Abstract
Plant defensins are small, highly stable, cysteine-rich antimicrobial peptides produced by the plants for inhibiting a broad-spectrum of microbial pathogens. Some of the well-characterized plant defensins exhibit potent antifungal activity on certain pathogenic fungal species only. We characterized a defensin, TvD1 from a weedy leguminous herb, Tephrosia villosa. The open reading frame of the cDNA was 228 bp, which codes for a peptide with 75 amino acids. Expression analyses indicated that this defensin is expressed constitutively in T. villosa with leaf, stem, root, and seed showing almost similar levels of high expression. The recombinant peptide (rTvD1), expressed in the Escherichia coli expression system, exhibited potent in vitro antifungal activity against several filamentous soil-borne fungal pathogens. The purified peptide also showed significant inhibition of root elongation in Arabidopsis seedlings, subsequently affecting the extension of growing root hairs indicating that it has the potential to disturb the plant growth and development.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Defensins represent a defense strategy in plants against invading pathogens, particularly in protein rich leguminous species (Osborn et al. 1995). The characteristic feature of plant defensins is the presence of four disulfide bridges between eight cysteine residues within the mature peptide. Generally, defensins have a net positive charge (Broekaert et al. 1995) and are implicated in the front line of host defense (Thomma et al. 2002). The expression of plant defensin is sometimes tissue specific with varied levels in different tissues like HaDef1in the roots of Helianthus spp. (Zélicourt et al. 2007); SD2 and SF18, in contrast, are localized in the inflorescences of the same plant (Urdangarín et al. 2000). Most of the defensins are seed specific, such as Pa-AMP of pokeweed (Liu et al. 2000), Mj-AMP-1 and Mj-AMP-2 of Mirabilis jalapa (De Bolle et al. 1995), SPE10 of Pachyrrhizus erosus (Song et al. 2005), Rs-AFP1 and Rs-AFP2 of Raphanus sativus (Terras et al. 1992). Some defensins like Rs-AFP3 and Rs-AFP4 together with Rs-AFP1 and Rs-AFP2, and Lm-def are expressed in leaves of R. sativus (Terras et al. 1995) and Lepidium meyenii (Solis et al. 2007), respectively. Arabidopsis plants express a leaf-specific defensin, PDF1.2 upon pathogen challenge (Thomma and Broekaert 1998).
It has also been demonstrated that defensins are induced by external stimuli such as biotic and abiotic stress signals including pathogen infection and methyl jasmonate treatment (Manners et al. 1998). In addition, some of the plant defensins are induced by environmental stimuli; cold in winter wheat induces the expression of defensin TAD1 (Koike et al. 2002) and the infestation of the parasitic plant Orobanche spp upregulates the gene for HaDef1 defensin in the roots of Helianthus spp (Zélicourt et al. 2007).
The mode of action of defensins is still not clear. Generally, leguminous defensins do not induce any morphological change, but the defensins of other families like Brassicaceae and Saxifragaceae may induce hyperbranching (Osborn et al. 1995). Defensins like MsDef1 inhibit the growth by blocking the mammalian L-type calcium ion channels, which are dependent on a calcium gradient. Although structurally similar to MsDef1, RsAFP2, and MtDef2 do not block the L-type calcium channels (Spelbrink et al. 2004). Defensins like DmAMP1 bind on the site M(IP)2C, which are present as patches on the hyphal membranes or alternatively required for anchoring of other membrane or cell-wall associated proteins, which themselves interact with DmAMP1 (Thevissen et al. 2000). Some similar antifungal peptides bind to the N-acetyl glucosylceramides present on the fungal plasma membrane (Thevissen et al. 2004). Fungal mutants lacking glucosylceramides are not sensitive to RsAFP-2. Recently, it was reported that plant defensins have roles in regulating plant growth and development. Defensin like MsDef1, MtDef2, and RsAFP2 inhibit the root growth in the germinating Arabidopsis at micromolar concentrations (Allen et al. 2007). In our reports also, we suggest the root growth inhibitory action of the defensin at the same concentration required for antifungal activity. Most of the bacterially expressed plant defensins were shown to be active as antifungal peptides at concentrations ranging from 100–200 μg/ml (Xu and Reddy 1997; Park et al. 2002; Olli and Kirti 2006; Hui et al. 2007; Olli et al. 2007; Solis et al. 2007).
