Abstract
Recent studies have suggested that a decrease in the specific activity of the 2-oxoglutarate dehydrogenase complex (ODHC) is important for glutamate overproduction by Corynebacterium glutamicum. To further investigate the role of the odhA gene and its product in this process, we constructed the recombinant strains of C. glutamicum in which the expression of the odhA and its product could be controlled by odhA overexpression and odhA antisense RNA expression. We examined changes in glutamate production and ODHC specific activity of the constructed strains during glutamate production triggered by Tween 40 addition. The ODHC specific activity increased with odhA overexpression, resulting in dramatically reduced glutamate production despite Tween 40 addition, indicating that a decrease in the specific activity of ODHC is required for glutamate production induced by Tween 40 addition. However, odhA antisense RNA expression alone did not result in glutamate overproduction in spite of the decrease in ODHC specific activity. Rather, it enhanced glutamate production triggered by Tween 40 addition due to the additional decrease in ODHC specific activity, suggesting that odhA antisense RNA expression is effective in enhancing Tween-40-triggered glutamate overproduction. Our results suggest that a change in ODHC specific activity is critical but is not the only factor responsible for glutamate overproduction by C. glutamicum.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
A coryneform bacterium, Corynebacterium glutamicum, is a Gram-positive and facultatively anaerobic bacterium with a high G + C content. It was originally isolated as a glutamate-producing bacterium (Kinoshita et al. 1957; Udaka 1960) and has been used for the industrial fermentation of various amino acids such as lysine and threonine (Nakayama et al. 1961; Shiio and Nakamori 1970). Recently, the genomic DNA sequences of C. glutamicum were determined by three independent research groups in Japan and Europe (Ikeda and Nakagawa 2003; Kalinowski et al. 2003; Yukawa et al. 2007).
Among those produced by microbial fermentation, glutamate is one of the prominent amino acids. Glutamate overproduction by C. glutamicum is induced by the depletion of biotin, which is an essential element for growth (Shiio et al. 1962); by the addition of fatty acid ester surfactants such as Tween 40 (polyoxyethylene sorbitan monopalmitate; Takinami et al. 1965); by the addition of penicillin, which inhibits peptidoglycan biosynthesis (Nara et al. 1964; Nunheimer et al. 1970); or by the addition of ethambutol, an antimycobacterial drug which inhibits the biosynthesis of mycolic acid in Mycobacteria (Radmacher et al. 2005). Because these triggers affect the cell surface structure of C. glutamicum, it was thought that glutamate passively leaks through the membrane and cell wall. In the late 1980s and 1990s, however, many researchers wondered “leak model” to explain glutamate secretion in C. glutamicum in terms of the material balance of extracellular and intracellular glutamate. However, leak model itself could not explain the high accumulation of extracellular glutamate. Therefore, many researchers (Hoischen and Krämer 1989; Gutmann et al. 1992) proposed the presence of a specific glutamate export system in the membrane, which differs from the earlier-mentioned leak model. Very recently, Nakamura et al. (2007) found a possible glutamate exporter that is coded by the NCgl1221 gene in C. glutamicum. During glutamate production, the membrane tension is altered, which triggers conformational change in the NCgl1221 protein and results in the export of glutamate. Furthermore, specific mutations in the NCgl1221 gene exhibit constitutive glutamate secretion without any triggers.
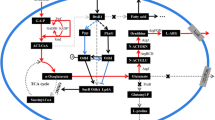
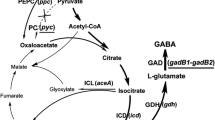
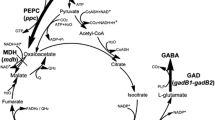
The 2-oxoglutarate dehydrogenase complex (ODHC) is a key enzyme in glutamate production, which is located at the branch point between the tricarboxylic acid (TCA) cycle and the glutamate biosynthesis pathway. ODHC consists of three subunits: 2-oxoglutarate dehydrogenase (E1o; EC 1.2.4.2), coded by odhA (Usuda et al. 1996); dihydrolipoamide S-succinyltransferase (E2; EC 2.3.1.61), coded by sucB (found from the genome sequencing); and dihydrolipoamide dehydrogenase (E3; EC 1.8.1.4), coded by lpd (Schwinde et al. 2001). Shingu and Terui (1971) and Kawahara et al. (1997) found that ODHC specific activity is dramatically reduced during glutamate overproduction. Moreover, metabolic flux analysis revealed that attenuation of ODHC activity has a great impact on glutamate production by C. glutamicum (Shimizu et al. 2003; Shirai et al. 2005). Asakura et al. (2007) reported that deletion of the odhA gene resulted in glutamate overproduction by C. glutamicum in the presence of excess biotin. These results suggest that decrease in ODHC specific activity is an important factor for producing glutamate by C. glutamicum, leading to increase in the metabolic flux for glutamate biosynthesis.
