawaguchi et al. (2009), the ara gene cluster might have been acquired from another high G + C content Gram-positive bacterium present in soil by a recent horizontal transfer event after the phylogenic branch containing the genus Corynebacterium.
The metabolic pathway for d-ribose catabolism is probably present in all strains of C. glutamicum. Formation of the PPP intermediate d-ribose-5-phosphate from internal d-ribose is catalyzed by two ribokinases: RbsK1 and RbsK2 (Brinkrolf et al. 2008).
The normal xylose catabolic pathway in bacteria consists of conversion of xylose to xylulose by xylose isomerase (xylA) and subsequent phosphorylation of xylulose by xylulose kinase (xylB) to the PPP intermediate d-xylulose-5-phosphate. But none of the C. glutamicum strains so far isolated including C. glutamicum ATCC13032 and C. glutamicum R (Ikeda and Nakagawa 2003; Kalinowski et al. 2003; Yukawa et al. 2007) were reported to utilize xylose, in the absence of xylA genes in them. The final gene for the xylose catabolism xylulokinase (xylB) has been found functional in C. glutamicum R (Kawaguchi et al. 2006). In general, pathways for pentose catabolism present in C. glutamicum strains finally yield pentose phosphate pathway (PPP) intermediates, which are metabolized through the non-oxidative part of the PPP (Blombach and Seibold 2010). Fungi more commonly transform d-xylose into xylitol by using xylose reductase and xylitol dehydrogenase with some exceptions like Piromyces which follows the bacterial pathway (Harhangi et al. 2003). Caulobacter crescentus, a freshwater bacterium expresses an NAD-dependent xylose dehydrogenase encoded by the xylose inducible xylXABCD operon, and follows a distinct xylose utilizing pathway yielding α-ketoglutarate as shown in Fig. 1 (Stephens et al. 2007).
Genetic engineering in C. glutamicum for pentose utilization
The bacterial pathway for arabinose utilization is charted in Fig. 2 (Sprenger 1995). In the fungal pathway like that of Pichia stipitis, Candida tenuis and Spathaspora passalidarum (Fig. 3), aldose reductase reduces l-arabinose to l-arabitol, which is further oxidized to l-xylulose by l-arabinitol 4-dehydrogenase. l-Xylulose is converted to xylitol by l-xylulose reductase and xylitol is then converted to d-xylulose by d-xylulose reductase. Finally xylulokinase will convert d-xylulose to PPP intermediate d-xylulose-5-P (Richard et al. 2001; Hahn-Hagerdal et al. 2007a, b). Surprisingly, both of these pathways are absent in all the C. glutamicum except in C. glutamicum ATCC31831.
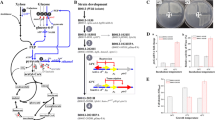
Genetically engineered arabinose utilizing C. glutamicum strains are constructed by introducing E. coli araBAD operon (Fig. 4a) to C. glutamicum R strain (Kawaguchi et al. 2008), glutamate producer ATCC13032 and lysine producer DM1729 (Schneider et al. 2010). The recombinant of C. glutamicum R, under oxygen-deprived condition with l-arabinose as the sole carbon source produced significant quantities of succinic acid and lactic acid together with small amounts of acetic acid. When a mixture of 5% d-glucose and 1% l-arabinose is used under the same condition, a constant metabolism of l-arabinose resulted in a combined organic acid yield based on the sugar mixture consumed. This showed that the engineered strain is able to utilize l-arabinose as a substrate for organic acid production even in the presence of d-glucose. Similarly, heterologous expression of the araBAD operon in the wild type and in l-lysine producing C. glutamicum (Schneider et al. 2010) resulted in a mutant strain for production of both l-glutamate and l-lysine from arabinose as sole carbon source. They also constructed l-ornithine and l-arginine producing C. glutamicum strains for arabinose utilization by expressing E. coli araBAD operon. They were able to produce l-glutamate, l-lysine, l-ornithine and l-arginine, respectively, from pure arabinose alone and also from pure glucose–arabinose blends as carbon sources.
a Arabinose operon of E. coli (Schleif 2000). Arabinose isomerase, encoded by araA, coverts arabinose to ribulose; ribulokinase, encoded by araB, phosphorylates ribulose; ribulose-5-phosphate epimerase, encoded by araD, converts ribulose-5-phosphate to xylulose-5-phosphate, which can then be metabolized via the pentose phosphate pathways. The three structural genes are arranged in an operon that is regulated by the araC gene product. b Xylose operon of E. coli (Song and Park 1997). The sugar is first isomerized into d-xylulose by xylose isomerase (XylA) and then phosphorylated by xylulokinase (XylB) to produce d-xylulose5-phosphate. XylF as the xylose-binding protein, XylG as an ATP-binding protein, and XylH as a membrane transporter.They are linked to xylAB but are oriented in the opposite direction. Expression of xylA, xylB, and the transport genes is under the positive control of xylR
The C. glutamicum usually do not use d-xylose as a carbon source; however, it harbours a gene encoding the xylulokinase (xylB) (Kawaguchi et al. 2006), which is capable of catalyzing phosphorylation of d-xylulose to the PPP intermediate d-xylulose-5-phosphate, the last step in the d-xylose metabolism. Kawaguchi et al. (2006) constructed two recombinant C. glutamicum strains with gene xylA encoding xylose isomerase alone (strain CRX1) and in combination with the E. coli gene xylB (strain CRX2) for the utilization of xylose. The genes in the xylose operon of E. coli (Fig. 4b) were provided on a high-copy-number plasmid and were expressed under the control of the constitutive promoter trc, derived from plasmid pTrc99A. Both recombinant strains were able to grow in mineral medium containing xylose as the sole carbon source. Strain CRX2 showed faster growth on xylose in comparison to CRX1 and produced lactic and succinic acids predominantly by consuming xylose under growth arrested condition. Moreover, in mineral medium containing a sugar mixture of 5% glucose and 2.5% xylose, strain CRX2 cells simultaneously consumed both sugars under oxygen-deprived conditions, demonstrating the absence of diauxic phenomena relative to the new xylA-xylB construct, albeit glucose-mediated regulation still exerted a measurable influence on xylose consumption kinetics.
