Abstract
Corynebacterium glutamicum wild type lacks the ability to utilize the pentose fractions of lignocellulosic hydrolysates, but it is known that recombinants expressing the araBAD operon and/or the xylA gene from Escherichia coli are able to grow with the pentoses xylose and arabinose as sole carbon sources. Recombinant pentose-utilizing strains derived from C. glutamicum wild type or from the l-lysine-producing C. glutamicum strain DM1729 utilized arabinose and/or xylose when these were added as pure chemicals to glucose-based minimal medium or when they were present in acid hydrolysates of rice straw or wheat bran. The recombinants grew to higher biomass concentrations and produced more l-glutamate and l-lysine, respectively, than the empty vector control strains, which utilized the glucose fraction. Typically, arabinose and xylose were co-utilized by the recombinant strains along with glucose either when acid rice straw and wheat bran hydrolysates were used or when blends of pure arabinose, xylose, and glucose were used. With acid hydrolysates growth, amino acid production and sugar consumption were delayed and slower as compared to media with blends of pure arabinose, xylose, and glucose. The ethambutol-triggered production of up to 93 ± 4 mM l-glutamate by the wild type-derived pentose-utilizing recombinant and the production of up to 42 ± 2 mM l-lysine by the recombinant pentose-utilizing lysine producer on media containing acid rice straw or wheat bran hydrolysate as carbon and energy source revealed that acid hydrolysates of agricultural waste materials may provide an alternative feedstock for large-scale amino acid production.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Lignocellulosic biomass is a cheap carbon source available from agricultural wastes. Currently, it is underutilized, and its proper recycling may even be an ecological problem. Lignocellulosic biomass and agricultural wastes thus represent viable candidates as alternate carbon sources for the large-scale biotechnological production of commodity chemicals. Lignocellulose is composed mainly of cellulose (40–50%), hemicellulose (25–30%), and lignin (10–20%) (Wyman 1999). Among them, hemicellulose is a heteropolymer of various hexose (glucose, galactose, and mannose) and pentose (xylose and arabinose) sugars. Lignocellulosic hydrolysates contain not only glucose as major component, but also a significant fraction of xylose (5–20%) and arabinose (1–5%) (Aristidou and Penttila 2000). Unfortunately, relatively few native strains of industrial microorganisms can utilize pentose sugars as fermentable substrates (Jeffries and Jin 2000). The lack of pentose-utilizing industrially relevant microorganisms often is a major bottleneck to the implementation of successful industrial processes based on lignocellulosic biomass, although successful examples of metabolic engineering, e.g., of Saccharomyces cerevisiae for ethanol production from pentoses, have been described (Becker and Boles 2003; Aristidou and Penttila 2000; Hahn-Hagerdal et al. 2007a; Hahn-Hagerdal et al. 2007b; Karhumaa et al. 2006).
Whereas considerable efforts have been made to utilize hemicellulose for the production of organic acids like lactic acid or succinic acid and of alcohols like ethanol (Hahn-Hagerdal et al. 2007a; Kawaguchi et al. 2006), amino acid fermentation using hemicellulosic biomasses has received much less attention. Corynebacterium glutamicum is widely used for the fermentative production of amino acids ever since its discovery five decades ago. C. glutamicum has also been engineered for production of compounds derived from amino acids and their precursors such as 1,4-diaminobutane (Schneider and Wendisch 2010) or 1,5-diaminopentane (Mimitsuka et al. 2007), 2-ketoisovalerate (Krause et al. 2010), and isobutanol (Blombach et al. 2011). Co-utilization of carbon sources is a typical characteristic of C. glutamicum (Eggeling and Bott 2005; Wendisch 2006). Technical substrates such as starch hydrolysates and molasses contain glucose, fructose, and sucrose, which are imported and phosphorylated by the phosphoenolpyruvate-dependent phosphotransferase (PTS) system or via the inositol permeases (Lindner et al. 2011) with subsequent phosphorylation by ATP- and/or polyphosphate-dependent glucokinases (Lindner et al. 2010). This amino acid producing bacterium can grow aerobically on a variety of sugars (e.g., glucose, fructose, sucrose, ribose, or maltose), alcohols (myo-inositol and ethanol) or organic acids (acetate, propionate, pyruvate, l-lactate, citrate, and l-glutamate) as sole or combined carbon and energy sources (Dominguez et al. 1998; Eikmanns 2005; Gerstmeir et al. 2003; Kiefer et al. 2002; Krämer et al. 1990; Moon et al. 2005; Polen et al. 2005; Frunzke et al. 2008; Stansen et al. 2005; Kato et al. 2010; Krings et al. 2006). By metabolic engineering, the carbon substrate spectrum of C. glutamicum could be broadened to enable growth on starch (Cadenas et al. 1992; Seibold et al. 2006; Tateno et al. 2007), on the whey sugars lactose and galactose (Brabetz et al. 1991; Barrett et al. 2004), on glycerol (Rittmann et al. 2008), the dicarboxylates succinate, fumarate or malate (Youn et al. 2008, 2009), and on cellobiose (Kotrba et al. 2003).
C. glutamicum wild type (WT), which is unable to utilize the pentose sugars xylose and arabinose, has also been engineered for growth on the pentoses xylose and arabinose (Kawaguchi et al. 2006, 2008; Schneider et al. 2011). Arabinose utilization by bacteria involves arabinose isomerase (AraA), which converts arabinose to ribulose, which is phosphorylated by ribulokinase (AraB), and ribulose-5-phosphate-4-epimerase (AraD) converts ribulose-5-phosphate to xylulose-5-phosphate, an intermediate of the pentose phosphate pathway (Hahn-Hagerdal et al. 2007a,b). Unlike Escherichia coli, which is capable of growth on l-arabinose as the sole carbon source (Lin 1996), none of the genes of the arabinose metabolic pathway mentioned above have been identified in any of the corynebacterial genomes sequenced so far (Cerdeno-Tarraga et al. 2003; Kalinowski et al. 2003; Nishio et al. 2003; Tauch et al. 2005). A typical bacterial xylose utilization pathway includes only two steps: first, xylose is converted to xylulose by xylose isomerase (XylA), before xylulokinase (XylB) phosphorylates xylulose to xylulose-5-phosphate, an intermediate of the pentose phosphate pathway. Most corynebacteria are unable to utilize xylose as a carbon source because no xylose isomerase encoding gene is present in any of the corynebacteria sequenced to date. The C. glutamicum R strain contains a functional xylulokinase (Kawaguchi et al. 2006), and heterologous expression of a xylose isomerase gene is sufficient to enable growth with xylose. C. glutamicum R strain has been shown to withstand pretreatment-derived inhibitors like furfural, hydroxymethyl furfural, and 4-hydroxybenzaldehyde under growth-arrested conditions (Sakai et al. 2007).
