Abstract
Penicillium chrysogenum npe10 (Δpen; lacking the 56.8-kbp amplified region containing the penicillin gene cluster), complemented with one, two, or three penicillin biosynthetic genes, was used for in vivo studies on transport of benzylpenicillin intermediates. 6-Aminopenicillanic acid (6-APA) was taken up efficiently by P. chrysogenum npe10 unlike exogenous δ(l-α-aminoadipyl)-l-cysteinyl-d-valine or isopenicillin N (IPN), which were not taken up or were taken up very poorly. Internalization of exogenous IPN and 6-APA inside peroxisomes was tested by quantifying their peroximal conversion into benzylpenicillin in strains containing only the penDE gene. Exogenous 6-APA was transformed efficiently into benzylpenicillin, whereas IPN was converted very poorly into benzylpenicillin due to its weak uptake. IPN was secreted to the culture medium. IPN secretion decreased when increasing levels of phenylacetic acid were added to the culture medium. The P. chrysogenum membrane permeability to exogenous benzylpenicillin was tested in the npe10 strain. Penicillin is absorbed by the cells by an unknown mechanism, but its intracellular concentration is kept low.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
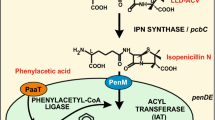
Penicillium chrysogenum is a filamentous fungus able to synthesize penicillins containing an aromatic side chain (Martín et al. 1990; Elander 2003). However, this fungus is known to also secrete isopenicillin N (IPN) and other penicillins containing polar side chains (Demain 1983). The ability to convert IPN into benzylpenicillins is exclusive of a few species of fungi that have acquired the penDE gene (Barredo et al. 1989; Gutiérrez et al. 1991). Penicillin biosynthesis starts with the nonribosomal condensation of the three amino acids l-α-aminoadipic acid, l-cysteine, and l-valine to form the tripeptide δ(l-α-aminoadipyl)-l-cysteinyl-d-valine (ACV) catalyzed by the multienzyme δ(α-aminoadipyl)-cysteinyl-valine synthetase (encoded by the pcbAB gene) (Díez et al. 1990; Byford et al. 1997; Martín 2000a). ACV is cyclized to IPN by the activity of IPN synthase (encoded by the pcbC gene) (Ramos et al. 1985; Müller et al.1991). In the last step of the pathway, the l-α-aminoadipyl side chain of IPN is exchanged for a more hydrophobic side chain in a reaction catalyzed by the IPN acyltransferase (IAT), which is encoded by the penDE gene (Álvarez et al. 1987; Martín et al. 1990). The aromatic acyl side chain has to be previously activated by a specific aryl-CoA ligase (Lamas-Maceiras et al. 2006). These two steps take place inside the peroxisomal matrix because IAT and phenylacetyl-CoA ligase are located inside the microbodies (peroxisomes) (Müller et al. 1991, 1992, 1995; Lamas-Maceiras et al. 2006).
It is well known that the three intermediates of the penicillin pathway are secreted to the culture broths. The tripeptide ACV is found in the culture broth of P. chrysogenum strains (López-Nieto et al 1985). Similarly, IPN is present in the culture broths of P. chrysogenum strains (Demain 1983), although nothing is known about its possible reabsorption by the cells or its in vivo conversion into aromatic penicillins. Finally, 6-aminopenicillanic acid (6-APA) is well known to accumulate in the P. chrysogenum culture broths (Demain 1983) and has been reported to be converted into aromatic penicillins (Meesschaert and Eyssen 1981), which would require an uptake system. In all these cases, it is unclear if the secretion of the intermediates is due to an overflow of the biosynthetic pathway in the high producing strains or whether it is a natural process with ecological significance.
The aromatic penicillins produced by P. chrysogenum are amphipatic compounds, are moderately hydrophobic and negatively charged at normal cytosolic pH values (Kirschbaum 1986), and might be secreted by diffusion across both the microbody and plasma membrane. However, the IPN is more hydrophilic, and its transport into peroxisomes remains a mystery. To date, no penicillin transporters have been identified in P. chrysogenum. However, in Aspergillus nidulans, it was found that an ABC transporter encoded by the atrD gene is involved, somehow, in penicillin secretion (Andrade et al. 2000). It was therefore of great interest to study the secretion and the uptake of the intermediates of the penicillin biosynthetic pathway.
