Abstract
Malolactic fermentation (MLF) is the bacterially driven decarboxylation of l-malic acid to l-lactic acid and carbon dioxide, and brings about deacidification, flavour modification and microbial stability of wine. The main objective of MLF is to decrease wine sourness by a small increase in wine pH via the metabolism of l-malic acid. Oenococcus oeni is the main lactic acid bacterium to conduct MLF in virtually all red wine and an increasing number of white and sparkling wine bases. Over the last decade, it is becoming increasingly recognized that O. oeni exhibits a diverse array of secondary metabolic activities during MLF which can modify the sensory properties of wine. These secondary activities include the metabolism of organic acids, carbohydrates, polysaccharides and amino acids, and numerous enzymes such as glycosidases, esterases and proteases, which generate volatile compounds well above their odour detection threshold. Phenotypic variation between O. oeni strains is central for producing different wine styles. Recent studies using array-based comparative genome hybridization and genome sequencing of three O. oeni strains have revealed the large genomic diversity within this species. This review will explore the links between O. oeni metabolism, genomic diversity and wine sensory attributes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In addition to the initial impact that wine colour has on the perception of wine quality, it is the aroma and flavour of a glass of red or white wine that has the greatest impact on the consumer. The formation of wine aroma is complex, with a combination of many factors contributing and interacting throughout grapegrowing, harvesting, winemaking and maturation. Grape variety and composition, viticultural practices, yeast and bacterial metabolism during fermentation, winemaking techniques, ageing in oak barrels and bottling with different closures (natural cork, synthetic closure or screw cap) all contribute to the sensory experience of wine.
Oenococcus oeni is the main lactic acid bacteria (LAB) involved in winemaking, and its major role is conducting malolactic fermentation (MLF), the decarboxylation of l-malic acid to l-lactic acid and CO2 (Henick-Kling 1993). The consequence of MLF is an increase in wine pH 0.2 to 0.5 units and a decrease in titratable acidity, which translates into a decrease in wine sourness (Amerine and Roessler 1983). There are two other consequences of MLF: increased microbial stability of wine through the removal of a potential carbon source (malic acid) which can be utilised by spoilage yeast and bacteria, and the bacterial production of various secondary metabolites, which can improve the organoleptic properties of wine. Whilst other wine-associated LAB species, from the Lactobacillus and Pediococcus genera, are able to conduct MLF, species from these groups are also associated with wine spoilage and therefore are generally regarded as undesirable.
Malolactic bacteria have had an interesting history in microbiology. A summary of this is shown in Fig. 1. Evidence of winemaking dates back almost 7,000 years, and it has long been recognised that bacteria make a positive contribution (Chambers and Pretorius 2010; Swiegers et al. 2005). Louis Pasteur first described the presence of bacteria in wine over 150 years ago, and despite their role in malic acid metabolism being elucidated only 50 years later, through the work of Müller-Thurgau, Berthelot, de Fleurieu, Ordonneau, Koch and Möslinger (Bartowsky 2005), the actual bacterial species responsible was not formally classified (originally as Leuconostoc oenos) until the mid-1960s by Garvie (1967). With the introduction of molecular biology techniques, a new genus, Oenococcus, was described, and L. oenos was reclassified as O. oeni (Dicks et al. 1995). O. oeni was the sole species in the genus until the mid-2000s when Oenococcus kitaharae was identified in composting distilled shochu residue (Endo and Okada 2006). The genome of O. oeni was mapped in the 1990s (Ze-Ze et al. 1998), and several strains have now been fully sequenced (Mills et al. 2005; Borneman et al. 2010; Guzzo, unpublished data).
This mini review aims to highlight how genomics is beginning to assist in connecting the sensory aspects of MLF and O. oeni metabolism with genetic diversity within the species.
O. oeni and malolactic fermentation
One of the most important aspects of MLF is to ensure that the process is reliably completed in a timely manner so that the wine can be stabilised as soon as possible and spoilage microorganisms do not proliferate; a prolonged or delayed MLF augments the risk of spoilage by microorganisms including Lactobacillus, Pediococcus, Acetobacter and Brettanomyces species (Gerbaux et al. 2009; Bartowsky and Pretorius 2008; Renouf et al. 2007). Spoilage by the yeast Brettanomyces bruxellensis can, for example, lead to production of 4-ethyl phenol and related compounds giving rise to undesirable sensory qualities (e.g. sweaty, barnyard, horsey, medicinal characters) (Loureiro and Malfeito-Ferreira 2003; Curtin et al. 2007). Bacteria-related spoilage includes mousy off flavour, geranium taint, acrolein, ropy wines, elevated acetic acid production (volatile acidity) and the production of biogenic amines (Bartowsky and Henschke 2008; Bartowsky and Pretorius 2008; Sponholz 1993; Lonvaud-Funel 2001).