Pentose uptake and transport engineering in C. glutamicum
Ribose uptake in C. glutamicum is facilitated by an ATP binding cassette transporter ABCrib (Nentwich 2009). RbsACBD-deletion mutant C. glutamicum SN1 completely lacks d-ribose uptake and growth when d-ribose is used as sole carbon source, explaining the role of rbsABCD (TC 3.A.1.2.1) in ribose transport. In C. glutamicum ATCC31831, the gene cluster for the arabinose metabolism includes an arabinose transporter gene (araE) and a metabolic operon, araBAD (Kawaguchi et al. 2009). The role of ara E (TC 2.A.1.1.55) found in Bacillus subtilis (Krispin and Allmansberger 1998), and C. glutamicum are found to be similar even though amino acid similarity between the two araE proteins found is only 34%. Growth experiments with an araE deletion mutant of C. glutamicum ATCC31831 revealed its functionality in high affinity uptake of l-arabinose. Low-affinity uptake systems for l-arabinose are probably present in C. glutamicum strains which are functional only at higher concentrations of l-arabinose (Kawaguchi et al. 2009). By expressing araE in C. glutamicum R harbouring arabinose-catabolising genes from E. coli, Sasaki and coworkers demonstrated a 3-fold increase in the arabinose consumption over a strain without araE (Sasaki et al. 2009).
Xylose assimilation in xylitol producing strain also suggested that the wild type possesses a transporter associated with xylose uptake. The mechanisms of pentose transport in C. glutamicum are still not yet clearly understood, and an unidentified transporter (IMxyl) was also reported for d-xylose uptake in C. glutamicum R (Blombach and Seibold 2010). E. coli genes for arabinose (araBAD) or xylose metabolism (xylAB) in C. glutamicum R strain were introduced for utilization of the respective sugars under both aerobic and oxygen-deprived conditions (Kawaguchi et al. 2006, 2008). Heterologous expression of araE from C. glutamicum ATCC31831 in C. glutamicum R also contributes to d-xylose uptake (Sasaki et al. 2009). It enhanced 3-fold xylose consumption at low concentration of xylose for its catabolising pathway (Sasaki et al. 2009). Particularly noteworthy was the observation in the studies of C. glutamicum that pentose consumption by either growing or oxygen-deprived cells of recombinant strains expressing araE was not repressed in the presence of glucose (Sasaki et al. 2009). Fast consumption of xylose under oxygen-deprived conditions when compared to fully aerobic conditions indicates the influence of molecular oxygen in xylose transport (Kim et al. 2010).
Chemical commodities produced by C. glutamicum from pentoses
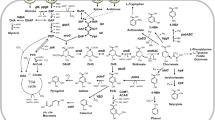
Since its discovery, C. glutamicum was manipulated as an efficient and robust biocatalyst for the manufacture of commodity chemicals including organic acids, amino acids, sugar alcohols. The central metabolic pathways have been engineered for pentose sugar utilization and to direct the flux towards the product of interest. Figure 5 summarises the flow chart indicating the value-added commodities obtained through the manipulation of Corynebacterium central metabolism and some the important commodities were analysed in detail.
Organic acids
Microbial organic acid production is attributed to excretion under particular physiological conditions or as intermediates of major metabolic pathways. The overproduction of any of the organic acids depends on the ability of the microorganism to reduce its by-product formation and these pathways can be diverted by metabolic engineering and gene manipulation to organic acids of interest. C. glutamicum is reported to produce high yields of various organic acids including l-lactic acid, succinic acid and acetic acid in mineral salts medium (Inui et al. 2004a). The crucial parameter leading to the production in this case is oxygen deprivation, achieved by high cell density and mild agitation without aeration. Okino and co-workers (2008) found that under this physical condition, l-lactic acid production is in high volumetric capacity in comparison with succinic acid and acetic acid. This high productivity of lactic acid was further refined by overexpression of d-LDH (lactate dehydrogenase) genes from E. coli and Lactobacillus delbrueckii in l-LDH (l -lactate dehydrogenase) encoding ldhA-null C. glutamicum mutants. Likewise the disruption of ldhA gene for l-LDH and overexpression of pyc gene for pyruvate carboxylase in another recombinant redirects the flux towards succinic acid with resultant enhancement in productivity in the presence of added bicarbonate (Okino et al. 2008a, 2008b). LDHs are enzymes that enable both production and utilization of lactate. LdhA, encoded by the ldhA gene, is the fermentative lactate producing LDH and is NAD-dependent. On the other end are the respiratory LDHs. LldD, encoded by the lldD gene, is the membrane-bound, quinone-dependent LDH essential for growth on l-lactate and Dld encoded by dld gene is the quinone dependent LDH responsible for growth on d-lactate (Stansen et al. 2005; Kato et al. 2010). Even though there are reports of ldh-encoded NAD+-dependent LDH from C. glutamicum ATCC 13032 operating unidirectionally for l-lactate formation, Sharkey et al. (2011) found that it can act in both directions with Vmax for the reduction of pyruvate being approximately 10-fold that of the value for l-lactate oxidation.