Recently, Niebisch et al. (2006) found that the phosphorylation state of a 15-kDa protein, OdhI, of C. glutamicum is responsible for the inhibition of ODHC specific activity by direct interaction with the E1o subunit of ODHC (OdhA). Moreover, deletion of the odhI gene abolishes glutamate overproduction by C. glutamicum (Schultz et al. 2007). These results suggest that the inhibition of ODHC specific activity by the OdhI protein might be one of the important factors in glutamate overproduction by C. glutamicum.
The antisense RNA technique is recognized as one of the effective methods for regulating gene expression at the posttranscriptional level. Almost all of the naturally occurring cases of antisense RNA have been found in bacteria (see Wagner and Simons 1994). It forms a complementary duplex between the antisense RNA and target messenger RNA (mRNA); thus, translation from the target mRNA is blocked. This technique can be applied in metabolic engineering to control the flow of specific metabolic pathways. Moreover, down-regulation of protein expression by antisense RNA offers ease of implementation and flexibility, which are not seen in gene deletion or knockout technologies. For example, studies have demonstrated the effectiveness of antisense RNA in the metabolic engineering of Clostridium acetobutylicum (Desai and Papoutsakis 1999; Tummala et al. 2003a; Tummala et al. 2003b).
In this study, to further investigate the role of the odhA gene and its product in glutamate overproduction by C. glutamicum, we examined the effect of modulation of ODHC specific activity by odhA overexpression and odhA antisense RNA expression on glutamate production by C. glutamicum. The relationship between changes in ODHC specific activity levels and glutamate production due to odhA overexpression or odhA antisense RNA expression was also investigated.
Materials and methods
Bacterial strains, plasmids, and media
All bacterial strains and plasmids used in this study are listed in Table 1. For the recombinant DNA techniques, Escherichia coli and C. glutamicum were cultivated in Lennox medium (1% polypeptone, 0.5% Bacto yeast extract, 0.5% NaCl, and 0.1% glucose; pH 7.2). To escape the restriction system of C. glutamicum, the plasmids obtained from the dam dcm mutant strains of E. coli SCS110 were used for the transformation (Vertès et al. 1993).
For glutamate production assay of C. glutamicum, two culture media were used. For seed preparation, a CM2B plate (1% polypeptone, 1% Bacto yeast extract, 0.5% NaCl, 10 μg/l d-biotin, and 1.5% agar; pH 7.2; Miwa et al. 1985) was used. For the preculture, the synthetic medium (Shirai et al. 2006) used was 80 g glucose, 30 g (NH4)2SO4, 3 g Na2HPO4, 6 g KH2PO4, 2 g NaCl, 3.9 mg FeCl3, 0.9 mg ZnSO4·7H2O, 0.3 mg CuCl2·2H2O, 5.56 mg MnSO4·5H2O, 0.1 mg (NH4)6Mo7O24·4H2O, 0.3 mg Na2B4O7·10H2O, 0.4 g MgSO4·7H2O, 40 mg FeSO4·7H2O, 84 mg CaCl2, 500 μg thiamin hydrochloride, 0.1 g ethylenediaminetetraacetic acid, and 10 μg biotin per liter of deionized water (pH 7.2). To avoid the decrease in pH throughout culture using Sakaguchi flask, 25 g/l of CaCO3 was added to the culture broth. The composition of the medium for the main culture was the same as that for the preculture except for the concentration of biotin (20 μg/l). When necessary, kanamycin (20 μg/ml for the E. coli strains and 10 μg/ml for the C. glutamicum strains) or ampicillin (50 μg/ml for E. coli) was added to the medium.
Construction of odhA gene and odhA antisense RNA expressing strains
The target regions for odhA overexpression and odhA antisense RNA expression were amplified by polymerase chain reaction (PCR) from the genomic DNA of C. glutamicum ATCC 13032 using Z-Taq DNA polymerase (Takara Bio, Shiga, Japan) and the primers shown in Table 2. The PCR products were cloned in a pGEM-T easy vector (Promega, Madison, WI, USA), and the sequences of the cloned PCR products were confirmed using a BigDye terminator cycle sequencing kit v1.1 (Applied Biosystems, Foster City, CA, USA) and an ABI PRISM 310 genetic analyzer (Applied Biosystems).
For the construction of the odhA overexpression plasmid pECt-odhA, the PCR product corresponding to the odhA gene on the pGEM-T easy vector was digested with PciI and BamHI and then cloned into the NcoI and BamHI sites of the E. coli–C. glutamicum shuttle expression plasmid pECt (Sato et al. 2008).