The cultivated peanuts suffer seriously from the attack by the late leaf spot pathogen, Pheaoisariopsis personata in many agro-climatic zones of the world and conventional crop improvement programs could not, so far, offer effective solutions to this problem. A defensin that could fight the late leaf spot infection in peanut is highly pertinent in plant biotechnology.
In the present study, we have observed the potent antifungal activity of the constitutively expressed defensin TvD1 purified from the Escherichia coli expression system against the target pathogen, P. personata and several other soil-borne fungal pathogens, which cause economically significant diseases such as Fusarium oxysporum f. sp. vasinfectum (vascular wilt of tomato), F. moniliforme (root rot of rice), Rhizoctonia solani (sheath blight of rice), Phytophthora parasitica f. sp. nicotianae (leaf rot of tobacco), Curvularia spp (leaf spots), Botrytis cinerea (grape rot) and Alternaria helianthi (black or gray leaf spot of sunflower). These observations are reported in this communication.
Materials and methods
Nucleic acid extraction, PCR amplification, and cloning
Genomic DNA was isolated from freshly frozen leaves of Tephrosia villosa plants collected from the field using the standard CTAB method (Doyle and Doyle 1990). Total RNA was also isolated from the leaves without any stress or pathogen treatment using Tri-reagent (Sigma-Aldrich, USA). Polymerase chain reaction was carried out for amplifying the genomic clone and the open reading frame (ORF) of the cDNA of TvD1 using primers designed on the basis of an alfalfa defensin (MsDef1). The primers used were forward: 5′GGGTACCATGGAGAAGAAATCACTAGC 3′ and reverse: 5′ GGGATCCTTTAACATCTTTTAGTACACCA 3′. KpnI and BamHI sites were included (italicized) for future cloning steps in the desired vectors. For RT-PCR, the first strand of the cDNA was synthesized by reverse transcription with an oligo-(dT) primer and M-MLV reverse transcriptase enzyme (Sigma-Aldrich, USA). Amplification conditions for PCR were: 94°C for 4 min, 35 cycles: 94°C for 1 min, 58°C for 55 s, 72°C for 1 min and final extension was 72°C for 10 min.
The gel-eluted (Gel Clean-up Kit, Eppendorf, Germany) amplification products were cloned into the T/A cloning vector, pTZ57R (MBI Fermentas) and sequenced. Nucleotide and deduced amino acid sequence comparisons were made using the BLAST (Basic Local Alignment Search Tool) programs such as BLASTN (Zhang et al. 2000) and BLASTX (Altschul et al. 1997), respectively, on the non-redundant database of the National Center for Biotechnology Information (http://www.ncbi.nlm.nih.gov/BLAST/). Expasy tool was used to deduce the amino acid sequence coded by the gene TvD1.
Expression and purification of Tephrosia defensin protein
The coding region of the TvD1 gene was cloned between KpnI and BamHI sites in pET32a vector (Novagen, USA). The cloned vector was transformed into E. coli BL21 (DE3) pLys S and the transformed cells were grown at 37°C in LB Broth (Himedia, India) overnight at 200 rpm in a shaker. A 10-ml aliquot from the overnight culture was added into 1 l of fresh LB medium for 1 h till the OD600 reached 0.6–0.8. These cells were induced using 0.4 mM IPTG (Fermentas, Germany). After 6 h incubation, the cells were harvested by centrifugation at 5,000 rpm for 5 min at 4°C. The pellet was washed with fresh 1× PBS buffer. Subsequently, the first lysis (prolysis) buffer (300 mM NaCl, 50 mM NaHPO4, 10 mM imidazole, 15% glycerol, and 0.1% SDS) was added and mixed homogeneously. The cells were lysed by sonication at 6× pulse for 5 min at 4°C (every 20-s pulse, a 10-s interval was given). The lysate was centrifuged at 12,000 rpm for 20 min to remove the debris. The supernatant was mixed with the second lysis buffer (300 mM NaCl, 50 mM NaHPO4, and 10 mM imidazole) along with 2 ml Ni-NTA (Qiagen, Germany), and the volume was made up to 50 ml/l culture of cells and mixed for 1 h at 4°C at 60 rpm in a rocker. The supernatant mixture was loaded on to the column (Sigma-Aldrich, USA) and the flow-through was collected at the rate of 0.5 ml/min, which was maintained till the end. Finally, the column was washed with ten volumes of wash buffer (300 mM NaCl, 50 mM NaHPO4, 20 mM imidazole, and 15% glycerol). The recombinant peptide rTvD1, was eluted with elution buffer (300 mM NaCl, 50 mM NaHPO4, and 250 mM imidazole), according to the manufacturer’s instructions (Qiagen, Germany).