C. glutamicum has been metabolically engineered for organic acid production from lignocellulosic hydrolysates containing pentoses like xylose and arabinose. C. glutamicum CRX2 was able to accumulate significant amounts of lactic acid (0.45 g/g) and succinic acid (0.2 g/g) under oxygen deprived conditions from mineral medium containing xylose as carbon source. The usefulness of the strain for organic acid production from glucose–xylose mixtures was also demonstrated without negative effect on organic acid production and xylose consumption (Kawaguchi et al. 2006). Organic acid production from arabinose was best demonstrated by recombinant C. glutamicum CRA1, developed by incorporation of E. coli genes araA, araB and ara C encoding arabinose catabolising enzymes. Succinic acid, lactic acid and acetic acid (0.45, 0.34 and 0.06 g/g) were respectively produced with arabinose as the carbon source under oxygen deprivation. The strain was versatile in utilizing mixed sugars (Kawaguchi et al. 2008)
Alcohol production
Besides the general consideration as GRAS, it was the properties like tolerance to ethanol, inhibitors and broad substrate utilization range of engineered C. glutamicum strains makes it as a potential ethanologen. C. glutamicum has been used as a versatile biocatalyst for alcohol production under growth arrested conditions and is engineered to broaden the substrate utilization range. Metabolic engineering for pentose utilizing C. glutamicum CRX2 and CRA1 (for xylose and arabinose metabolism) has been done by Kawaguchi et al. 2006 and Kawaguchi et al. 2008, respectively. Engineering for cellobiose utilization was also carried out in C. glutamicum R. This strain was capable of methyl β-glucoside and the natural aryl β-glucosides, e.g., salicin and arbutin utilization depending on the PTS (Phospho Transferase System). bglA and bglF are genes encoding PTS permease and β glucosidase, respectively, and are components of the β-glucoside PTS and was demonstrated that a point mutation in the codon 317 of the bglF(bglF317A) led to substitutions V317A and/or V317M near the putative PTS active-site H313 in the membrane-spanning IIC domain of bglF (TC 4.A.1.2.5) and allowed bglF to act on cellobiose (Kotrba et al. 2003). Substrate utilization range was extended to include d-xylose. For this another strain X5C1 was constructed by integration of E. coli xylA and xylB genes and the C. glutamicum R bglF 317A and bglA genes into the chromosomal DNA of C. glutamicum R, which can simultaneously utilize d-cellobiose, d-glucose, and d-xylose (Sasaki et al. 2008). The ethanol producing C. glutamicum was constructed by the introduction pdc and adhB genes of Zymomonas mobilis. Under the influence of IdhA promoter, these genes coding for enzymes pyruvate decarboxylase and alcohol dehydrogenase efficiently synthesized ethanol from pyruvate under oxygen-deprived conditions. The addition of small quantities of pyruvate and acetaldehyde is shown to increase the sugar consumption and ethanol production in strains deficient of organic acid synthesizing genes; illustrated with ethanol production reaching up to 642 mmol l h−1 in ldhA-deficient strain(R-ldhA-pCRA723) with pyruvate addition under oxygen deprivation (Inui et al. 2004a). Albeit the ethanol production capacity, it is capable of growing on ethanol with growth rates up to 0.24 h−1 and biomass yields up to 0.47 g dry weight (g ethanol)−1. The cells grown on ethanol showed increased expression of genes encoding phosphotransacetylase (PTA), isocitrate lyase (ICL) and malate synthase (MS) and acetate kinase (AK) (Arndt et al. 2008). Moreover ethanol producing C. glutamicum under the growth arrested conditions, showed tolerance towards inhibitors like furan, organic acids and phenolics, which can be found in various pretreated lignocellulosic hydrolysates. The C. glutamicum strain under study R-ldhA-pCRA723 retained almost 100% ethanol productivity with model composition of inhibitors simulating various pretreatment conditions.
Recent advances in C. glutamicum research showed their competence for alcohols other than ethanol. As C. glutamicum is an established amino acid producer and the 2-keto acid pathways for higher chain alcohol synthesis share common precursors with amino acids, this microbe shows excellent potential for higher chain alcohol. By manipulating the 2-keto acid pathways, it is able to produce isobutanol from pyruvate with a production reaching up to 0.1 g/g (Smith et al. 2010). The C. glutamicum also showed better isobutanol tolerance compared to E. coli. Overexpression of alsS (acetolactate synthase) from B. subtilis, ilvC (acetohydroxy acid isomer reductase) and ilvD (dihydroxy-acid dehydratase) of C. glutamicum, kivd (keto isovalerate decarboxylase) of Lactococcus lactis, and a native adhA (alcohol dehydrogenase), led to the production of 2.6 g/l isobutanol and 0.4 g/l 3-methyl-1-butanol in 48 h. In addition, other higher chain alcohols such as 1-propanol, 2-methyl-1-butanol, 1-butanol, and 2-phenylethanol were also detected as by products (Smith et al. 2010). Recently, Blombach et al. (2011) engineered C. glutamicum for the production of isobutanol from glucose under oxygen deprivation conditions by inactivation of l-lactate and malate dehydrogenases, implementation of keto acid decarboxylase from L. lactis, alcohol dehydrogenase 2 (ADH2) from Saccharomyces cerevisiae, and expression of the pntAB transhydrogenase genes from E. coli. Based on a 2-ketoisovalerate production strain engineered by inactivation of the pyruvate dehydrogenase complex, pyruvate: quinone oxidoreductase, transaminase B, and additional overexpression of the ilvBNCD genes, encoding acetohydroxyacid synthase, acetohydroxyacid isomeroreductase, and dihydroxyacid dehydratase. The resulting strain produced isobutanol with a substrate-specific yield (YP/S) of 0.60 mol per mol glucose. Overexpression of the corynebacterial chromosomal adhA gene increased the YP/S to 0.77 mol of isobutanol per mol of glucose. Inactivation of the malic enzyme significantly reduced the YP/S, indicating that the metabolic cycle consisting of pyruvate and/or phosphoenolpyruvate carboxylase, malate dehydrogenase, and malic enzyme is responsible for the conversion of NADH + H+ to NADPH + H+ (Blombach et al. 2011). By taking all these points into consideration, a cost-effective production of various alcohols from lignocellulosic biomass is well possible with a pentose utilizing mutant of C. glutamicum.
Sugar alcohols
Xylitol, a sugar alcohol derivative of xylose, possesses health beneficial properties which have resulted in a rapid expansion in xylitol consumption to the current annual global value of $340 million (Kadam et al. 2008). Being a valuable synthetic building block (Granstrom et al. 2007), it recently joined the top 12 value-added materials produced from biomass, thereby serving as a key economic driver for the biorefinery concept (Werpy et al. 2004). Wild-type C. glutamicum produced 0.019 g/g xylitol, indicating that it harbours a gene encoding enzymes for xylitol production but the enzyme activities are very low (Sasaki et al. 2010). As reported by Doo et al. (2009), C. glutamicum is a promising biocatalyst for biotransformation and it lacks the xylose isomerase gene (Kawaguchi et al. 2006), and hence is manipulated for the production of xylitol from d-xylose (Kim et al. 2010). The xylose reductase gene of P. stipitis was expressed in C. glutamicum, enabling the recombinant strain to convert d-xylose into xylitol efficiently with a molar yield over 0.97. The biotransformation was markedly influenced by the glucose concentration and oxygen availability in the medium, which appeared to cause lose catabolite repression on xylose transport and NADPH regeneration ability of the organism. By doing biotransformation under glucose and oxygen-limited conditions, xylitol was produced at a concentration of 34.4 g/l (226 mM) with an average specific productivity and molar yield to glucose of 0.092 g (g CDW)−1 h−1 and 1.6 mol/mol, respectively (Kim et al. 2010). In the view that the biocatalytic productivity could be further improved by enhancing the transport efficiency of xylose into the cells, Sasaki et al. (2010) incorporated the pentose transporter araE gene of C. glutamicum 31831 into C. glutamicum R strain (CtXR1) with xylose reductase gene to form CtXR2, which had 13-fold higher xylitol production than its parental strain. In C. glutamicum, xylitol is transported back in to the cells by a phosphoenolpyruvate dependent fructose phosphotransferase system (PTSfru) (Dominguez and Lindley 1996). The xylitol production was further increased by disrupting this phosphotransferase (PTSfru) and xylulose kinase (xylB) genes (Sasaki et al. 2010). For the efficient conversion of hemicellulosic portion of agro residues to xylitol, it is necessary to utilize arabinose also. Introduction of arabinose reductase and arabitol dehydrogenase to C. glutamicum may enable the conversion of arabinose into xylitol and need to be studied in detail.
