Abstract
The recent years have seen tremendous progress towards the understanding of microbial metabolism on a higher level of the entire functional system. Hereby, huge achievements including the sequencing of complete genomes and efficient post-genomic approaches provide the basis for a new, fascinating era of research—analysis of metabolic and regulatory properties on a global scale. Metabolic flux (fluxome) analysis displays the first systems oriented approach to unravel the physiology of microorganisms since it combines experimental data with metabolic network models and allows determining absolute fluxes through larger networks of central carbon metabolism. Hereby, fluxes are of central importance for systems level understanding because they fundamentally represent the cellular phenotype as integrated output of the cellular components, i.e. genes, transcripts, proteins, and metabolites. A currently emerging and promising area of research in systems biology and systems metabolic engineering is therefore the integration of fluxome data in multi-omics studies to unravel the multiple layers of control that superimpose the flux network and enable its optimal operation under different environmental conditions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
One of the past and future driving forces of systems-oriented microbiology is the increasing need for superior cell factories, requesting efficient design and optimization strategies. Previous strain improvement has enabled random mutagenesis and selection or targeted genetic engineering without consideration of its impact on the entire metabolism. This caused detrimental side effects so that industrial biocatalysts today suffer from slow growth, weak stress tolerance, or the formation of undesired by-products, meaning that the potential of the available biochemical reaction set is often not fully recruited (Wittmann 2010). This is one of the reasons why industrial strain development is shifted towards systems metabolic engineering approaches which rationally introduce beneficial genetic modifications based on a systems-wide understanding of the underlying metabolic and regulatory networks (Lee et al. 2009). Such multi-omics studies have been performed more and more in the recent years in systems biology and systems metabolic engineering (Ishii et al. 2007; Park et al. 2008; Zhang et al. 2010). The present contribution highlights the application of flux analysis for quantitative analysis and engineering of metabolic pathways and its integration into a systems-oriented framework (Fig. 1). The underlying concept as well as the potential of flux-based analysis and engineering of microbial metabolism is hereby illustrated by selected examples, not aiming at a full coverage of studies on microbial fluxomics. For a more complete literature overview on metabolic flux studies of microorganisms, the reader is directed to recent review articles (Iwatani et al. 2008; Kim et al. 2008; Blank and Kuepfer 2010).
Tools and concepts for 13C-fluxomics
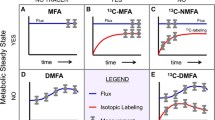
Metabolic flux analysis is about the quantification of intracellular fluxes, i.e., in vivo reaction rates through the different pathways within the intact living cell. Hereby, the level of substrates, cofactors, effectors, or regulators determine the in vivo flux through a particular enzyme as systems property. This differs in most cases dramatically from estimates on the in vitro activity for which enzymes are withdrawn from their physiological environment and analyzed under artificial assay conditions. In practice, flux analysis typically focuses on the 50–100 reactions of central metabolism which have particular relevance for biotechnology applications since they catalyze the major carbon flow and match with the reactions involved in biosynthesis of most industrial products.
Initially, flux studies recruited measured extracellular fluxes, i.e., substrate uptake or product secretion, and biosynthetic requirements entered into assumed stoichiometric reaction networks to derive certain intracellular fluxes (Shuichi and Masayoshi 1979). These pioneering studies provided first valuable insights into microbial metabolism on the flux level, but relied on uncertain assumptions, simplifications, and constraints such as balances of reduction equivalents or energy stoichiometry which unfortunately determine the actual flux result quite significantly (Wittmann 2002). Moreover, important fine structures as parallel or cyclic reactions which play an important role in microorganisms, e.g., in biosynthesis of biotechnology products such as lysine (Sonntag et al. 1993) or in central carbon metabolism (Hult et al. 1980; Sauer and Eikmanns 2005), were not accessible. In recent years, flux analysis has been extended by the integration of 13C labeling information from stable isotope experiments overcoming previous limitations. In these studies, 13C-labeled tracer substrates are fed to the examined cells until the 13C label has propagated through the metabolic network into metabolic products. Their 13C labeling pattern depends on the particular flux distribution and thus provides a sensitive “fingerprint” to calculate the fluxes. By means of mass spectrometry (MS) (Christensen and Nielsen 1999; Wittmann and Heinzle 1999; Dauner and Sauer 2000) and nuclear magnetic resonance spectroscopy (NMR) (Szyperski 1995), the labeling pattern can be quantified, whereby both methods ideally provide complementary data sets from the same sample. For flux calculation, the measured 13C labeling data, biosynthetic requirements and extracellular fluxes are integrated with a computer model which is an in silico representation of the metabolic network investigated. The basic workflow of the resulting comprehensive strategy, comprising experimental and computational steps, is given in Fig. 2. Metabolic flux analysis as outlined below has evolved into an advanced approach to assess steady-state fluxes of microbial cells (Sauer 2006). Inherently, fluxomics is laborious and requires a broad spectrum of quite diverse experimental and computational expertises, which is not easily brought together. Accordingly, and in contrast to other omics technologies, only a few leading laboratories have so far significantly contributed to its development and application.
Schematic work flow for 13C metabolic flux (fluxome) analysis comprising (I) experimental design to identify optimal substrates for a given flux problem, (II) isotope tracer experiments with measurement of 13C labeling patterns by MS together with extracellular fluxes and biosynthetic requirements, and (III) computational calculation of fluxes with models as in silico representation of the studied network. This basic concept is typically applied for routine quantification of steady-state fluxes. It can be varied in several aspects depending on the focus of the flux study
13C labeling analysis
Most common metabolic flux studies recruit mass spectrometric 13C labeling analysis of about 10–15 amino acids obtained from hydrolyzed cell protein (Christensen and Nielsen 1999; Dauner and Sauer 2000). They contain rich information for flux estimation, since they reflect the carbon backbone of eight key intermediates from different parts of central metabolism (Szyperski 1995). The most popular technique for amino acid labeling analysis today is GC/MS, since this approach requires only about 1 mg of cells, allows an excellent separation of the analytes within only about 20–30 min, and provides high precision data on mass isotopologue distribution with measurement errors of <0.5% (Wittmann 2007). Initially developed and applied mainly in the field of biomedicine (Lapidot and Nissim 1980; Nissim et al. 1981; Tsalikian et al. 1984; Nissim and Lapidot 1986), GC–MS-based metabolic flux approaches have been substantially extended and optimized and emerged as a key technology in metabolic physiology and biotechnology (Christensen and Nielsen 2000; Kelleher 2001; Wiechert 2001; Wittmann 2002; Des Rosiers et al. 2004; Des Rosiers and Chatham 2005).
