Abstract
l-2-Aminobutyric acid can be synthesized in a transamination reaction from l-threonine and l-aspartic acid as substrates by the action of threonine deaminase and aromatic aminotransferase, but the by-product l-alanine was produced simultaneously. A small amount of l-alanine increased the complexity of the l-2-aminobutyric acid recovery process because of their extreme similarity in physical and chemical properties. Acetolactate synthase has been introduced to remove the pyruvate intermediate for reducing the l-alanine concentration partially. To eliminate the remnant l-alanine, alanine racemase of Bacillus subtilis in combination with d-amino acid oxidase of Rhodotorula gracilis or Trigonopsis variabilis respectively was introduced into the reaction system for the l-2-aminobutyric acid synthesis. l-Alanine could be completely removed by the action of alanine racemase of B. subtilis and d-amino acid oxidase of R. gracilis; thereby, high-purity l-2-aminobutyric acid was achieved. The results revealed that alanine racemase could discriminate effectively between l-alanine and l-2-aminobutyric acid, and selectively catalyzed l-alanine to d-alanine reversibly. d-Amino acid oxidase then catalyzed d-alanine to pyruvate stereoselectively. Furthermore, this method was also successfully used to remove the by-product l-alanine in the production of other neutral amino acids such as l-tertiary leucine and l-valine, suggesting that multienzymatic whole-cell catalysis can be employed to provide high purity products.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
l-2-Aminobutyric acid (l-ABA), an unnatural amino acid, has been used as an important precursor for the synthesis of many chiral drugs such as antiepileptic levetiracetam, brivaracetam, and antituberculotic ethambutol. l-ABA can be synthesized in a transamination reaction from 2-oxobutyric acid and l-aspartic acid as substrates using aromatic aminotransferase (AAR; Fotheringham et al. 1999; Fig. 1) or produced from 2-oxobutyric acid and benzylamine using ω-aminotransferase (Shin and Kim 2009), and it could also be produced from the reduction of α-keto acids with leucine dehydrogenase (Krix et al. 1997) or glutamate dehydrogenase (Zhang et al. 2010). Besides the biosynthetic methods described above, l-ABA has been prepared by deracemization of d/l-ABA using chemical resolution or metal catalyst and whole cell d-amino acid oxidase (DAAO; Fotheringham et al. 2006). With broad substrate specificity and no requirement for external cofactor regeneration, bacterial AARs were usually applied in the large-scale biosynthesis of enantiomercally pure unnatural amino acids (Cho et al. 2003; Hwang et al. 2009; Lo et al. 2005). However, in the production of l-ABA using AAR, the by-product l-alanine (l-Ala) was produced in the transamination reaction from pyruvate as substrate (Fotheringham et al. 1999; Fig. 1). Acetolactate synthase has been introduced to remove the pyruvate intermediate and partially reduced the l-Ala concentration. l-Ala shares extreme similarity with l-ABA in physical and chemical properties, so it was intricate to eliminate the by-product l-Ala with the present methods for isolation and recovery of l-ABA.