Amino acids
Amino acids like l-glutamate, l-lysine, l-arginine, and l-ornithine were produced from arabinose as sole carbon source by genetically engineered C. glutamicum strains (Schneider et al. 2010). Arabinose utilizing amino acid secreting strains were constructed by the heterologous expression of the araBAD operon from E. coli into the corresponding amino acid producing strain of C. glutamicum. l-Glutamate and l-lysine producing arabinose utilizing strains were constructed by the recombination of E. coli araBAD operon in ATCC13032 and DM1729 strains, respectively. l-Ornithine production by the recombinant C. glutamicum strain was obtained by deletion of argR for pathway de-repression and deletion of argF to block l-ornithine conversion. Transformation of the ornithine producer with the plasmid pVWEx1-araBAD for arabinose utilization allowed for arabinose-based ornithine production with yields comparable to those obtained with glucose. l-Arginine production by a recombinant C. glutamicum strain carrying an argR deletion for pathway de-repression and expressing a feedback insensitive variant of endogenous N-acteylglutamate kinase from glucose was about twice as high as described for a similar strain (Ikeda et al. 2009). More importantly, l-arginine production from arabinose as sole carbon source and as combined carbon source was even slightly higher. While l-ornithine and l-arginine producing strains were constructed and shown to produce l-ornithine and l-arginine from arabinose when araBAD from E. coli was expressed. Moreover, the recombinant strains produced l-glutamate, l-lysine, l-ornithine and l-arginine, respectively, from arabinose also when glucose arabinose blends were used as carbon sources. This provided a basis for efficient amino acid production from sugars present in lignocellulosic hydrolysates.
While xylose utilizing C. glutamicum R strains were constructed by heterologous expression of E. coli xylose isomerase gene (xylA), which converts xylose to xylulose, the second enzyme xylulokinase (xylB) is already present in C. glutamicum R (Kawaguchi et al. 2006). Although in their studies, they focused on organic acid production, this strain is further engineered to produce the amino acid l-alanine. l-Alanine producing C. glutamicum R strain has been created by deleting the genes associated with production of organic acids and overexpressing alanine dehydrogenase gene (alaD) from Lysinibacillus sphaericus (Jojima et al. 2010). Thus, amino acids were successfully produced by engineered C. glutamicum by utilising pentose sugars. Very recently, the co-utilization of hexose and pentose sugars present in wheat bran and rice straw hydrolysates for the production of l-lysine and l-glutamate by genetically modified C. glutamicum was demonstrated by our group (Gopinath et al. 2011) and this will be a beginning for developing a new bioprocess for amino acid production from hemicellulosic biomass.
Outlook and prospects
Fermentations for the production of commodity chemicals and fuels have always been in the quest of cost effectiveness and sustainability. Lignocellulosics were then identified as prime feed stock for alternative cheap and sustainable sugar supply. This mini-review illustrated the broad product range of speciality and platform chemicals produced from pentose sugars by the work horse of amino acid industry, C. glutamicum. This industrial strain has been moulded to demonstrate its flexible product range from amino acids to alcohols with respect to the pentose uptake from the underutilized lignocellulosic biomass. Simultaneous utilization of various carbon sources by C. glutamicum (Dominguez et al. 1997; Eggeling and Bott 2005; Engels et al. 2008; Lee et al. 1998; Wendisch 2006; Wendisch et al. 2000) is a hallmark of this bacterium setting it apart from yeasts, E. coli and B. subtilis, which typically show sequential utilization of substrates present in blends, and this growth pattern is often accompanied by a diauxic growth lag. On the other hand, C. glutamicum shows very few exceptions to substrate co-utilization (e.g., glucose being utilized prior to ethanol (Arndt et al. 2008; Arndt and Eikmanns 2008) or prior to glutamate (Kronemeyer et al. 1995). The better inhibitor tolerance along with this co-utilization ability makes C. glutamicum superior to other microbes for pentose utilization from lignocellulosic biomass hydrolysates. In-depth knowledge of the metabolic pathways and regulatory mechanisms of the organism made genetic engineering easy, and the pathways for pentose sugar utilization were incorporated in the central metabolic pathway of C. glutamicum. The recent advances in the metabolic engineering paved the way for the beginning of an epoch of cost-effective and sustainable biotechnological production processes.
Earnest efforts were made for making C. glutamicum an efficient pentose utilizer by metabolic engineering and transport engineering. Even though advances in metabolic engineering and membrane transport engineering made the desired product yield very viable and economical to a great extent. Gene regulatory engineering is also conceivable to improve pentose utilization, as it was observed (Kawaguchi et al. 2006, 2008; Schneider et al. 2010; Gopinath et al. 2011) that glucose slowed pentose utilization, indicating some sort of glucose repression to the endogenous hitherto-unknown pentose transporter gene(s) rather than plasmid-borne expression of the heterologous genes for arabinose and xylose catabolism.
C. glutamicum has proved itself a versatile microorganism with multiple product range and productivity. Most of the work we have come across in our mini-review was focused on the utilization of pure chemicals with either arabinose or xylose alone. But for the effective utilization of the lignocellulosic biomass, the co-utilization of all pentose fractions—mainly arabinose and xylose—is very essential. It is necessary to find what happens when this engineered strains were introduced to real life situation, i.e., in the hydrolysates of various agro residues and its economic scale. Also, enough attention must be given to the recovery and purification of the fermentation products derived from lignocellulosic biomass hydrolysates. Future developments in these directions will definitely enhance the potential of C. glutamicum as an efficient pentose utilizing biocatalyst for value addition.