These characteristics and previous work demonstrating that genetically engineered C. glutamicum strains utilize arabinose and xylose when added as pure chemicals (Kawaguchi et al. 2006, 2008, 2009; Schneider et al. 2011) prompted us to analyze whether recombinant C. glutamicum strains may be used for amino acid production based on hemicellulosic hydrolysates. It is desirable that large-scale processes such as amino acid production make use of non-food carbon sources in order to avoid competition with the use of carbon sources such as starch and sugar in nutrition.
Materials and methods
Bacterial strains, plasmids, and culture inoculums
Bacterial strains and plasmids used in this study have been listed in Table 1. For growth experiments and l-glutamate production, C. glutamicum WT and derived strains were used, while C. glutamicum DM1729 and derived strains were used for l-lysine production (Georgi et al. 2005). A loopful of respective cells from fresh Luria–Bertani (LB) plates was inoculated into 50 ml brain heart infusion medium (Difco), and the culture was incubated at 30°C for 18 h, and the cells were removed from this preculture by centrifugation at 4°C and washed in CgXII medium (Eggeling and Reyes 2005) without carbon source, and was used to inoculate the CgXII production medium to an optical density at 600 nm (OD600) of 1, with either (blends of) glucose, arabinose, or xylose, or acid rice straw or wheat bran hydrolysates as carbon source(s). E. coli MG1655 and DH5α were cultivated in LB medium or on LB agar plates at 37°C. E. coli strain DH5α was used as host for cloning. When appropriate, kanamycin was used at a concentration of 25–50 μg/ml, spectinomycin at a concentration of 100 μg/ml, and isopropyl β-d-1-thiogalactopyranoside (IPTG) at concentrations up to 1 mM.
Chromosomal DNA from E. coli MG1655 was isolated as described (Eikmanns et al. 1994). Plasmid DNA was isolated with the QIAprep Spin Miniprep Kit (Qiagen, Hilden, Germany). Transformation of E. coli was performed using the rubidium chloride method (Hanahan 1983), while C. glutamicum was transformed by electroporation as described (Eggeling and Reyes 2005). The xylA gene used in this study was amplified from genomic DNA from E. coli K12 MG1655 using xylA_fw_SmaI_RBS_E.coli 5′-ACGCCCCGGG AAAGAGATATAGATATGCAAGCCTATTTTGAC-3′ (the SmaI restriction site is given in bold, the region containing the ribosomal binding site in italics) and xylA_rev_SmaI_E.coli 5′-CTGTTCGACAAATAACCCGGGACG-3′ (the SmaI restriction site is given in bold) primers. The overexpression vector pEKEx3-xylA was obtained by cloning of the resulting PCR product into the vector pEKEx3 (Stansen et al. 2005) using the SmaI restriction site within the multiple cloning site of the vector. Pentose-utilizing C. glutamicum strains were constructed by transforming C. glutamicum WT and the l-lysine producing strain DM1729 with plasmid pVWEx1-araBAD (Kawaguchi et al. 2006, 2008, 2009; Schneider et al. 2011) for heterologous expression of araBAD from E. coli and/or with plasmid pEKEx3-xylA. Plasmid pVWEx1-araBAD has previously been shown to be functional (Kawaguchi et al. 2006, 2008, 2009; Schneider et al. 2011) and complementation of E. coli JW3537-1, the xylA deletion mutant in the KEIO collection, revealed that plasmid pEKEx3-xylA is functional (data not shown). The respective empty vector control strains were constructed for comparison.
Screening of agricultural residues
In Kerala, India, locally available agricultural residual substrates, such as rice straw, rice husk, wheat bran, sugarcane bagasse, wood powder (Atrocarpus myrestus), reed, bamboo, and cotton stalk, were screened to find out the pentose sugar percentage in each of them and to use the best hemicellulolytic substrate for the hydrolysis to generate maximum pentose sugars and to use it for amino acid fermentation. Laboratory Analytical Protocol by National Renewable Energy Laboratory (NREL) (Sluiter et al. 2008) was used for the composition analysis of the above samples. In brief, 0.3 g of biomass was mixed with 72% H2SO4 and kept in shaking water bath at 30°C for 3 h. After incubation in the water bath, the samples were diluted to 4% H2SO4 with distilled water and autoclaved at 121°C for 1 h. The acid soluble portion was assayed for carbohydrate composition (glucose, xylose, and arabinose) by high-performance liquid chromatography (HPLC) (see below).
Acid hydrolysis of agricultural residues
Hydrolysis has been carried out with different pretreatment agents such as H2SO4, HCl, and NaOH. Out of this, H2SO4 was the most efficient pretreatment agent at 121°C. Other pretreatment parameters, like substrate loading percentage, temperature, retention time, and reagent concentration, were optimized with H2SO4 (data not shown). Optimized pretreatment conditions for rice straw and wheat bran hydrolysis were found to be with 4% (w/v) H2SO4 at 134°C for 35 min with 37.5% substrate loading. Under these conditions, 1.9 g solids remained when 7.5 g of wheat bran were hydrolysed and 4.7 g solids remained from 7.5 g rice straw. Under the chosen conditions, hemicellulose was readily hydrolysed, while sulfonation of cellulose precluded its efficient hydrolysis.
Fermentation
C. glutamicum cultivations for amino acid production were carried out in 500-ml Erlenmeyer flasks with 50 ml medium at 30°C and 120 rpm. Selective antibiotics were added to the production medium in case of plasmid harboring cultures. To trigger l-glutamate production in all strains, ethambutol (500 μg/ml) was added to the production medium (Radmacher et al. 2005). The genetically defined l-lysine-producing strain DM1729, which differs from C. glutamicum WT by point mutations in the aspartokinase gene lysC, the homoserine dehydrogenase gene hom, and the pyruvate carboxylase gene pyc (Georgi et al. 2005), was transformed with the following plasmid combinations pVWEx1 and pEKEx3, pVWEx1-araBAD and pEKEx3, pVWEx1 and pEKEx3-xylA, as well as pVWEx1-araBAD and pEKEx3-xylA. The resulting strains were cultured in CgXII minimal medium supplemented with the hydrolysates as carbon source for l-lysine production. Plasmids were induced with 1 mM IPTG, and the samples were removed at different intervals for the determination of biomass, amino acid production, and sugar utilization.