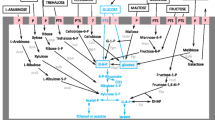
Metabolism of l-malic acid in O. oeni is via the malolactic enzyme (Kunkee 1991) and is a direct enzymatic decarboxylation, with NAD and Mn++ as cofactors and without free intermediates. The enzyme is composed of two identical subunits of 60 kDa and has been purified from several LAB species (Naouri et al. 1990; Lonvaud-Funel and Strasser de Saad 1982; Caspritz and Radler 1983; Spettoli et al. 1984). Genes encoding malolactic enzyme (mleA), malate permease (mleP) and a proposed regulatory protein (mleR) have been cloned, sequenced and mapped on the O. oeni chromosome (Labarre et al. 1996a, b; Mills et al. 2005; Ze-Ze et al. 2008).
MLF can occur spontaneously via indigenous O. oeni populations, but better control over the time of onset of MLF can be achieved by inoculating wines with a selected bacterial culture (Nielsen et al. 1996). However, efficient MLF is not easily achieved as the often nutritionally poor and harsh chemical composition of wine (high ethanol concentration [can be >15% v/v], low pH [can be <3.5] and high SO2 concentration [>50 mg/L]) are natural hindrances to the growth of O. oeni.
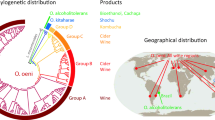
O. oeni strains vary in their ability to metabolise l-malic acid. However, strains selected for commercialisation are usually chosen for their ability to metabolise malic acid efficiently and confer desirable sensory properties on the wine. Recently there has been growing interest in characterising O. oeni strains that are unique to particular geographical wine regions in order to enhance regionality in the wines (Yanagida et al. 2008; Solieri et al. 2010; Ruiz et al. 2010; Capozzi et al. 2010; Vigentini et al. 2009; Canas et al. 2009; Sico et al. 2008; Li et al. 2006).
Aroma and flavour aspects of MLF
Wine is a highly complex mixture of compounds which largely define its appearance, aroma, flavour and mouthfeel properties. Grape-derived compounds provide varietal distinction and basic structure to wine, but it is largely the volatile metabolites that originate from yeast and bacterial metabolism of grape compounds that provide wine its individual character and shape wine style. O. oeni has an extensive suite of metabolic pathways and enzymes that generate volatile secondary compounds at concentrations well above their odour detection threshold, including ethyl and acetate esters, higher alcohols, carbonyls, volatile fatty acids and sulphur compounds (Siebert et al. 2005; Bartowsky 2005), and strain-to-strain variation in metabolic capabilities impact on the types and concentrations of compounds produced (Bartowsky 2005; Francis and Newton 2005; Matthews et al. 2004). For example, some O. oeni strains contribute neutral aroma–flavour to red and white wine (Bartowsky and Henschke 1995), whilst others enhance fruity (Laurent et al. 1994) or the buttery characters (Bartowsky and Henschke 2004; Martineau and Henick-Kling 1995).
Diacetyl is a major secondary metabolite associated with citric acid metabolism during O. oeni-driven MLF, and the kinetics of its production is well understood (Ramos et al. 1994, 1995; Bartowsky and Henschke 2004). Diacetyl imparts a buttery character which adds complexity to wine and is mostly present in concentrations well above its odour detection threshold (Bartowsky et al. 2002b; Martineau and Henick-Kling 1995; Martineau et al. 1995). The genetics of the diacetyl pathway are well described in the literature on dairy research (Cogan 1995; Smit et al. 2005). The two O. oeni genes (alsS and alsD) encode the main enzymes involved; α-acetolactate synthetase and α-acetolactate decarboxylase genes, respectively, have been cloned and characterised (Garmyn et al. 1996).