Antifungal activity assay
Antifungal activity of rTvD1 was tested by micro-spectrophotometry as well as an in vitro plate assay with the following fungal pathogens: F. oxysporum f. sp. vasinfectum, F. moniliforme, R. solani, A. helianthi, P. parasitica f. sp. nicotiana, B. cinerea, and Curvularia sp.
Briefly, 10 μl of protein diluted to different concentrations was pipetted into a well of a 96-well microtiter plate containing 90 μl of the test fungal spore suspension (~2.5 × 104 spores/ml) in potato dextrose broth (PDB), which was placed in an incubator at 28°C. Each protein concentration was tested for its antifungal activity in triplicate. Fungal spore germination and growth were observed microscopically and the optical density (OD) was measured with a microplate reader at a wavelength of 595 nm after inoculation for 30 min and 48 h. Controls were tested identically except that the protein was omitted. Value of the growth inhibition lower than 10% was not considered as significant (growth inhibition is defined as the ratio of the corrected absorbance at 595 nm of the control minus the corrected absorbance of the test sample, divided by the corrected absorbance of the control). The corrected absorbance is defined as the absorbance at 48 hr minus that at 30 min. IC50 is defined as the protein concentration at which 50% inhibitions was reached). Percent inhibition of fungal growth on the plate has been estimated as: area of mycelial growth in the absence of antifungal protein minus area of mycelial growth in the presence of antifungal protein / area of mycelial colony in absence of antifungal protein) × 100.
A graph plotting percent inhibition of fungal growth against the concentration of protein was used to determine the IC50 of antifungal activity, i.e., the concentration producing 50% inhibition.
For the in vitro plate assay, fungal discs of uniform size were inoculated at the center of the Potato Dextrose Agar (Himedia, India) carrying 25 ml of the medium and incubated at 28°C. When the mycelia reached 6 cm in diameter, four sterile Whatman no.1 filter paper discs (1 cm diameter) were placed on the plate at equal distance from the center. Purified rTvD1 was added at various concentrations, (ranging from 25–100 μg/μl) at the center of the disc on the plate. The elution buffer served as the control. The plates were incubated at 28°C and the plates were observed periodically till the mycelial mat covered the control discs.
For the sclerotium germination assay, the sclerotium was placed at four places at equal distance from the centre of the PDA plate. The protein at respective concentration was added directly over the sclerotia and its effect on their germination and growth was observed.
Arabidopsis germination assay
The purified protein was diluted to desired concentration in 10 mM Tris–HCl pH 7.6. This dilution was then mixed with an equal volume of Murashige and Skoog basal salts (Murashige and Skoog 1962) with B5 vitamins (Gamborg’s vitamins) in a 96-well assay plate. A single sterilized Columbia seed was placed in each well and grown at 24°C under a 16 h photoperiod for 7 days (Allen et al. 2007). The experiment was carried out in triplicates.
Results
Cloning of TvD1 coding region
The cDNA amplification from total RNA after reverse transcription resulted in a 228-bp product. No amplification was observed when the total RNA was used without reverse transcription indicating that the coding region was obtained from the amplification of the reverse transcribed cDNA. Amplification using the genomic DNA also resulted in a similar sized product indicating that the gene does not contain any intron(s).
TvD1 codes for a predicted 75-amino-acid-containing peptide of 8.2 kDa with the first 28 amino acids serving as signal peptide and the mature peptide of about 5.2 kDa (NCBI GenBank accession number AY907349 with an E value of 3.1e − 40). A BLASTX search in the GenBank protein database showed that the amino acid sequence has homology with other known defensins characterized from different plants with eight-conserved cysteine residues forming four disulfide bridges. Comparison of the deduced amino acid sequence of TvD1 (Fig. 1) with some of the well-characterized legume defensin mature peptides using alignment tool (CLUSTALW) showed that TvD1 had significant similarity to the defensins from Cicer arietinum (95%), Arachis hypogaea (93%), Vigna radiata defensin-2 (91%), Medicago sativa (87%), Trigonella foenum-graecum (82%), etc.