Analytical methods
Growth was followed by measuring the OD600 with UV 160A spectrophotometer (Shimadzu). The biomass concentration was calculated from OD600 values using an experimentally determined correlation factor of 0.25 g cell dry weight (DW) l−1for OD600 = 1 (Wendisch et al. 2000). The culture samples were centrifuged (10,000×g, 4°C; 10 min), and the resulting clear supernatant was used for various analysis. Quantitative determination of l-lysine and l-glutamate in the supernatants was carried out by reversed phase HPLC as described (Georgi et al. 2005). Various sugar concentrations were determined by high-performance liquid chromatography using Schimadzu HPLC system equipped with a refractive index detector and an HPX-87P column (Bio-Rad, USA) operating at 85°C with water as mobile phase at a flow rate of 0.6 ml/min.
Results
Screening and hydrolysis of agricultural residues
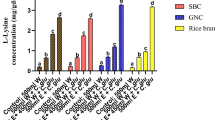
The composition analysis of several agricultural residues showed the highest xylose concentration in rice straw (37%) and the highest arabinose concentration in wheat bran (21%) (Table 2). Hemicellulosic hydrolysates were prepared by acid treatment of various agricultural residues (see “Materials and methods”). CgXII minimal medium with acid rice straw hydrolysate was prepared to contain 42 g/l of carbohydrate (40 mM glucose, 166 mM xylose, and 66 mM arabinose) as carbon source. Similarly, CgXII minimal medium with acid wheat bran hydrolysate was prepared to contain 41 g/l of carbohydrate (125 mM glucose, 62 mM xylose, and 64 mM arabinose) as carbon source. For comparison, a xylose/arabinose/glucose minimal medium was prepared by adding glucose, arabinose, and xylose as pure chemicals to CgXII medium (60 mM glucose, 200 mM xylose, and 120 mM arabinose). The xylose/arabinose/glucose minimal medium contained a higher total carbohydrate concentration (60 g/l) to compensate for osmotic effects due to the non-carbohydrate fractions of the acid rice straw and wheat bran hydrolysates. These media were used for growth and amino acid production experiments.
l-Glutamate production using acid rice straw and wheat bran hydrolysates
The empty vector control strain C. glutamicum WT(pEKEx3)(pVWEx1) and the pentose-utilizing recombinants WT(pEKEx3)(pVWEx1-araBAD), WT(pEKEx3-xylA)(pVWEx1), and WT(pEKEx3-xylA)(pVWEx1-araBAD) were grown in CgXII medium with either acid hydrolysates of rice straw or wheat bran or with the xylose/arabinose/glucose minimal medium (Table 3), and l-glutamate production was triggered by ethambutol addition as described previously (Radmacher et al. 2005; Stansen et al. 2005). All strains could grow with acid rice straw and wheat bran hydrolysates (Table 3). In all media tested, the empty vector control utilized glucose for biomass formation and l-glutamate production, while the pentose-utilizing recombinants grew to higher biomass concentrations and produced more l-glutamate as they utilized xylose and/or arabinose in addition to glucose (Table 3). In medium with acid rice straw hydrolysate, for example, the empty vector control strain grew to an OD of 12 ± 2 and produced 16 ± 3 mM l-glutamate, while the arabinose-utilizing strain grew to an OD of 20 ± 3 and produced 55 ± 4 mM l-glutamate, the xylose-utilizing strain grew to an OD of 33 ± 3 and produced 77 ± 5 mM l-glutamate, and the strain utilizing both pentoses grew to an OD of 42 ± 2 and produced 93 ± 4 mM l-glutamate (Table 3 and Fig. 1). In the experiments with acid rice straw hydrolysate, WT(pVWEx1-araBAD)(pEKEx3-xylA) showed a carbon-normalized product yield of 0.34 mol carbon in l-glutamate per mole of carbon in the sugars and a specific growth rate of 0.10 h−1 (compare to Fig. 1). A similar pattern was observed with respect to yields and growth rates using acid wheat bran hydrolysate (compare to Fig. S1).
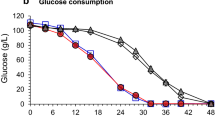
l-Glutamate production by different C. glutamicum strains with rice straw hydrolysate. a WT(pEKEx3)(pVWEx1); b WT(pEKEx3)(pVWEx1-araBAD); c WT(pEKEx3-xylA)(pVWEx1); d WT(pEKEx3-xylA)(pVWEx1-araBAD). Xylose concentrations in the culture supernatant are represented by filled triangles, arabinose concentrations by open squares, glucose concentrations by open circles, l-glutamate concentrations by filled circles, and OD600 by open diamonds. Ethambutol was added at the beginning of the experiment. Averages and standard deviations for three or more replicates are reported
l-Lysine production using acid rice straw and wheat bran hydrolysates
To characterize l-lysine production from hemicellulosic hydrolysates by derivatives of the l-lysine model producer strain DM1729, the empty vector control strain C. glutamicum DM1729(pEKEx3)(pVWEx1) and the pentose-utilizing recombinant strains DM1729(pEKEx3)(pVWEx1-araBAD), DM1729(pEKEx3-xylA)(pVWEx1), and DM1729(pEKEx3-xylA)(pVWEx1-araBAD) were grown in CgXII medium with either acid hydrolysates of rice straw or wheat bran or with the xylose/arabinose/glucose minimal medium (Table 3). As observed in the l-glutamate production experiments, growth and product formation from the pentose fractions of the hydrolysates depended on the presence of heterologously expressed araBAD and/or xylA whereby all pentose-utilizing strains reached higher final biomass and l-lysine concentrations than the empty vector controls strains (Table 3). For example, in medium with acid rice straw hydrolysate, the arabinose-utilizing strain grew to a higher OD (23 ± 2) than the empty vector control strain (OD of 15 ± 2) and produced more l-lysine (23 ± 2 mM as compared to 11 ± 2). As the xylose concentration in acid rice straw hydrolysate exceeded the arabinose concentration, the xylose-utilizing strain formed more biomass (OD of 33 ± 2) and produced more l-lysine (32 ± 2 mM), while the highest biomass formation (OD of 42 ± 2) and l-lysine production (42 ± 2 mM) was observed for the strain utilizing both xylose and arabinose (Table 3 and Fig. 2). The carbon-normalized product yields of the control and the arabinose-utilizing strains (0.27 ± 0.15 and 0.25 ± 0.06 mol-c l-lysine per mol-c monosaccharide) were higher than those of the strains utilizing xylose or both pentoses (0.18 ± 0.03 and 0.19 ± 0.02 mol-c l-lysine per mol-c monosaccharide, Fig. 3). However, the total amount of substrate consumed differed strikingly (7.2 g l−1 for the control and 41 g l−1 for the strain utilizing both pentoses), which may affect product yields.