For the construction of the odhA antisense RNA expression plasmid, the PCR product on the pGEM-T easy vector was digested using appropriate restriction enzymes, as shown in Table 2, and then inserted into pECt in an antisense orientation relative to the open reading frame of the odhA gene including the coding regions and ribosome binding site. The resulting plasmids, pECt-odhA for odhA overexpression, pECt-Anti-odhA-1 for odhA antisense RNA expression, were introduced into the C. glutamicum wild-type strain ATCC 13032. In the odhA overexpressing and odhA antisense RNA expressing strain, expressions of the odhA gene and odhA antisense RNA cloned under the trc promoter on pECt were induced by addition of isopropyl-β-d-thiogalactoside (IPTG).
Measurements of cell growth and glucose and glutamate concentrations
Cell growth was monitored by measuring the optical density of the culture at 660 nm (OD660) using a spectrophotometer (U-2000; Hitachi High-Technologies, Tokyo, Japan) after dilution of the culture with 0.2 M HCl to dissolve CaCO3. Glucose and glutamate in the supernatant were measured using a biochemical analyzer (2700; YSI, Yellow Springs, OH, USA) and F-kit glutamate (R-Biopharm AG, Darmstadt, Germany), respectively.
Culture conditions for glutamate production assay
In this study, Sakaguchi flask culture was used for preliminary evaluation of glutamate production because of its easy operation. However, since C. glutamicum requires a lot of oxygen supply for its growth and glutamate production, Sakaguchi flask culture is not suitable for detailed evaluation of glutamate production by C. glutamicum because of the disadvantage in supplying oxygen into the culture broth. Moreover, a large amount of C. glutamicum cells was necessary for measurement of ODHC specific activity. Therefore, for further evaluation of effects of odhA overexpression and odhA antisense RNA expression on glutamate production and ODHC specific activity, we used a jar bioreactor, which enables to supply enough amount of oxygen to the culture by changing the agitation speed and to obtain a large amount of C. glutamicum cells.
The C. glutamicum strains carrying pECt, pECt-odhA, and pECt-Anti-odhA-1, respectively, were grown at 30°C for 24 h on a CM2B plate for seed preparation. For the preculture, cells on the CM2B plate were collected and inoculated into 40 ml of the synthetic medium in a Sakaguchi flask and then incubated aerobically at 31.5°C. Seventeen hours after incubation, 1 ml of this preculture was inoculated into 40 ml of the synthetic medium in a Sakaguchi flask for glutamate production assay, and the cells were cultivated for 38 h at 31.5°C.
In jar bioreactor culture, 80 ml of the preculture was inoculated into 2 l of the synthetic medium in a 5-l jar bioreactor (KMJ-5B; Mitsuwa Rikagaku, Osaka, Japan), and the cells were aerobically cultivated at 31.5°C. The agitation speed was changed to allow enough oxygenation, and the airflow rate was maintained at 2 l/min. The pH was maintained at 7.2 by automatic addition of 14% (v/v) ammonia solution, which was also supplied as a nitrogen source. Foaming was controlled by the addition of an antifoaming agent (Disfoam GD-113K; NOF, Japan). After cell growth reached the early exponential phase (OD660 = 12), Tween 40 was added to the medium in order to achieve the final concentrations of 4 mg/ml for triggering glutamate production in both flask and jar bioreactor. Expressions of the odhA gene and odhA antisense RNA were induced by IPTG addition into the culture media, at the appropriate time, at a final concentration of 0.1 mM for odhA overexpression and 1 mM for odhA antisense RNA expression, respectively.
Measurement of ODHC specific activity
ODHC specific activity was measured according to the method described by Shiio and Ujigawa-Takeda (1980). C. glutamicum cells were harvested, washed twice with 0.2% KCl, and suspended in 3 ml of 0.1 M N-tris(hydroxymethyl) methyl-2-aminoethanesulfonic acid (TES)·NaOH buffer (pH 7.7) containing 30% (v/v) glycerol. Cells were disrupted by sonication and then centrifuged to remove the cell debris. One milliliter of the supernatant, applied to the gel filtration column (PD-10 Desalting Column; GE Healthcare UK, Buckinghamshire, UK) to remove low-molecular-weight compounds, was used for the enzyme assay.
The reaction mixture was prepared as follows: 100 mM TES·NaOH (pH 7.7), 5 mM MgCl2, 3 mM cysteine, 0.3 mM thiamine pyrophosphate, 0.2 mM coenzyme A, 1 mM 3-acetylpyridine adenine dinucleotide (APAD+) instead of oxidized form of nicotinamide adenine dinucleotide (NAD+), and 150 μl of the cell extract. After addition of 1 mM 2-oxoglutarate to the reaction mixture, the initial increase in the absorbance of APADH at 365 nm was measured at 31.5°C for 3 min with 15-s intervals. ODHC specific activity was defined as the amount of enzyme required for the conversion of an equivalent of 1 μmol of NAD+ into reduced form of nicotinamide adenine dinucleotide (NADH) in 1 min. The concentration of total protein in the crude extract was determined by Bradford’s method (Bradford 1976) using a Bio-Rad protein assay kit (Bio-Rad Laboratories, Hercules, CA, USA).