References
Akinterinwa O, Khankal R, Cirino PC (2008) Metabolic engineering for bioproduction of sugar alcohols. Curr Opin Biotechnol 19:461–467. doi:10.1016/j.copbio.2008.08.002
Anuj KC, Gajula C, Konakalla R, Rudravaram R, Pogaku R (2011) Bioconversion of pentose sugars into ethanol: a review and future direction. Biotechnol Mol Biol Rev 6:8–20
Arndt A, Eikmanns BJ (2008) Regulation of carbon metabolism in Corynebacterium glutamicum. In: Burkovski A (ed) Corynebacteria: genomics and molecular biology. Caister Academic, Wymondham, pp 155–182
Arndt A, Auchter M, Ishiqe T, Wendisch VF, Eikmanns BJ (2008) Ethanol catabolism in Corynebacterium glutamicum. J Mol Microbiol Biotechnol 15:222–233. doi:10.1159/000107370
Arslan Y, Eken-Saracogl N (2010) Effects of pretreatment methods for hazelnut shell hydrolysate fermentation with Pichia Stipitis to ethanol. Bioresour Technol 101:8664–8670. doi:10.1016/j.biortech.2010.05.085
Blombach B, Seibold GM (2010) Carbohydrate metabolism in Corynebacterium glutamicum and applications for the metabolic engineering of l-lysine production strains. Appl Microbiol Biotechnol 86:1313–1322. doi:1007/s00253-010-2537-z
Blombach B, Riester T, Wieschalka S, Ziert C, Youn J, Wendisch VF, Eikmanns BJ (2011) Corynebacterium glutamicum tailored for efficient isobutanol production. Appl Environ Microbiol 77:3300–3310. doi:10.1128/AEM.02972-10
Brinkrolf K, Plöger S, Solle S, Brune I, Nentwich SS, Hüser AT, Kalinowski J, Pühler A, Tauch A (2008) The LacI/GalR family transcriptional regulator UriR negatively controls uridine utilization of Corynebacterium glutamicum by binding to catabolite-responsive element (cre)-like sequences. Microbiol 154:1068–1081
Burkovski A (2008) Corynebacteria: genomics and molecular biology. Caister, Academic Press, Erlangen, Germany
Cimpeanu C, Campeanu G, Begea M, Vladescu M, Cornea CP (2010) Bio ethanol production by new thermotolerant Romanian yeast strains. Rom Biotechnol Lett 15(3):5310–5316
Cirino PC, Chin JW, Ingram LO (2006) Engineering Escherichia coli for xylitol production from glucose–xylose mixtures. Biotechnol Bioeng 95:1167–1176. doi:10.1002/bit.21082
Collins MD, Cummins CS (1984) Genus Corynebacterium. ed. Trans. In: Sneath PHA, Mair NS, Sharpe ME, Holt JG (eds) Bergey’s manual of systematic bacteriology. Williams & Wilkins, Baltimore, MD, pp 1266–1283
Date M, Yokoyama K, Umezawa Y, Matsui H, Kikuchi Y (2003) Production of native-type Streptoverticillium mobaraense transglutaminase in Corynebacterium glutamicum. Appl Environ Microbiol 69:3011–3014. doi:10.1128/AEM.69.5.3011-3014.2003
Date M, Itaya H, Matsui HY (2006) Secretion of human epidermal growth factor by Corynebacterium glutamicum. Lett Appl Microbiol 42:66–70. doi:10.1111/j.1472-765X.2005.01802.x
Dien BS, Cotta MA, Jeffries TW (2003) Bacteria engineered for fuel ethanol production: current status. Appl Microbiol Biotechnol 63:258–266. doi:10.1007/s00253-003-1444-y
Dominguez H, Lindley ND (1996) Complete sucrose metabolism requires fructose phosphotransferase activity in Corynebacterium glutamicum to ensure phosphorylation of liberated fructose. Appl Environ Microbiol 62:3878–3880
Dominguez H, Cocaign-Bousquet M, Lindley ND (1997) Simultaneous consumption of glucose and fructose from sugar mixtures during batch growth of Corynebacterium glutamicum. Appl Microbiol Biot 47(5):600–603
Doo EH, Lee WH, Seo HS, Seo JS, Park JB (2009) Productivity of cyclohexanone oxidation of the recombinant Corynebacterium glutamicum expressing chnB of Acinetobacter calcoaceticus. J Biotechnol 142:164–169. doi:10.1016/j.jbiotec.2009.04.008
Eggeling L, Bott M (2005). Handbook of Corynebacterium glutamicum. CRC Boca Raton
Engels V, Georgi T, Wendisch VF (2008) ScrB (Cg2927) is a sucrose-6-phosphate hydrolase essential for sucrose utilization by Corynebacterium glutamicum. FEMS Microbiol Lett 289(1):80–89
Fang X, Huang W, Xia L (2004) Xylitol production from corn cob hemicellulosic hydrolysate by Candida sp. Sheng Wu Gong Cheng Xue Bao 20:295–298
Gopinath V, Meiswinkel TM, Wendisch VF, Nampoothiri KM (2011) Amino acid production from rice straw and wheat bran hydrolysates by recombinant pentose-utilizing Corynebacterium glutamicum. Appl Microbiol Biotechnol. doi:10.1007/s00253-011-3478-x
Grabar TB, Zhou S, Shanmugam KT, Yomano LP, Ingram LO (2006) Methylglyoxal bypass identified as source of chiral contamination in l(+) and d(−)-lactate fermentations by recombinant Escherichia coli. Biotechnol Lett 28:1527–1535
Granstrom TB, Izumori K, Leisola M (2007) A rare sugar xylitol: Part II. Biotechnological production and future applications of xylitol. Appl Microbiol Biotechnol 74(2):273–276. doi:10.1007/s00253-006-0760-4
Gururajan VT, Van Rensburgi P, Hahn-Hagerdal B, Pretorius IS, Otero RRC (2007) Development and characterisation of a recombinant Saccharomyces cerevisiae mutant strain with enhanced xylose fermentation properties. Ann Microbiol 57(4):599–607
Hahn-Hagerdal B, Karhumaa K, Fonseca C, Spencer-Martinz I, Gorwa-Grauslund MF (2007a) Towards industrial pentose-fermenting yeast strains. Appl Microbiol Biotechnol 74(5):937–953. doi:10.1007/s00253-006-0827-2
Hahn-Hagerdal B, Karhumaa K, Jeppsson M, Gorwa-Grauslund MF (2007b) Metabolic engineering for pentose utilization in Saccharomyces cerevisiae. Adv Biochem Eng Biotechnol 108:147–177. doi:10.1007/10_2007_062
Harhangi HR, Akhmanova AS, Emmens R, Van der Drift C, De Laat WTAM, Van Dijken JP, Jetten MSM, Pronk JT, Op den Camp HJM (2003) Xylose metabolism in the anaerobic fungus Piromyces sp. strain E2 follows the bacterial pathway. Arch Microbiol 180:134–141. doi:10.1007/s00203-003-0565-0