Modeling and software packages
For flux calculation, computer-based models of the investigated metabolic network are usually utilized to globally fit the unknown flux parameters combining isotopomer and metabolite balancing (Wittmann 2007). Applying an optimization algorithm the deviation of the labeling data between computer model and experiment is minimized by iterative variation of the free fluxes until the flux distribution is identified. In combination with experimentally determined extracellular fluxes, absolute carbon fluxes throughout the network are derived. With certain constraints, GC–MS labeling data can also be used to calculate local flux ratios of converging reactions in the network (Fischer and Sauer 2003; Fischer et al. 2004). Although at first limited to only about 10–15 flux ratios directly accessible, this method has been recently combined with stoichiometric models and measured extracellular fluxes also providing a distribution of absolute fluxes (Zamboni et al. 2005). In recent years, general modeling frameworks have been developed for a general and systematic description of carbon transfer. These include based balancing of isotopomers (Schmidt et al. 1997), cumomers (Wiechert et al. 1999), and bondomers (van Winden et al. 2002) which display different representations of metabolic labeling states. The most efficient approach with regard to simulation speed is the recent introduction of elementary mode units (EMU) (Antoniewicz et al. 2007a). The different algorithms are implemented into convenient software tools which allow user-friendly flux calculation from measurement data as well as statistical treatment of flux data. Examples are OpenFlux using the EMU approach (Quek et al. 2009), 13C-Flux based on cumomers (Wiechert et al. 2001), Fiat-Flux involving flux ratio analysis (Zamboni et al. 2005), and a Matlab-based tool recruiting isotopomers (Wittmann and Heinzle 1999). Most of these packages support optimal experimental design of carbon labeling experiments by identifying optimal tracer substrates for a given flux problem as well as statistical treatment of flux data. First promising studies have recently shown dynamic labeling experiments for flux analysis under dynamic conditions, which open new possibilities to resolve fluxes in non-stationary systems (Nöh et al. 2006; Antoniewicz et al. 2007b; Nöh et al. 2007).
Application of 13C fluxomics
Pathway function
Extensive biochemical analysis carried out since the 1940s has basically provided detailed knowledge on the enzymes and pathways present in biological systems—including various microorganisms. With the extension of flux quantification from individual reactions to particular subsets of selected enzymes and finally to networks at the systems level, fascinating insights into metabolic properties of microorganisms could be obtained, especially through the recent years. Firstly, this has led to a new understanding of the role of certain pathways beyond their classically attributed function. An outstanding example is the pentose phosphate pathway, originally assumed to supply building blocks for biosynthesis. Metabolic flux analysis, however, demonstrated that this pathway carries flux far beyond the actual requirement for anabolism and fulfils a flexible role in catabolic breakdown and redox metabolism. Flexible partitioning of the flux between the glycolysis and the pentose phosphate pathway has been found in many prokaryotic and eukaryotic microorganisms (Blank et al. 2005b; Fuhrer et al. 2005). Illustrating examples include growth of Saccharomyces cerevisiae at high and low glucose concentration (Gombert et al. 2001) or at varied specific growth rate (Frick and Wittmann 2005), the utilization of alternative carbon sources by Corynebacterium glutamicum (Kiefer et al. 2004; Wittmann et al. 2004), or the production of antibiotics in Streptomyces coelicolor (Borodina et al. 2008). In cases of significantly changed metabolic burden, cells recruit their entire network and distribute the response on the flux level globally as observed for amino acid producing C. glutamicum (Marx et al. 1997). Even a single nucleotide exchange in the whole genome leading to deregulation of a biosynthetic enzyme may cause a systems-wide rearrangement of flux when cells switch from growth to production (Kim et al. 2006).
Also the flux through the tricarboxylic acid (TCA) cycle has been shown to be a function of the environmental conditions as exemplified for S. cerevisiae (Gombert et al. 2001; Blank and Sauer 2004). Interestingly, the slow-growing myxobacterium Sorangium cellulosum exhibits an enormously high relative TCA cycle flux far beyond typical values found in fast-growing microbial cells (Bolten et al. 2009). Calculation on energy stoichiometry based on the estimated fluxes indicated that such a high TCA cycle flux is needed to ensure growth in this microorganism, because about 90% of ATP produced is withdrawn for maintenance requirements.
Discovery of novel pathways
The rich information on metabolism by flux analysis, previously not accessible, led to the discovery of different novel pathways. Important findings comprise the simultaneous operation of glycolytic and gluconeogenetic reactions at the junction between glycolysis and TCA cycle crucial to anticipate changing environmental conditions at the expense of ATP in many microorganisms (Sauer and Eikmanns 2005), the PEP–glyoxylate pathway as novel catabolic route in Escherichia coli (Nanchen et al. 2006) or a novel biosynthetic route for isoleucine biosynthesis in C. glutamicum (Krömer et al. 2006).
More recently, 13C flux analysis was extended to investigate fluxes in more unexplored species. Among the various interesting findings obtained from such studies is the identification of the Entner–Doudoroff pathway as major catabolic route in the actinomycete Streptomyces tenebrarius (Borodina et al. 2005) or in the marine bacteria Dinoroseobacter shibae and Phaeobacter gallaeciensis (Fürch et al. 2009), the discovery of a novel route for biosynthesis of serine in lactate-grown Shewanella oneidensis (Tang et al. 2009) or of isoleucine in Geobacter metallireducens during growth on acetate (Tang et al. 2007).