The operon carrying alsS and alsD has been shown to be constitutively expressed in O. oeni BAA-1163 (previously referred to as Lo84.13), but there is strain-to-strain variation in the final concentration of diacetyl synthesised during MLF of the same wine (Martineau and Henick-Kling 1995; Bartowsky et al. 2002a). O. oeni strains that produce high concentrations of diacetyl can be encouraged to do so, and winemaking techniques can be used to maintain the desired diacetyl concentration to accentuate the buttery character of wine (Bartowsky and Henschke 2004).
Latent aroma and flavour compounds are often glycosylated and can be enzymatically liberated by microbial glycosidases (Gunata et al. 1988, 1990). While numerous fungi and bacteria produce glycosidases, there is variation in the abilities of these microbes to function efficiently under the high alcohol and low pH present in winemaking conditions. O. oeni has numerous glycosidases (Grimaldi et al. 2005, 2000), and their activities contribute to the release of numerous aroma compounds, including monoterpenes, norisoprenoids and aliphatic compounds, all of which contribute to fruity and floral wine attributes (D'Incecco et al. 2004; Williams et al. 1989, 1982; Ugliano et al. 2003; Ugliano and Moio 2005, 2006). As for diacetyl production, there is large variation in O. oeni strain capacity to release aroma compounds from grape glycosides (Grimaldi et al. 2005; Ugliano et al. 2003; Ugliano and Moio 2006).
Most commercial glycosidase preparations are crude extracts prepared from fungi rather than bacteria. However, there has recently been renewed interest in using wine LAB as potential sources of these enzymes as these bacteria are well adapted to the high ethanol and acid conditions encountered during winemaking, and therefore, their enzymes might be expected to perform efficiently under these conditions. Several research teams have cloned and characterised β-glucosidases from O. oeni and Lactobacillus brevis strains (Michlmayr et al. 2010a, b; Capaldo et al. 2011) and α-arabinofuranosidase from O. oeni and L. brevis (Michlmayr et al. 2011).
Many wine aromas are attributable to interacting compounds, which together confer the sensory impact of the aroma descriptor. For example, the red and black berry aromas of a red wine are made up of at least six different esters and volatile fatty acids (Pineau et al. 2009). Other compounds play a more complex role. Dimethyl sulphide, for example, has a cooked corn aroma; however, it also enhances fruity characters in red wine (Segurel et al. 2004).
Enhancement of the fruity and berry characters of red wine is an important goal for winemakers, and one approach involves using selected O. oeni strains to attain sought-after wine styles (Iland et al. 2009). Using sensory descriptive wine analysis and wine composition–ethyl ester concentrations, we are gaining a better understanding of which metabolites contribute to the fruitness of red wine (Pineau et al. 2009, 2010; Guth 1997a, b). A recent study in Cabernet Sauvignon wines has identified relationships between enhanced red berry sensory attributes and ester concentration (Fig. 2) (Bartowsky et al. 2010).
Example of sensory changes following MLF in Cabernet Sauvignon wines with different strains of O. oeni. I Radial plot of aroma and flavour sensory descriptors of Cabernet Sauvignon wine (Berri, Australia) following MLF with O. oeni strain AWRI B429 (adapted from Bartowsky et al. 2008). II Relative changes in sensory descriptor rating for Cabernet Sauvignon wines (sourced over three vintage from Clare Valley, Australia) and the concentration of fruity esters following MLF by three O. oeni strains (P. Costello and colleagues, 2009, unpublished). Total fruity esters are based on a combination of ethyl ester concentrations (Pineau et al. 2009)
Clearly, MLF is important for winemaking, and O. oeni strains make a significant contribution to the aroma and flavour of wine. The next phase to unravelling how malolactic bacteria enhance wine sensory characters is to link phenotypic with genotype characteristics. However, O. oeni is not very amenable to the uptake of DNA; even though there are transformation and conjugation methods cited, they are not very efficient (Dicks 1994; Beltramo et al. 2004; Assad-Garcia et al. 2008; Eom et al. 2010). Nonetheless, with the aid of modern ‘genomic’ technologies, we can now use comparative genomic approaches to enable the identification of genetic elements in O. oeni that shape wine flavour during MLF.
