Abstract
The identity of polyhydroxyalkanoates (PHA) storing bacteria selected under aerobic dynamic feeding conditions, using propionate as carbon source (reactor P), was determined by applying reverse transcriptase–polymerase chain reaction (RT-PCR) on micromanipulated cells and confirmed by fluorescence in situ hybridisation (FISH). Four genera, Amaricoccus, Azoarcus, Thauera and Paraccoccus were detected, the latter only rarely present. All the biomass was involved in PHA storage as shown by Nile Blue staining. By quantitative FISH, their specific amount was determined in this and two other systems using acetate as the carbon substrate (sequencing batch reactor [SBR] A and A1). SBR A and reactor P had the same sludge retention time (SRT, 10 days), while reactor A1 was operated with the SRT of 1 day and the double organic loading rate (OLR). Systems fed with acetate (41.1 ± 2.2 and 49.4 ± 1.4% total Bacteria, for A and A1, respectively) became enriched in Thauera independently on the SRT and OLR, while it was only present in a minor amount when propionate was used as a substrate (1.9 ± 0.2% total Bacteria). Amaricoccus was present in both reactors operated at 10 days SRT, favoured in the one fed with propionate (61.4 ± 1.9% total bacteria), and almost completely removed at the SRT of 1 day. Azoarcus cells were found in all the analysed systems (3.9 ± 0.3, 23.3 ± 1.5 and 45.9 ± 1.5 for P, A and A1, respectively), while Paracoccus was scarcely present.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Awareness concerning the environment and the use of clean technologies is growing worldwide. As a consequence, research on biodegradability and use of renewable resources for industrial processes has been intensive in the last years. Polyhydroxyalkanoates (PHA) are biodegradable biopolymers that have been recognised as good candidates for synthetic polymers partial replacement. PHA are accumulated intracellularly by a wide variety of microorganisms, acting as carbon and energy sources. The industrial production of PHA is achieved using pure or genetically engineered cultures with a high storage capacity (up to 90% of cell dry weight; Marchessault 1996). The greatest drawback for their use as petrol-based polymers substitutes is the production cost, which can be four times higher than the chemical synthesis (Direction Générale des Ressources Naturelles et de l’Environnement du Ministère de la Région du Valoni 2002). PHA production processes based on mixed microbial cultures (MMC) are being investigated as a possible technology to decrease production costs, as no sterilisation is required and bacteria can adapt quite well to the complex substrate present in low-cost substrates. Although many processes using MMC have been proposed to reach this goal, the most promising one is based on the utilisation of periodic supply of carbon substrates: a short period of excess carbon (feast) followed by a long period of starvation (famine; Majone et al. 1996). This process is currently known as ‘feast and famine’ or aerobic dynamic feeding (ADF). These conditions favour the development of bacterial populations with a high and stable PHA storage capacity. Under ADF conditions, with carbon addition (acetate) by sequential pulses, Serafim et al. (2004) showed that the intracellular PHA content reached 65.4% cell dry weight. The polymer produced was a 3-hydroxybutyrate (3HB) homopolymer.
For the majority of applications, a co-polymer of 3HB and another monomer, such as 3-hydroxyvalerate (3HV), is preferred because it presents a lower crystalinity and melting temperatures, making the polymer more flexible and with improved mechanical properties. The use of different carbon substrates allows modulating the polymer composition according to the desired polymer-specific applications. As an example, propionate can act as the precursor for the three-carbon unit (propionyl-CoA) needed for the hydroxyvalerate synthesis. PHA co-polymers and terpolymers production using propionate and mixed cultures are reported in the literature (Beccari et al. 1998; Dionisi et al. 2004; Lemos et al. 2006).
One critical factor on the development of a competitive process for PHA production with MMC is the selection of organisms with high storage capacity. Therefore, it is mandatory to monitor the population dynamics and to assess for different groups of organisms, in terms of PHA storage and cell growth kinetics. In terms of microbial population dynamics, scarce information is available about the organisms responsible for PHA accumulation under ADF and how they evolve in time. Recently, molecular methods had been applied to identify PHA-accumulating organisms in an ADF reactor (Dionisi et al. 2005, 2006). These authors used denaturing gradient gel electrophoresis (DGGE) and performed a 16S rRNA (ribosomal ribonucleic acid) clone library to characterise the evolution of the microbial community with high storage capacity. From the 100 clones obtained, the most important organisms belonged to the Betaproteobacteria (Thauera, Alcaligenes, Comamonas, Achromobacter), but also several Alphaproteobacteria (Xantobacter, Curtobacterium) and Gammaproteobacteria (Kluyvera, Pseudomonas, Acinetobacter) were present. The Thauera genus was assumed to be responsible for the PHA accumulation, although neither the abundance of this microorganism in the analysed sludge nor its involvement in PHA storage was confirmed in situ.