Biosynthesis of l-ABA using recombinant E. coli cells. Threonine deaminase (ilvA) and aromatic aminotransferase (tyrB) are obtained from E. coli, acetolactate synthase (alsS) and alanine racemase (Guillouet et al. 1999) from B. subtilis, and d-amino acid oxidase (daao) from R. gracilis or T. variabilis
To solve this problem, we try to find an enzyme which can effectively discriminate between l-Ala and l-ABA. As we know, there are six types of enzymes involved in the reaction with l-Ala, including alanine dehydrogenase, transaminase, 8-amino-7-oxononanoate synthase, alanine racemase, N-acetylmuramoyl-l-alanine amidase, and alanine-tRNA ligase. The former two enzymes have broad substrate specificity and can react with both l-Ala and l-ABA (Schroder et al. 2004; Han and Li 2002). The third one, 8-amino-7-oxononanoate synthase, only catalyzes d/l-Ala and 6-carboxyhexanoyl to 8-amino-7-oxononanoate (Bhor et al. 2006). The last two enzymes are specifically involved in the biosynthesis of cell wall (Pennartz et al. 2009) and peptides (Pleiss et al. 2000), and not used as catalysts. In contrast, the fourth type, alanine racemase (EC 5.1.1.1, ALR), could catalyze the interconversion between l-Ala and d-Ala with high specificity (Ju et al. 2005; Johnston et al. 1984). It was also reported that ALR of Schizosaccharomyces pombe (Uo et al. 2001) and Aquifex pyrophilus (Kim and Yu 2000) had lower activity on l-ABA than that on l-Ala (0.37% and 18% of l-Ala, respectively), while the same property was not reported on ALR of Bacillus subtilis (BsALR; Johnston et al. 1984). So in this study, BsALR was employed to catalyze l-Ala to d-Ala, which was further converted to pyruvate by d-amino acid oxidase (EC 1.4.3.3, DAAO; Fig. 1). Concerning DAAO, three available DAAO from Rhodotorula gracilis (RgDAAO), Trigonopsis variabilis (TvDAAO), and pig kidney were well known (Pollegioni et al. 2004), but enzymes from microorganisms appeared to be by far more suitable catalysts for bioconversion (Pollegioni et al. 2008). The ability of eliminating l-Ala from the reaction mixtures of l-ABA biosynthesis using BsALR in combination with RgDAAO and TvDAAO was described. We also attempted to apply this method to eliminate the by-product l-Ala in the producing process of other neutral amino acids.
Materials and methods
Materials
Restriction endonuclease and T4 DNA ligase were purchased from New England Biolabs, Inc. (Beijing, China), and Kod/plus DNA polymerase was from Toyobo (Shanghai, China). The bacterial strains and plasmids used in this study are listed in Table 1.
Plasmid construction
DNA manipulations and cloning steps were carried out according to standard protocols (Sambrook et al. 1989). Plasmid constructs were verified by DNA sequencing. Cloning primers used in the construction of plasmids are shown in Table 2.
The ilvA gene (Gene ID: 948287) and tyrB gene (Gene ID: 948563) were amplified from Escherichia coli genomic DNA and individually cloned into the NcoI/BamHI sites of pET28b to construct pET-ilvA and pET-tyrB. The alsS gene (Gene ID: 936852) and dal gene (Gene ID: 9721445) were amplified from the chromosome of B. subtilis and respectively inserted into the NcoI/BamHI sites of pET28b and the NdeI/BamHI sites of pET24a to create pET-alsS and pETALR. The Rg-daao gene was amplified from plasmid pRset-RgDAAO (Kim and Khang 1995) and then inserted into the NdeI/HindIII sites of pET28b to construct pETRD. The pMSS (Zheng et al. 2006) was digested with NcoI/EcoRI to have the Tv-daao gene, and ligated with the fragment resulting from similar cleavage of pET28b to construct pETW2. The resulting plasmids were used to transform chemically competent BL21 (DE3) E. coli cells, and all of genes were expressed under control of the T7 promoter.
Preparation of whole cell biocatalysts
The strain BL21 (DE3) containing pET-ilvA, pET-tyrB, pET-alsS, pETALR, pETRD, and pETW2 respectively were grown overnight with agitation 200 rpm at 37°C in 5 ml of Luria–Bertani (LB) medium plus kanamycin (50 μg/ml) in a 50-ml flask. These cultures were then diluted into 100 ml of Teriffic broth (TB) to an OD600 of 0.05 and grown at 37°C in a 1,000-ml flask with agitation 200 rpm to an OD600 of 0.5. Then, the cultures were induced with 1 mM IPTG and incubated for a further 4 h at 30°C. The cells were then recovered by centrifugation at 10,000×g for 15 min, washed in 50 mM Tris–HCl buffer (pH 7.5) and similarly pelleted. The cell pellets were frozen for use in the biotransformation.