In P. chrysogenum, the three genes responsible for penicillin biosynthesis are clustered in a DNA region that is amplified in tandem repeats bounded by a conserved hexanucleotide TGTAAA/T in penicillin-overproducing strains (Fierro et al. 1995; Newbert et al. 1997). The 56.8-kbp amplified region of P. chrysogenum contains 13 open reading frames (ORFs) linked to the penicillin gene cluster (Fierro et al. 2006), but the implication of the proteins encoded by these ORFs in penicillin metabolism or transport is not yet clear.
The hydrophobic penicillin biosynthesis is an excellent model to study the enzyme targeting and the transport systems involved in the localization of the pathway intermediates because there is good knowledge on the molecular genetics and the biochemistry of the biosynthetic enzymes (Aharonowitz et al. 1992; Martín 2000b). The lack of appropriate mutants to clarify the transport processes has prevented progress in this field. In the present work, we generate several tailored strains to obtain an insight on the in vivo secretion and uptake of benzylpenicillin and its intermediates.
Materials and methods
Strains and culture conditions
Penicillium chrysogenum Wis54-1255, which contains a single copy of the penicillin gene cluster (Fierro et al. 1995), and P. chrysogenum Wis54-1255 npe10 pyrG−(Δpen), an uridine auxotroph of Wis54-1255 npe10 lacking the whole amplified region including the penicillin gene cluster (Cantoral et al. 1993; Fierro et al. 1996), were used in this work. Tailored P. chrysogenum strains, such as npe10-AB·C, npe10-DE, npe10-C·DE, npe10-AB·C·DE, and npe10-C·DE-ACV, were generated by introducing the genes pcbAB, pcbC, or penDE in different combinations into the npe10 pyrG−(Δpen) strain as a host strain (Table 1). Fungal spores were plated on Power medium (Casqueiro et al. 1999) and grown for 5 days at 28°C. Penicillium chrysogenum liquid cultures were initiated inoculating fresh spores in 100 ml of defined double porosity (DP) medium (Casqueiro et al. 1999) without potassium phenylacetate. After incubation at 25°C for 20 h in an orbital shaker (250 rpm), a 10% aliquot was inoculated in DP medium (100 ml in 500-ml flasks) with or without phenylacetate (see “Results” section). Uridin auxotrophs were grown in the presence of 140 μg/ml of uridine.
For uptake experiments, ACV, 6-APA, IPN, or potassium penicillin were added to DP medium at this inoculation time. Penicillium chrysogenum phlAR+, a strain overexpressing the gene encoding the phenylacetyl Co-A ligase (phl gene), and P. chrysogenum DPhl1, a strain disrupted in the phl gene (Lamas-Maceiras et al. 2006), were also used for studies on the effect of phenylacetic acid on IPN secretion. Three different cultures were performed in duplicates to obtain the data from secretion and uptake experiments. Escherichia coli DH5α cells were used for plasmid amplification and were grown in Luria–Bertani medium supplemented with ampicillin (100 μg/ml).
Plasmid constructions
Plasmid pPyrAB·C, used for the expression in P. chrysogenum of both the pcbAB and the pcbC genes, was constructed as follows: Clone NA1 containing a 17-kbp recombinant fragment including both genes pcbAB and pcbC was isolated from a P. chrysogenum NRRL 1951 genomic library in the vector λGEM12 (Promega, Madison, WI, USA). The recombinant fragment was excised by digestion with NotI and ligated to NotI-digested pBG plasmid (a pBluescript® KS + derivative containing the P. chrysogenumpyrG gene in the HindIII site).