Connecting the O. oeni genome with wine
O. oeni has a compact genome of approximately 1.8 Mb which appears to be highly streamlined as a consequence of adaptation to its ecologically restricted niche, i.e. grape juice and wine. Prior to several strains having their genome fully sequenced, detailed genome mapping had been conducted on several strains, providing insight into the genetic organisation of the bacterium (Ze-Ze et al. 1998, 2000). Important phenotypes, such as malate degradation (MLF), citrate metabolism and diacetyl production, were mapped. Many of the stress response genes have been studied, and metabolic pathways relating to amino acid metabolism have also been well characterised (summarised in Bartowsky 2005). However, the genetics underpinning the generation of metabolites contributing to wine aroma have been less thoroughly investigated.
Comparison of the three available O. oeni genome sequences suggests the three strains share a group of conserved ORFs (52% of total ORFs), with up to 10% of the coding potential of any one strain being specific to that strain (Borneman et al. 2010). It is presumably this variation in the O. oeni species-wide (pan) protein coding capacity that will ultimately be the likely key to phenotypic differences between O. oeni strains.
In our recent array-based comparative genomics research, we analysed 11 O. oeni strains, all of which were either commercial or natural isolates that are routinely used in Australian winemaking. One of the most interesting observations from this work was that there was significant intra-specific genomic variation due to substantial insertions and deletions throughout the 11 O. oeni strains, including two large deletions in regions 45–65 and 1,400–1,450 kb (Borneman et al. 2010). However, as all of these strains are used in the wine industry, these rearrangements clearly do not have a significant impact on the MLF capability of the O. oeni strains, but may well contribute to the phenotypic differences between strains, resulting in wines with different sensory profiles.
A relationship between genome variation and efficient malate metabolism has been suggested by studying the MLF capacity of over 70 O. oeni strains in three wines and using comparative genome subtractive hybridization to propose that the presence of eight stress-responsive genes could be associated with high MLF performance (Bon et al. 2009).
The winemaking capabilities of O. oeni strain AWRIB429 have been well studied with observations that red wines produced with this O. oeni strain consistently lift red fruit–red berry aroma attributes (Schmid et al. 2007; Borneman et al. 2010; Bartowsky et al. 2010, 2008) (Fig. 2). Within the O. oeni AWRIB429 genome, numerous novel ORFs were identified, and amongst the annotated ORFs were several potential glycosidases (Borneman et al. 2010). Thus, in addition to other aroma-forming pathways, the acquisition of additional glycosidases by AWRIB429 might provide a genomic link to the consistent ability of this strain to enhance red fruit–red berry aroma attributes to red wines.
Conclusions and future outlook
O. oeni conducts malolactic fermentation, often under difficult environmental circumstances, to impart important sensory attributes to wine. It has a compact genome of approximately 1.8 Mb resulting from a high degree of genomic streamlining which occurred during its adaptation to the complex wine environment. Recent analysis of several O. oeni genomes has demonstrated a shared core genome with up to 10% variation in coding capacity between strains. This variation in coding capacity is likely to be responsible for observed differences in winemaking phenotypes. A link between the presence of several stress-related genes and MLF capability has been proposed, and the presence of additional glycosidases appears to enhance the fruity sensory attributes of an O. oeni strain. Even though O. oeni is not easily genetically manipulated, due to poor transformability, the inter-strain genetic variation of the O. oeni genome holds the secret to exploiting its potential to influence wine aroma and flavour.