Comparison of the mature peptide sequence of TvD1 with mature defensin peptides characterized in legumes. Alignment of the sequences of some of the legume defensins sequences available in the GenBank like 2GL1_A (V. radiata VrD2); DQ296045.1 (A. hypogaea); DQ342338.1 (C. arietinum); AY907349.1 (T. villosa); AAG40321 (M. sativa); AY313166.1 (Medicago truncatula); AY227192.1 (T. foenum-graecum); AY288448.1 (Arachis diogoi); P81929.1 (Pisum sativum PsD1) NM_128160.1 1TI5_A (V. radiata VrD1) was performed using CLUSTAL-W program available at http://www.ncbi.nlm.nih.gov. TvD1 shows 95% sequence identity with Cicer defensin, 92% identity with A. hypogaea defensin and 91% identity with VrD2 of V. radiata
Constitutive expression of TvD1
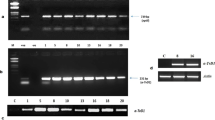
Using RT-PCR, the expression of TvD1 gene was analyzed in seeds, roots, stems, young, and mature leaves, and flowers of the plant (Fig. 2). Interestingly, this defensin is constitutively expressed in all the organs, however, with higher levels of expression in young and mature leaves.
RT-PCR of the total RNA isolated from different tissues such as (A) seed, (B) root, (C) stem, (D) inflorescence, (E) mature leaves and (F) young leaves for amplifying the open reading frame (ORF) of the cDNA for studying the expression of TvD1 in different tissues. The amplification of 18S rRNA (400 bp) was used as a control for equal loading of total RNA in the RT-PCR reactions. Minus RT control represents PCR for the cDNA of TvD1 using total RNA as template without reverse transcription
In silico characterization of TvD1 peptide
We constructed the three-dimensional structure of the mature portion of TvD1 (29–75 amino acid residue region) using standard protein modeling. The NMR structure of the defensin, VrD2 from V. radiata identified with an E value = 5e − 11 (PDB_ID: 2GL1, Lin et al. 2007) shared 91% pair-wise identity with the query sequence. The pair-wise sequence alignment was used for modeling the structure of the TvD1. The overall three-dimensional fold of the model defensin comprised three β-strands and one α-helix held together with four cysteine disulphide bridges as in the case of VrD2 (Fig. 3).
Homology modeling of TvD1 showed 91% similarity with that of VrD2. In the 3D model (a), the barrel-shaped blue-colored structure denotes the α-helix, pleated sheets represent β-strand and yellow-colored structures denote the cysteine bridges. In the stick model (b), the yellow-colored structures represent the cysteine bridges
Prokaryotic protein expression and purification
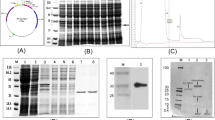
The rTvD1 protein from prokaryotic expression was purified and characterized further. The solubility characteristics of the recombinant protein were not largely affected either by increasing or reducing the IPTG concentration. The induction period also appeared to be crucial and maximum induction was observed after 6 h at 0.4 mM IPTG with no considerable variation afterwards. The recombinant protein had a molecular weight of approximately 25 kDa with approximately 8.2 kDa defensin of interest and the remaining peptide is the tag region of pET32a vector (Fig. 4). The use of prolysis buffer has increased the protein concentration to 1.8 mg/ml as against 0.7 mg/ml without this treatment. The protein concentration was observed to be similar to that of protein purified from the inclusion bodies after 10 h of IPTG induction (data not shown) using the method followed by Kirubakaran and Sakthivel (2007). This protein was used for in vitro antifungal assays.