Fluorescence in situ hybridisation (FISH) has been recently employed for the characterisation of the microbial population in two reactors operated under ADF conditions, one adapted to acetate (reactor A) and the other to propionate (reactor P; Serafim et al. 2006). Three major morphotypes, Gram-negative Zoogloea-like bacilli (morphotype I) and two similar tetrad-forming bacteria (morphotypes II and III), affiliated, respectively, to the Betaproteobacteria and the Alphaproteobacteria, were dominant in both reactors and were involved in PHA storage. The main responsible for PHA storage was affiliated to the genus Azoarcus belonging to Betaproteobacteria. These morphotypes were always present, but their relative abundance changed during the sequencing batch reactor (SBR) operation. However, variation in biomass composition did not affect the storage capacity and the type of polymer produced (Serafim et al. 2006). System A always produced an homopolymer P(3HB) from acetate and a terpolymer P(3HB-co-3HV-co-2M3HV) from propionate, while population P always produced a co-polymer P(3HB-co-3HV) from acetate and propionate. Differences in polymer composition suggested a diverse biomass composition in the two reactors, which was not evident by morphological observation. These results seem to indicate that reactor operational conditions as well as the kind of substrate used may have a great impact in population selection. The microbial population composition developed in these systems was likely different from that reported by Dionisi et al. (2004), in which P(3HV) was produced from propionate in a reactor operated with different conditions (cycle length, sludge age, organic loading rate).
In the present paper, the sorting/reverse transcriptase–polymerase chain reaction (RT-PCR) approach was utilised to reveal the main composition of the PHA-accumulating biomass present in the system fed with propionate (P). Using FISH, the relative abundance of the microorganisms identified by the sorting/RT-PCR approach was estimated for the biomass selected in reactors A and P. This comparison intended to determine the effect of carbon substrate on the biomass composition, as for both reactors, all the other operational conditions were the same. The influence of short sludge retention time (SRT) and double organic load rate (OLR), on the storage performance and biomass composition, was also investigated in the reactor fed with acetate (A1). Correlation between the specific abundance of the identified microbial populations in these systems and the reactor performance was established.
Materials and methods
Reactors operation
In this study, a SBR for the production of PHA, operating for 4 years and adapted to propionate as carbon source (reactor P), was investigated.
The reactor working volume was 1 L and the total SBR cycle length 12 h, consisting of 10.5 h of aerobiosis, 1 h of settling (mixing and air bubbling switched off) and 0.5 h withdrawing half of the volume of the supernatant, which was replaced by the same volume of fresh medium during the first 15 min at the beginning of the cycle. The hydraulic retention time was 1 day. At the end of each cycle, before settling, a defined volume of mixed liquor was removed to keep the SRT at 10 days. Oxygen was supplied by an air compressor through a ceramic membrane disperser introduced inside the reactor at an airflow rate of 1.0 vvm (vol air/vol reactor min), maintaining dissolved oxygen concentration around 80% of the saturation value. The reactor was operated without pH control, but its value was monitored online (ranging between 8.0 and 9.2); the temperature was kept at 22°C and the stirring rate at 250 rpm. Timers controlled the air compressor, the medium feed, waste pumps and stirring.
A second reactor operated under the same conditions but using acetate as carbon source (A reactor) was also investigated. A third reactor (A1 reactor), using acetate as substrate, was operated under the same conditions of reactor A except that SRT was shorter (1 day) and organic loading rate was doubled (OLR, 120 C mmol L−1 day−1).
Culture medium
The standard mineral salts medium used in the SBR P was composed of (per litre of distilled water): 1.269 g CH3CH2COOH, 600 mg MgSO4.7H2O, 160 mg NH4Cl, 100 mg EDTA ethylenediamine tetraacetic acid, 92 mg K2HPO4, 45 mg KH2PO4, 70 mg CaCl2 2H2O and 2 ml of trace elements solution. The trace solution consisted of (per litre of distilled water): 1,500 mg FeCl3.6H2O, 150 mg H3BO3, 150 mg CoCl2 6H2O, 120 mg MnCl2.4H2O, 120 mg ZnSO4.7H2O, 60 mg Na2MoO4.2H2O, 30 mg CuSO4 5H2O and 30 mg of KI. Thiourea (10 mg/l) was added to inhibit nitrification. The pH of the salt solution was adjusted to 7.2 and then sterilised. The phosphorus solution was sterilised separately. After sterilisation, the two solutions were allowed to cool and were mixed together. The same mineral medium was used for SBR A and A1 with acetate, respectively, 40.796 and 81.592 g of CH3COONa.3H2O, as the only carbon source.
Analytical techniques
Cell dry weight was determined as volatile suspended solids according to Standard Methods (APHA 1995).