Results
Verification of OdhA protein overexpression in odhA overexpressing strain of C. glutamicum
Overexpression of the OdhA protein in C. glutamicum ATCC 13032 carrying pECt-odhA was verified by 7.5% sodium dodecyl sulfate (SDS)-polyacrylamide gel electrophoresis. Cells of the ATCC 13032 strains carrying pECt-odhA and pECt, respectively, were cultured at 30°C and harvested 2, 4, and 6 h after IPTG addition. Proteins were obtained from the harvested cells by sonication and then separated by SDS-polyacrylamide gel.
As shown in Fig. 1, the protein band with a molecular mass of about 140 kDa, corresponding to the OdhA protein, was detected in ATCC 13032/pECt-odhA but not in ATCC 13032/pECt after visualization with Coomassie Brilliant Blue staining; therefore, OdhA overexpression was successfully exhibited in the odhA overexpressing strain.
Confirmation of OdhA overexpression in the odhA overexpressing strain. All the cellular proteins from the ATCC 13032/pECt-odhA and ATCC 13032/pECt strains separated by 7.5% SDS-polyacrylamide gel electrophoresis are shown. Lane 1 proteins from the cells of ATCC 13032/pECt-odhA before IPTG addition; lanes 2–4 proteins obtained from the cells of ATCC 13032/pECt-odhA 2, 4, and 6 h after 0.1 mM IPTG addition, respectively; lane 5 proteins obtained from the cells of ATCC 13032/pECt 6 h after IPTG addition; arrow position corresponding to the OdhA protein (138.8 kDa)
Effect of odhA overexpression on glutamate production triggered by Tween 40 addition
To investigate the effect of odhA overexpression on glutamate production, jar bioreactor cultures of ATCC 13032/pECt-odhA and ATCC 13032/pECt were carried out under Tween-40-triggered glutamate production. When cell growth reached the early exponential phase (OD660 = 12), Tween 40 for triggering glutamate production and IPTG for odhA overexpression were simultaneously added to the media.
As shown in Fig. 2, cell growth was reduced, and glutamate was produced in both ATCC 13032/pECt-odhA and ATCC 13032/pECt by Tween 40 addition. However, in the presence of IPTG, the ATCC 13032/pECt-odhA strain showed significantly low glutamate production (less than 4 g/l) and the increase in ODHC specific activity in spite of Tween 40 addition; on the other hand, the control strain ATCC 13032/ pECt showed decreased ODHC specific activity and glutamate production (about 18 g/l). These results indicate that the increase in ODHC specific activity by odhA overexpression reduces glutamate production and supports the view (see Shingu and Terui 1971; Kawahara et al. 1997; Shimizu et al. 2003; Shirai et al. 2005) that decrease in ODHC specific activity is an important factor for glutamate overproduction by C. glutamicum.
Cell growth (circles), glucose consumption (squares), glutamate production (triangles), and relative ODHC specific activity (diamonds) of the odhA overexpressing strain ATCC 13032/pECt-odhA (closed symbols) and the control strain ATCC 13032/pECt (open symbols) under Tween-40-triggered glutamate production in jar bioreactor culture. Arrows indicate both IPTG and Tween 40 addition for odhA overexpression and triggering glutamate overproduction, respectively
Effect of odhA antisense RNA expression on glutamate production in flask culture
It was assumed that controlling the expression level of the OdhA protein by odhA antisense RNA expression leads to glutamate production and understanding the glutamate production mechanism in C. glutamicum. Therefore, we developed C. glutamicum strain carrying the odhA antisense RNA expression plasmid pECt-Anti-odhA-1 and examined glutamate production by this strain. The target region for the antisense RNA on the pECt-Anti-odhA-1 includes the ribosome binding site and complete coding region of odhA gene. The odhA antisense RNA expression was induced by 1 mM IPTG addition at the early exponential growth phase.
In the flask culture, neither was cell growth and glucose consumption of C. glutamicum affected, nor was there any significant increase in glutamate production by odhA antisense RNA expression only in the ATCC 13032/pECt-Anti-odhA-1 strain (Fig. 3). Under Tween 40 addition condition, cell growth of the odhA antisense RNA expressing strain with IPTG addition was similar to that without IPTG addition; exponential growth was initially observed, and then the cell growth was reduced after Tween 40 addition (Fig. 3a). As expected, Tween 40 addition induced glutamate production in the odhA antisense RNA expressing strain (Fig. 3c). Moreover, the level of Tween-40-triggered glutamate production with IPTG addition was higher than that with no IPTG added (Fig. 3c). These results indicate that the expression of odhA antisense RNA can further enhance Tween-40-triggered glutamate production by C. glutamicum.