l-Lysine production by different C. glutamicum strains with rice straw hydrolysate. a DM1729(pEKEx3)(pVWEx1); b DM1729(pEKEx3)(pVWEx1-araBAD); c DM1729(pEKEx3-xylA)(pVWEx1); d DM1729(pEKEx3-xylA)(pVWEx1-araBAD). Xylose concentrations in the culture supernatant are represented by filled triangles, arabinose concentrations by open squares, glucose concentrations by open circles, l-lysine concentrations by filled squares, and OD600 by open diamonds. Averages and standard deviations for three or more replicates are reported
Yields and volumetric productivities of l-lysine production experiments in different media and with different C. glutamicum strains. Minimal medium with hydrolysates of rice straw or wheat bran as well as xylose/arabinose/glucose minimal medium (mock) has been used. a Product yields expressed in mmol carbon (c) in l-lysine per mmol-c substrate. b Volumetric productivities expressed in g h−1 l−1. Averages and standard deviations from triplicate experiments are given in white columns for DM1729(pEKEx3)(pVWEx1); in hatched columns for DM1729(pEKEx3)(pVWEx1-araBAD); in gray columns for DM1729(pEKEx3-xylA)(pVWEx1); and in black columns for DM1729(pEKEx3-xylA)(pVWEx1-araBAD)
The maximal specific growth rates in acid rice straw hydrolysate determined in the experiments depicted in Fig. 2 were 0.11 h−1 for the empty vector control strain, 0.10 h−1 for DM1729(pVWEx1-araBAD), 0.09 h−1 for DM1729(pEKEx3-xylA), and 0.07 h−1 for DM1729(pVWEx1-araBAD)(pEKEx3-xylA). The volumetric productivities observed with rice straw hydrolysate were about twofold higher for the strain utilizing both pentoses (0.085 ± 0.004 g h−1 l−1) than that of the control strain (0.043 ± 0.004 g h−1 l−1), while the volumetric productivities of the strains utilizing only a single pentose were intermediate (Fig. 3). Taken together, the l-lysine production experiments (Figs. 2 and S2) revealed that the recombinant pentose-utilizing strains are superior to the parent strain with respect to production of l-lysine from acid rice straw and wheat bran hydrolysates.
Compared to l-lysine production from a blend of glucose, arabinose, and xylose as pure chemicals (xylose/arabinose/glucose minimal medium; Fig. S2), hemicellulosic hydrolysate-based l-lysine production was slower. This may be exemplified by growth and l-lysine production of strain DM1729(pVWEx1-araBAD)(pEKEx3-xylA) on the synthetic medium with the blend of glucose, arabinose, and xylose (Fig. 4a), with acid rice straw hydrolysate (Fig. 4b) and with acid wheat bran hydrolysate (Fig. 4c). The stationary growth phase was reached earlier (30 h) in the xylose/arabinose/glucose minimal medium than with the acid rice straw and wheat bran hydrolysates (48 h, Fig. 4). Similarly, the final l-lysine concentrations were reached earlier in the xylose/arabinose/glucose minimal medium than with the acid rice straw and wheat bran hydrolysates (50 h as compared to 72 h, Fig. 4). In addition, the substrates were exhausted earlier in the xylose/arabinose/glucose minimal medium than with the acid rice straw and wheat bran hydrolysates (Fig. 4). As a consequence, the volumetric l-lysine productivity for DM1729(pEKEx3-xylA)(pVWEx1-araBAD) (Fig. 3b) was higher in the xylose/arabinose/glucose minimal medium (0.170 ± 0.001 g h−1 l−1) than in media with acid hydrolysates of rice straw (0.085 ± 0.004 g h−1 l−1) and of wheat bran (0.085 ± 0.004 g h−1 l−1).
l-lysine production with xylose/arabinose/glucose minimal medium (a), rice straw hydrolysate (b; data taken from Fig. 2d) and wheat bran hydrolysate (c) with C. glutamicum DM1729(pEKEx3-xylA)(pVWEx1-araBAD). Xylose concentrations in the culture supernatant are represented by filled triangles, arabinose concentrations by open squares, glucose concentrations by open circles, l-lysine concentrations by filled squares, and OD600 by open diamonds. Averages and standard deviations for three or more replicates are reported
Discussion
Minimal media with acid hydrolysates of, e.g., rice straw or wheat bran support growth of C. glutamicum WT with the glucose fraction of the hydrolysates being utilized completely for biomass formation. However, the maximal specific growth rate of 0.38 h−1 for C. glutamicum WT in glucose minimal medium (Wendisch et al. 2000) is clearly higher than that of the empty vector control strains used here in hydrolysate-based media (Figs. 1, 2, S1 and S2), which were about 0.10–0.20 h−1. This might be due in part to the higher osmolality of the hydrolysate-based media as compared to glucose minimal medium because the growth rate of the control strain in the xylose/arabinose/glucose minimal medium (i.e., when arabinose and xylose were present besides glucose, but could not be utilized) was also lower than in the glucose minimal medium. In addition, the presence of inhibitors in hemicellulosic hydrolysates may as well play a role leading to reduced growth and production rates in hydrolysate-based media (see below).