Robustness and rigidity of metabolic networks
The successful miniaturization of flux analysis approaches, enabling the broad investigation of mutant libraries, has manifested flux analysis as a key technology to unravel control mechanisms in biological systems. In a large-scale study, the relative flux distribution in 137 null mutants of Bacillus subtilis was found rather invariant (Fischer and Sauer 2005). Flux profiling of different S. cerevisiae deletion mutants demonstrated that network redundancy through duplicate genes was the major (75%) and alternative pathways the minor (25%) molecular mechanism of genetic network robustness (Blank et al. 2005a). Hereby, local flux rerouting seems one key mechanism of cellular response to maintain major growth characteristics upon gene deletion. As prominent example, the lack of the genes encoding for pyruvate kinase, a central glycolytic enzyme, is compensated in E. coli (Emmerling et al. 2002) and in C. glutamicum (Becker et al. 2008) by activation of a by-pass recruiting PEP carboxylase, malate dehydrogenase, and malic enzyme to convert PEP into pyruvate, whereas other fluxes in the network remain almost unaffected.
Engineering of metabolic pathway fluxes for biotechnology
It lies at hand that metabolic flux analysis has become a core methodology in metabolic engineering which particularly aims at the optimization of flux (Iwatani et al. 2008; Park et al. 2008). With regard to metabolic flux analysis, C. glutamicum can probably be considered as the most important model organism (Wittmann and de Graaf 2005; Wittmann 2010) so that the following examples taken from investigation of this microorganism might give a flavor to which level of understanding and targeted metabolic engineering flux analysis can contribute. To date, the most valuable flux data towards rational strain optimization have certainly been obtained by routine stationary metabolic flux analysis (Wittmann 2010). The systematic quantification of flux in different lysine-producing strains (Marx et al. 1997, 1999; Wittmann and Heinzle 2001, 2002; Marx et al. 2003) or under different environmental conditions (Wendisch et al. 2000; Klapa et al. 2003; Kiefer et al. 2004; Wittmann et al. 2004; Shirai et al. 2006) has unraveled key pathways to be modified for improved production performance (Ikeda 2003; Wittmann and Becker 2007). Based on the predictions by 13C fluxomics, superior lysine production strains could be created which exhibited improved performance via single gene modifications. Prominent examples are an increased supply of NADPH, required as cofactor for lysine biosynthesis, by amplification of the pentose phosphate pathway (Ohnishi et al. 2005; Becker et al. 2007) or the gluconeogenetic enzyme fructose 1,6-bisphosphatase (Becker et al. 2005; Georgi et al. 2005). Similarly, the targeted attenuation of competing enzymes and pathways, including the TCA cycle (Becker et al. 2009), pyruvate dehydrogenase (Becker et al. 2010) as well as phosphoenolpyruvate carboxykinase (Riedel et al. 2001) successfully increased product formation. Also, glutamate production by C. glutamicum has been in the focus of flux analysis. As for lysine, the biosynthesis of glutamate demands for efficient anaplerotic replenishment (Peters-Wendisch et al. 2001; Shirai et al. 2007). Based on flux predictions, deletion of pyruvate kinase improved glutamate production under biotin-limited conditions (Sawada et al. 2010). Flux analysis further revealed that the flux through ODHC significantly decreased after induction of glutamate production (Shirai et al. 2005) which could be utilized for subsequent strain engineering (Asakura et al. 2007; Kim et al. 2009). In addition, flux analysis has been applied to a number of other microbial production strains and processes, providing extensive insights into the underlying metabolism, as recently summarized in excellent overviews (Iwatani et al. 2008; Blank and Kuepfer 2010).
Towards systems-level understanding of metabolism
Integration of multi omics data sets with fluxes
Single omics analyses, which are used quite frequently, provide information about only one layer of a cellular system, but fail to capture the interaction between the different layers essential to understand the system properties. To this end, we see more and more multi-omics studies in recent years, often involving the alliance of specialized research groups due to high costs and efforts (Sauer et al. 2007). Most of these studies combine the most frequently used tools for transcriptomics and proteomics in order to obtain complementary coverage of metabolism, perform cross-validation, or obtain novel biological insights into post-transcriptional regulation mechanisms (Zhang et al. 2010). As described above, flux analysis of microbial cells is a rather advanced and routine method in itself, but has not been as broadly used, as genomics, transcriptomics, and proteomics which have advanced into widely applied commercially available technologies. The analysis of the metabolome is not as mature and yet not fully comprehensive, which is inherently caused by the different chemical nature, high turnover rate and large concentration difference of the analytes (Zhang et al. 2010), and persisting difficulties with appropriate sampling and cell quenching hampering quantitative metabolomics (Bolten et al. 2007). This is one explanation for the tremendous difference of several orders of magnitude to which extent the different omics technologies are applied today (Fig. 3). It appears obvious that in many cases, the interaction between components of the network involved cannot be understood without knowledge on in vivo flux. To unravel and quantify the different layers of control superimposing the flux network, the flux state has to be linked to the set of cellular components, i.e., genes, transcripts, proteins, and metabolites. The integration of 13C fluxomics in multi-omics experiments, however, has special requirements. Care has to be taken concerning the label introduced in 13C fluxomics experiments which typically interferes with conventional MS-based technologies for identification and quantification of proteins as well as metabolites. In such cases, a parallelized setup is needed. The isotope labeling experiments for the fluxome have to be run under identical conditions in parallel to those for the other omics, including a thorough check for key physiological parameters such as substrate uptake, growth, product formation, respiration, or intracellular fingerprints which are not affected by the 13C label (Krömer et al. 2004). This might include additional controls via certain intracellular metabolites accessible by enzyme assays or HPLC as well as transcription profiling, which are not biased by the presence of label. In addition, financial constraints have to be considered with respect to the high price of 13C substrates, typically demanding for small cultivation volume.