Helle SS, Murray A, Lam J, Cameron DR, Duff SJB (2004) Xylose fermentation by genetically modified Saccharomyces cerevisiae 259ST in spent sulfite liquor. Bioresour Technol 92(2):163–171. doi:10.1016/j.biortech.2003.08.011
Ikeda M, Katsumata R (1992) Metabolic engineering to produce tyrosine or phenylalanine in a tryptophan-producing Corynebacterium glutamicum strain. Appl Environ Microbiol 58(3):781–785
Ikeda M, Nakagawa S (2003) The Corynebacterium glutamicum genome: features and impacts on biotechnological processes. Appl Microbiol Biotechnol 62(2–3):99–109. doi:10.1007/s00253-003-1328-1
Ikeda M, Ohnishi J, Hayashi M, Mitsuhashi S (2006) A genome-based approach to create a minimally mutated, Corynebacterium glutamicm strain for efficient l-lysine production. J Ind Microbiol Biotechnol 33(7):610–615. doi:10.1007/s10295-006-0104-5
Ikeda M, Mitsuhashi S, Tanaka K, Hayashi M (2009) Reengineering of a Corynebacterium glutamicum l-arginine and l-citrulline producer. Appl Environ Microbiol 75(6):1635–1641. doi:10.1128/AEM.02027-08
Inui M, Kawaguchi H, Murakami S, Vertes AA, Yukawa H (2004a) Metabolic engineering of Corynebacterium glutamicum for fuel ethanol production under oxygen-deprivation conditions. J Mol Microbiol Biotechnol 8(4):243–254. doi:10.1159/000086705
Inui M, Murakami S, Okino S, Kawaguchi H, Vertes AA, Yukawa H (2004b) Metabolic analysis of Corynebacterium glutamicum during lactate and succinate productions under oxygen deprivation conditions. J Mol Microbiol Biotechnol 7(4):182–196. doi:10.1159/000079827
Jantama K, Zhang X, Moore JC, Shanmugam KT, Svoronos SA, Ingram LO (2008) Eliminating side products and increasing succinate yields in engineered strains of Escherichia coli C. Biotechnol Bioeng 101:881–893
Jeffries TW, Shi NQ, Jin YS, Cruz J (2000) metabolic pathway engineering for improved xylose fermentation by yeasts. Abstracts of Papers of the American Chemical Society, 219, 32-BTEC
Jo S, Maeda M, Ooi TS (2006) Production system for biodegradable polyester polyhydroxybutyrate by Corynebacterium glutamicum. J Biosci Bioeng 102:233–236. doi:10.1263/jbb.102.233
Jojima T, Fujii M, Mori E, Inui M, Yukawa H (2010) Engineering of sugar metabolism of Corynebacterium glutamicum for production of amino acid l-alanine under oxygen deprivation. Appl Microbiol Biotechnol 87(1):159–165. doi:10.1007/s00253-010-2493-7
Kadam KL, Chin CY, Brown LW (2008) Flexible biorefinery for producing fermentation sugars, lignin and pulp from corn stover. J Ind Microbiol Biotechnol 35(5):331–341. doi:10.1007/s10295-008-0322-0
Kalinowski J, Bathe B, Bartels D, Bischoff N, Bott M, Burkovski A, Dusch N, Eggeling L, Eikmanns BJ, Gaigalat L, Goesmann A, Hartmann M, Huthmacher K, Kramer R, Linke B, McHardy AC, Meyer F, Mockel B, Pfefferle W, Puhler A, Rey DA, Ruckert C, Rupp O, Sahm H, Wendisch VF, Wiegrabe I, Tauch A (2003) The complete Corynebacterium glutamicum ATCC 13032 genome sequence and its impact on the production of l-aspartate-derived amino acids and vitamins. J Biotechnol 104(1–3):5–25. doi:10.1016/S0168-1656(03)00154-8
Kato O, Youn JW, Stansen KC, Matsui D, Oikawa TVF (2010) Quinone-dependent d-lactate dehydrogenase Dld(Cg1027) is essential for growth of Corynebacterium glutamicum on d-lactate. BMC Microbiol 10:321
Kawaguchi H, Vertes AA, Okino S, Inui M, Yukawa H (2006) Engineering of a xylose metabolic pathway in Corynebacterium glutamicum. Appl Environ Microbiol 72(5):3418–3428. doi:10.1128/AEM.72.5.3418-3428.2006
Kawaguchi H, Sasaki M, Vertes AA, Inui M, Yukawa H (2008) Engineering of an l-arabinose metabolic pathway in Corynebacterium glutamicum. Appl Microbiol Biotechnol 77(5):1053–1062. doi:10.1007/s00253-007-1244-x
Kawaguchi H, Sasaki M, Vertes AA, Inui M, Yukawa H (2009) Identification and functional analysis of the gene cluster for l-arabinose utilization in Corynebacterium glutamicum. Appl Environ Microbiol 75:3419–3429. doi:10.1128/AEM.02912-08
Kikuchi Y, Date M, Yokoyama K, Umezawa Y (2003) Secretion of active-form Streptoverticillium mobaraense transglutaminase by Corynebacterium glutamicum: processing of the pro-domain by a co-secreted subtilisin-like protease from Streptomycesalbo griseolus. Appl Environ Microbiol 69:358–366. doi:10.1128/AEM.69.1.358366.2003
Kim TB, Lee YJ, Kim P, Kim CS, Oh DK (2004) Increased xylitol production rate during long-term cell recycle fermentation of Candida tropicalis. Biotechnol Lett 26(8):623–627. doi:10.1023/B:BILE.0000023019.02411.54
Kim SH, Yun JY, Kim SG, Seo JH, Park JB (2010) Production of xylitol from d-xylose and glucose with recombinant Corynebacterium glutamicum. Enzyme Microb Technol 46(5):366–371. doi:10.1016/j.enzmictec.2009.12.012
Kinoshita S (1985) Glutamic acid bacteria. In: Demain AL, Solomon NA (eds) Biology of industrial microorganisms. Benjamin Cummings Pub, London, pp 115–146
Ko CH, Chun P, Lin C, Hao K (2008) Xylitol conversion by fermentation using five yeast strains and polyelectrolyte-assisted ultrafiltration. Biotechnol Lett 30:81–86. doi:10.1007/s10529-007-9507-2
Kotrba P, Inui M, Yukawa H (2003) A single V317A or V317M substitution in Enzyme II of a newly identified β-glucoside phosphotransferase and utilization system of Corynebacterium glutamicum R extends its specificity towards cellobiose. Microbiology 149:1569–1580. doi:10.1099/mic.0.26053-0
Krispin O, Allmansberger R (1998) The Bacillus subtilis galE gene is essential in the presence of glucose and galactose. J Bacteriol 180:2265–2270
