Abstract
Antibiotic-resistant bacteria are an increasing source of concern in all environments in which these drugs have been used. More stringent regulations have led to a slow but sure decrease in antibiotic use in the food industry worldwide, but have also stimulated the search for alternative antibacterial agents. In medicine, the number of people infected with pan-resistant bacteria is driving research to develop new treatments. Within these contexts, studies on the use of bacteriophages in both medicine and the food industry have recently flourished. This renewed interest has coincided with the demonstration that these viruses are involved in geochemical cycles, revolutionizing our vision of their ecological role on our planet. Bacteriophages have co-evolved with bacteria for billions of years and retain the ability to infect bacteria efficiently. They are undoubtedly one of the best potential sources of new solutions for the management of undesirable bacteria.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
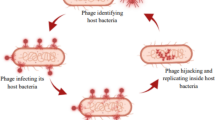
Bacteriophages are viruses that specifically infect bacteria. The infection process starts with bacteriophage adsorption to the bacterial cell surface, followed by the injection of the viral genetic material into the cytoplasm. Virulent bacteriophages infect bacterial cells by immediately hijacking host cell replication, transcription, and translation machineries for the production of new virions, thus converting the host cell into a virus factory (Lwoff 1967). By contrast, temperate bacteriophages may delay these steps by inserting their genome into the host chromosome, remaining in this dormant state as long as bacteria do not perceive any signal leading to the resumption of their infectious cycle. The last step of the infectious cycle is the release of new virions, through either lysis of the cell wall or extrusion without cell wall disruption (Fig. 1).
In 1917, Félix d’Herelle coined the term “bacteriophage” for the agent responsible for the bacterial lysis he observed. It was subsequently recognized that Twort had already, in 1915, described the same phenomenon, and that the bactericidal effect on Vibrio cholerae in the Ganges river reported by Hankin in 1896 was also due to bacteriophages (d’Herelle 1917; Hankin 1896; Twort 1915). A century later, Faruque et al. observed an inverse relationship between the likelihood of cholera epidemics and the presence of bacteriophages in the concerned area (Faruque et al. 2005). Both environmental and medical aspects were considered in this study, corresponding to the two fields of research on bacteriophages that have recently expanded considerably.
Research on bacteriophages over the last century may be divided into three periods. The first of these periods immediately followed d’Herelle’s publication relating the isolation of bacteriophages from the stools of patients recovering from dysentery. During this period, d’Herelle developed a medical application for bacteriophages, inventing a discipline known as “phage therapy” (Summers 2001). This application expanded worldwide, but as both successes and failures were reported, it was unable to resist the onslaught of antibiotics, which began to come into widespread use in the 1940s. At about the same time, phage therapy was also developed for plants (Balogh et al. 2010).
The second phase began in the 1940s, with the identification of bacteriophages as model organisms for experiments leading to major fundamental discoveries, paving the way for a new scientific discipline known as “molecular biology” (Summers 1999). For example, the ability of temperate bacteriophages to insert their genome into that of the host formed the basis of the development of genetic tools in microbiology (Shapiro et al. 1969). Until the 1990s, bacteriophages were mostly studied for the development of molecular tools for advancing our understanding of many aspects of biology, from the detailed regulation of gene expression to the three-dimensional structures of proteins. One key example of this is the phage display techniques currently widely used in biological studies of eukaryotes, an application far removed from the primary role of bacteriophages as infectious agents of bacteria (Bratkovic 2010).
The third phase began some 20 years ago and involves studies of bacteriophages focusing on their ecological role in the biosphere and on their use to manage undesirable bacterial populations.
Ecological role of bacteriophages
Studies on aquatic viruses have highlighted the tremendously high abundance of these organisms on Earth (1031 particles versus only 107 humans; Fuhrman 1999; Bergh et al. 1989). The stability of these viruses is affected by various environmental factors, including UV light. Their maintenance at such large numbers thus requires a dynamic renewal of viral populations, involving the massive infection of many bacterial hosts. It is estimated that bacteriophages account for 20% to 50% of bacterial mortality. These infections release large amounts of cellular compounds, greatly increasing the pool of dissolved and particulate organic matter involved in biogeochemical cycles (Wommack and Colwell 2000). However, the renewal of these viral populations does not lead to a global decrease in bacterial biomass. Nevertheless, some local variations have been observed, indicating that bacterium and bacteriophage populations co-exist through dynamic evolution (Faruque et al. 2005). The so-called “arms race” between bacteriophages and bacteria is a major driving force behind the evolution of both types of organisms, based on bacterial resistance to bacteriophage infection and the ways in which bacteriophages can overcome it (Pal et al. 2007; Suttle 2007; Rohwer et al. 2009; Rodriguez-Brito et al. 2010). The molecular basis of this competition between bacteriophages and bacteria has been studied for decades, but the recent discovery of the CRISPR system demonstrates that there may still be other as yet undiscovered mechanisms (Labrie et al. 2010; Barrangou et al. 2007). Bacteriophage genomes are much smaller than bacterial genomes and might therefore be thought unlikely to contain much new information. However, this assumption has proved very wrong, given that most of the bacteriophage sequences recently obtained display only limited identity to previously sequenced viruses, with up to 80% of their open reading frames (ORFs) displaying no similarity to sequences in current databases (Abedon 2009; Thurber 2009). Moreover, we have yet to find, and are indeed unlikely to find, any ORF conserved in all bacteriophages. Only at the structural level can some resemblances between distantly related viruses be found (Krupovic et al. 2010). Bacteriophages should therefore be considered as a continually evolving reservoir of genes (Paterson et al. 2010). They can provide new functions that are selected or lost during evolution (Forterre 2010; Forterre and Prangishvili 2009). This pool of constantly evolving functions probably constitutes one of the best resources for potential applications (Schoenfeld et al. 2010).