Acetate and propionate were analysed by high-performance liquid chromatography using a BioRad Aminex HPX-87H column, with 0.01 N sulphuric acid as eluent, an elution rate of 0.6 ml/min and an operating temperature of 50°C. A UV detector (Merck) set at 210 nm was used. Before injection, samples were filtered using a 0.2-μm membrane. PHA were determined by gas chromatography after acidic estherification according to Serafim et al. (2004) and Lemos et al. (2006). Ammonia was analysed using an ammonia gas-sensing combination electrode ThermoOrion 9512. Exopolymeric substances (EPS) were analysed according to Frølund et al. (1996).
Morphological characterisation and FISH analysis of the biomass
SBR biomass samples were collected regularly, at the end of the feeding period, for microscopic observation and staining. An aliquot of each sample was fixed for later FISH analysis, as described by Amann (1995). The protocol described in Jenkins et al. (1993) was utilised for Gram, Neisser and India Ink stainings. Intracellular PHA granules were shown by Nile Blue staining (Ostle and Holt 1982).
FISH was performed following Amann (1995). According to Maszenan et al. (2000), a pre-enzymatic treatment was applied on sample before FISH to increase permeability of Gram-negative tetrad-forming organisms. The oligonucleotide probes utilised are reported in Table 1. An Olympus BX51 microscope was used for the observations of the biomass in phase contrast, bright field (after Gram, Neisser and Nile blue staining) and epifluorescence (after Nile blue staining and FISH). Images were collected and analysed with F-View Soft Image System video camera and AnalySIS software.
Quantification was carried out on the fixed biomass according to previously published methods (Bouchez et al. 2000; Crocetti et al. 2002), and results were visualised on a Zeiss LSM 510 Meta confocal laser-scanning microscope. Specific probes used were all CY3 labelled and used in combination with CY5-labelled EUBmix (EUB338+EUB 338-II+EUB338-III). Image analysis of 40 randomly selected fields was performed using the free ImageJ software package (version 1.37b, Wayne Rasband, National Institute of Health, USA). The abundance of the diverse groups was expressed as percentage of all bacteria (as area occupied by probe-binding cells). The calculation was performed from the images where the number of pixels with a positive signal from both the specific probe and the EUBmix probes was compared to the number of pixels with a positive signal from the EUBmix probe. The standard error of the mean was calculated as the standard deviation divided by the square root of the number of images.
Cell sorting–RT-PCR approach
The sorting–RT-PCR approach, described hereafter, was applied on ethanol fixed samples. For the specific amplification of the targeted bacteria 16S rRNA, a small cluster of cells with distinctive morphology was collected by micromanipulation from the mixed biomass spread on an agar plate. Small homogenous clusters composed mostly of a single morphotype with abundant 16S rRNA were necessary for the specific micromanipulation and the amplification. By microscopic observation and FISH analysis with the EUBmix probe, a suitable sample for the sorting/RT-PCR was selected. The cells collected were transferred from the micromanipulation hook directly in a PCR tube, containing 10 μl of high purity distilled water, and then amplified by RT-PCR. RT-PCR was performed using the Invitrogen SuperscriptTM One RT-PCR kit (Invitrogen, Carlsbad, USA) with the universal primers U27f and U1492r.
Cloning and sequence analysis
Since the micromanipulation was carried out on clusters of bacteria selected solely on the basis of their morphology, it could not be guaranteed that all the collected cells belonged to the same species. Amplified rDNAs (ribosomal deoxyribonucleic acids) pooled from the three reactions were purified by the QIAquick PCR purification kit (Quiagen) according to the manufacturer’s instructions. Amplicons were then ligated into pcr®4-TOPO® vector (Invitrogen) and transformed into One Shot® TOP10 chemically competent E. coli cells according to the manufacturer’s instructions. The obtained clones were partially sequenced with T3 forward primer, while the 16S rRNAs of clone 11 from morphotype I and clone 5 from morphotypes II–III were completely sequenced with T3 and T7 primers (MWG biotech). The compiled 16S rRNA sequences were manually aligned, and the organism identity was deduced by standard programs including BLASTN and BLAST2 for most likely identities. The sequences were deposited in GenBank (accession numbers are EF367341 for clone 11 and EF367342 for clone 5).
ProbeBase online tool (Loy et al. 2003) was utilised for the selection of probes matching the retrieved 16S rRNA sequences.
Results
Reactors performance
The microbial characterisation of the PHA-accumulating microorganisms was performed in three reactors, SBRs P, A and A1, operated under different conditions.