References
Amerine MA, Roessler EB (1983) Wines—their sensory evaluation, 2nd edn. Freeman, San Francisco
Assad-Garcia JS, Bonnin-Jusserand M, Garmyn D, Guzzo J, Alexandre H, Grandvalet C (2008) An improved protocol for electroporation of Oenococcus oeni ATCC BAA-1163 using ethanol as immediate membrane fluidizing agent. Lett Appl Microbiol 47(4):333–338
Bartowsky EJ (2005) Oenococcus oeni and malolactic fermentation—moving into the molecular arena. Aust J Grape Wine Res 11(2):174–187
Bartowsky EJ, Henschke PA (1995) Malolactic fermentation and wine flavour. Aust Grapegrower Winemaker 378a:83–94
Bartowsky EJ, Henschke PA (2004) The ‘buttery’ attribute of wine—diacetyl—desirability, spoilage and beyond. Int J Food Microbiol 96:235–252
Bartowsky EJ, Henschke PA (2008) Acetic acid bacteria spoilage of bottled red wine—a review. Int J Food Microbiol 125:60–70
Bartowsky EJ, Pretorius IS (2008) Microbial formation and modification of flavour and off-flavour compounds in wine. In: König H, Unden G, Fröhlich J (eds) Biology of microorganisms on grapes, in must and wine. Springer, Heidelberg, pp 211–233
Bartowsky E, Costello P, Henschke P (2002a) Management of malolactic fermentation—wine flavour manipulation. Aust Grapegrower Winemaker 461a:7–12
Bartowsky EJ, Francis IL, Bellon JR, Henschke PA (2002b) Is buttery aroma perception in wines predictable from diacetyl concentration? Aust J Grape Wine Res 8:180–185
Bartowsky E, Costello P, McCarthy J (2008) MLF—adding an ‘extra dimension’ to wine flavour and quality. Aust NZ Grapegrower Winemaker 533a:60–65
Bartowsky E, Costello P, Krieger-Weber S, Markides A, Francis L, Travis B (2010) Influence of malolactic fermentation on the fruity characters of red wine—bringing wine chemistry and sensory together. In: International Intervitis Interfructa Congress 2010, Stuttgart, Germany, 24–28 March 2010. pp 52–59
Beltramo C, Oraby M, Bourel G, Garmyn D, Guzzo J (2004) A new vector, pGID052, for genetic transfer in Oenococcus oeni. FEMS Microbiol Lett 236(1):53–60
Bon E, Delaherche A, Bilhere E, De Daruvar A, Lonvaud-Funel A, Le Marrec C (2009) Oenococcus oeni genome plasticity is associated with fitness. App Environ Microbiol 75(7):2079–2090
Borneman AR, Bartowsky EJ, McCarthy J, Chambers PJ (2010) Genotypic diversity in Oenococcus oeni by high-density microarray comparative genome hybridization and whole genome sequencing. Appl Microbiol Biotechnol 86(2):681–691. doi:10.1007/s00253-009-2425-6
Canas PMI, Perez PR, Prieto SS, Herreros MLP (2009) Ecological study of lactic acid microbiota isolated from Tempranillo wines of Castilla-La Mancha. J Biosci Bioeng 108(3):220–224
Capaldo A, Walker ME, Ford CM, Jiranek V (2011) beta-Glucoside metabolism in Oenococcus oeni: cloning and characterisation of the phospho-beta-glucosidase bgID. Food Chem 125(2):476–482
Capozzi V, Russo P, Beneduce L, Weidmann S, Grieco F, Guzzo J, Spano G (2010) Technological properties of Oenococcus oeni strains isolated from typical southern Italian wines. Lett Appl Microbiol 50(3):327–334
Caspritz G, Radler F (1983) Malolactic enzyme of Lactobacillus plantarum. Purification, properties, and distribution among bacteria. J Biol Chem 258:4907–4910
Chambers PJ, Pretorius IS (2010) Fermenting knowledge: the history of winemaking, science and yeast research. EMBO Rep 11(12):914–920
Cogan TM (1995) Flavour production by dairy starter cultures. J Appl Bacteriol Symp Suppl 79:49S–64S
Curtin CD, Bellon JR, Henschke PA, Godden P, de Barros LM (2007) Genetic diversity of Dekkera bruxellensis yeasts isolated from Australian wineries. FEMS Yeast Res 7(3):471–481
Dicks LMT (1994) Transformation of Leuconostoc oenos by electroporation. Biotechnol Tech 8(12):901–904
Dicks LMT, Dellaglio F, Collins MD (1995) Proposal to reclassify Leuconostoc oenos as Oenococcus oeni [corrig.] gen. nov., comb. nov. Int J Sys Evol Bacteriol 45(2):395–397