Determination of enzyme activity
The cell pellets were suspended in lysis buffer (100 mM Tris–HCl, pH 7.5), and lysed using sonication. The extracts were clarified by centrifugation (120,000×g), and the resulting supernatants were subjected to assay enzyme activity. IlvA activity was measured by determining the diminishment of threonine. The reaction mixture (2 ml) contained 400 mM threonine, 100 mM Tris–HCl (pH 7.5), and an appropriate amount of the crude extracts. Reactions were carried out at 37°C for 10 min and then terminated by adding 2 ml 10% HCl. The amounts of threonine were determined by HPLC. One unit of IlvA corresponds to 1 μmol threonine decreased per min at 37°C.
TyrB activity assay was carried out at 37°C in 100 mM Tris–HCl (pH 7.5). One unit of the TyrB was defined as the amount of enzyme necessary to catalyze the formation of 1 μM l-ABA from 500 mM l-aspartic acid and 500 mM 2-oxobutyric acid per minute.
AlsS activity was measured by monitoring the diminishment of sodium pyruvate. The assay was carried out in 100 mM Tris–HCl (pH 7.5) supplemented with 200 mM sodium pyruvate and the crude extracts in a final volume of 2 ml. After incubation for 10 min at 37°C, the reaction was stopped by adding 2 ml 10% HCl. The sodium pyruvate was measured by HPLC, and one unit of AlsS was defined as the amount of enzyme necessary to achieve 1 μM diminishment of sodium pyruvate per minute.
The activity of BsALR was determined by measuring the amounts of d- and l-alanine by HPLC (Ju et al. 2005). The reaction mixture (2 ml) was composed of 100 mM Tris–HCl (pH 7.5), 200 mM l-Ala, and an appropriate amount of the crude extracts. The reaction was started by the addition of the crude extracts, continued for 10 min at 37°C, and then terminated by adding 2 ml 10% HCl. One unit of BsALR was defined as the quantity of enzyme that catalyzed the formation of 1 μM of d-alanine from l-Ala per minute.
The activity of DAAO was measured as follows: The reaction mixture contained 100 mM d-Ala, 100 mM Tris–HCl (pH 7.5), 80,000 IU catalase, and an appropriate amount of enzyme in a final volume of 198 ml. After incubation at 37°C for 20 min with continuous supply of 30% H2O2 at rate of 0.1 ml/min, 1-ml reaction mixture was taken and terminated by adding 1 ml 10% HCl, and the d-Ala was determined by HPLC. One unit of DAAO corresponds to 1 μM d-Ala decreased per minute at 37°C.
Biosynthesis of l-ABA
The 0.35 M l-aspartic acid and 0.45 M l-threonine dissolved in a 500-ml solution containing 50 mM Tris–HCl (pH 7.5), 2.4 mM MnSO4·5H2O, and 0.8 mM MgSO4·7H2O (adjusted to pH 7.4 with NaOH) were prepared in a 2,000-ml flask. The wet cells of 1 g/l BL21 (DE3)/pET-ilvA, 3 g/l BL21 (DE3)/pET-tyrB, and 2 g/l BL21 (DE3)/pET-alsS were resuspended in the freshly prepared solution, and the mixture was incubated at 40°C with agitation at 300 rpm for about 24 h until l-aspartic acid was completely consumed. The sample of the first reaction stage was taken, and the cells were removed by centrifugation. The resulting supernatant was diluted 100-fold and subjected to analysis by HPLC.
Elimination of l-Ala from the l-ABA production solution
The cell pellets of 0.1 g/l BL21 (DE3)/pETALR (BsALR), 5 g/l BL21 (DE3)/pETRD (RgDAAO), or 7 g/l BL21 (DE3)/pETW2 (TvDAAO) were suspended in the l-ABA production solution described above and supplemented with 5,000 U/l catalase. The mixture was adjusted to pH 8.0 with NaOH and incubated at 37°C with continuous supply of 20% H2O2 at rate of 0.5 ml/min and agitation at 300 rpm. Samples were drawn regularly and centrifuged to remove the cells. The second transformed broth was diluted 100-fold and subjected to analysis by HPLC.