Plasmid pPenIAT was constructed to express the acyltransferase (IAT) in the tailored strains of P. chrysogenum as follows. The cDNA of the penDE gene was cloned in the NcoI–StuI sites of pIBRC43 (Cardoza et al. 1998) between the Aspergillus awamorigdh gene promoter (a very efficient promoter in ascomycetes) and the Saccharomyces cerevisiaecyc1 transcriptional terminator, thus generating plasmid pIBRC43-penDE. Digestion with KpnI and HindIII released the cassette Pgdh-penDE-Tcyc1, which was inserted in the same restriction sites of plasmid pJL43Trp, which contains the ble gene (for phleomycin resistance) as selectable marker under the control of the gpdA promoter and the trpC terminator.
Plasmid pPenC·DE, derived from plasmid pIBRC43 (Cardoza et al. 1998), includes the phleomycin-resistant cassette and a 5-kb SalI genomic fragment carrying the pcbC and penDE genes. Plasmid pACV, containing the 12.1 kb SpeI fragment that includes the complete pcbAB gene (with its native promoter and terminator regions) cloned into a SpeI-digested pBluescript® KS+(Stratagene, La Jolla, CA, USA), was used to cotransform together with plasmid pBG (which includes the pyrG gene as indicated above) the npe10-C·DE strain.
Transformation of P. chrysogenum protoplasts
Protoplasts were obtained and transformed as previously described (Cantoral et al. 1987; Díez et al. 1987). Transformant clones were selected by either complementation of the uridine auxotrophy or by resistance to phleomycin.
Southern, Northern, and Western blotting
DNA and RNA isolation Southern and Northern blotting hybridizations were carried out as indicated previously (Lamas-Maceiras et al. 2006; Fierro et al. 2006). Immunological detection of IAT was performed as previously described (Fernández et al. 1994). “Precision Plus Protein All Blue Standards” (Bio-Rad, Hercules, CA, USA), ranging in size between 10 and 250 kDa, were used as molecular weight markers. Polyclonal antibodies against P. chrysogenum IAT were obtained previously (Fernández et al. 2003).
Penicillin intermediates
BisACV (disulfide form) was obtained from Bachem Fein Chemicalien AG (Bubendorf, Switzerland), and monomeric ACV was purchased from Sigma-Aldrich (St. Louis, MO, USA). Crude ACV preparations (obtained from A. chrysogenum N2) were a gift from Jorge Martín (INBIOTEC, Leon, Spain). Potassium penicillin was kindly provided by Antibióticos S.A. (Leon, Spain), and 6-APA was a gift from Gist-Brocades (Delft, the Netherlands). IPN was purified in our laboratory. Stock solutions of potassium benzylpenicillin, IPN, and monomeric ACV were prepared in Milli-Q water. BisACV was dissolved directly in DP medium. 6-APA was dissolved in 50 mM sodium borate pH 7.7.
Bioassays and cell extracts for high-performance liquid chromatography analysis
Agar plugs containing sporulated P. chrysogenum strains on solid Power medium and bioassays with Micrococcus luteus ATCC 9341 were performed as described by Casqueiro et al. (1999). Mycelia samples were centrifuged for 10 min at 4,400×g and washed three times in 0.9% NaCl. Then, they were resuspended in 0.05 M borate pH 7.5, sonicated on ice (six pulses of 10 s with 30-s pauses between each pulse) and centrifuged at 4,400×g for 10 min at 4°C. The extract supernatant was used for further high-performance liquid chromatography (HPLC) analysis of the intermediates or the penicillin itself.
Extraction and HPLC analyses of penicillin from filtrates and intracellular extracts
Filtrates or cell extracts (3 ml) were acidified until pH 2.0 with 0.1 N HCl. Benzylpenicillin was extracted by adding n-butyl acetate (3 × 1 ml) and re-extracted from the organic phase with 10 mM phosphate buffer pH 7.5 (3 × 1 ml). This procedure was performed at 4°C. The aqueous phase was lyophilized and resuspended in 300 μl of Milli-Q water for filtrates (in 100 μl for intracellular extracts), which were analyzed by HPLC and bioassay.