D'Incecco N, Bartowsky E, Kassara S, Lante A, Spettolli P, Henschke P (2004) Release of glycosidically bound flavour compounds of Chardonnay by Oenococcus oeni during malolactic fermentation. Food Microbiol 21(3):257–265
Endo A, Okada S (2006) Oenococcus kitaharae sp nov., a non-acidophilic and non-malolactic-fermenting oenococcus isolated from a composting distilled shochu residue. Int J Sys Evol Microbiol 56:2345–2348. doi:10.1099/ijs.0.64288-0
Eom HJ, Cho SK, Park MS, Ji GE, Han NS (2010) Characterization of Leuconostoc citreum plasmid pCB18 and development of broad host range shuttle vector for lactic acid bacteria. Biotechnol Bioprocess Eng 15(6):946–952. doi:10.1007/s12257-010-0089-9
Francis IL, Newton JL (2005) Determining wine aroma from compositional data. Aust J Grape Wine Res 11(2):114–126
Garmyn D, Monnet C, Martineau B, Guzzo J, Cavin J-F, Diviès C (1996) Cloning and sequencing of the gene encoding α-acetolactate decarboxylase from Leuconostoc oenos. FEMS Microbiol Lett 145:445–450
Garvie EI (1967) Leuconostoc oenos sp.nov. J Gen Microbiol 48(3):431–438. doi:10.1099/00221287-48-3-431
Gerbaux V, Briffox C, Dumont A, Krieger S (2009) Influence of inoculation with malolactic bacteria on volatile phenols in wines. Am J Enol Vitic 60(2):233–235
Grimaldi A, McLean H, Jiranek V (2000) Identification and partial characterization of glycosidic activities of commercial strains of the lactic acid bacterium, Oenococcus oeni. Am J Enol Vitic 51(4):362–369
Grimaldi A, Bartowsky E, Jiranek V (2005) A survey of glycosidase activities of commercial wine strains of Oenococcus oeni. Int J Food Microbiol 105(2):233–244
Gunata Z, Bitteur S, Brillouet J-M, Bayonove C, Cordonnier R (1988) Sequential enzymatic hydrolysis of potentially aromatic glycosides from grape. Carbohyd Res 184:139–149
Gunata YZ, Bayonove CL, Tapiero C, Cordonnier RE (1990) Hydrolysis of grape monoterpenyl β-D-glucosides by various β-glucosidases. J Agric Food Chem 38:1232–1236
Guth H (1997a) Identification of character impact odorants of different white wine varieties. J Agric Food Chem 45(8):3022–3026
Guth H (1997b) Quantitation and sensory studies of character impact odorants of different white wine varieties. J Agric Food Chem 45(8):3027–3032
Henick-Kling T (1993) Malolactic fermentation. In: Fleet GH (ed) Wine microbiology and biotechnology. Harwood Academic Publisher, Amsterdam, pp 289–326
Iland P, Gago P, Caillard A, Dry P (2009) A taste of the world of wine. Patrick Iland Wine Promotions Pty Ltd, Campbelltown
Kunkee RE (1967) Malo-lactic fermentation. Adv Appl Microbiol 9:235–279
Kunkee RE (1991) Some roles of malic acid in the malolactic fermentation in wine making. FEMS Microbiol Rev 88:55–72
Labarre C, Divies C, Guzzo J (1996a) Genetic organization of the mle locus and identification of a mleR-like gene from Leuconostoc oenos. App Environ Microbiol 62(12):4493–4498
Labarre C, Guzzo J, Cavin JF, Divies C (1996b) Cloning and characterization of the genes encoding the malolactic enzyme and the malate permease of Leuconostoc oenos. App Environ Microbiol 62(4):1274–1282
Laurent M-H, Henick-Kling T, Acree TE (1994) Changes in the aroma and odor of Chardonnay wine due to malolactic fermentation. Vitic Enol Sci 49:3–10
Li H, Zhang CH, Liu YL (2006) Species attribution and distinguishing strains of Oenococcus oeni isolated from Chinese wines. World J Microb Biot 22(5):515–518
Lonvaud-Funel A (2001) Biogenic amines in wines: role of lactic acid bacteria. FEMS Microbiol Lett 199(1):9–13
Lonvaud-Funel A, Strasser de Saad AM (1982) Purification and properties of a malolactic enzyme from a strain of Leuconostoc mesenteroides isolated from grapes. App Environ Microbiol 43:357–361
Loureiro V, Malfeito-Ferreira M (2003) Spoilage yeasts in the wine industry. Int J Food Microbiol 86:23–50
Martineau B, Henick-Kling T (1995) Performance and diacetyl production of commercial strains of malolactic bacteria in wine. J Appl Bacteriol 78:526–536