Frequency of the use of the terms “genome”, “transcriptome”, “proteome”, “metabolome”, and “fluxome” in scientific publications. Since the terms “metabolome” and “fluxome” are not always used in the relevant studies, the search included “intracellular metabolite” or “metabolic flux” as alternative terms to represent the corresponding field. The number of appearance was extracted from the PubMed database (August, 2010)
Unraveling of regulation networks
The resulting picture immediately indicates that only a small set of multi-omics studies so far includes fluxes—and thus allows to really bridging cellular components with network function, which is a key goal of systems biology and systems metabolic engineering. Out of the few studies performed so far, most investigated biotechnologically relevant microorganisms, underlining systems metabolic engineering as one of the major drivers. One of the pioneering examples focused on C. glutamicum for production of the feed amino acid lysine (Krömer et al. 2004). Through integration of transcriptomics with fluxomics, correlation between gene expression and in vivo flux could be quantified for various enzymes in carbon core metabolism providing novel insights into regulation of this microorganism. It became obvious that the different reactions and pathways are controlled at different levels. As example, reactions of glucose uptake, pentose phosphate pathway, and TCA cycle were found closely regulated on the transcriptional level (Fig. 4a), whereas genes in lysine biosynthesis showed a constant expression level, despite a marked change of the metabolic flux, indicating that they are strongly regulated at the post-transcriptional level (Fig. 4b). Similar studies on E. coli (Shimizu 2004; Fong et al. 2006; Hua et al. 2007) and B. subtilis (Schilling et al. 2007) confirmed that transcriptional control is involved, but not sufficient to account for metabolic regulation, indicating other important mechanisms such as metabolite–protein interactions or protein phosphorylation (Heinemann and Sauer 2010). A comprehensive insight into the production of pigmented antibiotics in S. coelicolor was gained by coupling the analysis of fluxes with profiling of intracellular metabolites and gene expression (Borodina et al. 2008). It could be shown that the increased pentose phosphate pathway flux, linked to enhanced production, appeared largely because of accumulation of glucose 6-phosphate and fructose 6-phosphate. Integrated analysis of gene expression data further revealed transcriptional changes in genes encoding redox co-factor-dependent enzymes as well as those encoding pentose phosphate pathway enzymes.
Integration of 13C fluxome and transcriptome data to classify regulation for metabolic pathways into transcriptional (a) and post-transcriptional control (b). The data are taken from a multi-omics study on lysine production by Corynebacterium glutamicum (Krömer et al. 2004). Gene expression and flux for each pathway are given here as mean value for the corresponding genes and reaction steps involved which all showed the same trend. The normalization of flux and expression is described in the original manuscript
In E. coli, the combined analysis of transcriptome, metabolome, and fluxome confirmed the existence of hidden reactions in the central metabolism of its cells (Nakahigashi et al. 2009). One of the most massive data sets was recently generated for a systematic analysis of E. coli cells to genetic and environmental perturbation (Ishii et al. 2007). This included the quantification of 4,000 mRNA transcripts, 2,000 proteins, 600 metabolites as well as pathway fluxes. The study revealed only few and local responses to gene deletions, whereas the metabolism responded highly flexible and globally to change in the growth rate. E. coli thus seems to use complementary strategies that result in a metabolic network robust against perturbations. While many questions remain unanswered from the above studies, their data sets provide a rich source for future computational analysis to extract novel biological insights (Zamboni and Sauer 2009).
Understanding network-wide balancing of redox and energy status
The combined use of different omics tools, especially metabolomics and fluxomics, has been applied to understand how metabolic networks organize the overall adjustment to optimal redox and energy levels and distribute the give and take among the various reactions involved. In a remarkable recent study, using a combination of 13C fluxomics, metabolomics, and in vitro enzyme assays, the formation of NADPH was found to mismatch the anabolic demand in different microbial species, which could be attributed to the transhydrogenase activity in Paracoccus versutus, B. subtilis and E. coli, whereas other species recruit other mechanisms, possibly also a certain promiscuity in cofactor specificity of catabolic enzymes, surprisingly also in the pentose phosphate pathway (Fuhrer and Sauer 2009). Interesting insights into the balancing of redox metabolism could be also obtained for C. glutamicum combining proteomics, metabolomics, and 13C fluxomics under conditions of oxidative stress (Krömer et al. 2008). The results are highlighted by the identification of a global redistribution of NADPH supplying pathways as response to the expression of protective NADPH-consuming proteins and an imbalance in the redox status. Interestingly, the TCA cycle, but not the normally observed pentose phosphate pathway, was found as the major NADPH supplying pathway under conditions of oxidative stress corresponding to a low NADPH/NADP+ ratio (Fig. 5). This obviously leads to suboptimal supply of NADPH at substantially increased loss of CO2. The flux redistribution, however, was not sufficient to fully re-adjust the redox state to the normal value found in the wild type. Clearly, this opens promising possibilities to engineer the metabolism for superior performance under conditions of oxidative stress, a phenomenon often observed in biotechnological production processes. Similarly to the redox metabolism, also the maintenance of the energy status of the cell seems to include network-wide tasks. Combined metabolomics and 13C fluxomics showed that cells of E. coli respond to a decrease in the adenylate energy charge by an increase of the glycolytic flux upon a temperature shift from 37 to 42 °C for induction of recombinant protein production (Wittmann et al. 2007). It is interesting to note that the flux response also involved the activation of the previously discovered PEP–glyoxylate pathway (see above) as new catabolic route, probably induced by a bottleneck at the entry to the TCA cycle.