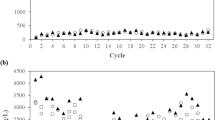
The SBR A and P have been simultaneously operated for several years under dynamic feeding conditions, with a SRT of 10 days and OLR of 60 C mmol L−1 day−1 (30 C mmol/cycle) using, respectively, acetate and propionate as carbon source. Reactor A was fully described in Serafim et al. (2004). Figure 1 shows the profiles for carbon and nitrogen concentrations and for oxygen uptake rate during a batch experiment (one cycle) of the reactor P fed with propionate. The propionate consumption was linear with time, and the oxygen uptake rate was almost stable, dropping sharply after carbon depletion. P(3HB-co-3HV) was produced with a higher fraction of 3HV. Polymer production occurred simultaneously with cell growth, as shown by the ammonia consumption. When no external carbon source was available, the formed polymer was consumed for cellular maintenance. The kinetic and stoichiometric parameters calculated for these systems are reported in Table 2.
A new reactor for PHA production, SBR A1, was set with biomass of reactor A as inoculum. This reactor was operated for 101 days, fed with acetate, a SRT of 1 day and a OLR of 120 C mmol L−1 day−1, while the C/N ratio was the same as in reactors A and P. Figure 2 presents the profiles of acetate, P(3HB) and ammonia during a feast and famine cycle of this system. This culture performed similarly to that of reactor A, where a homopolymer was produced. During the analysed operation period, the biomass from this system showed an average PHA cell content of 27.9% cell dry weight (maximum PHA storage capacity not optimised). Kinetic and stoichiometric parameters of this system were compared with the same data from reactors A and P (Table 2).
Sorting/RT-PCR approach
Cell sorting/RT-PCR was applied to determine the composition of the reactor P biomass at the genus level. By this method, bacteria with a distinctive morphology were selectively separated from the microbial community by micromanipulation and soon after the collected cells were utilised for successive 16S rRNA gene amplification by RT-PCR.
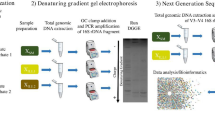
Because cell sorting is performed on clusters of bacteria selected on the basis of their morphology, the collected cells could belong to different bacterial species. Therefore, a small 16S rRNA gene clone library was constructed for each amplificate, and clones were sequenced to analyse all amplified 16S rDNAs. The obtained clones were sequenced, and the organism’s identity was ascertained. The approach validation was obtained by FISH analysis through the application of specific oligonucleotide probes fully matching all the retrieved 16S rDNA sequences. A general scheme of the experimental procedure is given in Fig. 3.
Sorting/reverse transcriptase–PCR strategy applied for the identification of PHA accumulation organisms. Morphologically distinguishable cell clusters were micromanipulated and transferred to a PCR tube containing high-purity water for successive 16S rRNA gene amplification by RT-PCR. The pool of amplified rDNAs were then ligated and transformed into chemically competent E. coli cells. The obtained 16S rRNA gene clones were sequenced, and the organism identities were ascertained. FISH was used to validate the approach: Oligonucleotide probes fully matching the retrieved 16S rRNA gene sequences were selected and applied on the original sample
The main morphotypes present in SBR P biomass (Fig. 4), their affiliation to Proteobacteria classes and their role in PHA storage were previously described (Serafim et al. 2006). To identify the major component of this biomass, in the present study, one cluster of bacilli (Morphotype I) was collected by micromanipulation and directly utilised to 16S rRNA gene amplification by RT-PCR. The same procedure was applied to one cluster of tetrad-forming organisms, corresponding to morphotypes II and III that, being morphologically very similar, could not be differentiated at the microscopic magnification utilised for micromanipulation (×350). In both cases, amplificates were obtained from a single cluster.
Morphotypes composing reactors A, A1 and P biomass (I morphotype I Betaproteobacteria, bacilli (1 × 1.2–1.5 μm), II morphotype II Alphaproteobacteria, coccobacilli (1 × 2 μm) in pairs or loosely aggregated tetrads, III morphotype III Alphaproteobacteria, large cocci (2 × 2 μm) in tightly packed tetrads)
Four 16S rRNA gene plasmid inserts, two from each micromanipulated bacterial aggregate, were sequenced. In Table 3, the closest relative of the obtained sequences are reported together with the available genus- and species-specific probes targeting these 16S rRNA.
16S rRNA gene sequences deriving from morphotype I cells (clones 11 and 2) belonged to the Betaproteobacteria, being almost identical to an uncultured bacterium affiliated to deeply branching clade in the Azoarcus (Juretschko et al. 2002). The 16S rDNA sequences obtained from morphotype II/III cells were, instead, affiliated to two different taxa in the order Rhodobacterales within the Alphaproteobacteria, one of them belonging to the Amaricoccus genus.