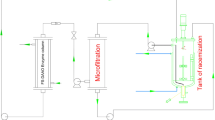
The reaction was also carried out in a reactor. Pure oxygen was supplied continuously at 30 ml/min with an agitation at 100 rpm, and the reactor pressure was 0.05 MPa. Samples were drawn regularly and centrifuged to remove the cells. The supernatant was analyzed by HPLC following the procedure described above.
Large-scale preparation of l-ABA using five enzymes in whole cell
For large-scale study, a reaction (1,500 l) was carried out in a 2,000-l jar fermentor. The cultivation mixture containing 0.35 M l-aspartic acid, 0.45 M l-threonine, 50 mM Tris–HCl (pH 7.5), 2.4 mM MnSO4·5H2O, 0.8 mM MgSO4·7H2O, 1 g/l BL21 (DE3)/pET-ilvA, 3 g/l BL21 (DE3)/pET-tyrB, and 2 g/l BL21 (DE3)/pET-alsS (adjusted to pH 7.4 with NH3·H2O and 30% acetic acid) was incubated at 40°C with agitation at 600 rpm for 24 h to synthesize l-ABA. Then, 5,000 U/l catalase and cell pellets of 0.1 g/l BL21 (DE3)/pETALR (BsALR) and 5 g/l BL21 (DE3)/pETRD (RgDAAO) were added, and the resultant mixture (adjusted to pH 8.0 with NH3·H2O and 30% acetic acid) was incubated at 37°C with continuous supply of 20% H2O2 at rate of 0.5 ml/min for 8 h. The supernatant was analyzed by HPLC following the procedure described above.
Isolation and purification of l-ABA
l-Ala shares extreme similarity with l-ABA in physical and chemical properties: they both dissolve in water (l-Ala 127.3 g/l, l-ABA 210.5 g/l at 25°C) and sparingly dissolve in ethanol (l-Ala 0.2% in cold 80% ethanol, l-ABA 1.8 g/l in boiling ethanol). The isolation and purification of l-ABA were described as follows: The reaction mixture was heated to 70°C, and then 1% activated charcoal and 2% diatomite were added. The precipitation was removed by filtration with a frame filter and an ultrafilter with a 5,000 molecular weight cut-off, and the l-ABA product was concentrated to 150 g/l and purified one more time. The l-ABA product was further concentrated to 450 g/l, and two volumes of ethanol were then added for crystallization.
Elimination of l-Ala from the l-tertiary leucine and l-valine solution
The cell pellets of 0.1 g/l BL21 (DE3)/pETALR (BsALR) and 5 g/l BL21 (DE3)/pETRD (RgDAAO) were added into a 100-ml solution containing 50 mM Tris–HCl (pH 7.5), 5,000 U/l catalase, 30 g/l l-tertiary leucine (L-Terleu), and 6 g/l l-Ala or 30 g/l l-valine (l-Val) and 6 g/l l-Ala. The mixture was adjusted to pH 8.0 with NaOH and incubated at 37°C with continuous supply of 20% H2O2 at rate of 0.5 ml/min and agitation at 300 rpm for 8 h. Samples were drawn and centrifuged to remove the cells. The samples were then diluted 100-fold and subjected to analysis by HPLC.
HPLC analysis of amino acids
The amino acids were derivatized with OPA/BOC-Cys and determined by HPLC using a LC-18DB column (5 μm, 4.6 × 250 mm, Agilent) at 338 nm with a flow rate of 1.0 ml/min. The gradient elution profile was as follows: of 100% A at 0–17 min, 40% A and 60% B at 17–18.1 min, 100% B at 18.1–25 min (A = 0.02 M sodium acetate, with triethylamine 180 μl/l and 3 ml/l tetrahydrofuran, pH 7.2; B = 0.02 M sodium acetate:acetonitrile:MeOH = 1:2:2 (v/v/v), pH 7.2).