Cell growth (a), glucose consumption (b), and glutamate production (c) of the odhA antisense RNA expressing strain ATCC 13032/pECt-Anti-odhA-1 in Sakaguchi flask culture. The representative results of three independent experiments under each culture condition are shown. Arrows Tween 40 and IPTG addition, open circles neither IPTG nor Tween 40 addition, closed circles IPTG addition, open triangles Tween 40 addition, closed triangles both Tween 40 and IPTG addition
Effect of odhA antisense RNA expression on glutamate production without any triggers in jar bioreactor cultures
Decrease in ODHC specific activity in the TCA cycle is one of the important factors for producing glutamate (Shingu and Terui 1971; Kawahara et al. 1997). To further investigate the effect of odhA antisense RNA expression on glutamate production and ODHC specific activity in C. glutamicum without any triggers, the odhA antisense RNA expressing strain was cultivated in a jar bioreactor.
As shown in Fig. 4, ODHC specific activity in this strain in the presence of IPTG (i.e., odhA antisense RNA expressing condition) was lower than that in the absence of IPTG, suggesting that odhA antisense RNA expression can indeed decrease in ODHC specific activity. However, no significant glutamate production was achieved by expression of odhA antisense RNA only (less than 1 g/l). In addition, in the odhA antisense RNA expressing strain, cell density became significantly high (OD660 = over 140) in a short time, and glucose consumption was also high in both the absence and presence of IPTG (Fig. 4). From the mid-exponential to the early stationary phases, cell density and glucose consumption in the presence of IPTG were slightly lower than those in the absence of IPTG. It was assumed that the decreased cell density and glucose consumption by IPTG addition results from the malfunction of TCA cycle due to the decrease in ODHC specific activity, leading to the decreased cell growth by expression of odhA antisense RNA. These results suggest that expression of odhA antisense RNA constructed in this study alone is able to decrease ODHC specific activity, leading to slight decrease in cell growth and substrate consumption; however, it does not directly lead to glutamate overproduction by C. glutamicum. This means that decrease in ODHC specific activity might not be the only factor in glutamate overproduction by C. glutamicum.
Cell growth (circles), glucose consumption (squares), glutamate production (triangles), and relative ODHC specific activity (diamonds) of the odhA antisense RNA expressing strain ATCC 13032/pECt-Anti-odhA-1 by odhA antisense RNA expression without any triggers for glutamate overproduction in jar bioreactor culture. Arrows IPTG addition for odhA antisense RNA expression, open symbols without IPTG addition, closed symbols with IPTG addition
Effect of odhA antisense RNA expression on glutamate production triggered by Tween 40 addition
For further investigation on the effect of odhA antisense RNA expression under Tween-40-triggered glutamate production, jar bioreactor cultures of the odhA antisense RNA expressing strain was also carried out.
Figure 5 shows the time course for cell growth, glucose consumption, glutamate production, and ODHC specific activity of the odhA antisense RNA expressing strain under Tween-40-triggered glutamate production. Similar to results obtained by the Sakaguchi flask culture experiments (Fig. 3), Tween 40 addition resulted in reduced cell growth and glutamate overproduction; Tween-40-triggered glutamate production was enhanced in the ATCC 13032/pECt-Anti-odhA-1 strain by IPTG-induced odhA antisense RNA expression as shown in Fig. 5. Moreover, ODHC specific activity in the ATCC 13032/pECt-Anti-odhA-1 was further lowered by odhA antisense RNA expression under Tween-40-triggered glutamate production, leading to enhanced glutamate production in jar bioreactor culture (Fig. 5).
Cell growth (circles), glucose consumption (squares), glutamate production (triangles), and relative ODHC specific activity (diamonds) of the odhA antisense RNA expressing strain ATCC 13032/pECt-Anti-odhA-1 under Tween-40-triggered glutamate production in jar bioreactor culture. Average with standard deviation in three independent experiments is shown. Arrows Tween 40 and/or IPTG addition, open symbols with Tween 40 addition only, closed symbols with both Tween 40 and IPTG addition
Our results shown in this paper suggest the expression of odhA antisense RNA is effective for the enhancement of glutamate production triggered by Tween 40 addition.
Discussion
It has already been demonstrated that decrease in ODHC specific activity is one of the key factors in glutamate overproduction by C. glutamicum, which is triggered by biotin depletion, penicillin addition, or Tween 40 addition (Shingu and Terui 1971; Kawahara et al. 1997). Therefore, we focused on the relationship between change in ODHC specific activity and glutamate production triggered by Tween 40 addition. In this study, the effects of odhA overexpression and the odhA antisense RNA expression were analyzed to further investigate the role of the odhA gene and its product in glutamate production by C. glutamicum.