FISH analysis was performed on the biomass sample utilised for the sorting/RT-PCR approach, to ascertain the presence and the importance of the identified taxa. Probes available in literature for the genera Azoarcus (AZA645; Hess et al. 1997), Amaricoccus (AMAR839; Maszenan et al. 2000) and Paracoccus (PAR651; Neef et al. 1996), which have a full match with the retrieved 16S rRNA sequences, were used. Although no sequence belonging to the genus Thauera was obtained, their presence was also investigated with probe THAU832 (Loy et al. 2005). This microorganism was indicated as the most abundant and likely responsible for the storage phenomena in a system operated under ADF conditions for PHA production (Dionisi et al. 2006).
Probes AZA645- and AMAR839-positive cells, respectively, morphotype I and II cells (Fig. 5), were present. Azoarcus cells were dominating, and the two species accounted for most of the biomass. The genus-specific probe THAU832 detected only a small portion of the reactor’s bacterial population that could not be morphologically discriminated from Azoarcus cells. Probe PAR651 hybridised to thin bacilli (0.5 × 0.8 μm), rarely present and usually disperse in flocs, while morphotype III cells, also present in a minor amount, were not recognised with the applied probes. The results did not allow the identification of this component of the biomass but confirmed that, although similar, the two morphotypes (II and III) belong to different taxons. The Amaricoccus cells could be differentiated from morphotype III also by the yellow autofluorescence of the cell wall when they were observed with a fluorescein filter (U-MWB2 Olympus).
Quantitative FISH analysis
The presence and abundance of the genera Amaricoccus, Azoarcus, Thauera and Paracoccus in the population selected in reactors P, A and A1 was determined by FISH.
A sample from reactor P, different from the one used for the sorting RT-PCR approach, was analysed to determine the relative abundance of the microorganisms and to validate the previous identifications. The population of reactor A was also characterised. The effect of different carbon sources on microbial population dynamics was evaluated by comparing the composition of reactors A- and P-selected biomasses (Table 4).
FISH analysis confirmed that Amaricoccus spp. and Azoarcus spp. were still present in reactor P biomass after 2 years of operation. A major change in their relative abundance was observed in the present sample compared with that used for sorting/RT-PCR. Amaricoccus spp. was in this case dominant (61.4%), and Azoarcus was present in minor amount (3.9%). In this reactor, only a few THAU832-positive cells were detected (1.9%). The biomass of reactor P was only partially identified (67.2% of the EUBmix-positive cells) with the genus-specific probes applied. This result was consistent with the abundant presence of morphotype III cells that hybridised only with the general probe for the Alphaproteobacteria.
Differently from what was observed in rector P, almost all the biomass from reactor A was identified with the probes THAU832, AMAR839 and AZA645. All the three genera were present in a large amount: Thauera spp. cells composed approximately half of the biomass (41.1%), but also Azoarcus (23.3%) and Amaricoccus (28.8%) were quite abundant in the system. Morphotype III was not detected in reactor A, and PAR651-positive cells belonging to the Paracoccus genus were only rarely observed in both reactors.
Nile blue staining for lipid detection was performed on these biomasses before feeding and just after the substrate depletion. All the biomass was involved in PHA storage, as shown by the increase of fluorescence from the beginning to the end of the feast phase. However, morphotype I cells always contained the higher amount of PHA (higher fluorescence intensity) and seemed to be the main responsible of the storage phenomena.
The microbial composition of reactor A1 was monitored by phase-contrast observation of the morphotypes present. This biomass appeared as large compact flocs composed of morphotype I cells, and rarely, tetrad-forming bacteria corresponding to morphotypes II/III were observed. No significant variations in the morphotypes were detected along the time of reactor operation. The presence and abundance of microorganisms belonging to the genera Azoarcus, Thauera, Paracoccus and Amaricoccus were investigated in this biomass by FISH analysis. Thauera and Azoarcus cells (morphotype I) constituted most of the microbial community being, respectively, 49.5 and 45.9% of EUBmix-positive cells. The genus Paracoccus was detected in low amount (data not shown). The main difference from reactor A was the almost complete disappearance of Amaricoccus genus (<l% EUBmix-positive cells) from this system. Nile blue staining also confirmed that, in this reactor, all the biomass was involved in PHA storage.
Discussion
The use of acetate or propionate as a carbon substrate and the application of diverse operating conditions (SRT and OLR) selected microbial populations with different composition and PHA storage performances.
In particular, the utilisation of different substrates led to the production of P(3HB) in SBR A and A1 and P(3HB-co-3HV) in SBR P. Acetate is the precursor of 3HB, while propionate, depending on the pathway, can originate monomers of 3HB and 3HV (Lemos et al. 2006). Biomass from reactor A presented substrate uptake rate (q S, three times), PHA storage rate (q P, 12 times) and polymers storage yield (Y P/S, 2.5 times) higher than biomass from reactor P. The growth yield was slightly higher for sludge from reactor P (Y X/S, 1.25 times) than for reactor A. The higher storage yield from acetate compared to that from propionate is partially explained by the fact that 3HB and 3HV synthesis from propionate requires decarboxilation, while 3HB synthesis from acetate does not. This was experimentally proven by feeding either acetate or propionate to the same enriched population (Lemos et al. 2006). Reactor A showed high PHA-specific productivities and PHA cell content. The latter (65.4% of cell dry weight) was obtained when acetate was supplied in three pulses of 60 C mmol/L each, being the highest value so far reported for mixed cultures (Serafim et al. 2004). In reactor P, the maximum PHA cell content obtained was lower, 41.2% of cell dry weight (unpublished results).