Results
Expression of cloned genes
Plasmids pET-ilvA, pET-tyrB, pET-alsS, pETALR, pETW2, and pETRD carrying the cloned ilvA and tyrB genes of E. coli, alsS, and dal genes of B. subtilis, daao genes of T. variabilis and R. gracilis were respectively transformed into E. coli BL21 (DE3). These genes were successfully overexpressed as soluble proteins, and 20–25-g/l wet cells were obtained by culturing the cells in TB medium for 4 h at 30°C after adding IPTG. When the activity values were calculated, the IlvA, TyrB, AlsS, and BsALR were obtained (1,000, 100, 400, and 30,000 U/g cell, respectively) in E. coli cells. The level of RgDAAO obtained 57.6 U/g was higher than that of TvDAAO 40.9 U/g cell in E. coli cells.
Elimination of l-Ala from the biosynthesis mixtures of l-ABA using E. coli BL21 (DE3)/pETALR, BL21 (DE3)/pETW2, and BL21 (DE3)/pETRD
The synthesis of l-ABA, as the first reaction stage, was catalyzed by three enzymes: threonine deaminase (IlvA), aromatic aminotransferase (TyrB), and acetolactate synthase (AlsS; Fotheringham et al. 1999). After 24-h incubation at 40°C, 29.8 g/l l-ABA (290 mM) and 4.7 g/l l-Ala (53 mM) were produced. Following the biosynthesis of l-ABA, two reaction schemes with DAAO from different microorganisms were carried out to remove the remnant l-Ala from the reaction mixtures of the l-ABA biosynthesis. The results showed that, with the action of BsALR and TvDAAO, l-Ala was decreased quickly from 4.7 to 1.25 g/l after 8 h and then slowly to 0.32 g/l till 20 h, and l-ABA was slightly decreased from 29.8 to 27.2 g/l after 8 h and then to 24.9 g/l till 20 h (Fig. 2a). When using the cells expressing BsALR and RgDAAO, l-Ala was completely removed from the reaction mixtures after 8 h. At the same time, l-ABA was slightly decreased from 29.8 to 27.2 g/l (Fig. 2b). E. coli BL21 (DE3), which carried the only plasmid pET28b, was used as the control. We found that l-ABA was decreased by 2.1 g/l and the l-Ala was almost not changed (decreased 0.34 g/l) after 20 h.
Time courses of l-Ala elimination from biosynthesis mixtures of l-ABA catalyzed by BsALR combined with two different DAAOs in whole cell. Concentrations of D/l-Ala (triangles) and l-ABA (squares) in reaction mixtures were determined by HPLC. a, b The reaction was carried out in a flask with agitation at 300 rpm. c, d The reaction was carried out in a reactor with a supply of pure oxygen and agitation at 100 rpm
Additionally, to increase the bioproduct yield, industrial processes using DAAO are usually performed with a supply of pure oxygen and frequently under high-pressure conditions (Pilone and Pollegioni 2002). Here, we evaluated the activity of these two different combined enzymes on l-Ala removal with a supply of pure oxygen. l-Ala can be removed with the action of BsALR and TvDAAO after 80 min (Fig. 2c). At the same condition, l-Ala was completely removed with the action of BsALR and RgDAAO only for 40 min (Fig. 2d). The results showed that the high concentration of oxygen could greatly enhance the activity of DAAO. With a shorter reaction time, the l-ABA was almost not changed during both reaction processes (Fig. 2c, d). Therefore, with the l-Ala eliminated completely by the concert action of BsALR and DAAO in whole cell, the purification of l-ABA will be easier than before.