Kronemeyer W, Peekhaus N, Kramer R, Sahm H, Eggeling L (1995) Structure of the gluABCD cluster encoding the glutamate uptake system of Corynebacterium glutamicum. J Bacteriol 177(5):1152–1158
Lee J (1997) Biological conversion of lignocellulosic biomass to ethanol. J Biotechnol 56:1–24
Lee HW, Pan JG, Lebeault JM (1998) Enhanced l-lysine production in threonine-limited continuous culture of Corynebacterium glutamicum by using gluconate as a secondary carbon source with glucose. Appl Microbiol Biotechnol 49(1):9–15
Leuchtenberger W, Huthmacher K, Drauz K (2005) Biotechnological production of amino acids and derivatives: current status and prospects. Appl Microbiol Biotechnol 69(1):1–8. doi:10.1007/s00253-005-0155-y
Liebl W (1992) The genus Corynebacterium—nonmedical. In: Balows A, Truper HG, Dworkin M, Harder W, Schleifer KH (eds) The prokaryotes: a handbook on the biology of bacteria: ecophysiology, isolation, identification, applications, Volume 1, 2nd edn. Springer- Verlag, New York, pp 1157–1172
Liebl W (2005) Corynebacterium taxonomy. In: Eggeling L (ed) Bott M. Handbook of Corynebacterium glutamicum.CRC Press, Boca Raton, pp 9–34
Marx A, de Graaf AA, Wiechert W, Eggeling L, Sahm H (1996) Determination of the fluxes in the central metabolism of Corynebacterium glutamicum by nuclear magnetic resonance spectroscopy combined with metabolite balancing. Biotechnol Bioeng 49(2):111–129. doi:10.1002/(SICI)1097-0290(19960120)49:2<111::AID-BIT1>3.0.CO;2-T
Marx A, Striegel K, deGraaf AA, Sahm H, Eggeling L (1997) Response of the central metabolism of Corynebacterium glutamicum to different flux burdens. Biotechnol Bioeng 56(2):168180. doi:10.1002/(SICI)1097-0290(19971020)56:2<168::AID-BIT6>3.0.CO;2-N
Mimitsuka T, Sawai H, Hatsu MK (2007) Metabolic engineering of Corynebacterium glutamicum for cadaverine fermentation. Biosci Biotechnol Biochem 71:2130–2135. doi:10.1271/bbb.60699
Nentwich SS (2009) Characterization of the LacI-type transcriptional repressor RbsR controlling ribose transport in Corynebacterium glutamicum ATCC 13032. Microbiology 155:150–164
Nigam JN (2001) Development of xylose-fermenting yeast Pichia stipitis for ethanol production through adaptation on hardwood hemicellulose acid prehydrolysate. J Appl Microbiol 90:208–215
Okino S, Noburyu R, Suda M, Jojima T, Inui M, Yukawa H (2008a) An efficient succinic acid production process in a metabolically engineered Corynebacterium glutamicum strain. Appl Microbiol Biotechnol 81(3):459–464. doi:10.1007/s00253-008-1668-y
Okino S, Suda M, Fujikura K, Inui M, Yukawa H (2008b) Production of d-lactic acid by Corynebacterium glutamicum under oxygen deprivation. Appl Microbiol Biotechnol 78(3):449–454. doi:10.1007/s00253-007-1336-7
Patel M, Ou M, Ingram LO, Shanmugam KT (2004) Fermentation of sugar cane bagasse hemicellulose hydrolysate to l(+)-lactic acid by a thermotolerant acidophilic Bacillus sp. Biotechnol Lett 26(11):865–868
Pereira N Jr, Penuela V, de Souza MB Jr (2006) RSM Analysis of the effects of the oxygen transfer coefficient and size inoculum on Xylitol production by Candida guilliermondii. Appl Biochem Biotechnol 129:256–264
Rangaswamy S, Agblevor FA (2002) Screening of facultative anaerobic bacteria utilizing d-xylose for xylitol production. Appl Microbiol Biotechnol 60:88–93. doi:10.1007/s00253-002-1067-8
Rao RS, Prakasham RS, Prasad KK, Rajesham S, Sarma PN, Rao LV (2004) Xylitol production by Candida sp.: parameter optimization using Taguchi approach. Process Biochem 39(8):951–956. doi:10.1016/S0032-9592(03)00207-3
Richard P, Londesborough J, Putkonen M, Kalkkinen N, Penttila M (2001) Cloning and expression of a fungal l-arabinitol 4-dehydrogenase gene. J Biol Chem 276(44):40631–40637. doi:10.1074/jbc.M104022200
Rybak KV, Slivinskaya EA, Savrasova EA, Akhverdian VZ, Klyachko EV, Mashko SV, Doroshenko VG, Airikh LG, Leonova TV, Gusyatiner MM, Voroshilova EB, Kozlov YI, Hara Y, Ueda T (2011) Method for producing l-amino acids using bacteria of the Enterobacteriaceae family. United States Patent Application 20110143403. Ajinomoto Co., Inc, Tokyo
Sakai S, Tsuchida Y, Okino S, Ichihashi O, Kawaguchi H, Watanabe T, Inui M, Yukawa H (2007) Effect of lignocellulose-derived inhibitors on growth of and ethanol production by growth arrested Corynebacterium glutamicum R. Appl Environ Microbiol 73(7):2349–2353. doi:10.1128/AEM.02880-06
Sasaki M, Jojima T, Inui M, Yukawa H (2008) Simultaneous utilization of d-cellobiose, d-glucose, and d-xylose by recombinant Corynebacterium glutamicum under oxygen-deprived conditions. Appl Microbiol Biotechnol 81(4):691–699. doi:10.1007/s00253-008-1703-z
Sasaki M, Jojima T, Kawaguchi H, Inui M, Yukawa H (2009) Engineering of pentose transport in Corynebacterium glutamicum to improve simultaneous utilization of mixed sugars. Appl Microbiol Biotechnol 85(1):105–115. doi:10.1007/s00253-009-2065-x
Sasaki M, Jojima T, Inui M, Yukawa H (2010) Xylitol production by recombinant Corynebacterium glutamicum under oxygen deprivation. Appl Microbiol Biotechnol 86(4):1057–1066.doi: 10.1007/s00253-009-2372-2
Schleif R (2000) Regulation of the l-arabinose operon of Escherichia coli. Trends Genet 16:559–565. doi:10.1016/S0168-9525(00)02153-3
Schneider J, Wendisch VF (2010) Putrescine production by engineered Corynebacterium glutamicum. Appl Microbiol Biotechnol 88:859–868. doi:10.1007/s00253-010-2778-x