HPLC analysis of benzylpenicillin was performed in an Agilent 1100 HPLC system (Santa Clara, CA, USA) with an analytical 4.6 × 250 mm (5 μm) RPC18 Lichrospher® 100 (Merck, Darmstadt, Germany) column, a flow rate of 1 ml/min, and a detector wavelength of 214 nm. Samples (20 μl) were injected and eluted using as mobile phase buffer A (30 mM ammonium formate pH 5.0 and 5% acetonitrile) and buffer B (same as buffer A plus acetonitrile 20:80, v/v) with an isocratic method (85% of A). Benzylpenicillin showed a retention time of 15.39 ± 0.17 min and its detection limit was 0.1 μg/ml.
Derivatization of IPN and 6-APA and HPLC analysis
Culture supernatants or cell extract samples (1 ml) were deproteinized by adding 2 ml of acetone and centrifugation at 4,400×g for 30 min at 4°C. After centrifugation, 300 μl of 0.2 M sodium-borate buffer pH = 7.7 and 400 μl of 15 mM 9-fluoronylmethyl chloroformate (FMOC) (Sigma-Aldrich) diluted in acetone were added to each sample and vortexed for 1 min. After a 4-min incubation at room temperature, the excess FMOC was removed by extraction with hexane (3 × 4 ml). The derivatizated IPN and 6-APA samples were analyzed in the same HPLC system as described above but using an analytical 4.6 × 150 mm (3.5 μm) Eclipse XDB-C18 (Agilent) column. Samples (10 μl) were injected in the HPLC using 50 mM sodium acetate pH 4.2 as solvent A and acetonitrile as solvent B. The elution gradient for IPN was as follows: 20%B→25%B linear over 7 min, 25%B→20%B linear over 8 min, 20%B→45%B for 15 min. For 6-APA, the elution gradient was 20%B→30%B linear over 10 min, isocratic for 8 min, 30%B→35%B linear over 2 min, isocratic for 10 min. The flow used was 1 ml/min. Detection was performed at 254 nm. Under these conditions, the retention time for derivatized IPN was 22.24 ± 0.08 min and for derivatized 6-APA was 24.78 ± 0.03 min. Detection limits of 0.1 and 0.01 μg/ml were achieved for IPN and 6-APA, respectively.
Results
Complementation of the P. chrysogenum npe10 strain (Δpen) with the pcbAB and pcbC genes restores the biosynthesis of IPN
The 56.8-kbp amplified region of P. chrysogenum, which bears the three genes responsible for penicillin biosynthesis (pcbAB, pcbC, and penDE) was found to contain 13 ORFs linked to the penicillin gene cluster (Fierro et al. 2006). To test whether any of these ORFs is essential for IPN production or transport, a tailored P. chrysogenum strain containing only the pcbAB and pcbC genes and lacking the other 13 ORFs present in the amplified region was obtained by introducing pPyrAB·C (see “Materials and methods” section and Table 1), carrying the two early genes (pcbAB and pcbC) of the pathway, into P. chrysogenum npe10 pyrG−(Δpen). Transformants were selected based on their ability to produce IPN, which was confirmed in one of the randomly selected transformants (transformant T3) by bioassay (Fig. 1a) and HPLC analysis (Fig. 1b); the transformant tested showed the production of a β-lactamase-sensitive compound coincident with a control of IPN. This strain, which only contains the first two penicillin biosynthesis genes, is referred to hereafter as P. chrysogenum npe10-AB·C. The IPN synthesized was secreted in both solid and liquid cultures of P. chrysogenum npe10-AB·C. These findings confirm that IPN is secreted without being converted to benzylpenicillin.