Large-scale preparation of l-ABA
To apply this method to l-ABA production in factory, the large-scale reaction (1,500 l) was carried out in the 2,000-l jar fermentor. At the first stage of the reaction, 28.74 g/l (279 mM) l-ABA and 4.02 g/l l-Ala (45 mM) were obtained with the action of three enzymes after incubation for 24 h at 40°C. The aspartate amino donor was almost entirely consumed in the reaction. Subsequently, the l-Ala was completely removed by the action of BsALR and RgDAAO after incubation for 8 h at 37°C, resulting in the l-ABA concentration of 25.38 g/l (246 mM) at the second stage of the reaction.
Isolation and purification of l-ABA
High-quality l-ABA product (content > 98.5%, purity 99%, single impurity < 0.1%, ee > 99%) can be used as the synthetic precursor for pharmaceuticals. To obtain a high-quality l-ABA, the reaction mixture containing 29.8 g/l l-ABA and 4.7 g/l l-Ala at the first reaction stage was purified and crystallized once as methods, and the product containing 97.76% l-ABA and 0.59% l-Ala was obtained. To further increase the purity of l-ABA, the crystal was redissolved in water at 70°C, and then two volumes of ethanol were added for recrystallization. The content of l-ABA was increased up to 99.2%, but l-Ala was still high of 0.47%. The level of l-Ala could not be reduced to lower than 0.1%, although the crystallization process was repeated for several times. As for the second reaction mixture, in which BsALR and RgDAAO was introduced to remove the by-product l-Ala completely, the high-quality l-ABA with the content 99.8% was obtained by crystallization once after simple purification as the methods.
Elimination of l-Ala from mixtures of l-Ala and l-Terleu or l-Val using BsALR and RgDAAO
We also investigated the ability of BsALR and RgDAAO to eliminate the l-Ala from the mixtures of 30 g/l l-Terleu and 6 g/l l-Ala or 30 g/l l-Val and 6 g/l in a stirred reactor, where the enzymes were used in the cell form. The products of the bioconversion reaction were quantified by HPLC. In both cases, l-Ala could be completely removed from the mixtures with the action of BsALR and RgDAAO after 8 h, and l-Terleu and l-Val were both retained at high concentration of over 26 g/l, although they were inevitably partially catabolized by the host cells.
Discussion
In the present study, we achieved to remove completely the by-product l-alanine from the reaction mixtures of l-2-aminobutyric acid biosynthesis by the concerted action of BsALR and RgDAAO. The overall yield of l-ABA represented was over 97% of the total amino acids at the end of the reaction which was higher than before (92%; Fotheringham et al. 1999). More importantly, the complexity of purification of l-ABA was decreased dramatically after the removal of the by-product l-Ala, and the high-quality l-ABA (content 99.8%) was prepared by crystallization once after simple purification. These results suggested that enzymatic methods could resolve the problems in the isolation and purification of compounds.
The results demonstrated that BsALR can selectively act on l-Ala in spite of l-ABA sharing high similarity with l-Ala in structure and chemical properties, so high-yield l-ABA was retained but l-Ala diminished, which suggested that it was important to consider the substrate preference of the enzymes we used. Moreover, this method was also successfully used to remove the l-Ala from the reaction mixtures of other neutral amino acids such as l-Terleu and l-Val which further indicated that BsALR had a strict specificity to l- and d-Ala.
However, in this study, the combination of BsALR and RgDAAO showed higher activities than that of BsALR and TvDAAO on the removal of l-Ala even at high oxygen concentration which should be attributed to the different catalytic efficiency (as expressed by the ratio kcat, app/km, app) of TvDAAO and of RgDAAO on d-alanine by more than 1 order of magnitude (Pollegioni et al. 2004). Therefore, the high catalytic efficiency of RgDAAO for d-alanine could accelerate the conversion of l-alanine to d-alanine and lead to the complete removal of l-Ala ultimately.