Glucose is co-utilized with arabinose and xylose by the recombinant C. glutamicum strains not only when the substrates were present as pure chemicals but also when present in acid hydrolysates from agricultural residues (Figs. 1, 2, S1 and S2). Co-utilization of pentoses and hexoses by C. glutamicum has been observed in this and previous studies (Kawaguchi et al. 2006, 2008; Sasaki et al. 2008; Schneider et al. 2011) for several but not all (Buschke et al. 2011) strains tested. Efforts towards engineering co-utilization of glucose, arabinose, and xylose by S. cerevisiae were made for a long time and have led to recent successes (Karhumaa et al. 2006; van Maris et al. 2006), but until today, alternative solutions are followed as well, e.g., sequential fermentation of rice straw hydrolysate first by S. cerevisiae, followed by heat inactivation, and finally by fermentation of the xylose fraction by Pichia stipitis (Li et al. 2011). Simultaneous utilization of various carbon sources by C. glutamicum (Dominguez et al. 1997; Eggeling and Bott 2005; Engels et al. 2008; Lee et al. 1998; Wendisch 2006; Wendisch et al. 2000) is a hallmark of this bacterium setting it apart from yeasts, E. coli and B. subtilis, which typically show sequential utilization of substrates present in blends and this growth pattern is often accompanied by a diauxic growth lag. Very few exceptions to substrate co-utilization have been described for C. glutamicum (e.g., glucose being utilized prior to ethanol (Arndt et al. 2008; Arndt and Eikmanns 2008) or prior to glutamate (Kronemeyer et al. 1995)). Thus, C. glutamicum lends itself as ideal candidate for processes based on lignocellulosic or hemicellulosic hydrolysates and other second-generation substrate blends.
While co-utilization of pentoses and glucose was observed using acid hydrolysates as well as a blend of the pure chemicals, glucose, arabinose, and xylose growth and substrate utilization were slower in media based on the acid hydrolysates, which might contain growth inhibitors. It is known for E. coli and yeasts that compounds present in lignocellulosic hydrolysates such as acetic acid, hydroxymethylfurfural, or furfural inhibit growth (Klinke et al. 2004; Palmqvist et al. 1999; Zaldivar and Ingram 1999; Zaldivar et al. 1999, 2000; Heer and Sauer 2008). It has been described for C. glutamicum that a number of organic acids, furan, and phenolic inhibitors did not affect ethanol production by growth-arrested C. glutamicum notably, which was contributed primarily to the growth-arrested conditions (Sakai et al. 2007). While it is known that acetic acid reduces the growth rate of C. glutamicum to some extent when added to glucose minimal medium, C. glutamicum can utilize acetic acid efficiently as sole or combined source of carbon and energy (Gerstmeir et al. 2003; Wendisch et al. 2000). By contrast, the growth sensitivities of C. glutamicum to furfural and hydroxymethylfurfural were found to be similar to those of yeasts (e.g., S. cerevisiae CBS 1200, Candida shehatae ATCC 22984 or P. stipitis NRRL Y 7124) (Sakai et al. 2007). Thus, the slower growth and amino acid production observed using media based on hemicellulosic hydrolysates may at least in part be due to the presence of furfural and/or hydroxymethylfurfural. Recently, a (hydroxymethyl)furfural degrading bacterium was isolated, and the genes for enzymes of the involved pathway, which had been proposed previously based on biochemical evidence (Koenig and Andreesen 1990), have been identified (Koopman et al. 2010). The latter study opened the way to in situ detoxification of lignocellulosic hydrolysates either using sequential or co-fermentation of Cupriavidus basilensis with the biotechnologically relevant microorganisms (yeast, E. coli, and C. glutamicum) or by metabolic engineering of the biotechnologically relevant microorganisms using the genes for 5-(hydroxymethyl)furfural degradation from C. basiliensis (Koopman et al. 2010).
To increase growth and amino acid production based on hemicellulosic hydrolysates by recombinant C. glutamicum, the capacities to import arabinose and/or xylose and to catabolize them to intermediates of the central metabolism may be engineered. Faster pentose catabolic flux may be obtained by increasing and/or balancing heterologous gene expression levels, by making use of genes taken from pentose-utilizing bacteria other than E. coli or by implementing the fungal pentose utilization pathways as, e.g., present in P. stipitis. Increased pentose uptake rates may be achieved by transport engineering, e.g., by employing the l-arabinose transporter AraE from C. glutamicum strain ATCC31831 (Sasaki et al. 2009). This uptake system apparently accepts both l-arabinose and xylose as substrates (Sasaki et al. 2009). However, as AraE is not encoded in the genome of C. glutamicum WT, other hitherto-unknown uptake systems ensure at least a basal pentose transport capacity. Gene regulatory engineering is also conceivable to improve pentose utilization, as it was observed in this and previous studies (Kawaguchi et al. 2006, 2008; Schneider et al. 2011) that glucose slowed pentose utilization, indicating some sort of glucose repression. Presumably, this type of glucose repression affects expression of the endogenous hitherto-unknown pentose transporter gene(s) rather than plasmid-borne expression of the heterologous genes for arabinose and xylose catabolism.
Notwithstanding the potential to further improve recombinant pentose-utilizing C. glutamicum strains, this study has shown that C. glutamicum, a work-horse of biotechnology (Eggeling and Bott 2005; Wendisch et al. 2006), may be used for biotechnological processes based on agricultural residues by capitalizing on its property to simultaneously utilize several carbon substrates such as the pentose and hexose fractions of hemicellulosic hydrolysates.