Martineau B, Acree TE, Henick-Kling T (1995) Effect of wine type on the detection threshold for diacetyl. Food Res Int 28(2):139–143
Matthews A, Grimaldi A, Walker M, Bartowsky E, Grbin P, Jiranek V (2004) Lactic acid bacteria as a potential source of enzymes for use in vinification. App Environ Microbiol 70(10):5715–5731. doi:10.1128/aem.70.10.5715-5731.2004
Michlmayr H, Schumann C, da Silva N, Kulbe KD, del Hierro AM (2010a) Isolation and basic characterization of a beta-glucosidase from a strain of Lactobacillus brevis isolated from a malolactic starter culture. J Appl Microbiol 108(2):550–559. doi:10.1111/j.1365-2672.2009.04461.x
Michlmayr H, Schumann C, Wurbs P, da Silva N, Rogl V, Kulbe KD, del Hierro AM (2010b) A beta-glucosidase from Oenococcus oeni ATCC BAA-1163 with potential for aroma release in wine: cloning and expression in E. col. World J Microb Biot 26(7):1281–1289
Michlmayr H, Schumann C, Kulbe KD, del Hierro AM (2011) Heterologously expressed family 51 α-L-arabinofurnaosidases from Oenococcus oeni and Lactobacillus brevis. App Environ Microbiol 77(4):1528–1531
Mills DA, Rawsthorne H, Parker C, Tamir D, Makarova K (2005) Genomic analysis of Oenococcus oeni PSU-1 and its relevance to winemaking. FEMS Microbiol Rev 29:465–475
Möslinger (1901) Über die Säuren des Weines und den Säurerückgang. Zeitschr f Untersuchung d Nahr Genussmittel 4:1120–1130
Müller-Thurgau H (1891) Über die Ergebnisse neuer Untersuchungen auf den Gebiete der Weinbereitung. Ber XII Dtsch Weinbaukong (Worms):128
Müller-Thurgau H, Osterwalder A (1913) Die Bakterien im Wein und Obstwein und die dadurch verursachten Veränderungen. Zentr Bakt Parasitenk Hyg Abt 2(36):129–338
Naouri P, Chagnaud P, Arnaud A, Galzy P (1990) Purification and properties of a malolactic enzyme from Leuconostoc oenos ATCC-23278. J Basic Microb 30(8):577–585
Nielsen JC, Prahl C, Lonvaud-Funel A (1996) Malolactic fermentation in wine by direct inoculation with freeze-dried Leuconostoc oenos cultures. Am J Enol Vitic 47(1):42–48
Pasteur L (1873) Études sur le Vin, 2nd edn. Savy, Paris
Pineau B, Barbe JC, Van Leeuwen C, Dubourdieu D (2009) Examples of perceptive interactions involved in specific “red-” and “black-berry” aromas in red wines. J Agric Food Chem 57(9):3702–3708
Pineau B, Barbe JC, van Leeuwen C, Dubourdieu D (2010) Olfactory specificity of red- and black-berry fruit aromas in red wines and contribution to the red Bordeaux wine concept. Journal International Des Sciences De La Vigne Et Du Vin 44(1):39–49
Ramos A, Poolman B, Santos H, Lolkema JS, Konings WN (1994) Uniport of anionic citrate and proton consumption in citrate metabolism generates a proton motive force in Leuconostoc oenos. J Bacteriol 176(16):4899–4905
Ramos A, Lolkema JS, Konings WN, Santos H (1995) Enzyme basis for pH regulation of citrate and pyruvate metabolism by Leuconostoc oenos. App Environ Microbiol 61(4):1303–1310
Renouf V, Claisse O, Lonvaud-Funel A (2007) Inventory and monitoring of wine microbial consortia. Appl Microbiol Biotechnol 75(1):149–164
Ruiz P, Izquierdo PM, Sesena S, Palop ML (2010) Selection of autochthonous Oenococcus oeni strains according to their oenological properties and vinification results. Int J Food Microbiol 137(2–3):230–235
Schmid F, Li Y, Liebich B, Culbert J, Day C, Jiranek V (2007) Evaluation of red wine made on a small scale utilizing frozen must. J Agric Food Chem 55(17):7156–7161
Segurel MA, Razungles AJ, Riou C, Salles M, Baumes RL (2004) Contribution of dimethyl sulfide to the aroma of Syrah and Grenache Noir wines and estimation of its potential in grapes of these varieties. J Agric Food Chem 52(23):7084–7093
Sico MA, Bonomo MG, Salzano G (2008) Isolation and characterization of Oenococcus oeni from Aglianico wines. World J Microb Biot 24(9):1829–1835
Siebert TE, Smythe HE, Capone DL, Neuwöhner C, Pardon KH, Skouroumounis GK, Herderich MJ, Sefton MA, Pollnitz AP (2005) Stable isotope dilution analysis of wine fermentation products by HS = SPME-GC-MS. Anal Bioanal Chem 381:937–947