In vitro antifungal assay
Antifungal activity of the rTvD1 was studied using various assays. It was observed that the antifungal activity of this protein was strongly dependent on the target fungus. Even though the peptide showed superior antifungal effect in comparison to the known defensins, the concentration at which the peptide inhibited the growth of the fungus was not uniform and varied with the fungal pathogen under consideration. In the spore germination assay (Fig. 5), the growth of some fungal pathogens was inhibited at very low concentrations (10–25 μg/ml) and a moderately higher concentration was needed for inhibition of other pathogens (50 μg/ml) (Table 1). Among the fungal species tested, the peanut late leaf spot fungus P. personata appeared to be the most sensitive with IC50 < 10 μg/ml. The antifungal activity was assessed in other fungal pathogens like F. oxysporum, F. moniliforme, P. parasitica, B. cinerea, A. helianthi, and Curvularia sp with IC50 < 25 μg/ml. Hyperbranching of the mycelium was also observed, when the spores of F. oxysporum were germinated in the presence of 10 μg/ml of rTvD1 (Fig. 5). The growth of R. solani sclerotium was also tested; rTvD1 showed its detrimental effect on the growth of mycelium. Its IC50 value was determined as 38 μg/ml (Fig. 6, Table 1), and, also, it was observed that the formation of sclerotia was greatly affected by forming a clear zone of inhibition near the disc where rTvD1 was added.
Effect of TvD1 on growth of R. solani sclerotia. C, 1, 2, and 3 represent control, 25 μg/ml, 50 μg/ml, and 100 μg/ml, respectively. a Protein at respective concentration was directly added over the sclerotium and photograph was taken after 48 h of growth and b protein was added on discs at respective concentrations and formation of sclerotia was observed after 3 weeks
In the plate assays, there were varied zones of inhibition in the test fungal species depending on the peptide concentration used, but at 100 μg/ml, a distinct inhibition zone was noted in all the species tested, R. solani, B. cinerea, F. moniliforme, and P. parasitica (Fig. 7).
Arabidopsis seed germination assay
We have also seen whether the rTvD1 was able to inhibit root elongation in Arabidopsis thaliana. Root growth in Arabidopsis was largely affected by TvD1 (Fig. 8), and it was inhibited in a dose-dependent manner (Table 2). At 10 μg/ml of recombinant TvD1, there was about 50% reduction in root length and further development was not evident. There was no root growth at concentrations of 50 μg/ml and above. It has also been noticed that the protein affected the number and extent of lateral root formation.
Discussion
Defensins are amongst the highly potent antimicrobial peptides advanced by the plant to protect itself against the invading pathogens. Hence, characterization of effective defensins from several sources is an emerging tool in plant biotechnology for enhancing disease resistance in crop plants (Osusky et al. 2000; Punja 2001).
From the pair-wise alignment, it was inferred that almost all the reported legume defensins have highly conserved domains within the protein. Homology modeling of the TvD1 indicated that it has a three-dimensional structure similar to the characterized defensins like VrD2 (Lin et al. 2007) and PsD1 (Almeida et al. 2002). These secondary structural elements are held together by four disulfide bridges, thus, forming a cysteine-stabilized α,β-fold. This fold belongs to the Knot-1 superfamily (INTERPRO ID: IPR003614). Examination of the model structure indicated that more than 95% residues were in the allowed regions of the phi–psi Ramachandran map indicating the confidence measure of the 3-D with a score of 17.8/21. The C-alpha traces of template and target structures were superimposed with 0.15 Å RMSD indicating high conservation in their structures. Two conserved regions distinguish plant defensins with or without antifungal activity (Almeida et al. 2002). From the alignment, it can be observed that His residue at position 29 in Psd1 is also conserved in TvD1 and the conserved Phe residue at position 41 is conservatively mutated to Trp. These structural features indicate that TvD1 would also exhibit strong antifungal activity similar to Psd1. The phylogenic tree indicated that TvD1 showed a common evolutionary origin based on conserved sequence and structural characteristics such as amino acid homology and conserved motifs from the legumes.
TvD1 is expressed constitutively to high levels in all the tissues of the plant, namely seed, leaf, root, stem, and flower. A similar constitutive expression of the defensin PDF2.3 was also observed in Arabidopsis in all organs except in roots (Manners et al. 1998). But, the latter protein was not shown to exhibit any antifungal activity.