Applications of bacteriophages
Bacteria resistant to multiple antibiotics, the famous “superbugs”, have become an increasing concern for human health in recent years, with the pipeline of drugs for dealing with such bacteria almost empty (Kumarasamy et al. 2010; Nordmann et al. 2007; Taubes 2008). As a consequence, the medical application of bacteriophages is now being reconsidered. The food industry is also concerned by these “superbugs” in two ways. Firstly, the lack of efficient antibiotics increases the risk of considerable losses in food production systems (livestock, fisheries, agriculture). Secondly, stronger regulation has been imposed on the food industry, forcing the industry to abandon the use of antibiotics as growth promoters, to limit the risk of transmitting drug resistance to human pathogens (regulation IP/03/1058 adopted by the EU on July 2003). In this particular context, research into bacteriophages and their applications in both food and medicine has recently expanded, as pointed out by numerous reviews (Sulakvelidze et al. 2001; Merril et al. 2003; Housby and Mann 2009; O'Flaherty et al. 2009).
Three major issues relating to the use of bacteriophages should be taken into account when developing bacteriophage applications. The first issue is the emergence of resistant bacterial clones. However, this may be limited by the use of cocktails that have been shown in vitro to prevent the rapid emergence of resistant strains (Tanji et al. 2004). Nevertheless, it may not be entirely possible to prevent the emergence of resistant bacteria, so bacterial resistance to bacteriophages should be closely monitored during any application of bacteriophages.
The second issue is the specificity of bacteriophages, which may limit potential applications in situations in which the specific bacterial strain cannot be identified. Consequently, preventive applications of bacteriophages are based on the likelihood of efficacy. The use of bacteriophages with a broad host spectrum, such as Staphylococcus aureus phi812 or Listeria P100, should, however, increase the probability of success (Goodridge 2010; Pantucek et al. 1998; Carlton et al. 2005). In other cases, the use of bacteriophage cocktails to increase host range may increase overall potential efficiency, although efficiency is unlikely to reach 100% (Denou et al. 2009).
The third issue is the safety of bacteriophages. Genome sequencing can help to rule out the use of bacteriophages containing undesirable genes encoding toxins or integrases, for example (Denou et al. 2009). However, bacteriophage genomes contain many ORFans, some of which may be classified as undesirable genes in the future. Nevertheless, genes of unknown function are also present in the genomes of probiotic bacteria classified as safe for human consumption. The safety of bacteriophage preparations is also a matter of concern. However, purification techniques have evolved considerably in the last few years and standard procedures may eventually be developed, improving the homogeneity and safety of these preparations (Kramberger et al. 2010; Merabishvili et al. 2009; Gill and Hyman 2010; Courchesne et al. 2009).
Bacteriophage development for the food industry
Bacteriophages represent a serious threat in the dairy industry, causing losses in the production of cheese and fermented products (Brussow 2001). However, they are also seen as positive agents, replacing antibiotics in the control of pathogens. Two main applications have been developed to prevent food contamination and/or to treat bacterial infections in animals or plants.
Prevention
There are two principal sources of contamination in the production of food products: the raw material, which may contain bacteria pathogenic to humans, and humans, who may contaminate products during their processing. The use of bacteriophages before the initiation of a process should reduce the probability of contamination originating from the initial product, and their use during processing could also potentially prevent recontamination. Such strategies for targeting single pathogens, such as Listeria in cheese production and Escherichia coli in the meat industry are currently being developed (Guenther et al. 2009; Rivas et al. 2010). In 2006, the US Food and Drug Administration approved a product based on bacteriophages targeting Listeria as an additive for use in cheese production. This unique industrial application should be followed by others, provided that the scaling-up process from laboratory to industry takes into account factors associated with food processing itself, such as temperature, pH, and humidity, which may affect the efficacy of preventive applications.
Treatment
In 2005, the US Environmental Protection Agency approved a bacteriophage-based product for the treatment of bacterial spot on pepper and tomato and bacterial speck on tomato (EPA Registration No 67986–1). As in the meat industry, bacteriophage treatments for plants are confronted with problems associated with scaling up. The use of bacteriophages in open fields is hindered by several factors: uncontrolled environments (temperature, sun exposure, humidity etc.), uneven dispersion, and rapid inactivation. However, these environmental problems can be limited by the use of greenhouses. In addition, protective formulations have been shown to reduce the inactivation of bacteriophages by sunlight and rain washout (Balogh et al. 2010).