References
Aristidou A, Penttila M (2000) Metabolic engineering applications to renewable resource utilization. Curr Opin Biotechnol 11(2):187–198
Arndt A, Auchter M, Ishige T, Wendisch VF, Eikmanns BJ (2008) Ethanol catabolism in Corynebacterium glutamicum. J Mol Microbiol Biotechnol 15(4):222–233
Arndt A, Eikmanns BJ (2008) Regulation of carbon metabolism in Corynebacterium glutamicum. In: Burkovski A (ed) Corynebacteria: genomics and molecular biology. Caister Academic, Wymondham, pp 155–182
Barrett E, Stanton C, Zelder O, Fitzgerald G, Ross RP (2004) Heterologous expression of lactose- and galactose-utilizing pathways from lactic acid bacteria in Corynebacterium glutamicum for production of lysine in whey. Appl Environ Microbiol 70(5):2861–2866
Becker J, Boles E (2003) A modified Saccharomyces cerevisiae strain that consumes L-arabinose and produces ethanol. Appl Environ Microbiol 69(7):4144–4150
Blombach B, Riester T, Wieschalka S, Ziert C, Youn JW, Wendisch VF, Eikmanns BJ (2011) Corynebacterium glutamicum tailored for efficient isobutanol production. Appl Environ Microbiol 77(10):3300–3310
Brabetz W, Liebl W, Schleifer KH (1991) Studies on the utilization of lactose by Corynebacterium glutamicum, bearing the lactose operon of Escherichia coli. Arch Microbiol 155(6):607–612
Buschke N, Schroder H, Wittmann C (2011) Metabolic engineering of Corynebacterium glutamicum for production of 1,5-diaminopentane from hemicellulose. Biotechnol J 6(3):306–317
Cadenas RF, Gil JA, Martin JF (1992) Expression of Streptomyces genes encoding extracellular enzymes in Brevibacterium lactofermentum: secretion proceeds by removal of the same leader peptide as in Streptomyces lividans. Appl Microbiol Biotechnol 38(3):362–369
Cerdeno-Tarraga AM, Efstratiou A, Dover LG, Holden MT, Pallen M, Bentley SD, Besra GS, Churcher C, James KD, De Zoysa A, Chillingworth T, Cronin A, Dowd L, Feltwell T, Hamlin N, Holroyd S, Jagels K, Moule S, Quail MA, Rabbinowitsch E, Rutherford KM, Thomson NR, Unwin L, Whitehead S, Barrell BG, Parkhill J (2003) The complete genome sequence and analysis of Corynebacterium diphtheriae NCTC13129. Nucleic Acids Res 31(22):6516–6523
Dominguez H, Cocaign-Bousquet M, Lindley ND (1997) Simultaneous consumption of glucose and fructose from sugar mixtures during batch growth of Corynebacterium glutamicum. Appl Microbiol Biot 47(5):600–603
Dominguez H, Rollin C, Guyonvarch A, Guerquin-Kern JL, Cocaign-Bousquet M, Lindley ND (1998) Carbon-flux distribution in the central metabolic pathways of Corynebacterium glutamicum during growth on fructose. Eur J Biochem 254(1):96–102
Eggeling L, Bott M (2005) Handbook of Corynebacterium glutamicum. CRC, USA
Eggeling L, Reyes O (2005) Experiments. In: Eggeling L, Bott M (eds) Handbook of Corynebacterium glutamicum. CRC Press, USA, pp 3535–3566
Eikmanns BJ (2005) Central metabolism: tricarboxylic acid cycle and anaplerotic reactions. In: Eggeling L, Bott M (eds) Handbook on Corynebacterium glutamicum. CRC Press, USA, pp 241–276
Eikmanns BJ, Thum-Schmitz N, Eggeling L, Lüdtke KU, Sahm H (1994) Nucleotide sequence, expression and transcriptional analysis of the Corynebacterium glutamicum gltA gene encoding citrate synthase. Microbiology 140(Pt 8):1817–1828
Engels V, Georgi T, Wendisch VF (2008) ScrB (Cg2927) is a sucrose-6-phosphate hydrolase essential for sucrose utilization by Corynebacterium glutamicum. FEMS Microbiol Lett 289(1):80–89
Frunzke J, Engels V, Hasenbein S, Gatgens C, Bott M (2008) Co-ordinated regulation of gluconate catabolism and glucose uptake in Corynebacterium glutamicum by two functionally equivalent transcriptional regulators, GntR1 and GntR2. Mol Microbiol 67(2):305–322
Georgi T, Rittmann D, Wendisch VF (2005) Lysine and glutamate production by Corynebacterium glutamicum on glucose, fructose and sucrose: roles of malic enzyme and fructose-1,6-bisphosphatase. Metab Eng 7(4):291–301
Gerstmeir R, Wendisch VF, Schnicke S, Ruan H, Farwick M, Reinscheid D, Eikmanns BJ (2003) Acetate metabolism and its regulation in Corynebacterium glutamicum. J Biotechnol 104(1–3):99–122
Hahn-Hagerdal B, Karhumaa K, Fonseca C, Spencer-Martins I, Gorwa-Grauslund MF (2007a) Towards industrial pentose-fermenting yeast strains. Appl Microbiol Biotechnol 74(5):937–953
Hahn-Hagerdal B, Karhumaa K, Jeppsson M, Gorwa-Grauslund MF (2007b) Metabolic engineering for pentose utilization in Saccharomyces cerevisiae. Adv Biochem Eng Biotechnol 108:147–177
Hanahan D (1983) Studies on transformation of Escherichia coli with plasmids. J Mol Biol 166(4):557–580
Heer D, Sauer U (2008) Identification of furfural as a key toxin in lignocellulosic hydrolysates and evolution of a tolerant yeast strain. Microb Biotechnol 1(6):497–506
Jeffries TW, Jin YS (2000) Ethanol and thermotolerance in the bioconversion of xylose by yeasts. Adv Appl Microbiol 47:221–268
Kalinowski J, Bathe B, Bartels D, Bischoff N, Bott M, Burkovski A, Dusch N, Eggeling L, Eikmanns BJ, Gaigalat L, Goesmann A, Hartmann M, Huthmacher K, Kramer R, Linke B, McHardy AC, Meyer F, Mockel B, Pfefferle W, Puhler A, Rey DA, Ruckert C, Rupp O, Sahm H, Wendisch VF, Wiegrabe I, Tauch A (2003) The complete Corynebacterium glutamicum ATCC 13032 genome sequence and its impact on the production of L-aspartate-derived amino acids and vitamins. J Biotechnol 104(1–3):5–25
Karhumaa K, Wiedemann B, Hahn-Hagerdal B, Boles E, Gorwa-Grauslund MF (2006) Co-utilization of L-arabinose and D-xylose by laboratory and industrial Saccharomyces cerevisiae strains. Microb Cell Fact 5:18