For the same substrate (acetate), changing the reactor SRT and organic loading rate resulted in different behaviour. In reactor A1, acetate and ammonia profiles along the feast and famine cycle were different from those observed for the SBR A (Serafim et al. 2004). In the former, ammonia was consumed at twice the rate observed in the reactor A. Despite the growth yield being slightly higher for A1 than for A, ammonia uptake rate (q N) was faster. Operation of the reactor at SRT of 1 day selected faster-growing organisms than at SRT of 10 days. Specific acetate uptake and specific PHA storage rates were, respectively, three and four times lower with 1 day of SRT than with 10 days. The storage yield, for 1 day of SRT, was almost half of the value observed for 10 days. These results show that low values of SRT select biomass with a high specific growth rate but with low storage rate. Microscopic observation of biomass produced in SBR A1 appeared very slimy and resistant to India Ink diffusion, which are characteristics of EPS. Analytical determination of EPS showed a EPS yield on acetate of 0.07 C mmol glucose/C mmol S, which could explain the lower PHA polymer yield obtained in the biomass of reactor A1 compared with that of reactor A, where no EPS were detected. According to Cho et al. (2005), low values of SRT may favour EPS production by sludge.
The use of a short SRT (1 day against 10 days) and high OLR did not improve the biomass PHA storage capacity. Similar results were obtained by Beun et al. (2002), who estimated a decrease of the storage yield for SRT lower than 2 days.
The efficiency of the sorting/RT-PCR approach, previously applied for filamentous bacteria (Levantesi et al. 2006), was confirmed on these floc-former morphotypes. The most abundant of the targeted bacteria, morphotypes I and II, could be successfully identified analysing a restricted number of clones. Both Thauera and morphotype III cells, not retrieved with sorting/RT-PCR, were present in low number in the biomass.
The main microbial components of the analysed SBRs biocenosis were all previously found in activated sludge environment, and their storage capacities were reported. In particular, Amaricoccus spp. was detected in large number in wastewater treatment plants (WWTPs; Maszenan et al. 2000), and the high PHA storage capacity was confirmed (Falvo et al. 2001; Aulenta et al. 2003). The Thauera/Azoarcus group constituted 16% of the activated sludge biomass in a nitrifying–denitrifying industrial WWTP (Juretschko et al. 2002). The capacity of these bacteria to store PHA was reported in (Bergey’s Manual 2005), and the presence of Thauera was already ascertained in a SBR for PHA production, although its involvement in lipid storage was not shown (Dionisi et al. 2006).
FISH analysis showed that bacteria belonging to the genera Amaricoccus, Thauera and Azoarcus were always present in reactors A and P. Clear differences in the abundance of these genera, which could be attributed to the kind of carbon source utilised, were, however, observed between the two reactors. As reported by Lemos et al. (2006), the different polymer composition produced as well as the kinetic and stoichiometric parameters calculated in these reactors, when the substrates supplied were shifted, could be explained on the basis of a diverse population composition of reactor A and P: propionate to population A and acetate to population P. Results indicated that the selection of Thauera spp. was favoured under ADF when acetate was present as a carbon source. While this genus was rarely observed in SBR P, it constituted almost half of the biomass in the two acetate reactors. Thauera was also enriched, in a SBR operated at a SRT of 1 day and under ADF conditions fed with acetate (Dionisi et al. 2005, 2006).
The use of propionate seemed, instead, to favour the development of Amaricoccus spp. Most of the species belonging to this genus are known to be able to use acetate and, a minor proportion, propionate as carbon source (Bergey’s Manual 2005). Amaricoccus kaplicensis and A. macauensis are among the species able to use both substrates. As observed in the two samples of P reactor and as previously reported by Serafim et al. (2006), the bacterial composition of biomass was constant along the time of reactor operation, but the relative abundance of each organisms varied. The dominance of the diverse genera in SBR A and P, in particular the selection between Azoarcus or Amaricoccus, seemed to be dependent not only on the substrate utilised but also on the reactor operational conditions. The major change in biomass composition was observed when the reactor SRT was reduced to 1 day and the OLR was duplicated. The latter conditions selected faster-growing organisms with lower storage capacity than those obtained at a SRT of 10 days. Amaricoccus was therefore washed out of SBR A1 (μmax of A. kaplicensis is 2.1–2.5 day−1, Falvo et al. 2001).