IPN production after complementation of the P. chrysogenum npe10 strain (Δpen) with the pcbAB and pcbC genes. a Bioassays using filtrates taken at 72 h from cultures of the Wis54-1255 (control), the npe10, and the npe10-AB·C strains (transformant T3). b HPLC of culture filtrates (72 h) of the transformant T3 of the npe10-AB·C strain (npe10-AB·C T3). Pure IPN was used as standard
Complementation of the P. chrysogenum npe10-AB·C strain with the penDE gene restores benzylpenicillin biosynthesis and secretion
When the npe10-AB·C strain was transformed with plasmid pPenIAT, containing the cDNA of the third gene of the penicillin pathway (penDE) encoding the IAT, under the control of the strong A. awamorigdh gene promoter (Table 1), all the transformants produced benzylpenicillin as shown by bioassay (Fig. 2a). Different transformants produced distinct levels of benzylpenicillin due to integration of the penDE gene in different places in the genome as seen by Southern blotting (data not shown). Transformant T5 was selected for further studies and will be referred to hereafter as P. chrysogenum npe10-AB·C·DE. Benzylpenicillin formation in this strain was confirmed by HPLC (Fig. 2b). Benzylpenicillin specific production of the npe10-AB·C·DE and the parental Wis54-1255 strains was compared in liquid cultures in defined DP medium. Results (Fig. 2c) showed that the Wis54-1255 strain produces, in liquid cultures, approximately four times more benzylpenicillin than the npe10-AB·C·DE strain. To test whether this difference was due to a lower production of IPN in the tailored strains, this compound was measured by HPLC in the npe10-AB·C, npe10-AB·C·DE, and Wis54-1255 strains cultured in defined DP medium without phenylacetic acid (to optimize IPN production). Results showed that the Wis54-1255 strain also produces approximately three to four times more extracellular IPN than the npe10-AB·C or npe10-AB·C·DE strains at 72 h (Fig. 2d). This result indicates that differences in benzylpenicillin production observed between the parental (Wis54-1255) and the engineered strain might be due to a lower ability of the latter to synthesize IPN. The intracellular IPN levels were also analyzed, being approximately twofold lower in the Wis54-1255 strain than in the npe10-AB·C or npe10-AB·C·DE strains at 72 h (Fig. 2e), even though the Wis54-1255 strain synthesizes and secretes much higher levels of IPN than the engineered strain. This result suggests that the engineered strain npe10-AB·C·DE is less efficient in secreting IPN, which might cause a feedback inhibition of its biosynthesis.
Complementation of the P. chrysogenum npe10-AB·C strain with the penDE gene. a Bioassay using sporulated agar plugs with different transformants of the npe10-AB·C·DE strain. The Wis54-1255, npe10, and npe10-AB·C strains were used as controls. b Chromatogram showing the benzylpenicillin extracted at 72 h from filtrates taken from cultures of the npe10-AB·C·DE strain (transformant 5). Potassium benzylpenicillin (PenG) was used as control at a concentration of 10 μg/ml. c Relative benzylpenicillin specific production between the Wis54-1255 strain (black bars) and the complemented npe10-AB·C·DE strain (gray bars). Filtrates were taken at 30, 48, and 72 h and bioassayed. The penicillin production corresponding to the Wis54-1255 strain at 72 h was set as 100%. d IPN production of the Wis54-1255 strain (black bars), the npe10-AB·C·DE strain (gray bars), and the npe10-AB·C strain (striped bars). Filtrates were taken at 30, 48, and 72 h and IPN was determined by HPLC. Results were expressed as micrograms of IPN/milligram of dry weight. IPN production at 48 and 72 h is also plotted as percent (inset), referred to the value corresponding to the Wis54-1255 strain as 100%. e Intracellular levels of IPN in the Wis54-1255 strain (black bars), the npe10-AB·C·DE strain (gray bars), and the npe10-AB·C strain (striped bars). Cellular extracts from samples taken at 30, 48, and 72 h were analyzed by HPLC. Results were expressed as micrograms of IPN/milligram of dry weight. Samples were taken in duplicate from three different cultures. Vertical lines above the bars indicate standard deviation of the mean value
Taken together, these results indicate that, although the presence of the pcbAB, pcbC, and penDE genes is sufficient for basic IPN and benzylpenicillin production, some of the missing 13 genes in npe10-AB·C·DE may contain information required for enhanced IPN and penicillin production in liquid cultures.