Kato O, Youn JW, Stansen KC, Matsui D, Oikawa T, Wendisch VF (2010) Quinone-dependent D-lactate dehydrogenase Dld (Cg1027) is essential for growth of Corynebacterium glutamicum on D-lactate. BMC Microbiol 10:321
Kawaguchi H, Sasaki M, Vertes AA, Inui M, Yukawa H (2008) Engineering of an L-arabinose metabolic pathway in Corynebacterium glutamicum. Appl Microbiol Biotechnol 77(5):1053–1062
Kawaguchi H, Sasaki M, Vertes AA, Inui M, Yukawa H (2009) Identification and functional analysis of the gene cluster for L-arabinose utilization in Corynebacterium glutamicum. Appl Environ Microbiol 75(11):3419–3429
Kawaguchi H, Vertes AA, Okino S, Inui M, Yukawa H (2006) Engineering of a xylose metabolic pathway in Corynebacterium glutamicum. Appl Environ Microbiol 72(5):3418–3428
Kiefer P, Heinzle E, Wittmann C (2002) Influence of glucose, fructose and sucrose as carbon sources on kinetics and stoichiometry of lysine production by Corynebacterium glutamicum. J Industrial Microbiol Biotechnol 28:338–343
Kinoshita S, Udaka S, Shimono M (1957) Studies on the amino acid fermentation. Production of L-glutamic acid by various microorganisms. J Gen Appl Microbiol 3:193–205
Klinke HB, Thomsen AB, Ahring BK (2004) Inhibition of ethanol-producing yeast and bacteria by degradation products produced during pre-treatment of biomass. Appl Microbiol Biotechnol 66(1):10–26
Koenig K, Andreesen JR (1990) Xanthine dehydrogenase and 2-furoyl-coenzyme A dehydrogenase from Pseudomonas putida Fu1: two molybdenum-containing dehydrogenases of novel structural composition. J Bacteriol 172(10):5999–6009
Koopman F, Wierckx N, de Winde JH, Ruijssenaars HJ (2010) Identification and characterization of the furfural and 5-(hydroxymethyl)furfural degradation pathways of Cupriavidus basilensis HMF14. Proc Natl Acad Sci USA 107(11):4919–4924
Kotrba P, Inui M, Yukawa H (2003) A single V317A or V317M substitution in enzyme II of a newly identified beta-glucoside phosphotransferase and utilization system of Corynebacterium glutamicum R extends its specificity towards cellobiose. Microbiology 149(Pt 6):1569–1580
Krämer R, Lambert C, Hoischen C, Ebbighausen H (1990) Uptake of glutamate in Corynebacterium glutamicum. 1. Kinetic properties and regulation by internal pH and potassium. Eur J Biochem 194(3):929–935
Krause FS, Blombach B, Eikmanns BJ (2010) Metabolic engineering of Corynebacterium glutamicum for 2-ketoisovalerate production. Appl Environ Microbiol 76(24):8053–8061
Krings E, Krumbach K, Bathe B, Kelle R, Wendisch VF, Sahm H, Eggeling L (2006) Characterization of myo-inositol utilization by Corynebacterium glutamicum: the stimulon, identification of transporters, and influence on L-lysine formation. J Bacteriol 188(23):8054–8061
Kronemeyer W, Peekhaus N, Kramer R, Sahm H, Eggeling L (1995) Structure of the gluABCD cluster encoding the glutamate uptake system of Corynebacterium glutamicum. J Bacteriol 177(5):1152–1158
Lee HW, Pan JG, Lebeault JM (1998) Enhanced L-lysine production in threonine-limited continuous culture of Corynebacterium glutamicum by using gluconate as a secondary carbon source with glucose. Appl Microbiol Biotechnol 49(1):9–15
Li Y, Park JY, Shiroma R, Tokuyasu K (2011) Bioethanol production from rice straw by a sequential use of Saccharomyces cerevisiae and Pichia stipitis with heat inactivation of Saccharomyces cerevisiae cells prior to xylose fermentation. J Biosci Bioeng 111(6):682–686
Lin ECC (1996) Dissimilatory pathways for sugars, polyols and carboxylates. In: Neidhardt FC, Curtiss R III, Ingraham JL, Lin ECC, Low KB, Magasanik B, Reznikoff WS, Riley M, Schaechter M, Umbarger HE (eds) Escherichia coli and Salmonella: cellular and molecular biology, 2nd edn. ASM, Washington, pp 307–342
Lindner SN, Knebel S, Pallerla SR, Schoberth SM, Wendisch VF (2010) Cg2091 encodes a polyphosphate/ATP-dependent glucokinase of Corynebacterium glutamicum. Appl Microbiol Biotechnol 87(2):703–713
Lindner SN, Seibold GM, Henrich A, Kramer R, Wendisch VF (2011) Phosphotransferase system-independent glucose utilization in Corynebacterium glutamicum by inositol permeases and glucokinases. Appl Environ Microbiol 77(11):3571–3581
Mimitsuka T, Sawai H, Hatsu M, Yamada K (2007) Metabolic engineering of Corynebacterium glutamicum for cadaverine fermentation. Biosci Biotechnol Biochem 71(9):2130–2135
Moon MW, Kim HJ, Oh TK, Shin CS, Lee JS, Kim SJ, Lee JK (2005) Analyses of enzyme II gene mutants for sugar transport and heterologous expression of fructokinase gene in Corynebacterium glutamicum ATCC 13032. FEMS Microbiol Lett 244(2):259–266
Nishio Y, Nakamura Y, Kawarabayasi Y, Usuda Y, Kimura E, Sugimoto S, Matsui K, Yamagishi A, Kikuchi H, Ikeo K, Gojobori T (2003) Comparative complete genome sequence analysis of the amino acid replacements responsible for the thermostability of Corynebacterium efficiens. Genome Res 13(7):1572–1579
Palmqvist E, Grage H, Meinander NQ, Hahn-Hagerdal B (1999) Main and interaction effects of acetic acid, furfural, and p-hydroxybenzoic acid on growth and ethanol productivity of yeasts. Biotechnol Bioeng 63(1):46–55
Peters-Wendisch PG, Schiel B, Wendisch VF, Katsoulidis E, Mockel B, Sahm H, Eikmanns BJ (2001) Pyruvate carboxylase is a major bottleneck for glutamate and lysine production by Corynebacterium glutamicum. J Mol Microbiol Biotechnol 3(2):295–300
Polen T, Kramer M, Bongaerts J, Wubbolts M, Wendisch VF (2005) The global gene expression response of Escherichia coli to L-phenylalanine. J Biotechnol 115(3):221–237
Radmacher E, Stansen KC, Besra GS, Alderwick LJ, Maughan WN, Hollweg G, Sahm H, Wendisch VF, Eggeling L (2005) Ethambutol, a cell wall inhibitor of Mycobacterium tuberculosis, elicits L-glutamate efflux of Corynebacterium glutamicum. Microbiology 151(Pt 5):1359–1368