Smit G, Smit BA, Engels WJM (2005) Flavour formation by lactic acid bacteria and biochemical flavour profiling of cheese products. FEMS Microbiol Rev 29(3):591–610
Solieri L, Genova F, De Paola M, Giudici P (2010) Characterization and technological properties of Oenococcus oeni strains from wine spontaneous malolactic fermentations: a framework for selection of new starter cultures. J Appl Microbiol 108(1):285–298
Spettoli P, Nuti MP, Zamorani A (1984) Properties of malolactic activity purified from Leuconostoc oenos ML34 by affinity chromatography. App Environ Microbiol 9:184–189
Sponholz W-R (1993) Wine spoilage by microorganisms. In: Fleet GH (ed) Wine microbiology and technology. Harwood Academic Publishing, Amsterdam, pp 395–420
Swiegers JH, Bartowsky EJ, Henschke PA, Pretorius IS (2005) Yeast and bacterial modulation of wine aroma and flavour. Aust J Grape Wine Res 11(2):139–173
Ugliano M, Moio L (2005) Changes in the concentration of yeast-derived volatile compounds of red wine during malolactic fermentation with four commercial starter cultures of Oenococcus oeni. J Agric Food Chem 53(26):10134–10139
Ugliano M, Moio L (2006) The influence of malolactic fermentation and Oenococcus oeni strain on glycosidic aroma precursors and related volatile compounds of red wine. J Sci Food Agric 86(14):2468–2476. doi:10.1002/jsfa.2650
Ugliano M, Genovese A, Moio L (2003) Hydrolysis of wine aroma precursors during malolactic fermentation with four commercial starter cultures of Oenococcus oeni. J Agric Food Chem 51(17):5073–5078. doi:10.1021/jf0342019
Vigentini I, Picozzi C, Tirelli A, Giugni A, Foschino R (2009) Survey on indigenous Oenococcus oeni strains isolated from red wines of Valtellina, a cold climate wine-growing Italian area. Int J Food Microbiol 136(1):123–128
Williams PJ, Strauss CR, Wilson B, Massy-Westropp RA (1982) Novel monoterpene disaccharide glycosides of Vitis vinifera grapes and wines. Phytochemistry 21:2013–2020
Williams PJ, Sefton MA, Wilson B (1989) Nonvolatile conjugates of secondary metabolites as precursors of varietal grape flavour components. In: Teranishi R, Buttery RG, Shahisi R (eds) Flavour chemistry, trends and developments, ACS symposium series no 388. American Chemical Society, Washington DC, pp 35–48
Yanagida F, Srionnual S, Chen YS (2008) Isolation and characteristics of lactic acid bacteria from koshu vineyards in Japan. Lett Appl Microbiol 47(2):134–139. doi:10.1111/j.1472-765X.2008.02398.x
Ze-Ze L, Teneiro R, Brito L, Santos MA, Paveia H (1998) Physical map of the genome of Oenococcus oeni PSU-1 and localization of genetic markers. Microbiology 144:1145–1156
Ze-Ze L, Teneiro R, Paveia H (2000) The Oenococcus oeni genome: physical and genetic map of strain GM and comparison with the genome of a ‘divergent’ strain, PSU-1. Microbiology 146(12):3195–3204
Ze-Ze L, Chelo IM, Tenreiro R (2008) Genome organization in Oenococcus oeni strains studied by comparison of physical and genetic maps. Int Microbiol 11(4):237–244
Acknowledgements
The authors thank Dr. Paul Chambers for discussions during the preparation of this mini review and Lallemand SA for allowing us to use data presented in Fig. 2 from a project for which they provided financial support. The Australian Wine Research Institute (AWRI) is supported by Australian grapegrowers and winemakers through their investment agency, the Grape and Wine Research and Development Corporation, with matching funds from the Australian Government. The AWRI is a member of the Wine Innovation Cluster in Adelaide.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Bartowsky, E.J., Borneman, A.R. Genomic variations of Oenococcus oeni strains and the potential to impact on malolactic fermentation and aroma compounds in wine. Appl Microbiol Biotechnol 92, 441–447 (2011). https://doi.org/10.1007/s00253-011-3546-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-011-3546-2