In this work, l-ABA slightly diminished in the second reaction stage when the reaction was not performed with a supply of pure oxygen. The same phenomena appeared in the first stage because the theoretical yield of l-ABA should be 350 mM according to the supply of aspartate. We considered the loss of l-ABA was mainly to be due to partial catabolism by the host cells and the action of five enzymes. Therefore, to study the mechanism of l-ABA diminishment, the reaction mixture of the control and that of the two reaction stages would be analyzed, and then, we can disrupt the catabolism pathway and develop the enzymes to reduce the l-ABA diminishment. However, this problem also can be solved if the reaction was carried out with a supply of pure oxygen which significantly reduced the reaction time and did not affect the concentration of l-ABA.
Here, large-scale preparation (1,500 l) of high purity l-ABA was successfully carried out with five enzymes at two stages of bioprocesses. However, preparation of these five enzymes will increase the cost of l-ABA production. This problem will be solved if the enzymes used at the two stages were coexpressed in the recombinant E. coli. Despite of the current shortcomings, this study showed that multienzymatic whole-cell catalysis not only is milder than the conventional chemical catalysis but also can provide higher purity of different types of unnatural amino acids. With the availability of abundant genomic DNA sequences and the development of DNA synthesis technology by chemical methods, it becomes much easier to obtain the desired genes. What is more is that we can improve the activities and properties of enzymes by means of protein engineering. All these technologies mentioned above will speed up the development of multienzyme-catalyzed production of natural and nonnatural compounds.
References
Bhor VM, Dev S, Vasanthakumar GR, Kumar P, Sinha S, Surolia A (2006) Broad substrate stereospecificity of the Mycobacterium tuberculosis 7-keto-8-aminopelargonic acid synthase: Spectroscopic and kinetic studies. J Biol Chem 281:25076–25088
Cho BK, Seo JH, Kang TW, Kim BG (2003) Asymmetric synthesis of L-homophenylalanine by equilibrium-shift using recombinant aromatic L-amino acid transaminase. Biotechnol Bioeng 83:226–234
Fotheringham IG, Grinter N, Pantaleone DP, Senkpeil RF, Taylor PP (1999) Engineering of a novel biochemical pathway for the biosynthesis of L-2-aminobutyric acid in Escherichia coli K12. Bioorg Med Chem 7:2209–2213
Fotheringham I, Archer I, Carr R, Speight R, Turner NJ (2006) Preparative deracemization of unnatural amino acids. Biochem Soc Trans 34:287–290
Guillouet S, Rodal AA, An G, Lessard PA, Sinskey AJ (1999) Expression of the Escherichia coli catabolic threonine dehydratase in Corynebacterium glutamicum and its effect on isoleucine production. Appl Environ Microbiol 65:3100–3107
Han Q, Li J (2002) Comparative characterization of Aedes 3-hydroxykynurenine transaminase/alanine glyoxylate transaminase and Drosophila serine pyruvate aminotransferase. FEBS Lett 527:199–204
Hwang JY, Park J, Seo JH, Cha M, Cho BK, Kim J, Kim BG (2009) Simultaneous synthesis of 2-phenylethanol and L-homophenylalanine using aromatic transaminase with yeast Ehrlich pathway. Biotechnol Bioeng 102:1323–1329
Johnston RB, Schreiber EC, Davis MP, Jillson L, Sorrell WT, Kirker ME (1984) Catalytic properties of the active site of alanine racemase from B. subtilis. Prog Clin Biol Res 144A:339–350
Ju J, Yokoigawa K, Misono H, Ohnishi K (2005) Cloning of alanine racemase genes from Pseudomonas fluorescens strains and oligomerization states of gene products expressed in Escherichia coli. J Biosci Bioeng 100:409–417