Rittmann D, Lindner SN, Wendisch VF (2008) Engineering of a glycerol utilization pathway for amino acid production by Corynebacterium glutamicum. Appl Environ Microbiol 74(20):6216–6222
Sakai S, Tsuchida Y, Okino S, Ichihashi O, Kawaguchi H, Watanabe T, Inui M, Yukawa H (2007) Effect of lignocellulose-derived inhibitors on growth of and ethanol production by growth-arrested Corynebacterium glutamicum R. Appl Environ Microbiol 73(7):2349–2353
Sasaki M, Jojima T, Inui M, Yukawa H (2008) Simultaneous utilization of D-cellobiose, D-glucose, and D-xylose by recombinant Corynebacterium glutamicum under oxygen-deprived conditions. Appl Microbiol Biotechnol 81(4):691–699
Sasaki M, Jojima T, Kawaguchi H, Inui M, Yukawa H (2009) Engineering of pentose transport in Corynebacterium glutamicum to improve simultaneous utilization of mixed sugars. Appl Microbiol Biotechnol 85(1):105–115
Schneider J, Niermann K, Wendisch VF (2011) Production of the amino acids l-glutamate, l-lysine, l-ornithine and l-arginine from arabinose by recombinant Corynebacterium glutamicum. J Biotechnol 154(2–3):191–198
Schneider J, Wendisch VF (2010) Putrescine production by engineered Corynebacterium glutamicum. Appl Microbiol Biotechnol 88(4):859–868
Seibold G, Auchter M, Berens S, Kalinowski J, Eikmanns BJ (2006) Utilization of soluble starch by a recombinant Corynebacterium glutamicum strain: growth and lysine production. J Biotechnol 124(2):381–391
Sluiter A, Hames B, Ruiz R, Scarlata C, Sluiter J, Templeton D (2008) Determination of sugars, byproducts, and degradation products in liquid fraction process samples. NREL Report No TP-510-42623 (January 2008)
Stansen C, Uy D, Delaunay S, Eggeling L, Goergen JL, Wendisch VF (2005) Characterization of a Corynebacterium glutamicum lactate utilization operon induced during temperature-triggered glutamate production. Appl Environ Microbiol 71(10):5920–5928
Tateno T, Fukuda H, Kondo A (2007) Production of L-lysine from starch by Corynebacterium glutamicum displaying alpha-amylase on its cell surface. Appl Microbiol Biotechnol 74(6):1213–1220
Tauch A, Kaiser O, Hain T, Goesmann A, Weisshaar B, Albersmeier A, Bekel T, Bischoff N, Brune I, Chakraborty T, Kalinowski J, Meyer F, Rupp O, Schneiker S, Viehoever P, Puhler A (2005) Complete genome sequence and analysis of the multiresistant nosocomial pathogen Corynebacterium jeikeium K411, a lipid-requiring bacterium of the human skin flora. J Bacteriol 187(13):4671–4682
van Maris AJ, Abbott DA, Bellissimi E, van den Brink J, Kuyper M, Luttik MA, Wisselink HW, Scheffers WA, van Dijken JP, Pronk JT (2006) Alcoholic fermentation of carbon sources in biomass hydrolysates by Saccharomyces cerevisiae: current status. Antonie Van Leeuwenhoek 90(4):391–418
Wendisch VF (2006) Genetic regulation of Corynebacterium glutamicum metabolism. J Microbiol Biotechnol 16(7):999–1009
Wendisch VF, Bott M, Eikmanns BJ (2006) Metabolic engineering of Escherichia coli and Corynebacterium glutamicum for biotechnological production of organic acids and amino acids. Curr Opin Microbiol 9(3):268–274
Wendisch VF, de Graaf AA, Sahm H, Eikmanns BJ (2000) Quantitative determination of metabolic fluxes during coutilization of two carbon sources: comparative analyses with Corynebacterium glutamicum during growth on acetate and/or glucose. J Bacteriol 182(11):3088–3096
Wyman CE (1999) Production of low cost sugars from biomass: progress, opportunities, and challenges. In: Overend RP, Chornet E (eds) Biomass: a growth opportunity in green energy and value added products, vol 1. Pergamon Press, United Kingdom, pp 867–872
Youn JW, Jolkver E, Kramer R, Marin K, Wendisch VF (2008) Identification and characterization of the dicarboxylate uptake system DccT in Corynebacterium glutamicum. J Bacteriol 190(19):6458–6466
Youn JW, Jolkver E, Kramer R, Marin K, Wendisch VF (2009) Characterization of the dicarboxylate transporter DctA in Corynebacterium glutamicum. J Bacteriol 191(17):5480–5488
Zaldivar J, Ingram LO (1999) Effect of organic acids on the growth and fermentation of ethanologenic Escherichia coli LY01. Biotechnol Bioeng 66(4):203–210
Zaldivar J, Martinez A, Ingram LO (1999) Effect of selected aldehydes on the growth and fermentation of ethanologenic Escherichia coli. Biotechnol Bioeng 65(1):24–33
Zaldivar J, Martinez A, Ingram LO (2000) Effect of alcohol compounds found in hemicellulose hydrolysate on the growth and fermentation of ethanologenic Escherichia coli. Biotechnol Bioeng 68(5):524–530
Acknowledgements
Work in the laboratories of the corresponding authors was supported in part by grants from the Department of Biotechnology (DBT), New Delhi, India, and from the International Bureau (IB) of the Federal Ministry of Education and Research (BMBF), Germany (IND 07/030) under the Indo-German bilateral program.
Author information
Authors and Affiliations
Corresponding authors
Additional information
V. Gopinath and T. M. Meiswinkel contributed equally to this work.
Electronic supplementary materials
Below is the link to the electronic supplementary material.
ESM1
(PDF 693 kb)
Rights and permissions
About this article
Cite this article
Gopinath, V., Meiswinkel, T.M., Wendisch, V.F. et al. Amino acid production from rice straw and wheat bran hydrolysates by recombinant pentose-utilizing Corynebacterium glutamicum . Appl Microbiol Biotechnol 92, 985–996 (2011). https://doi.org/10.1007/s00253-011-3478-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-011-3478-x