Extracellular ACV is not significantly taken up by P. chrysogenum
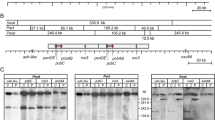
ACV is known to be partially secreted to the culture broth. The ability of P. chrysogenum to take up this tripeptide was assessed by quantifying the conversion of extracellullar ACV to benzylpenicillin by an engineered strain containing only the last two genes involved in penicillin biosynthesis (npe10 strain transformed with plasmid pPenC·DE; Table 1). Three positive transformants (T1, T2, and T3) were analyzed by Northern blotting using total RNA from these transformants to assure the expression of both pcbC and penDE genes. The three positive transformants were able to express these genes at levels similar to those generated by the P. chrysogenum wild-type strain (Fig. 3a). Transformant T1 was selected and named P. chrysogenum npe10-C·DE strain. To be sure about the ability of this strain to synthesize benzylpenicillin when ACV is provided, it was cotransformed with plasmids pACV (carrying the pcbAB gene; Table 1) and pBG (carrying the pyrG gene). Several transformants were analyzed by bioassay, and the production of benzylpenicillin in one of the transformants (named npe10-C·DE-ACV) was confirmed by HPLC (Fig. 3b). This result indicates that the npe10-C·DE strain is able to synthesize benzylpenicillin when the tripeptide ACV is supplied intracellularly (provided in the npe10-C·DE-ACV strain by the ACV synthetase coded by the pcbAB gene). Therefore, ACV uptake was tested adding to cultures of the npe10-C·DE strain two different concentrations of crude ACV preparations. No benzylpenicillin was detected at any time with any of the two concentrations of crude ACV used (Fig. 3c). The same experiment was repeated, but 0, 50, and 250 μg/ml of pure dimeric ACV (Bachem) were added to the culture medium. As shown in Fig. 3d, penicillin was not detected at any time, even when 250 μg/ml of ACV was added to the culture media. The same results were obtained when 50 μg/ml of pure monomeric ACV (Sigma-Aldrich) was added to the culture medium (data not shown). These results indicate that extracellular ACV is not significantly taken up in P. chrysogenum.
Characterization of the npe10-C·DE strain and transport of exogenous ACV inside P. chrysogenum. a Northern blot analysis performed with total RNA obtained from three transformants of the P. chrysogenum npe10-C·DE strain (T1, T2, and T3) and the Wis54-1255 strain (W). The membrane was hybridized to probes containing the penDE, the pcbC, and the β-actin genes (the latter was used as internal control). b Chromatogram showing the benzylpenicillin extracted from 72 h culture filtrates of the npe10-C·DE-ACV strain, confirming the ability of the npe10-C·DE strain to synthesize benzylpenicillin when the tripeptide ACV is provided. Potassium benzylpenicillin (PenG) was used as control at a concentration of 5 μg/ml. c Bioassays using culture filtrates obtained at different time points from the npe10-C·DE strain supplemented with two different concentrations of crude ACV. Filtrates obtained from the Wis54-1255 strain were used as control. d Bioassays using culture filtrates obtained at different time points from the npe10-C·DE strain treated with 0 (left petri dish), 50 (middle), and 250 μg/ml (right) of pure dimeric ACV. Filtrates from the Wis54-1255 strain were used as control. Note the lack of conversion of exogenous dimeric ACV into penicillin
Uptake of 6-APA and IPN in P. chrysogenum
To test whether the secreted penicillin precursors IPN and 6-APA are able to be taken up again from the culture medium, the P. chrysogenum npe10 strain was cultured in the presence of 0.05, 0.5, and 2.5 mg/ml (0.214, 2.14, and 10.7 mM, respectively) 6-APA or 0.05 and 0.5 mg/ml (0.139 and 1.39 mM) IPN. External and intracellular IPN and 6-APA were quantified by HPLC in filtrates and cellular extracts taken at 12, 24, 36, 48, and 72 h after addition of these compounds. Results (Fig. 4a) showed that 6-APA was accumulating inside P. chrysogenum along the incubation process when added at 0.5 and 2.5 mg/ml to the culture. This finding indicates that 6-APA is well transported inside P. chrysogenum, and this uptake is dose-dependent.