Animals could also be given bacteriophage-based treatments rather than antibiotics (Johnson et al. 2008). Bacteriophages active against three major human pathogens that are commensal in animals—E. coli O157:H7, Salmonella and Campylobacter—have been identified and are sometimes used in experimental treatments (Callaway et al. 2006; Rozema et al. 2009; Loc Carrillo et al. 2005; Wall et al. 2010). A recent study in an industrial poultry system showed that E. coli infection in chicken could be reduced by bacteriophage treatment (Oliveira et al. 2010). The authors of this study showed that low doses of bacteriophages were more effective than initially predicted on the basis of in vitro results. As the transmission of the disease from one infected chicken to another takes days to weeks, the early initiation of treatment with low doses of bacteriophages may be sufficient to prevent the development of advanced infections. Several studies on experimental farms have generated promising results requiring confirmation at the industrial scale (Jamalludeen et al. 2009; Rivas et al. 2010). Bacteriophage treatments have also been tested for aquaculture, with some success in the treatment and prevention of fish infections. These studies, using bacteriophages specific to Lactococcus garviaea and Pseudomonas plecoglossicida, pathogens of Yellowtail (Seriola quinqueradiata) and Ayu (Plecoglossus altivelis), respectively, suggest that bacteriophages could be useful for the fish industry (Almeida et al. 2009).
Recent advances in human phage therapy
A few years after the invention of phage therapy by d’Herelle, a Georgian researcher, Georgyi Eliava, worked at the Institut Pasteur in Paris for a few months as a visiting scientist. On his return to Georgia, he convinced the Soviet authorities to invest in phage therapy by building a dedicated research center. The resulting institute, the Eliava Institute for Bacteriophages, Microbiology, and Virology was created more than 80 years ago. This center has a long history in the domains of bacteriophage research and human phage therapy (Kutateladze and Adamia 2008).
A similar center at Wroclaw in Poland is also carrying out both human phage therapy and bacteriophage research (Gorski et al. 2009). These two centers have provided a clear demonstration that phage therapy can be useful for human patients, but there is not yet sufficient experimental proof to respond to the legitimate questions asked by public health authorities in Western countries. Nevertheless, in light of some recent work, the situation is clearly improving, paving the way for the promotion of additional research and clinical studies in this field.
Experimental models
Over the last 30 years, studies in several animal models have shown that bacteriophages can infect pathogenic bacteria in animals. Many reviews summarizing these data have been published (Gorski et al. 2009; O’Flaherty et al. 2009; Sulakvelidze and Kutter 2005). Bacteriophage treatments administered by intravenous or intraperitoneal injections have frequently been successful. However, these administration routes are the most difficult to use in humans, because it is not yet possible to ensure that bacteriophage solutions are absolutely pyrogen-free. There are two key aspects to be considered concerning the safety of bacteriophage preparations. Firstly, as mentioned above, no standard protocol has yet been validated by medical authorities. Secondly, as bacteriophages are very different in nature from currently authorized medical products, it may be necessary to adapt or to create regulations for full clinical assessment of the potential of these viruses (Verbeken et al. 2007).
As intravenous and intraperitoneal administrations are unlikely to be the first choices for the treatment of humans, some research groups have studied the administration of bacteriophages by more “natural” routes. The oral administration of bacteriophages to target intestinal pathogens is the most widely studied (Chibani-Chennoufi et al. 2004; Tanji et al. 2005). These studies have yielded promising results, but the only mouse model available in which a diarrheal disease (caused by Citrobacter rodentium) is reproduced has not yet been used to test the efficacy of phage therapy (Mundy et al. 2005). The lack of such data is currently making it difficult to study the consequences of bacteriophage treatments for both the host flora and the immune response. Skin infections can be treated topically by the direct application of bacteriophages to the surface of the skin, and formulations have been developed in which bacteriophages are incorporated into a cream (O’Flaherty et al. 2005). However, such modes of treatment have not yet been investigated in depth in animals and humans (Marza et al. 2006). Finally, the treatment of lung infections by the intranasal administration of bacteriophages in mice has been the object of two recent publications (Carmody et al. 2009; Debarbieux et al. 2010). Carmody et al. compared the intranasal and intraperitoneal treatments of infections caused by Burkholderia cenocepacia and suggested that intraperitoneal treatment was more effective. Studies by other groups have confirmed the efficacy of intraperitoneal treatment of lung infections caused by Pseudomonas aeruginosa or Klebsiella pneumoniae with bacteriophages (Chhibber et al. 2008; McVay et al. 2007). By contrast, Debarbieux et al. demonstrated the rapid efficacy of an intranasal instillation of a bacteriophage to treat an acute P. aeruginosa lung infection in mouse. In vivo imaging technology with a bioluminescent P. aeruginosa strain (expressing lux genes) was used to follow infection and treatment kinetics in live animals. A single intranasal application of one bacteriophage was found to be effective for the treatment of the lungs, nose, and throat. This result suggests that the nasal carriage of pathogens in healthy subjects could be reduced by the inhalation of appropriate bacteriophages. Furthermore, a single bacteriophage application was shown to be effective at preventing P. aeruginosa infection over a period of 24 h. This result was confirmed and extended, with another bacteriophage, to a period of 4 days of protection against an acute lung infection caused by a clinical strain of P. aeruginosa that was mucoid, multidrug-resistant, and isolated from a cystic fibrosis patient (Morello et al. 2011). The prophylactic use of bacteriophages is not a new idea. However, the results obtained for the intravenous injection of bacteriophages in animal models suggested a half-life too short for use in prophylaxis, unless long-circulating variants were selected (Uchiyama et al. 2009; Merril et al. 1996; 2003). If the results for lung treatment obtained in mice are confirmed in humans, then the incidence of lung infections, and particularly of those acquired in hospitals, could be reduced by preventive treatment over several days. Current monitoring procedures for hospital-acquired infections include identification of the principal pathogens involved. It should, therefore, theoretically, be possible to select bacteriophages effective against these pathogens for use in preventive treatments in patients due to attend the affected hospital or for direct treatment on admission.