The presence of a typical secretion signal peptide is one of the characteristic features of plant defensins (Broekaert et al. 1997). Xu and Reddy (1997) reported that bacteria failed to produce the pre-protein of a PR5-like protein of Arabidopsis, because its N-terminus affected E. coli growth. Similarly, no fusion protein could be induced under the same conditions, when the Trichosanthes kirilowii defensin (TDEF1) gene with its signal peptide-coding region was inserted into pET32a(+). Therefore, the partial TDEF1 cDNA, corresponding to the mature peptide, was inserted into the expression vector, and TDEF1 was produced as the fusion protein in E. coli without the N-terminal signal. However, the antifungal activity of the expressed protein was very low, requiring a dose up to 250 μg/ml of TDEF1 to have an effect on F. oxysporum (Hui et al. 2007). In the present study, the E. coli-expressed TvD1 defensin protein with its signal peptide was purified efficiently as a recombinant protein with significant antifungal activity at comparatively lower concentrations.
The assay of antifungal activity of the rTvD1 has shown that the 100-μg/ml concentration was sufficient to form an inhibitory zone for all the tested fungi. It was 100 and 150 μg/ml for Tfgd1 and Tfgd2 defensins, respectively, from T. foenum-graecum against R. solani and F. moniliforme (Olli and Kirti 2006; Olli et al. 2007). The IC50 value of Lm-Def was observed to be 100 μg/ml against P. infestans (Solis et al. 2007). The other E. coli-expressed PR proteins such as a chitinase from Sorghum required higher concentration such as 300 μg/ml against five fungal species, Viz., B. cinerea, R. solani, A. alternata, F. oxysporum, and wheat chitinase against Colletotrichum falcatum, Pestalotia theae, R. solani, Sarocladium oryzae, Alternaria sp., and Fusarium sp. (Kirubakaran and Sakthivel 2007; Singh et al. 2007).
In order to investigate the effect of recombinant TvD1 on fungal spore germination, we have tested some filamentous fungal pathogens. The conidia of P. personata failed to germinate at 10 μg/ml concentration. It was observed that 100 μg/ml concentration of the recombinant protein (Tfgd1) was an inhibitory factor for spore germination and hyphal growth even after 48 h for the conidia of P. personata (Olli and Kirti 2006).
However, similar growth with slight inhibitory effect was observed in the case of F. oxysporum, B. cinerea, F. moniliforme, A. helianthi, and Curvularia sp. at 10 μg/ml of TvD1, but higher concentrations were needed for 50% inhibition. The IC50 value of R. solani was 38 μg/ml and growth of sclerotium was distinctly arrested at higher concentration. Generally, defensins inhibit fungal growth by inducing hyperbranching (Brassicaceae) or the growth was arrested without hyperbranching (Fabaceae, Asteraceae, and Hippocastanaceae; Osborn et al. 1995). Also, it depended upon the fungal species tested (Spelbrink et al. 2004). Our studies showed that the defensin TvD1 from a member of Fabaceae, apart from inhibitory activity against spore germination also induced morphogenic changes like hyperbranching and changes in cell-wall morphology in the fungal pathogens like F. oxysporum and F. moniliforme. Such an activity was similar to Rs-AFP1 and Rs-AFP2 (Terras et al. 1995) and HaDef1 (Zélicourt et al. 2007) indicating that the TvD1 possesses both types of antifungal activities.
As in fungal hyphae, the root hair tip growth is also associated with an apex-high cytosolic-free-calcium gradient generated by a local calcium influx at the tip (Schiefelbein et al. 1992; Felle and Hepler 1997). Recombinant TvD1 was able to inhibit root elongation and lateral root formation at 10 μg/ml. The effect was more prominent at higher concentrations. Interestingly, this effect was observed at approximately the same concentration that was necessary for antifungal activity in some species. Antifungal defensins like MsDef1, MtDef2, RsAFP2, and KP4 also blocked the whole root development causing rapid RabA4b depolarization and halted the extension of growing root hairs (Allen et al. 2007). Hence, the present defensin has the potential to control the development and growth of the plant.
Defensins with antimicrobial activity are potent candidate genes for deployment in transgenic crops for protecting them against pathogens. With the inhibitory concentration as low as 10 μg/ml against some fungal pathogens, TvD1 appears to be a potent defensin gene for fungal disease resistance in transgenic crops.