A major challenge in the optimisation of the mixed culture PHA production process is the effective selection of cultures with high PHA storage capacity (high impact on PHA cell content) and growth rate (effect on the volumetric PHA production rate). In this work, the microbial population present in PHA accumulation processes was identified and correlated with the reactor-operating conditions. The application of microbiological and molecular techniques showed to be a useful tool to monitor the selection of target microorganisms. Cell sorting/RT-PCR was a valuable technique for the determination of the main species responsible for PHA production in the ADF SBRs used, and it can also be easily transferred for the analysis of any enrichment culture characterised by low biodiversity. Differently from other cell-sorting procedures (i.e. flow cytometry), it does not require sophisticated and expensive laboratory equipments and strongly limits the number of analysis to perform (i.e. low number of clones to screen).
The abundance of the different genera, Amaricoccus, Thauera and Azoarcus, where for some the presence was revealed by the cell sorting/RT-PCR approach, was indeed confirmed by quantitative FISH analysis. This is an important step forwards in the knowledge of this complex but most promising process.
References
Amann RI (1995) In situ identification of microorganisms by whole cell hybridization with rRNA-targeted nucleic acid probes. In: Akkermans A, van Elsas J, de Bruijn F (eds) Molecular microbial ecology manual. Kluwer, London, pp MMEM–3.3.6/1–MMEM-3.3.6/15
Amann RI, Binder BJ, Olson RJ, Chisholm SW, Devereux R, Stahl DA (1990) Combination of 16S rRNA-targeted oligonucleotide probes with flow cytometry for analyzing mixed microbial populations. Appl Environ Microbiol 56:1919–1925
APHA, AWWA, WPCF (1995) Standard methods for the examination of water and wastewater. Port City Press, Baltimore (edited by Eaton AD, Clesceri LS, Greenberg AE)
Aulenta F, Dionisi D, Majone M, Parisi A, Ramadori R, Tandoi V (2003) Effect of periodic feeding in SBR reactor on substrate uptake and storage rates by a pure culture of Amaricoccus kaplicensis. Water Res 37:2764–27
Beccari M, Majone M, Massanisso P, Ramadori R (1998) A bulking sludge with high storage response selected under intermittent feeding. Water Res 32:3403–3413
Bergey’s Manual of Systematic Bacteriology Volume Two: the Proteobacteria (Part C): the Alpha-, Beta-, Delta-, and Epsilonproteobacteria (2005) Garrity G, Brenner DJ, Krieg NR, Staley JT (eds) 2nd edn. Williams and Wilkins, Springer
Beun JJ, Dircks K, van Loosdrecht MCM, Heijnen JJ (2002) Poly-β-hydroxybutyrate metabolism in dynamically fed mixed cultures. Water Res 36:1167–1180
Bouchez T, Patureau D, Dabert P, Wagner M, Delgenes JP, Moletta R (2000) Successful and unsuccessful bioaugmentation experiments monitored by fluorescent in situ hybridization. Water Sci Technol 41:61–68
Cho J, Song KJ, Yun H, Ahn KH, Kim JY, Chung TH (2005) Quantitative analysis of biological effect on membrane fouling in submerged membrane bioreactor. Water Sci Technol 51(6–7):9–18
Crocetti GR, Banfield JF, Keller J, Bond PL, Blackall LL (2002) Glycogen-accumulating organisms in laboratory-scale and full-scale wastewater treatment processes. Microbiology 148:3353–3364
Daims H, Bruehl A, Amann R, Schleifer KH, Wagner M (1999) The domain-specific probe EUB338 is insufficient for the detection of all Bacteria: development and evaluation of a more comprehensive probe set. Syst Appl Microbiol 22:434–444
Dionisi D, Majone M, Papa V, Beccari M (2004) Biodegradable polymers from organic acids by using activated sludge enriched by aerobic periodic feeding. Biotechnol Bioeng 85:569–579
Dionisi D, Beccari M, Di Gregorio S, Majone M, Petrangeli Papini M, Vallini G (2005) Storage of biodegradable polymers by an enriched microbial community in a sequencing batch reactor operated at high organic load rate. J Chem Technol Biotechnol 80:1306–1318
Dionisi D, Majone M, Vallini G, Di Gregorio S, Beccari M (2006) Effect of applied organic load rate on biodegradable polymer production by mixed microbial cultures in a sequencing batch reactor. Biotechnol Bioeng 93:76–88
Direction Générale des Ressources Naturelles et de l’Environnement du Ministère de la Région du Valoni (2002) Technologies propres, no. 2. Kluwer, Dordrecht