Human trials
Only one human phase II trial has been carried out, in accordance with current European regulations (Wright et al. 2009). The patients involved in this study had developed chronic otitis caused by P. aeruginosa and previous treatments with antibiotics had been unsuccessful. The levels of bacteria recovered at the end of the treatment were significantly lower in the bacteriophage-treated group than in the untreated group. This treatment appeared slightly less effective than that in a previous study on dogs, and the results obtained were very different from those obtained in mice models, in which 100% of the animals could be cured (Hawkins et al. 2010). This is not surprising, because animal models only partly reflect the clinical situation of the human disease (Lecuit 2007). Animal models of infection are usually developed such that most of the animals die rapidly, making it possible to identify pathogens or virulence factors. There is now an urgent need to develop more clinically relevant models, particularly for chronic infections, for more careful evaluation of the efficacy of bacteriophage treatments.
A few phase I studies have also been reported (Bruttin and Brussow 2005; Kutter et al. 2010; Rhoads et al. 2009). These studies precede future phase II trials, as exemplified by the ongoing trial in Bangladesh focusing on the treatment of diarrheal infections in infants (clinical trial identifier NCT00937274). Future data from these trials, together with the data available from the centers in Poland and Georgia, should make it possible to determine both the potential and limitations of human phage therapy (Letkiewicz et al. 2010; Kutateladze and Adamia 2010).
Perspectives
After a long period of waning interest, the future of bacteriophage research is now looking up and, for the first time in a century, there seems to be a unique opportunity for both fundamental and applied scientists to join forces. We provide below some examples likely to benefit greatly from this renewed interest in bacteriophages.
Ecology
The ecological impact of the use of bacteriophages in industry and medicine has not been studied, so this area remains somewhat speculative. If the use of bacteriophages becomes widespread in medicine, there will probably be little impact on the total population of 1031 units versus 107 humans worldwide. As bacteriophages will be selected to be species-specific for human pathogens, they are likely to have smaller ecological consequences than broad-spectrum antibiotics (Barc et al. 2004; Denou et al. 2009). The use of high doses of bacteriophages, particularly in agriculture (directly on fields), may affect local ecological niches, but the extent of this impact remains to be determined. In aquatic environments, bacteriophage predation is such a widespread natural event that the emergence in a local area of a dominant population of specific bacteriophages is frequent and forms part of the natural cycle of co-evolution of bacteriophages and their hosts. Will the use of bacteriophages by the fish industry affect this evolution? Additional data are required to answer this question and many others.
The human gut shelters one of the most studied microbial communities. This community is regularly exposed to various bacteriophages found in food and water (Lodder et al. 2010; Garcia et al. 2009; Tsuei et al. 2007). A recent study on the viriome of human feces showed that most of the bacteriophages present are prophages with limited variability over time (Reyes et al. 2010). These data are clearly different from those obtained in aquatic environments, in which the dynamic renewal of bacterial populations has been shown to be driven mostly by virulent bacteriophages (Rodriguez-Valera et al. 2009). Has evolution of the gut microbial community resulted in the selection of a new type of defense against bacteriophage predation, keeping virulent bacteriophages under control? Future studies combining both microbiome and viriome analysis should shed more light on this particular ecological niche.
Technology
The possible technological applications of bacteriophages are so broad that we present here only few that are directly linked to the management of undesirable bacteria. Lysins, the bacteriophage-encoded enzymes implicated in the release of virion progeny, have been shown to cure animals infected with gram-positive pathogens (Fischetti 2008). These enzymes have opened the way to antimicrobial drug discovery through bacteriophage genomics. Engineered bacteriophages have been developed (1) to infect biofilms more efficiently; (2) to enhance the efficacy of antibiotics, and (3) to increase bacteriophage host range (Lu and Collins 2007, 2009; Pouillot et al. 2010). Recent advances in synthetic biology and the in vitro synthesis of large genomes in particular, have opened up new possibilities for designing “customized” bacteriophages (Burbelo et al. 2010). Further studies are required to determine whether it is likely to be possible to build a synthetic bacteriophage for a specific goal. Meanwhile, several biotechnology companies are currently selling bacteriophage-based products and the number of such companies will probably increase in the next few years with, for example, the development of bionanotechnologies (Housby and Mann 2009; Hemminga et al. 2010; Soto and Ratna 2010).
Medicine
The concept of “chemical medicine”, which is based on drugs, does not apply to phage therapy. Bacteriophages are much more complex than chemically defined molecules, but this complexity is actually an advantage when considering their specificity. They target only permissive hosts, which constitute only a small proportion of the bacterial populations they encounter, even in humans. As phage therapy is expected to have no adverse effects on the commensal flora, it is thought that phage therapy would result in a smaller number of secondary infections than antibiotic treatments (Croswell et al. 2009). Nevertheless, antibiotics are still routinely used in hospitals and will continue to be so in the near future, because they are reliably effective. The discovery of new drugs can also breathe new life into efforts to combat drug-resistant bacteria. Antibiotics and bacteriophages should therefore not be seen as conflicting and exclusive approaches. Their combined use is supported by the synergy between bacteriophages and antibiotics (Comeau et al. 2007). The development of such combination treatments requires more research, but seems promising.
One of the major obstacles to the development of phage therapy in humans is the lack of a specific regulatory framework. However, ways to adapt phage therapy to current regulations have been suggested (Verbeken et al. 2007; Pirnay et al. 2010).