First, the odhA overexpressing strain was constructed by introducing a plasmid pECt-odhA carrying the C. glutamicum odhA gene cloned under trc promoter on E. coli–C. glutamicum shuttle expression plasmid pECt (Sato et al. 2008). As expected, ODHC specific activity was increased, and glutamate production was concomitantly decreased by the odhA overexpression under Tween-40-triggered glutamate production condition (Fig. 2). This result supports the view that the decrease in ODHC specific activity is important for glutamate production by C. glutamicum (see “Introduction”). In spite of the increase in ODHC specific activity and low glutamate production, the cell growth and glucose consumption in odhA-overexpressing strain was lower than those in Tween-40-treated ATCC 13032/pECt strain. Since ODHC catalyzes the formation of NADH as well as succinyl-CoA, ODHC activity due to overexpression of odhA may increase the level of these metabolites. The relationship among changes in the balance of metabolites level, redox, and cell growth should be carefully examined in the future.
Asakura et al. (2007) reported that the odhA deletion strain could efficiently produce glutamate without any triggers; however, this strain showed a severe growth defect under the aerobic growth conditions needed the odhA gene in C. glutamicum. To improve cell growth, mutant strains carrying the missense mutation in the odhA gene were constructed using PCR-based in vitro mutagenesis (Nakamura et al. 2006). Although these mutants could produce high amounts of glutamate with less than half of the ODHC specific activity of the wild-type strain in the presence of excess biotin, they also showed a moderate growth defect. Therefore, odhA deletion strains might not be suitable for industrial glutamate production. As an alternative method for down-regulation of the expression of the odhA gene and its product, antisense RNA technology was considered for this study. This technology shows promise for application in industrial production processes. Among several antisense RNA methods such as antisense oligonucleotides, antisense RNA expression, and small interfering RNA (Lee and Roth 2003), the antisense RNA expression system was used because of its advantages: it is possible to control antisense RNA expression in trans at a certain time according to the requirements.
In this study, we attempted to decrease the ODHC specific activity through odhA antisense RNA expression. It was expected that the expressed odhA antisense RNA hybridizes to the complementary sequence of the target odhA mRNA and leads to translation stoppage from the odhA mRNA, resulting in decrease in ODHC specific activity. To evaluate the effect of odhA antisense RNA expression on glutamate production, the C. glutamicum strain carrying the odhA antisense RNA expression plasmid was constructed and cultivated by both Sakaguchi flask and jar bioreactor methods. It has been assumed that decrease in ODHC specific activity leads to malfunction of TCA cycle, and as a result, cell growth and substrate uptake decrease and probably glutamate production is induced. As shown in Fig. 4, we could confirm that odhA antisense RNA expression could decrease ODHC specific activity. Although the level of decrease in ODHC specific activity in the odhA antisense RNA-expressing strain was similar to those in the ATCC 13032/pECt treated with Tween 40, cell growth and glucose consumption in the odhA antisense RNA-expressing strain became higher than those in the Tween-40-treated ATCC 13032/pECt strain (Figs. 2 and 4). As is well known, Tween 40 affects the biosynthesis of fatty acids, particularly mycolic acid that is one of the components in cell wall structure of C. glutamicum. Therefore, the cell growth and glucose consumption might be decreased in the Tween-40-treated C. glutamicum strain due to the malfunction of TCA cycle and fatty acid biosynthesis.
Moreover, odhA antisense RNA expression alone could not achieve glutamate production (Fig. 4), but it could enhance glutamate production triggered by Tween 40 addition (Fig. 5). No glutamate production in the odhA antisense RNA-expressing strain without triggers suggests that not only decrease in ODHC specific activity but other factors as well might contribute to glutamate production by C. glutamicum. It can be thought that other factor(s) triggered by Tween 40 addition is important for glutamate overproduction in the odhA antisense RNA-expressing strain. Therefore, other factors have to be considered to explain the precise mechanism of glutamate overproduction by C. glutamicum. Niebisch et al. (2006) identified the OdhI protein that directly binds to the OdhA protein (one of the subunits of ODHC) and inhibits ODHC activity. The phosphorylation state of OdhI is controlled by serine/threonine protein kinase G (PknG) and the phosphatase (Ppp; Schultz et al. 2007). The unphosphorylated OdhI can bind to the OdhA protein and thus inhibit ODHC activity. Moreover, it was shown that the deletion strain of the odhI gene impaired glutamate production under biotin depletion, penicillin addition, and Tween 40 addition, indicating that inhibition of ODHC activity by the OdhI protein is essential for glutamate production (Schultz et al. 2007). Further experiments need to be performed to elucidate the precise mechanism of glutamate overproduction by C. glutamicum with regard to the phosphorylation of the OdhI protein in the odhA antisense RNA expressing strain. In addition, it is also important to investigate the other factor(s) necessary for glutamate production by odhA antisense RNA-expressing C. glutamicum related to cell membrane or transport/export systems, such as change in the level of intracellular metabolites, and structural proteins in the cell membrane concerning export systems, which were reported by Nakamura et al. (2007), necessary for glutamate production by C. glutamicum.