Integration of 13C fluxome and metabolome data to understand network-wide redox balancing. The data are taken from a multi-omics study on Corynebacterium glutamicum under oxidative stress (Krömer et al. 2008). The cellular redox state is expressed as ratio of NADPH/NADP+. The fluxes are given as relative flux normalized to the glucose uptake rate
Outlook
The increasing need for flux studies as key technology in future systems biology and systems metabolic engineering will clearly intensify efforts to apply and extend this technology. Concerning the integration with other omics approaches, promising future developments comprise the coupling of dynamic 13C labeling experiments with metabolomics. The increasing metabolite coverage in metabolomics provides direct access to labeling patterns of many more pathway intermediates than routinely available, greatly extending the resolution power of flux studies. This has been impressively shown by first examples, e.g., in assembling potentially involved metabolites into a sequence of reactions functioning in vivo, exemplified by the discovery of the ethylmalonyl–CoA pathway in Methylobacterium extorquens during the assimilation of C1 carbon (Peyraud et al. 2009). Beyond such studies, focusing on particular parts of the network, global multi-omics studies will require intelligent software tools that extract true novel biology out of the expected massive data sets. Current developments of statistical methods such as unsupervised learning, correlation network analysis, pattern recognition, or principal component analysis as well as dynamic Bayesian networks, or dynamic control theory appear quite useful, but will surely not suffice to fully unravel the complexity of the systems studied. Here, further exploration of novel concepts integration the heterogeneous data sets is still needed.
To study cells under conditions different from routine chemostat or balanced batch cultures will require the development of novel flux approaches including specific strategies for labeling input, cultivation, labeling analysis, and modeling. First interesting examples extend flux analysis to previously inaccessible conditions. This comprises the quantification of flux dynamics in non-stationary systems (Antoniewicz et al. 2007b), or of fluxes in non-growing cells (Yang et al. 2005, 2006). An interesting concept for flux analysis in large scale, involving precise labeling quantification at low enrichment by GC–IR–MS (Yuan et al. 2010) opens the possibility to perform flux studies in real case production processes. The coming years will hopefully see more conceptional extension of flux analysis so that it becomes applicable to the level of single cells towards unraveling of population heterogeneity, as well as to the flux cross-talk within consortia of different cell types, the dominating form of microbial life—both promising fascinating novel biology and useful information towards superior biocatalysts.
References
Antoniewicz MR, Kelleher JK, Stephanopoulos G (2007a) Elementary metabolite units (EMU): a novel framework for modeling isotopic distributions. Metab Eng 9:68–86
Antoniewicz MR, Kraynie DF, Laffend LA, Gonzalez-Lergier J, Kelleher JK, Stephanopoulos G (2007b) Metabolic flux analysis in a nonstationary system: fed-batch fermentation of a high yielding strain of E. coli producing 1, 3-propanediol. Metab Eng 9:277–292
Asakura Y, Kimura E, Usuda Y, Kawahara Y, Matsui K, Osumi T, Nakamatsu T (2007) Altered metabolic flux due to deletion of odhA causes L-glutamate overproduction in Corynebacterium glutamicum. Appl Environ Microbiol 73:1308–1319
Becker J, Klopprogge C, Zelder O, Heinzle E, Wittmann C (2005) Amplified expression of fructose 1, 6-bisphosphatase in Corynebacterium glutamicum increases in vivo flux through the pentose phosphate pathway and lysine production on different carbon sources. Appl Environ Microbiol 71:8587–8596
Becker J, Klopprogge C, Herold A, Zelder O, Bolten CJ, Wittmann C (2007) Metabolic flux engineering of L-lysine production in Corynebacterium glutamicum-over expression and modification of G6P dehydrogenase. J Biotechnol 132:99–109
Becker J, Klopprogge C, Wittmann C (2008) Metabolic responses to pyruvate kinase deletion in lysine producing Corynebacterium glutamicum. Microb Cell Fact 7:8
Becker J, Klopprogge C, Schröder H, Wittmann C (2009) Tricarboxylic acid cycle engineering for improved lysine production in Corynebacterium glutamicum. Appl Environ Microbiol 75:7866–7869
Becker J, Buschke N, Bücker R, Wittmann C (2010) Systems level engineering of Corynebacterium glutamicum—reprogramming translational efficiency for superior production. Chem Eng Life Sci 10:1–9
Blank LM, Kuepfer L (2010) Metabolic flux distributions: genetic information, computational predictions, and experimental validation. Appl Microbiol Biotechnol 86:1243–1255
Blank LM, Sauer U (2004) TCA cycle activity in Saccharomyces cerevisiae is a function of the environmentally determined specific growth and glucose uptake rates. Microbiology 150:1085–1093
Blank LM, Kuepfer L, Sauer U (2005a) Large-scale 13C-flux analysis reveals mechanistic principles of metabolic network robustness to null mutations in yeast. Genome Biol 6:R49
Blank LM, Lehmbeck F, Sauer U (2005b) Metabolic-flux and network analysis in fourteen hemiascomycetous yeasts. FEMS Yeast Res 5:545–558
Bolten CJ, Kiefer P, Letisse F, Portais J-C, Wittmann C (2007) Sampling for metabolome analysis of microorganisms. Anal Chem 79:3843–3849