Assuming that the use of bacteriophages in medicine is eventually accepted, how could this approach be applied? In cases of acute infection, the timing of treatment is crucial and bacteriophages are unlikely to be of great benefit unless the pathogen responsible for the infection has already been identified. By contrast, timing is less crucial in chronic infections, and phage therapy would therefore be more likely to succeed, as time could be devoted to selecting the most effective bacteriophages. However, such selection should ideally be carried out on a collection of bacteriophages approved for use in medicine (bacteriophages fully sequenced and prepared according to established standard procedures).
Clearly the concept of the biocontrol of undesirable bacteria by bacteriophages is now well established, but the extensive use of bacteriophages in the twenty-first century in the domains of human health and food requires additional research and appropriate evaluation in clinical trials.
References
Abedon ST (2009) Phage evolution and ecology. Adv Appl Microbiol 67:1–45
Almeida A, Cunha A, Gomes NC, Alves E, Costa L, Faustino MA (2009) Phage therapy and photodynamic therapy: low environmental impact approaches to inactivate microorganisms in fish farming plants. Mar Drugs 7(3):268–313
Balogh B, Jones JB, Iriarte FB, Momol MT (2010) Phage therapy for plant disease control. Curr Pharm Biotechnol 11(1):48–57
Barc MC, Bourlioux F, Rigottier-Gois L, Charrin-Sarnel C, Janoir C, Boureau H, Dore J, Collignon A (2004) Effect of amoxicillin–clavulanic acid on human fecal flora in a gnotobiotic mouse model assessed with fluorescence hybridization using group-specific 16 S rRNA probes in combination with flow cytometry. Antimicrob Agents Chemother 48(4):1365–1368
Barrangou R, Fremaux C, Deveau H, Richards M, Boyaval P, Moineau S, Romero DA, Horvath P (2007) CRISPR provides acquired resistance against viruses in prokaryotes. Science (New York, NY) 315(5819):1709–1712
Bergh O, Borsheim KY, Bratbak G, Heldal M (1989) High abundance of viruses found in aquatic environments. Nature 340(6233):467–468
Bratkovic T (2010) Progress in phage display: evolution of the technique and its application. Cell Mol Life Sci 67(5):749–767
Brussow H (2001) Phages of dairy bacteria. Annu Rev Microbiol 55:283–303
Bruttin A, Brussow H (2005) Human volunteers receiving Escherichia coli phage T4 orally: a safety test of phage therapy. Antimicrob Agents Chemother 49(7):2874–2878
Burbelo PD, Ching KH, Han BL, Klimavicz CM, Iadarola MJ (2010) Synthetic biology for translational research. Am J Transl Res 2(4):381–389
Callaway TR, Edrington TS, Brabban AD, Keen JE, Anderson RC, Rossman ML, Engler MJ, Genovese KJ, Gwartney BL, Reagan JO, Poole TL, Harvey RB, Kutter EM, Nisbet DJ (2006) Fecal prevalence of Escherichia coli O157, Salmonella, Listeria, and bacteriophage infecting E. coli O157:H7 in feedlot cattle in the Southern Plains region of the United States. Foodborne Pathog Dis 3(3):234–244
Carlton RM, Noordman WH, Biswas B, de Meester ED, Loessner MJ (2005) Bacteriophage P100 for control of Listeria monocytogenes in foods: genome sequence, bioinformatic analyses, oral toxicity study, and application. Regul Toxicol Pharmacol 43(3):301–312
Carmody LA, Gill JJ, Summer EJ, Sajjan US, Gonzalez CF, Young RF, LiPuma JJ (2009) Efficacy of bacteriophage therapy in a model of Burkholderia cenocepacia pulmonary infection. J Infect Dis 201(2):264–271
Chhibber S, Kaur S, Kumari S (2008) Therapeutic potential of bacteriophage in treating Klebsiella pneumoniae B5055-mediated lobar pneumonia in mice. J Med Microbiol 57(Pt 12):1508–1513
Chibani-Chennoufi S, Sidoti J, Bruttin A, Kutter E, Sarker S, Brussow H (2004) In vitro and in vivo bacteriolytic activities of Escherichia coli phages: implications for phage therapy. Antimicrob Agents Chemother 48(7):2558–2569
Comeau AM, Tetart F, Trojet SN, Prere MF, Krisch HM (2007) Phage-Antibiotic Synergy (PAS): beta-lactam and quinolone antibiotics stimulate virulent phage growth. PLoS ONE 2(8):e799
Courchesne NM, Parisien A, Lan CQ (2009) Production and application of bacteriophage and bacteriophage-encoded lysins. Recent Pat Biotechnol 3(1):37–45
Croswell A, Amir E, Teggatz P, Barman M, Salzman NH (2009) Prolonged impact of antibiotics on intestinal microbial ecology and susceptibility to enteric Salmonella infection. Infect Immun 77(7):2741–2753