References
Allen A, Snyder AK, Preuss M, Nielsen EE, Shah DM, Smith TJ (2007) Plant defensins and virally encoded fungal toxin KP4 inhibit plant root growth. Planta 227:331–339
Almeida MS, Cabral KMS, Kurtenbach E, Almeida FCL, Valente AP (2002) Solution structure of Pisum sativum defensin 1 by high resolution NMR: plant defensins, identical backbone with different mechanisms of action. J Mol Biol 315:749–757
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Broekaert WF, Terras FRG, Cammune BPA, Osborn RW (1995) Plant defensins: novel antimicrobial peptides components of the host defense systems. Plant Physiol 108:1353–1358
Broekaert WF, Cammue BPA, De Bolle MFC, Thevissen K, De Samblanx GW, Osborn RW (1997) Antimicrobial peptides in plants. Crit Rev Plant Sci 16:297–323
De Bolle MFC, Eggermont K, Duncan RE, Osborn RW, Terras FRG, Broekaert WF (1995) Cloning and characterization of two cDNA clones encoding seed-specific antimicrobial peptides from Mirabilis jalapa L. Plant Mol Biol 28:713–721
Doyle JJ, Doyle JL (1990) Isolation of plant DNA from fresh tissues. Focus 12:13–15
Felle HH, Hepler PK (1997) The cytosolic Ca2+ concentration gradient of Sinapsis alba root hairs as revealed by Ca2+-selective microelectrode tests and fur-dextran ratio imaging. Plant Physiol 114:39–45
Hui LD, Liang JG, Tao ZY, Min AT (2007) Bacterial expression of a Trichosanthes kirilowii defensin (TDEF1) and its antifungal activity on Fusarium oxysporum. Appl Microbiol Biotechnol 74:146–151
Kirubakaran SI, Sakthivel N (2007) Cloning and overexpression of antifungal barley chitinase gene in Escherichia coli. Protein Express Purif 52:159–166
Koike M, Okamoto T, Tsuda S, Imai R (2002) A novel plant defensin-like gene of winter wheat is specifically induced during cold acclimation. Biochem Biophys Res Commun 298:46–53
Lin KF, Lee TR, Tsai PH, Hsu MP, Chen CS, Lyu PC (2007) Structure-based protein engineering for a-Amylase inhibitory activity of plant defensin. Proteins 68:530–540
Liu Y, Luo J, Xu C, Ren F, Peng C, Wu G, Zhao J (2000) Purification, characterization, and molecular cloning of the gene of a seed-specific antimicrobial protein from poke weed. Plant Physiol 122:1015–1024
Manners JM, Penninckx IA, Vermaere K, Kazan K, Brown RL, Morgan A, Maclean DJ, Curtis MD, Cammue BP, Broekaert WF (1998) The promoter of the plant defensin gene PDF1.2 from Arabidopsis is systemically activated by fungal pathogen and responds to methyl jasmonate but not to salicylic acid. Plant Mol Biol 38:1071–1080
Murashige T, Skoog FA (1962) A revised medium for rapid growth and bioassays with tobacco tissue cultures. Plant Physiol 15:473–497
Olli S, Kirti PB (2006) Cloning, characterization and antifungal activity of defensin Tfgd1 from Trigonella foenum-graecum L. J Biochem Mol Biol 39:278–283
Olli S, Guruprasad L, Kirti PB (2007) Characterization of defensin (Tfgd2) from Trigonella foenum-graecum. Curr Sci 93:365–369
Osborn RW, De Samblanx GW, Thevissen K, Goderis I, Torrekens S, Van Leuven F, Attenborough S, Rees SB, Broekaert WF (1995) Isolation and characterization of plant defensins from the seeds of Asteraceae, Hippocastanaceae and Saxifragaceae. FEBS Letters 368:257–262
Osusky M, Zhou G, Osuska L, Hancock RE, Kay WW, Misra S (2000) Transgenic plants expressing cationic peptide chimeras exhibit broad-spectrum resistance to phytopathogens. Nature/Biotechnol 18:1162–1166
Park HC, Kang YH, Chun HJ, Koo JC, Cheong YH, Kim CY, Kim MC, Chung WS, Kim JC, Yoo JH, Koo YD, Koo SC, Lim CO, Lee SY, Cho MJ (2002) Characterization of a stamen-specific cDNA encoding a novel plant defensin in Chinese cabbage. Plant Mol Biol 50:59–69