Bolten CJ, Heinzle E, Muller R, Wittmann C (2009) Investigation of the central carbon metabolism of Sorangium cellulosum: metabolic network reconstruction and quantification of pathway fluxes. J Microbiol Biotechnol 19:23–36
Borodina I, Scholler C, Eliasson A, Nielsen J (2005) Metabolic network analysis of Streptomyces tenebrarius, a Streptomyces species with an active Entner–Doudoroff pathway. Appl Environ Microbiol 71:2294–2302
Borodina I, Siebring J, Zhang J, Smith CP, van Keulen G, Dijkhuizen L, Nielsen J (2008) Antibiotic overproduction in Streptomyces coelicolor A3 2 mediated by phosphofructokinase deletion. J Biol Chem 283:25186–25199
Christensen B, Nielsen J (1999) Isotopomer analysis using GC–MS. Metab Eng 1:282–290
Christensen B, Nielsen J (2000) Metabolic network analysis. A powerful tool in metabolic engineering. Adv Biochem Eng Biotechnol 66:209–231
Dauner M, Sauer U (2000) GC–MS analysis of amino acids rapidly provides rich information for isotopomer balancing. Biotechnol Prog 16:642–649
Des Rosiers C, Chatham JC (2005) Myocardial phenotyping using isotopomer analysis of metabolic fluxes. Biochem Soc Trans 33:1413–1417
Des Rosiers C, Lloyd S, Comte B, Chatham JC (2004) A critical perspective of the use of 13C-isotopomer analysis by GCMS and NMR as applied to cardiac metabolism. Metab Eng 6:44–58
Emmerling M, Dauner M, Ponti A, Fiaux J, Hochuli M, Szyperski T, Wuthrich K, Bailey JE, Sauer U (2002) Metabolic flux responses to pyruvate kinase knockout in Escherichia coli. J Bacteriol 184:152–164
Fischer E, Sauer U (2003) Metabolic flux profiling of Escherichia coli mutants in central carbon metabolism using GC–MS. Eur J Biochem 270:880–891
Fischer E, Sauer U (2005) Large-scale in vivo flux analysis shows rigidity and suboptimal performance of Bacillus subtilis metabolism. Nat Genet 37:636–640
Fischer E, Zamboni N, Sauer U (2004) High-throughput metabolic flux analysis based on gas chromatography–mass spectrometry derived 13C constraints. Anal Biochem 325:308–316
Fong SS, Nanchen A, Palsson BO, Sauer U (2006) Latent pathway activation and increased pathway capacity enable Escherichia coli adaptation to loss of key metabolic enzymes. J Biol Chem 281:8024–8033
Frick O, Wittmann C (2005) Characterization of the metabolic shift between oxidative and fermentative growth in Saccharomyces cerevisiae by comparative 13C flux analysis. Microb Cell Fact 4:30
Fuhrer T, Sauer U (2009) Different biochemical mechanisms ensure network-wide balancing of reducing equivalents in microbial metabolism. J Bacteriol 191:2112–2121
Fuhrer T, Fischer E, Sauer U (2005) Experimental identification and quantification of glucose metabolism in seven bacterial species. J Bacteriol 187:1581–1590
Fürch T, Preusse M, Tomasch J, Zech H, Wagner-Dobler I, Rabus R, Wittmann C (2009) Metabolic fluxes in the central carbon metabolism of Dinoroseobacter shibae and Phaeobacter gallaeciensis, two members of the marine Roseobacter clade. BMC Microbiol 9:209
Georgi T, Rittmann D, Wendisch VF (2005) Lysine and glutamate production by Corynebacterium glutamicum on glucose, fructose and sucrose: roles of malic enzyme and fructose-1, 6-bisphosphatase. Metab Eng 7:291–301
Gombert AK, Moreira dos Santos M, Christensen B, Nielsen J (2001) Network identification and flux quantification in the central metabolism of Saccharomyces cerevisiae under different conditions of glucose repression. J Bacteriol 183:1441–1451
Heinemann M, Sauer U (2010) Systems biology of microbial metabolism. Curr Opin Microbiol 13:337–343
Hua Q, Joyce AR, Palsson BO, Fong SS (2007) Metabolic characterization of Escherichia coli strains adapted to growth on lactate. Appl Environ Microbiol 73:4639–4647
Hult K, Veide A, Gatenbeck S (1980) The distribution of the NADPH regenerating mannitol cycle among fungal species. Arch Microbiol 128:253–255
Ikeda M (2003) Amino acid production processes. Adv Biochem Eng Biotechnol 79:1–35
Ishii N, Nakahigashi K, Baba T, Robert M, Soga T, Kanai A, Hirasawa T, Naba M, Hirai K, Hoque A, Ho PY, Kakazu Y, Sugawara K, Igarashi S, Harada S, Masuda T, Sugiyama N, Togashi T, Hasegawa M, Takai Y, Yugi K, Arakawa K, Iwata N, Toya Y, Nakayama Y, Nishioka T, Shimizu K, Mori H, Tomita M (2007) Multiple high-throughput analyses monitor the response of E. coli to perturbations. Science 316:593–597
Iwatani S, Yamada Y, Usuda Y (2008) Metabolic flux analysis in biotechnology processes. Biotechnol Lett 30:791–799
Kelleher JK (2001) Flux estimation using isotopic tracers: common ground for metabolic physiology and metabolic engineering. Metab Eng 3:100–110
Kiefer P, Heinzle E, Zelder O, Wittmann C (2004) Comparative metabolic flux analysis of lysine-producing Corynebacterium glutamicum cultured on glucose or fructose. Appl Environ Microbiol 70:229–239
Kim HM, Heinzle E, Wittmann C (2006) Deregulation of aspartokinase by single nucleotide exchange leads to global flux rearrangement in the central metabolism of Corynebacterium glutamicum. J Microbiol Biotechnol 16:1174–1179
Kim HU, Kim TY, Lee SY (2008) Metabolic flux analysis and metabolic engineering of microorganisms. Mol Biosyst 4:113–120
Kim J, Hirasawa T, Sato Y, Nagahisa K, Furusawa C, Shimizu H (2009) Effect of odhA overexpression and odhA antisense RNA expression on Tween-40-triggered glutamate production by Corynebacterium glutamicum. Appl Microbiol Biotechnol 81:1097–1106
Klapa MI, Aon JC, Stephanopoulos G (2003) Systematic quantification of complex metabolic flux networks using stable isotopes and mass spectrometry. Eur J Biochem 270:3525–3542