Debarbieux L, Leduc D, Maura D, Morello E, Criscuolo A, Grossi O, Balloy V, Touqui L (2010) Bacteriophages can treat and prevent Pseudomonas aeruginosa lung infections. J Infect Dis 201(7):1096–1104
Denou E, Bruttin A, Barretto C, Ngom-Bru C, Brussow H, Zuber S (2009) T4 phages against Escherichia coli diarrhea: potential and problems. Virology 388(1):21–30
d’Herelle F (1917) Sur un microbe invisible antagoniste des bacilles dysenteriques. Comptes Rendus Acad Sci Paris 165:373–375
Faruque SM, Islam MJ, Ahmad QS, Faruque AS, Sack DA, Nair GB, Mekalanos JJ (2005) Self-limiting nature of seasonal cholera epidemics: role of host-mediated amplification of phage. Proc Natl Acad Sci USA 102(17):6119–6124
Fischetti VA (2008) Bacteriophage lysins as effective antibacterials. Curr Opin Microbiol 11(5):393–400
Forterre P (2010) Defining life: the virus viewpoint. Orig Life Evol Biosph 40(2):151–160
Forterre P, Prangishvili D (2009) The origin of viruses. Res Microbiol 160(7):466–472
Fuhrman JA (1999) Marine viruses and their biogeochemical and ecological effects. Nature 399(6736):541–548
Garcia P, Madera C, Martinez B, Rodriguez A, Evaristo Suarez J (2009) Prevalence of bacteriophages infecting Staphylococcus aureus in dairy samples and their potential as biocontrol agents. J Dairy Sci 92(7):3019–3026
Gill JJ, Hyman P (2010) Phage choice, isolation, and preparation for phage therapy. Curr Pharm Biotechnol 11(1):2–14
Goodridge LD (2010) Designing phage therapeutics. Curr Pharm Biotechnol 11(1):15–27
Gorski A, Miedzybrodzki R, Borysowski J, Weber-Dabrowska B, Lobocka M, Fortuna W, Letkiewicz S, Zimecki M, Filby G (2009) Bacteriophage therapy for the treatment of infections. Curr Opin Investig Drugs 10(8):766–774
Guenther S, Huwyler D, Richard S, Loessner MJ (2009) Virulent bacteriophage for efficient biocontrol of Listeria monocytogenes in ready-to-eat foods. Appl Environ Microbiol 75(1):93–100
Hankin ME (1896) L’action bactericide des eaux de la Jumna et du Gange sur le microbe du cholera. Ann Inst Pasteur 10:511–523
Hawkins C, Harper D, Burch D, Anggard E, Soothill J (2010) Topical treatment of Pseudomonas aeruginosa otitis of dogs with a bacteriophage mixture: a before/after clinical trial. Vet Microbiol 146(3–4):309–313
Hemminga MA, Vos WL, Nazarov PV, Koehorst RB, Wolfs CJ, Spruijt RB, Stopar D (2010) Viruses: incredible nanomachines. New advances with filamentous phages. Eur Biophys J 39(4):541–550
Housby JN, Mann NH (2009) Phage therapy. Drug Discov Today 14(11–12):536–540
Jamalludeen N, Johnson RP, Shewen PE, Gyles CL (2009) Evaluation of bacteriophages for prevention and treatment of diarrhea due to experimental enterotoxigenic Escherichia coli O149 infection of pigs. Vet Microbiol 136(1–2):135–141
Johnson RP, Gyles CL, Huff WE, Ojha S, Huff GR, Rath NC, Donoghue AM (2008) Bacteriophages for prophylaxis and therapy in cattle, poultry and pigs. Animal Health Research Reviews/Conference of Research Workers in Animal Diseases 9(2):201–215
Kramberger P, Honour RC, Herman RE, Smrekar F, Peterka M (2010) Purification of the Staphylococcus aureus bacteriophages VDX-10 on methacrylate monoliths. J Virol Methods 166(1–2):60–64
Krupovic M, Forterre P, Bamford DH (2010) Comparative analysis of the mosaic genomes of tailed archaeal viruses and proviruses suggests common themes for virion architecture and assembly with tailed viruses of bacteria. J Mol Biol 397(1):144–160
Kumarasamy KK, Toleman MA, Walsh TR, Bagaria J, Butt F, Balakrishnan R, Chaudhary U, Doumith M, Giske CG, Irfan S, Krishnan P, Kumar AV, Maharjan S, Mushtaq S, Noorie T, Paterson DL, Pearson A, Perry C, Pike R, Rao B, Ray U, Sarma JB, Sharma M, Sheridan E, Thirunarayan MA, Turton J, Upadhyay S, Warner M, Welfare W, Livermore DM, Woodford N (2010) Emergence of a new antibiotic resistance mechanism in India, Pakistan, and the UK: a molecular, biological, and epidemiological study. Lancet Infect Dis 10(9):597–602
Kutateladze M, Adamia R (2008) Phage therapy experience at the Eliava Institute. Méd Mal Infect 38(8):426–430
Kutateladze M, Adamia R (2010) Bacteriophages as potential new therapeutics to replace or supplement antibiotics. Trends Biotechnol 28(12):591–595
Kutter E, De Vos D, Gvasalia G, Alavidze Z, Gogokhia L, Kuhl S, Abedon ST (2010) Phage therapy in clinical practice: treatment of human infections. Curr Pharm Biotechnol 11(1):69–86
Labrie SJ, Samson JE, Moineau S (2010) Bacteriophage resistance mechanisms. Nat Rev 8(5):317–327
Lecuit M (2007) Human listeriosis and animal models. Microbes Infect 9(10):1216–1225