Schneider J, Niermann K, Wendisch VF (2010) Production of the amino acids l-glutamate, l-lysine, l-ornithine and l-arginine from arabinose by recombinant Corynebacterium glutamicum. J Biotechnol 154:191–198. doi:10.1016/j.jbiotec.2010.07.009
Sharkey MA, Maher MA, Guyonvarch APC (2011) Kinetic characterisation of recombinant Corynebacterium glutamicum NAD + −dependent LDH over-expressed in E. coli and its rescue of an lldD2 phenotype in C. glutamicum: the issue of reversibility re-examined. Arch Microbiol. doi: 10.1007/s00203-011-0711-z
Shin SJ, Cho DH, Kim YH, Cho NS (2010) Bioethanol fermentation of acid hydrolysates from yellow poplar using Pichia stipitis. Res Prog Pap Ind Biorefinery (4th ISETPP) 1–3:1344–1347
Silva JPA, Mussatto SI, Roberto IC, Teixeira JA (2011) Ethanol production from xylose by Pichia stipitis NRRL Y-7124 in a stirred tank bioreactor. Braz J Chem Eng in Life Sci 28:151–156. doi:10.1590/S0104-66322011000100016
Smith KM, Cho KM, Liao JC (2010) Engineering Corynebacterium glutamicum for isobutanol production. Appl Microbiol Biotechnol 87(3):1045–1055. doi:10.1007/s00253-010-2522-6
Sonderegger M, Jeppsson M, Larsson C, Gorwa-Grauslund M, Boles E, Olsson L, Spencer-Martins I, Hahn-Hagerdal B, Sauer U (2004) Fermentation performance of engineered and evolved xylose-fermenting Saccharomyces cerevisiae strains. Biotechnol Bioeng 87:90–98. doi:10.1002/bit.20094
Song S, Park C (1997) Organization and regulation of the d-xylose operons in E. coli K-12: Xyl R acts as a transcriptional activator. J Bacteriol 179(22):7025–7035
Sonntag K, Schwinde J, deGraaf AA, Marx A, Eikmanns BJ, Wiechert W, Sahm H (1995) C-13 NMR studies of the fluxes in the central metabolism of Corynebacterium glutamicum during growth and overproduction of amino acids in batch cultures. Appl Microbiol Biotechnol 44:489–495
Sprenger GA (1995) Genetics of pentose–phosphate pathway enzymes of Escherichia coli K-12. Arch Microbiol 164(5):324–330. doi:10.1007/BF02529978
Stansen C, Uy D, Delaunay S, Eggeling L, Goergen JL, Wendisch VF (2005) Characterization of a Corynebacterium glutamicum lactate utilization operon induced during temperature-triggered glutamate production. Appl Environ Microbiol 71:5920–5928. doi:10.1128/aem.71.10.5920-5928.2005
Stephens C, Christen B, Fuchs T, Sundaram V, Watanabe K, Jenal U (2007) Genetic Analysis of a novel pathway for d-xylose metabolism in Caulobacter crescentus. J Bacteriol 189:2181–2185. doi:10.1128/JB.01438-06
Takors R, Bathe B, Rieping M, Hans S, Kelle R, Huthmacher K (2007) Systems biology for industrial strains and fermentation processes — example: amino acids. J Biotechnol 129(2):181–190. doi:10.1016/j.jbiotec.2007.01.031
Terasawa M, Yukawa H (1993) Industrial production of biochemicals by native immobilization. In: Tanaka A, Tosa T, Kobayashi T (eds) Industrial application of immobilized biocatalysts. Marcel Dekker, New York, pp 37–52
Van Arsdell SW, Perkins JB, Yocum RR, Luan L, Howitt CL, Chatterjee NP, Pero JG (2005) Removing a bottleneck in the Bacillus subtilis biotin pathway: BioA utilizes lysine rather than S-adenosylmethionine as the amino donor in the KAPA-to-DAPA reaction. Biotechnol Bioeng 91(1):75–83. doi:10.1002/bit.20488
Wendisch VF (2006) Genetic regulation of Corynebacterium glutamicum metabolism. J Microbiol Biotechnol 16(7):999–1009
Wendisch VF (2007). Amino acid biosynthesis—pathways, regulation and metabolic engineering. In: A. Steinbuchel (Ed.) Microbiology Monograph.
Wendisch VF, de Graaf AA, Sahm H, Eikmanns BJ (2000) Quantitative determination of metabolic fluxes during coutilization of two carbon sources: comparative analyses with Corynebacterium glutamicum during growth on acetate and/or glucose. J Bacteriol 182(11):3088–3096
Werpy T, Petersen G, Aden A, Bozell J, Holladay J, White J, Manheim A (2004) Top value added chemicals from biomass, volume 1: Results of screening for potential candidates from sugars and synthesis gas. In Werpy T, Petersen G (eds) Biomass. US Department of Energy
Wyman CE (1999) Biomass ethanol: technical progress, opportunities, and commercial challenges. Annu Rev Energy Environ 24:189–226
Yukawa H, Omumasaba CA, Nonaka H, Kos P, Okai N, Suzuki N, Suda M, Tsuge Y, Watanabe J, Ikeda Y, Vertes AA, Inui M (2007) Comparative analysis of the Corynebacterium glutamicum group and complete genome sequence of strain R. Microbiol 153(4):1042–1058. doi:10.1099/mic.0.2006/003657-0
Zhang X, Jantama K, Moore JC, Shanmugam KT, Ingram LO (2007) Production of l-lanine by metabolically engineered Escherichia coli. Appl Microbiol Biotechnol 77:355–366. doi:10.1007/s00253-007-1170-y
Acknowledgements
Our sincere thanks to the Department of Biotechnology (DBT), New Delhi and BMBF, Germany, for the Indo-German bilateral collaborative programme and also to Colgate Palmolive (USA) for financial support allowing us to work on amino acids. Special thanks to Prof. Volker Wendisch, Bielefeld University, Germany, with whom we collaborated on the topic. We deeply acknowledge the efforts put forward by the reviewers for improving this manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
An erratum to this article can be found at http://dx.doi.org/10.1007/s00253-011-3789-y.
Rights and permissions
About this article
Cite this article
Gopinath, V., Murali, A., Dhar, K.S. et al. Corynebacterium glutamicum as a potent biocatalyst for the bioconversion of pentose sugars to value-added products. Appl Microbiol Biotechnol 93, 95–106 (2012). https://doi.org/10.1007/s00253-011-3686-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-011-3686-4