References
Asakura Y, Kimura E, Usuda Y, Kawahara Y, Matsui K, Osumi T, Nakamatsu T (2007) Altered metabolic flux due to deletion of odhA causes l-glutamate overproduction in Corynebacterium glutamicum. Appl Environ Microbiol 73:1308–1319
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Desai RP, Papoutsakis ET (1999) Antisense RNA strategies for the metabolic engineering of Clostridium acetobutylicum. Appl Environ Microbiol 65:936–945
Gutmann M, Hoischen C, Krämer R (1992) Carrier-mediated glutamate secretion by Corynebacterium glutamicum under biotin limitation. Biochim Biophys Acta 1112:115–123
Hoischen C, Krämer R (1989) Evidence for an efflux carrier system involved in the secretion of glutamate by Corynebacterium glutamicum. Arch Microbiol 151:342–347
Ikeda M, Nakagawa S (2003) The Corynebacterium glutamicum genome: features and impacts on biotechnological processes. Appl Microbiol Biotechnol 62:99–109
Kalinowski J, Bathe B, Bartels D, Bischoff N, Bott M, Burkovski A, Dusch N, Eggeling L, Eikmanns BJ, Gaigalat L, Goesmann A, Hartmann M, Huthmacher K, Krämer R, Linke B, McHardy AC, Meyer F, Möckel B, Pfefferle W, Pühler A, Rey DA, Rückert C, Rupp O, Sahm H, Wendisch VF, Wiegräbe I, Tauch A (2003) The complete Corynebacterium glutamicum ATCC 13032 genome sequence and its impact on the production of l-aspartate-derived amino acids and vitamins. J Biotechnol 104:5–25
Kawahara Y, Takahashi-Fuke K, Shimizu E, Nakamatsu T, Nakamori S (1997) Relationship between the glutamate production and the activity of 2-oxoglutarate dehydrogenase in Brevibacterium lactofermentum. Biosci Biotechnol Biochem 61:1109–1112
Kinoshita S, Udaka S, Shimono M (1957) Studies on the amino acid fermentation. Part 1. Production of l-glutamic acid by various microorganisms. J Gen Appl Microbiol 3:193–205
Lee LK, Roth CM (2003) Antisense technology in molecular and cellular bioengineering. Curr Opin Biotechnol 14:505–511
Miwa K, Matsui K, Terabe M, Ito K, Ishida M, Takagi H, Nakamori S, Sano K (1985) Construction of novel shuttle vectors and a cosmid vector for the glutamic acid-producing bacteria Brevibacterium lactofermentum and Corynebacterium glutamicum. Gene 39:281–286
Nakamura J, Hirano S, Yamaguchi M, Ito H (2006) Japan Patent 2006-101875
Nakamura J, Hirano S, Ito H, Wachi M (2007) Mutations of the Corynebacterium glutamicum NCgl1221 gene, encoding a mechanosensitive channel homolog, induce l-glutamic acid production. Appl Environ Microbiol 73:4491–4498
Nakayama K, Kitada S, Kinoshita S (1961) Studies on lysine fermentation I. The control mechanism of lysine accumulation by homoserine and threonine. J Gen Appl Microbiol 7:145–154
Nara T, Samejima H, Kinoshita S (1964) Effect of penicillin on amino acid fermentation. Agric Biol Chem 28:120–124
Niebisch A, Kabus A, Schultz C, Weil B, Bott M (2006) Corynebacterial protein kinase G controls 2-oxoglutarate dehydrogenase activity via the phosphorylation status of the OdhI protein. J Biol Chem 281:12300–12307
Nunheimer TD, Birnbaum J, Ihnen ED, Demain AL (1970) Product inhibition of the fermentative formation of glutamic acid. Appl Microbiol 20:215–217
Radmacher E, Stansen KC, Besra GS, Alderwick LJ, Maughan WN, Hollweg G, Sahm H, Wendisch VF, Eggeling L (2005) Ethambutol, a cell wall inhibitor of Mycobacterium tuberculosis, elicits l-glutamate efflux of Corynebacterium glutamicum. Microbiology 151:1359–1368
Sato H, Orishimo K, Shirai T, Hirasawa T, Nagahisa K, Shimizu H, Wachi M (2008) Distinct roles of two anaplerotic pathways in glutamate production induced by biotin limitation in Corynebacterium glutamicum. J Biosci Bioeng. 106:51–58
Schultz C, Niebisch A, Gebel L, Bott M (2007) Glutamate production by Corynebacterium glutamicum: dependence on the oxoglutarate dehydrogenase inhibitor protein OdhI and protein kinase PknG. Appl Microbiol Biotechnol 76:691–700
Schwinde JW, Hertz PF, Sahm H, Eikmanns BJ, Guyonvarch A (2001) Lipoamide dehydrogenase from Corynebacterium glutamicum: molecular and physiological analysis of the lpd gene and characterization of the enzyme. Microbiology 147:2223–2231
Shiio I, Nakamori S (1970) Microbial production of l-threonine. Part II. Production by α-amino-β-hydroxyvaleric acid resistant mutants of glutamate producing bacteria. Agric Biol Chem 34:448–456