Letkiewicz S, Miedzybrodzki R, Klak M, Jonczyk E, Weber-Dabrowska B, Gorski A (2010) The perspectives of the application of phage therapy in chronic bacterial prostatitis. FEMS Immunol Med Microbiol 60(2):99–112
Loc Carrillo C, Atterbury RJ, el-Shibiny A, Connerton PL, Dillon E, Scott A, Connerton IF (2005) Bacteriophage therapy to reduce Campylobacter jejuni colonization of broiler chickens. Appl Environ Microbiol 71(11):6554–6563
Lodder WJ, van den Berg HH, Rutjes SA, de Roda Husman AM (2010) Presence of enteric viruses in source waters for drinking water production in The Netherlands. Appl Environ Microbiol 76(17):5965–5971
Lu TK, Collins JJ (2007) Dispersing biofilms with engineered enzymatic bacteriophage. Proc Natl Acad Sci USA 104(27):11197–11202
Lu TK, Collins JJ (2009) Engineered bacteriophage targeting gene networks as adjuvants for antibiotic therapy. Proc Natl Acad Sci USA 106(12):4629–4634
Lwoff A (1967) Principles of classification and nomenclature of viruses. Nature 215(5096):13–14
Marza JA, Soothill JS, Boydell P, Collyns TA (2006) Multiplication of therapeutically administered bacteriophages in Pseudomonas aeruginosa infected patients. Burns 32(5):644–646
McVay CS, Velasquez M, Fralick JA (2007) Phage therapy of Pseudomonas aeruginosa infection in a mouse burn wound model. Antimicrob Agents Chemother 51(6):1934–1938
Merabishvili M, Pirnay JP, Verbeken G, Chanishvili N, Tediashvili M, Lashkhi N, Glonti T, Krylov V, Mast J, Van Parys L, Lavigne R, Volckaert G, Mattheus W, Verween G, De Corte P, Rose T, Jennes S, Zizi M, De Vos D, Vaneechoutte M (2009) Quality-controlled small-scale production of a well-defined bacteriophage cocktail for use in human clinical trials. PLoS ONE 4(3):e4944
Merril CR, Biswas B, Carlton R, Jensen NC, Creed GJ, Zullo S, Adhya S (1996) Long-circulating bacteriophage as antibacterial agents. Proc Natl Acad Sci USA 93(8):3188–3192
Merril CR, Scholl D, Adhya SL (2003) The prospect for bacteriophage therapy in Western medicine. Nat Rev Drug Discov 2(6):489–497
Morello E, Saussereau E, Maura D, Huerre M, Touqui L, Debarbieux L (2011) Pulmonary bacteriophage therapy on Pseudomonas aeruginosa cystic fibrosis strains: first steps towards treatment and prevention. PLoS ONE 6(2):e16963
Mundy R, MacDonald TT, Dougan G, Frankel G, Wiles S (2005) Citrobacter rodentium of mice and man. Cell Microbiol 7(12):1697–1706
Nordmann P, Naas T, Fortineau N, Poirel L (2007) Superbugs in the coming new decade; multidrug resistance and prospects for treatment of Staphylococcus aureus, Enterococcus spp. and Pseudomonas aeruginosa in 2010. Curr Opin Microbiol 10(5):436–440
O’Flaherty S, Ross RP, Meaney W, Fitzgerald GF, Elbreki MF, Coffey A (2005) Potential of the polyvalent anti-Staphylococcus bacteriophage K for control of antibiotic-resistant staphylococci from hospitals. Appl Environ Microbiol 71(4):1836–1842
O’Flaherty S, Ross RP, Coffey A (2009) Bacteriophage and their lysins for elimination of infectious bacteria. FEMS Microbiol Rev 33(4):801–819
Oliveira A, Sereno R, Azeredo J (2010) In vivo efficiency evaluation of a phage cocktail in controlling severe colibacillosis in confined conditions and experimental poultry houses. Vet Microbiol 146(3–4):303–308
Pal C, Macia MD, Oliver A, Schachar I, Buckling A (2007) Coevolution with viruses drives the evolution of bacterial mutation rates. Nature 450(7172):1079–1081
Pantucek R, Rosypalova A, Doskar J, Kailerova J, Ruzickova V, Borecka P, Snopkova S, Horvath R, Gotz F, Rosypal S (1998) The polyvalent staphylococcal phage phi 812: its host-range mutants and related phages. Virology 246(2):241–252
Paterson S, Vogwill T, Buckling A, Benmayor R, Spiers AJ, Thomson NR, Quail M, Smith F, Walker D, Libberton B, Fenton A, Hall N, Brockhurst MA (2010) Antagonistic coevolution accelerates molecular evolution. Nature 464(7286):275–278
Pirnay JP, De Vos D, Verbeken G, Merabishvili M, Chanishvili N, Vaneechoutte M, Zizi M, Laire G, Lavigne R, Huys I, Van den Mooter G, Buckling A, Debarbieux L, Pouillot F, Azeredo J, Kutter E, Dublanchet A, Gorski A, Adamia R (2010) The phage therapy paradigm: pret-a-porter or sur-mesure? Pharm Res 1–4. doi:https://doi.org/10.1007/s11095-010-0313-5
Pouillot F, Blois H, Iris F (2010) Genetically engineered virulent phage banks in the detection and control of emergent pathogenic bacteria. Biosecur Bioterror 8(2):155–169
Reyes A, Haynes M, Hanson N, Angly FE, Heath AC, Rohwer F, Gordon JI (2010) Viruses in the faecal microbiota of monozygotic twins and their mothers. Nature 466(7304):334–338