Punja ZK (2001) Genetic engineering of plants to enhance resistance to fungal pathogens—a review of progress and future prospects. Can J Plant Pathol 23:216–235
Schiefelbein JW, Shipley A, Rowse P (1992) Calcium influx at the tip of growing root-hair cells of Arabidopsis thaliana. Planta 187:455–459
Singh A, Kirubakaran SI, Sakthivel N (2007) Heterologous expression of a new antifungal chitinase from wheat. Protein Express Purif 56:100–109
Solis J, Medrano GG, Ghislain M (2007) Inhibitory effect of a defensin gene from the Andean crop maca (Lepidium meyenii) against Phytophthora infestans. J Plant Physiol 164:1071–1082
Song X, Wang J, Wu F, Li X, Teng M, Gong W (2005) cDNA cloning, functional expression and antifungal activities of a dimeric plant defensin SPE10 from Pachyrrhizus erosus seeds. Plant Mol Biol 57:13–20
Spelbrink RG, Dilmac N, Allen A, Smith TJ, Shah DM (2004) Differential antifungal and calcium channel-blocking activity among structurally related plant defensins. Plant Physiol 135:2055–2067
Terras FRG, Schoofs HME, De Bolle MFC, Van Leuven F, Rees SB, Vanderleyden J, Cammue BPA, Broekaert WF (1992) Analysis of two novel classes of antifungal proteins from radish (Raphanus sativus L.) seeds. J Biol Chem 267:15301–15309
Terras FRG, Eggermont K, Kovaleva V, Raikhel NV, Osborn RW, Kester A, Rees SB, Torrekens S, Van Leuven F, Vanderleyeden J, Cammune BPA, Broekaert WF (1995) Small cysteine-rich antifungal proteins from radish: their role in host defence. Plant Cell 7:573–588
Thevissen K, Osborn RW, Acland DP, Broekaert WF (2000) Specific binding sites for an antifungal plant defensin from dahlia (Dahlia merckii) on fungal cells are required for antifungal activity. Mol Plant Microbe Interact 13:54–61
Thevissen K, Waranecke DC, Francois IE, Leipelt M, Heinz E, Ott C, Zahringer U, Thomma BP, Ferket KK, Cammune BP (2004) Defensins from insects and plant interact with fungal glucosylceramides. J Biol Chem 279:3900–3905
Thomma BP, Broekaert WF (1998) Tissue-specific expression of plant defensin genes PDF1.2 and PDF2.2 in Arabidopsis thaliana. Plant Physiol Biochem 36:533–537
Thomma BPHJ, Camme BPA, Thevissen K (2002) Plant defensins. Planta 216:193–202
Urdangarín MC, Norero NS, Broekaert WF, Canal LDL (2000) A defensin gene expressed in sunflower inflorescence. Plant Physiol Biochem 38:253–258
Xu H, Reddy ASN (1997) Cloning and expression of a PR5-like protein from Arabidopsis: inhibition of fungal growth by bacterially expressed protein. Plant Mol Biol 34:949–959
Zélicourt AD, Letousey P, Thoiron S, Campion C, Simoneau P, Elmorjani K, Marion D, Simier P, Delavault P (2007) Ha-DEF1, a sunflower defensin, induces cell death in Orobanche parasitic plants. Planta 226:591–600
Zhang Z, Schwartz S, Wagner L, Miller W (2000) A greedy algorithm for aligning DNA sequences. J Comput Biol 7:203–214
Acknowledgments
SV is grateful to the University Grant Commission, Government of India for financial support in the form of a senior research fellowship. The work is supported by a research grant from the Andhra Pradesh–Netherlands Biotechnology Program administered by the Institute of Public Enterprise, Hyderabad, India and by an exchange grant of the Department of Science and Technology, Government of India and the German Academic Exchange Program. The authors thank the Head, Department of Plant Sciences for allowing the use of the facilities under the UGC-SAP, COSIST, DST-FIST, etc.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Vijayan, S., Guruprasad, L. & Kirti, P.B. Prokaryotic expression of a constitutively expressed Tephrosia villosa defensin and its potent antifungal activity. Appl Microbiol Biotechnol 80, 1023–1032 (2008). https://doi.org/10.1007/s00253-008-1648-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-008-1648-2