Kim I-W, Khang Y-H (1995) Simple and rapid determination of the activity of recombinant D-amino acid oxidase in cephalosporin C bioconversion with use of a micro pO2 probe. Biotechnol Tech 9:863–868
Kim SS, Yu YG (2000) Molecular cloning of an extremely thermostable alanine racemase from Aquifex pyrophilus and enzymatic characterization of the expressed protein. J Biochem Mol Biol 33:82–88
Krix G, Bommarius AS, Drauz K, Kottenhahn M, Schwarm M, Kula MR (1997) Enzymatic reduction of alpha-keto acids leading to L-amino acids, D- or L-hydroxy acids. J Biotechnol 53:29–39
Lo HH, Hsu SK, Lin WD, Chan NL, Hsu WH (2005) Asymmetrical synthesis of L-homophenylalanine using engineered Escherichia coli aspartate aminotransferase. Biotechnol Prog 21:411–415
Pennartz A, Genereux C, Parquet C, Mengin-Lecreulx D, Joris B (2009) Substrate-induced inactivation of the Escherichia coli AmiD N-acetylmuramoyl-L-alanine amidase highlights a new strategy to inhibit this class of enzyme. Antimicrob Agents Chemother 53:2991–2997
Pilone MS, Pollegioni L (2002) D-Amino acid oxidase as an industrial biocatalyst. Biocatal Biotransform 20:145–159
Pleiss JA, Wolfson AD, Uhlenbeck OC (2000) Mapping contacts between Escherichia coli alanyl tRNA synthetase and 2' hydroxyls using a complete tRNA molecule. Biochemistry 39:8250–8258
Pollegioni L, Caldinelli L, Molla G, Sacchi S, Pilone MS (2004) Catalytic properties of D-amino acid oxidase in cephalosporin C bioconversion: a comparison between proteins from different sources. Biotechnol Prog 20:467–473
Pollegioni L, Molla G, Sacchi S, Rosini E, Verga R, Pilone MS (2008) Properties and applications of microbial D-amino acid oxidases: current state and perspectives. Appl Microbiol Biotechnol 78:1–16
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Schroder I, Vadas A, Johnson E, Lim S, Monbouquette HG (2004) A novel archaeal alanine dehydrogenase homologous to ornithine cyclodeaminase and mu-crystallin. J Bacteriol 186:7680–7689
Shin JS, Kim BG (2009) Transaminase-catalyzed asymmetric synthesis of L-2-aminobutyric acid from a chiral reactants. Biotechnol Lett 31:1595–1599
Uo T, Yoshimura T, Tanaka N, Takegawa K, Esaki N (2001) Functional characterization of alanine racemase from Schizosaccharomyces pombe: a eucaryotic counterpart to bacterial alanine racemase. J Bacteriol 183:2226–2233
Zhang KC, Li H, Cho KM, Liao JC (2010) Expanding metabolism for total biosynthesis of the nonnatural amino acid L-homoalanine. Proc Natl Acad Sci USA 107:6234–6239
Zheng H, Wang X, Chen J, Zhu K, Zhao Y, Yang Y, Yang S, Jiang W (2006) Expression, purification, and immobilization of His-tagged D-amino acid oxidase of Trigonopsis variabilis in Pichia pastoris. Appl Microbiol Biotechnol 70:683–689
Acknowledgments
This work was supported by the Knowledge Innovation Program of the Chinese Academy of Sciences (No. KSCX2-EW-G-7, KSCX2-YW-G-075-14, KSCX2-EW-G-8), the Huzhou Municipal Science and Technology Project (2010ZD1006), the “365” Outstanding Scientific and Technological Innovation Team of Huzhou (2010KC01), and the Hi-Tech industrialized seed fund projects by Pudong New Area and Chinese Academy of Sciences (No. PKC2010-03). This work was also supported in part by National Basic Research Program of China (973: 2007CB707803, 2011CBA00806).
Author information
Authors and Affiliations
Corresponding author
Additional information
Li Zhu and Rongsheng Tao contributed equally to this work.
Rights and permissions
About this article
Cite this article
Zhu, L., Tao, R., Wang, Y. et al. Removal of l-alanine from the production of l-2-aminobutyric acid by introduction of alanine racemase and d-amino acid oxidase. Appl Microbiol Biotechnol 90, 903–910 (2011). https://doi.org/10.1007/s00253-011-3127-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-011-3127-4