Shiio I, Ujigawa-Takeda K (1980) Presence and regulation of ioketoglutarate dehydrogenase complex in a glutamate producing bacterium, Brevibacterium flavum. Agric Biol Chem 44:1897–1904
Shiio I, Otsuka SI, Takahashi M (1962) Effect of biotin on the bacterial formation of glutamic acid. Part I. Glutamate formation and cellular permeability of amino acids. J Biochem 51:56–62
Shimizu H, Tanaka H, Nakato A, Nagahisa K, Kimura E, Shioya S (2003) Effects of the changes in enzyme activities on metabolic flux redistribution around the 2-oxoglutarate branch in glutamate production by Corynebacterium glutamicum. Bioprocess Biosyst Eng 25:291–298
Shingu H, Terui G (1971) Studies on process of glutamic acid fermentation at the enzyme level. Part I. On the change of α-ketoglutaric acid dehydrogenase in the course of culture. J Ferment Technol 49:400–405
Shirai T, Nakato A, Izutani N, Nagahisa K, Shioya S, Kimura E, Kawarabayasi Y, Yamagishi A, Gojobori T, Shimizu H (2005) Comparative study of flux redistribution of metabolic pathway in glutamate production by two coryneform bacteria. Metab Eng 7:59–69
Shirai T, Matsuzaki K, Kuzumoto M, Nagahisa K, Furusawa C, Shioya S, Shimizu H (2006) Precise metabolic flux analysis of coryneform bacteria by gas chromatography-mass spectrometry and verification by nuclear magnetic resonance. J Biosci Bioeng 102:413–424
Takinami K, Yoshii H, Tsuri H, Okada H (1965) Biochemical effects of fatty acid and its derivatives on l-glutamic acid fermentation. Part III. Biotin-Tween 60 relationship in the accumulation of l-glutamic acid and the growth of Brevibacterium lactofermentum. Agric Biol Chem 29:351–359
Tummala SB, Junne SG, Papoutsakis ET (2003a) Antisense RNA downregulation of coenzyme A transferase combined with alcohol-aldehyde dehydrogenase overexpression leads to predominantly alcohologenic Clostridium acetobutylicum fermentations. J Bacteriol 185:3644–3653
Tummala SB, Welker NE, Papoutsakis ET (2003b) Design of antisense RNA constructs for downregulation of the acetone formation pathway of Clostridium acetobutylicum. J Bacteriol 185:1923–1934
Udaka S (1960) Screening method for microorganisms accumulating metabolites and its use in the isolation of Micrococcus glutamicus. J Bacteriol 79:754–755
Usuda Y, Tujimoto N, Abe C, Asakura Y, Kimura E, Kawahara Y, Kurahashi O, Matsui H (1996) Molecular cloning of the Corynebacterium glutamicum (‘Brevibacterium lactofermentum’ AJ12036) odhA gene encoding a novel type of 2-oxoglutarate dehydrogenase. Microbiology 142:3347–3354
Vertès AA, Inui M, Kobayashi M, Kurusu Y, Yukawa H (1993) Presence of mrr- and mcr-like restriction systems in coryneform bacteria. Res Microbiol 144:181–185
Wagner EG, Simons RW (1994) Antisense RNA control in bacteria, phages, and plasmids. Annu Rev Microbiol 48:713–742
Westmoreland J, Porter G, Radman M, Resnick MA (1997) Highly mismatched molecules resembling recombination intermediates efficiently transform mismatch repair proficient E. coli. Genetics 145:29–38
Yanisch-Perron C, Vieira J, Messing J (1985) Improved M13 phage cloning vectors and host strains: nucleotide sequences of the M13mp18 and pUC19 vectors. Gene 33:103–119
Yukawa H, Omumasaba CA, Nonaka H, Kós P, Okai N, Suzuki N, Suda M, Tsuge Y, Watanabe J, Ikeda Y, Vèrtes AA, Inui M (2007) Comparative analysis of the Corynebacterium glutamicum group and complete genome sequence of strain R. Microbiology 153:1042–1058
Acknowledgments
This work was supported by Grant-in-Aids for Young Scientists (B) to TH (19780061) and CF (20700270), respectively, from the Ministry of Education, Culture, Sports, Science, and Technology of Japan. This work was also partly supported by the Global COE program and the Special Coordination Funds for Promoting Science and Technology: Yuragi Project of the Ministry of Education, Culture, Sports, Science, and Technology of Japan.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kim, J., Hirasawa, T., Sato, Y. et al. Effect of odhA overexpression and odhA antisense RNA expression on Tween-40-triggered glutamate production by Corynebacterium glutamicum . Appl Microbiol Biotechnol 81, 1097–1106 (2009). https://doi.org/10.1007/s00253-008-1743-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-008-1743-4