Rhoads DD, Wolcott RD, Kuskowski MA, Wolcott BM, Ward LS, Sulakvelidze A (2009) Bacteriophage therapy of venous leg ulcers in humans: results of a Phase I safety trial. J Wound Care 18(6):237–238, 240–233
Rivas L, Coffey B, McAuliffe O, McDonnell MJ, Burgess CM, Coffey A, Ross RP, Duffy G (2010) In vivo and ex vivo evaluations of bacteriophages e11/2 and e4/1c for use in the control of Escherichia coli O157:H7. Appl Environ Microbiol 76(21):7210–7216
Rodriguez-Brito B, Li L, Wegley L, Furlan M, Angly F, Breitbart M, Buchanan J, Desnues C, Dinsdale E, Edwards R, Felts B, Haynes M, Liu H, Lipson D, Mahaffy J, Martin-Cuadrado AB, Mira A, Nulton J, Pasic L, Rayhawk S, Rodriguez-Mueller J, Rodriguez-Valera F, Salamon P, Srinagesh S, Thingstad TF, Tran T, Thurber RV, Willner D, Youle M, Rohwer F (2010) Viral and microbial community dynamics in four aquatic environments. ISME J 4(6):739–751
Rodriguez-Valera F, Martin-Cuadrado AB, Rodriguez-Brito B, Pasic L, Thingstad TF, Rohwer F, Mira A (2009) Explaining microbial population genomics through phage predation. Nat Rev 7(11):828–836
Rohwer F, Prangishvili D, Lindell D (2009) Roles of viruses in the environment. Environ Microbiol 11(11):2771–2774
Rozema EA, Stephens TP, Bach SJ, Okine EK, Johnson RP, Stanford K, McAllister TA (2009) Oral and rectal administration of bacteriophages for control of Escherichia coli O157:H7 in feedlot cattle. J Food Prot 72(2):241–250
Schoenfeld T, Liles M, Wommack KE, Polson SW, Godiska R, Mead D (2010) Functional viral metagenomics and the next generation of molecular tools. Trends Microbiol 18(1):20–29
Shapiro J, Machattie L, Eron L, Ihler G, Ippen K, Beckwith J (1969) Isolation of pure lac operon DNA. Nature 224(5221):768–774
Soto CM, Ratna BR (2010) Virus hybrids as nanomaterials for biotechnology. Curr Opin Biotechnol 21(4):426–438
Sulakvelidze A, Kutter E (2005) Bacteriophage therapy in humans. In: Kutter E, Sulakvelidze A (eds) Bacteriophages: biology and applications. CRC, Boca Raton, pp 381–436
Sulakvelidze A, Alavidze Z, Morris JG Jr (2001) Bacteriophage therapy. Antimicrob Agents Chemother 45(3):649–659
Summers WC (1999) Félix d’Herelle and the origins of molecular biology. Yale University Press, New Haven
Summers WC (2001) Bacteriophage therapy. Annu Rev Microbiol 55:437–451
Suttle CA (2007) Marine viruses—major players in the global ecosystem. Nat Rev 5(10):801–812
Tanji Y, Shimada T, Yoichi M, Miyanaga K, Hori K, Unno H (2004) Toward rational control of Escherichia coli O157:H7 by a phage cocktail. Appl Microbiol Biotechnol 64(2):270–274
Tanji Y, Shimada T, Fukudomi H, Miyanaga K, Nakai Y, Unno H (2005) Therapeutic use of phage cocktail for controlling Escherichia coli O157:H7 in gastrointestinal tract of mice. J Biosci Bioeng 100(3):280–287
Taubes G (2008) The bacteria fight back. Science 321(5887):356–361
Thurber RV (2009) Current insights into phage biodiversity and biogeography. Curr Opin Microbiol 12(5):582–587
Tsuei AC, Carey-Smith GV, Hudson JA, Billington C, Heinemann JA (2007) Prevalence and numbers of coliphages and Campylobacter jejuni bacteriophages in New Zealand foods. Int J Food Microbiol 116(1):121–125
Twort F (1915) An investigation on the nature of ultramicroscopic viruses. Lancet 186(4814):1241–1243
Uchiyama J, Maeda Y, Takemura I, Chess-Williams R, Wakiguchi H, Matsuzaki S (2009) Blood kinetics of four intraperitoneally administered therapeutic candidate bacteriophages in healthy and neutropenic mice. Microbiol Immunol 53(5):301–304
Verbeken G, De Vos D, Vaneechoutte M, Merabishvili M, Zizi M, Pirnay JP (2007) European regulatory conundrum of phage therapy. Future Microbiol 2(5):485–491
Wall SK, Zhang J, Rostagno MH, Ebner PD (2010) Phage therapy to reduce preprocessing Salmonella infections in market-weight swine. Appl Environ Microbiol 76(1):48–53
Wommack KE, Colwell RR (2000) Virioplankton: viruses in aquatic ecosystems. Microbiol Mol Biol Rev 64(1):69–114
Wright A, Hawkins CH, Anggard EE, Harper DR (2009) A controlled clinical trial of a therapeutic bacteriophage preparation in chronic otitis due to antibiotic-resistant Pseudomonas aeruginosa; a preliminary report of efficacy. Clin Otolaryngol 34(4):349–357
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Maura, D., Debarbieux, L. Bacteriophages as twenty-first century antibacterial tools for food and medicine. Appl Microbiol Biotechnol 90, 851–859 (2011). https://doi.org/10.1007/s00253-011-3227-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-011-3227-1