Krömer JO, Sorgenfrei O, Klopprogge K, Heinzle E, Wittmann C (2004) In-depth profiling of lysine-producing Corynebacterium glutamicum by combined analysis of the transcriptome, metabolome, and fluxome. J Bacteriol 186:1769–1784
Krömer JO, Heinzle E, Schröder H, Wittmann C (2006) Accumulation of homolanthionine and activation of a novel pathway for isoleucine biosynthesis in Corynebacterium glutamicum McbR deletion strains. J Bacteriol 188:609–618
Krömer JO, Bolten CJ, Heinzle E, Schröder H, Wittmann C (2008) Physiological response of Corynebacterium glutamicum to oxidative stress induced by deletion of the transcriptional repressor McbR. Microbiology 154:3917–3930
Lapidot A, Nissim I (1980) Regulation of pool sizes and turnover rates of amino acids in humans: 15N-glycine and 15N-alanine single-dose experiments using gas chromatography–mass spectrometry analysis. Metabolism 29:230–239
Lee SY, Kim HU, Park JH, Park JM, Kim TY (2009) Metabolic engineering of microorganisms: general strategies and drug production. Drug Discov Today 14:78–88
Marx A, Striegel K, de Graaf A, Sahm H, Eggeling L (1997) Response of the central metabolism of Corynebacterium glutamicum to different flux burdens. Biotechnol Bioeng 56(2):168–180
Marx A, Eikmanns BJ, Sahm H, de Graaf AA, Eggeling L (1999) Response of the central metabolism in Corynebacterium glutamicum to the use of an NADH-dependent glutamate dehydrogenase. Metab Eng 1:35–48
Marx A, Hans S, Mockel B, Bathe B, de Graaf AA (2003) Metabolic phenotype of phosphoglucose isomerase mutants of Corynebacterium glutamicum. J Biotechnol 104:185–197
Nakahigashi K, Toya Y, Ishii N, Soga T, Hasegawa M, Watanabe H, Takai Y, Honma M, Mori H, Tomita M (2009) Systematic phenome analysis of Escherichia coli multiple-knockout mutants reveals hidden reactions in central carbon metabolism. Mol Syst Biol 5:306
Nanchen A, Schicker A, Sauer U (2006) Nonlinear dependency of intracellular fluxes on growth rate in miniaturized continuous cultures of Escherichia coli. Appl Environ Microbiol 72:1164–1172
Nissim I, Lapidot A (1986) Dynamic aspects of amino acid metabolism in alloxan-induced diabetes and insulin-treated rabbits: in vivo studies with 15N and gas chromatography–mass spectrometry. Biochem Med Metab Biol 35:88–100
Nissim I, Yudkoff M, Yang W, Terwilliger T, Segal S (1981) Rapid gas chromatographic–mass spectrometric analysis of [15N]urea: application to human metabolic studies. Clin Chim Acta 109:295–304
Nöh K, Wahl A, Wiechert W (2006) Computational tools for isotopically instationary 13C labeling experiments under metabolic steady state conditions. Metab Eng 8:554–577
van Winden WA, Heijnen JJ, Verheijen PJ (2002) Cumulative bondomers: a new concept in flux analysis from 2D [13C, 1H] COSY NMR data. Biotechnol Bioeng 80:731–745
Nöh K, Grönke K, Luo B, Takors R, Oldiges M, Wiechert W (2007) Metabolic flux analysis at ultra short time scale: isotopically non-stationary 13C labeling experiments. J Biotechnol 129:249–267
Ohnishi J, Katahira R, Mitsuhashi S, Kakita S, Ikeda M (2005) A novel gnd mutation leading to increased L-lysine production in Corynebacterium glutamicum. FEMS Microbiol Lett 242:265–274
Park JH, Lee SY, Kim TY, Kim HU (2008) Application of systems biology for bioprocess development. Trends Biotechnol 26:404–412
Peters-Wendisch PG, Schiel B, Wendisch VF, Katsoulidis E, Mockel B, Sahm H, Eikmanns BJ (2001) Pyruvate carboxylase is a major bottleneck for glutamate and lysine production by Corynebacterium glutamicum. J Mol Microbiol Biotechnol 3:295–300
Peyraud R, Kiefer P, Christen P, Massou S, Portais JC, Vorholt JA (2009) Demonstration of the ethylmalonyl–CoA pathway by using 13C metabolomics. Proc Natl Acad Sci U S A 106:4846–4851
Quek LE, Wittmann C, Nielsen LK, Kromer JO (2009) OpenFLUX: efficient modelling software for 13C-based metabolic flux analysis. Microb Cell Fact 8:25
Riedel C, Rittmann D, Dangel P, Mockel B, Petersen S, Sahm H, Eikmanns BJ (2001) Characterization of the phosphoenolpyruvate carboxykinase gene from Corynebacterium glutamicum and significance of the enzyme for growth and amino acid production. J Mol Microbiol Biotechnol 3:573–583
Sauer U (2006) Metabolic networks in motion: 13C-based flux analysis. Mol Syst Biol 2:62
Sauer U, Eikmanns BJ (2005) The PEP–pyruvate–oxaloacetate node as the switch point for carbon flux distribution in bacteria. FEMS Microbiol Rev 29:765–794
Sauer U, Heinemann M, Zamboni N (2007) Genetics. Getting closer to the whole picture. Science 316:550–551
Sawada K, Zen-In S, Wada M, Yokota A (2010) Metabolic changes in a pyruvate kinase gene deletion mutant of Corynebacterium glutamicum ATCC 13032. Metab Eng 12:401–407
Schilling O, Frick O, Herzberg C, Ehrenreich A, Heinzle E, Wittmann C, Stülke J (2007) Transcriptional and metabolic responses of Bacillus subtilis to the availability of organic acids: transcription regulation is important but not sufficient to account for metabolic adaptation. Appl Environ Microbiol 73:499–507
Schmidt K, Carlsen M, Nielsen J, Villadsen J (1997) Modeling isotopomer distributions in biochemical networks using isotopomer mapping matrices. Biotechnol Bioeng 55(6):831–840
Shimizu K (2004) Metabolic flux analysis based on 13C-labeling experiments and integration of the information with gene and protein expression patterns. Adv Biochem Eng Biotechnol 91:1–49
Shirai T, Nakato A, Izutani N, Nagahisa K, Shioya S, Kimura E, Kawarabayasi Y, Yamagishi A, Gojobori T, Shimizu H (2005) Comparative study of flux redistribution of metabolic pathway in glutamate production by two coryneform bacteria. Metab Eng 7:59–69