Falvo A, Levantesi C, Rossetti S, Seviour RJ, Tandoi V (2001) Synthesis of intracellular storage polymers by Amaricoccus kaplicensis, a tetrads forming bacterium present in activated sludge. J Appl Microbiol 91:299–305
Frølund B, Palmgren R, Keiding K, Nielsen PH (1996) Extraction of exopolymers from activated sludge using a cation exchange resin. Water Res 30(8):1749–1758
Hess A, Zarda B, Hahn D, Haner A, Stax D, Hohener P, Zeyer J (1997) In situ analysis of denitrifying toluene- and m-xylene-degrading bacteria in a diesel fuel-contaminated laboratory aquifer column. Appl Environ Microbiol 63:2136–2141
Jenkins D, Richards MG, Daigger GT (1993) Manual on the causes and control of activated sludge bulking and foaming, 2nd edn. Lewis, London
Juretschko S, Loy A, Lehner A, Wagner M (2002) The microbial community composition of a nitrifying-denitrifying activated sludge from an industrial sewage treatment plant analyzed by the full-cycle rRNA approach. Syst Appl Microbiol 25:84–99
Lemos PC, Serafim LS, Reis MAM (2006) Synthesis of polyhydroxyalkanoates from different short-chain fatty acids by mixed cultures submitted to aerobic dynamic feeding. J Biotechnol 122:226–238
Levantesi C, Rossetti S, Beimfohr C, Thelen K, Krooneman J, van der Waarde J, Tandoi V (2006) Description of filamentous bacteria present in industrial activated sludge WWTPs by conventional and molecular methods. Water Sci Technol 54:129–137
Loy A, Horn M, Wagner M (2003) probeBase—an online resource for rRNA-targeted oligonucleotide probes. Nucleic Acids Res 3:514–516
Loy A, Schulz C, Lücker S, Schöpfer-Wendels A, Stoecker K, Baranyi C, Lehner A, Wagner M (2005) 16S rRNA gene-based oligonucleotide microarray for environmental monitoring of the betaproteobacterial order Rhodocyclales. Appl Environ Microbiol 71:1373–1386
Majone M, Massanisso P, Carucci A, Lindrea K, Tandoi V (1996) Influence of storage on kinetic selection to control aerobic filamentous bulking. Water Sci Technol 34:223–232
Manz W, Amann R, Ludwig W, Wagner M, Schleifer K-H (1992) Phylogenetic oligodeoxynucleotide probes for the major subclasses of Proteobacteria: problems and solutions. Syst Appl Microbiol 15:593–600
Marchessault RH (1996) Tender morsels for bacteria: recent developments in microbial polyesters. Trends Polym Sci 4:163–168
Maszenan A-M, Seviour RJ, Patel BKC, Wanner J (2000) A fluorescently-labelled r-RNA targeted oligonucleotide probe for the in situ detection of G-bacteria of the genus Amaricoccus in activated sludge. J Appl Microbiol 88:826–835
Neef A (1997) Anwendung der in situ Einzelzell-Identifizierung von Bakterien zur Populationsanalyse in komplexen mikrobiellen Biozönosen. Doctoral thesis, Technische Universität München, München
Neef A, Zaglauer A, Meier H, Amann R, Lemmer H, Schleifer K-H (1996) Population analysis in a denitrifying sand filter: conventional and in situ identification of Paracoccus spp. in methanol-fed biofilms. Appl Environ Microbiol 62:4329–4339
Ostle AG, Holt JG (1982) Nile Blue A as a fluorescent stain for poly-β-hybroxybutyrate. Appl Environ Microbiol 44:238–241
Serafim LS, Lemos PC, Oliveira RF, Reis MAM (2004) Optimisation of polyhydroxybutyrate production by mixed cultures submitted to aerobic dynamic feeding conditions. Biotechnol Bioeng 87:145–160
Serafim LS, Lemos PC, Rossetti S, Levantesi C, Tandoi V, Reis MAM (2006) Microbial community analysis with a high PHA storage capacity. Water Sci Tech 54:183–188
Acknowledgements
This research was developed within the agreement for Scientific and Technological Cooperation between the Gabinete de Relações Internacionais da Ciência e do Ensino Superior—GRICES (Portugal) and National Research Council—CNR (Italy). The authors acknowledge the financial support of the Fundação para a Ciência e Tecnologia (FCT) through the project POCI/BIO/55789/2004 and EU Integrated Project Contract no. 026515. Luísa S. Serafim acknowledges Fundação para a Ciência e Tecnologia for grant SFRH/BPD/14663/2003.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Lemos, P.C., Levantesi, C., Serafim, L.S. et al. Microbial characterisation of polyhydroxyalkanoates storing populations selected under different operating conditions using a cell-sorting RT-PCR approach. Appl Microbiol Biotechnol 78, 351–360 (2008). https://doi.org/10.1007/s00253-007-1301-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-007-1301-5