Shirai T, Matsuzaki K, Kuzumoto M, Nagahisa K, Furusawa C, Shioya S, Shimizu H (2006) Precise metabolic flux analysis of coryneform bacteria by gas chromatography–mass spectrometry and verification by nuclear magnetic resonance. J Biosci Bioeng 102:413–424
Shirai T, Fujimura K, Furusawa C, Nagahisa K, Shioya S, Shimizu H (2007) Study on roles of anaplerotic pathways in glutamate overproduction of Corynebacterium glutamicum by metabolic flux analysis. Microb Cell Fact 6:19
Shuichi A, Masayoshi M (1979) Identification of metabolic model: citrate production from glucose by Candida lipolytica. Biotechnol Bioeng 21:1373–1386
Sonntag K, Eggeling L, De Graaf AA, Sahm H (1993) Flux partitioning in the split pathway of lysine synthesis in Corynebacterium glutamicum. Quantification by 13C- and 1H-NMR spectroscopy. Eur J Biochem 213:1325–1331
Szyperski T (1995) Biosynthetically directed fractional 13C-labeling of proteinogenic amino acids. An efficient analytical tool to investigate intermediary metabolism. Eur J Biochem 232:433–448
Tang YJ, Hwang JS, Wemmer DE, Keasling JD (2007) Shewanella oneidensis MR-1 fluxome under various oxygen conditions. Appl Environ Microbiol 73:718–729
Tang YJ, Martin HG, Dehal PS, Deutschbauer A, Llora X, Meadows A, Arkin A, Keasling JD (2009) Metabolic flux analysis of Shewanella spp. reveals evolutionary robustness in central carbon metabolism. Biotechnol Bioeng 102:1161–1169
Tsalikian E, Howard C, Gerich JE, Haymond MW (1984) Increased leucine flux in short-term fasted human subjects: evidence for increased proteolysis. Am J Physiol 247:E323–E327
Wendisch VF, de Graaf AA, Sahm H, Eikmanns BJ (2000) Quantitative determination of metabolic fluxes during coutilization of two carbon sources: comparative analyses with Corynebacterium glutamicum during growth on acetate and/or glucose. J Bacteriol 182:3088–3096
Wiechert W (2001) 13C metabolic flux analysis. Metab Eng 3:195–206
Wiechert W, Möllney M, Isermann N, Wurzel M, de Graaf AA (1999) Bidirectional reaction steps in metabolic networks: III. Explicit solution and analysis of isotopomer labeling systems. Biotechnol Bioeng 66:69–85
Wiechert W, Möllney M, Petersen S, de Graaf AA (2001) A universal framework for 13C metabolic flux analysis. Metab Eng 3:265–283
Wittmann C (2002) Metabolic flux analysis using mass spectrometry. Adv Biochem Eng Biotechnol 74:39–64
Wittmann C (2007) Fluxome analysis using GC–MS. Microb Cell Fact 6:6
Wittmann C (2010) Analysis and engineering of metabolic pathway fluxes in Corynebacterium glutamicum. Adv Biochem Eng Biotechnol 120:21–49
Wittmann C, Becker J (2007) The L-lysine story: from metabolic pathways to industrial production. Microbiol Monogr 5:39–70
Wittmann C, de Graaf A (2005) Metabolic flux analysis in Corynebacterium glutamicum. In: Eggeling L, Bott M (eds) Handbook of Corynebacterium glutamicum. CRC, Boca Raton, pp 277–304
Wittmann C, Heinzle E (1999) Mass spectrometry for metabolic flux analysis. Biotechnol Bioeng 62:739–750
Wittmann C, Heinzle E (2001) Application of MALDI-TOF MS to lysine-producing Corynebacterium glutamicum: a novel approach for metabolic flux analysis. Eur J Biochem 268:2441–2455
Wittmann C, Heinzle E (2002) Genealogy profiling through strain improvement by using metabolic network analysis: metabolic flux genealogy of several generations of lysine-producing corynebacteria. Appl Environ Microbiol 68:5843–5859
Wittmann C, Kiefer P, Zelder O (2004) Metabolic fluxes in Corynebacterium glutamicum during lysine production with sucrose as carbon source. Appl Environ Microbiol 70:7277–7287
Wittmann C, Weber J, Betiku E, Kromer J, Bohm D, Rinas U (2007) Response of fluxome and metabolome to temperature-induced recombinant protein synthesis in Escherichia coli. J Biotechnol 132:375–384
Yang TH, Heinzle E, Wittmann C (2005) Theoretical aspects of 13C metabolic flux analysis with sole quantification of carbon dioxide labeling. Comput Biol Chem 29:121–133
Yang TH, Wittmann C, Heinzle E (2006) Respirometric 13C flux analysis, Part I: design, construction and validation of a novel multiple reactor system using on-line membrane inlet mass spectrometry. Metab Eng 8:417–431
Yuan Y, Yang TH, Heinzle E (2010) 13C metabolic flux analysis for larger scale cultivation using gas chromatography-combustion-isotope ratio mass spectrometry. Metab Eng 12:392–400
Zamboni N, Sauer U (2009) Novel biological insights through metabolomics and 13C-flux analysis. Curr Opin Microbiol 12:553–558
Zamboni N, Fischer E, Sauer U (2005) FiatFlux—a software for metabolic flux analysis from 13C-glucose experiments. BMC Bioinform 6:209
Zhang W, Li F, Nie L (2010) Integrating multiple ‘omics’ analysis for microbial biology: application and methodologies. Microbiology 156:287–301
Acknowledgments
All authors acknowledge financial support by the BMBF grant 0315784F within the research initiative “Systems biology of Microorganisms 2” as part of the research consortium BaCell-SysMo2. Christoph Wittmann and Judith Becker further acknowledge financial support by the BMBF-Grant “Biobased Polyamides through Fermentation” (No 0315239A) within the initiative Bioindustry21.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kohlstedt, M., Becker, J. & Wittmann, C. Metabolic fluxes and beyond—systems biology understanding and engineering of microbial metabolism. Appl Microbiol Biotechnol 88, 1065–1075 (2010). https://doi.org/10.1007/s00253-010-2854-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-010-2854-2