Uptake of exogenous 6-APA by P. chrysogenum npe10. a Intracellular levels of 6-APA (as μg/mg dry weight) in cultures supplemented with 0.5 (continuous line) or 2.5 mg/ml (dotted line) of 6-APA. b Extracellular/intracellular ratio of 6-APA in cultures supplemented with 0.5 (left panel) or 2.5 mg/ml (right panel) of 6-APA
The internalization ratio of 6-APA was calculated. Plots (Fig. 4b) showed that, at 12 h, only a small fraction of 6-APA had been taken up into the cells. After this time, the internalization of 6-APA gradually increased.
Unlike 6-APA, which is clearly taken up by the cell, no IPN was detected by HPLC in cell extracts of the P. chrysogenumnpe10 strain at any time and at any dose added to the culture medium (data not shown); this finding indicates that IPN is poorly or not transported inside P. chrysogenum (see “Discussion” section).
Conversion of IPN and 6-APA into benzylpenicillin: internalization of exogenous 6-APA and IPN into peroxisomes
Because IPN is synthesized in the cytosol and the conversion of IPN to benzylpenicillin occurs inside the peroxisomal matrix (Müller et al. 1991, 1992, 1995), this precursor has to enter the peroxisome to form penicillin. 6-APA is formed by the IAT, and this reaction also takes place inside peroxisomes. Because 6-APA is able to be internalized by P. chrysogenum, as seen above, the ability of this compound to be transported into peroxisomes was also tested by following its conversion to benzylpenicillin in a P. chrysogenum strain containing only the penDE gene (Table 1). Expression and correct processing of IAT resulting in the formation of the 11 + 29 α–β heterodimer was confirmed by Western blotting (Fig. 5a) in one of the transformants (referred to hereafter as P. chrysogenum npe10-DE). This strain expresses the penDE gene, but it is unable to synthesize penicillin because it lacks the pcbAB and pcbC genes. Cultures of P. chrysogenum npe10-DE were supplemented with 10 or 100 μg/ml IPN (0.0278 and 0.278 mM, respectively) or 6.5 and 65 μg/ml 6-APA (0.0278 and 0.278 mM, respectively), and culture filtrates were taken at 0, 24, 36, 48, 60, 72, and 84 h. After acidic extraction, samples were analyzed by HPLC for benzylpenicillin detection. High benzylpenicillin levels were formed (Fig. 5b) when exogenous 6-APA was added to cultures of the npe10-DE strain. However, when IPN was added to the culture medium, only trace levels of benzylpenicillin were detected, indicating important differences in the transport of these two intermediates and confirming our findings on the poor uptake of IPN (see above). Quantitative data obtained with bioassay determinations that are very sensitive (Fig. 5c) indicate that benzylpenicillin production at 72 h is approximately 40-fold higher when 0.278 mM exogenous 6-APA was added than when the same concentration of IPN was used. These results indicate that 6-APA is transported through both the cellular and peroxisomal membranes of P. chrysogenum and converted into benzylpenicillin, unlike IPN, which is very poorly taken up into the cell.
Characterization of the npe10-DE strain and transport studies of exogenous 6-APA and IPN showing their peroxisomal conversion into penicillin. a Western blotting showing the correct expression and processing of IAT in one of the transformants of the npe10-DE strain at 48 h (T). Note that the antibodies only recognize the β-subunit (29-kDa) of the active α–β heterodimer. Protein extracts taken at 48 h from cultures of the npe10 strain were used as control (C). b Chromatogram showing the benzylpenicillin extracted at 72 h from culture filtrates of the npe10-DE strain supplemented with 0.278 mM of IPN and 6-APA. Potassium penicillin was used as internal control at a concentration of 10 μg/ml. c Representative bioassays performed using filtrates taken at different time points from cultures of the npe10-DE strain supplemented with 0.0278 and 0.278 mM of IPN and 6-APA. As control, a solution of diluted potassium penicillin (0.1 μg/ml) was used (C). Benzylpenicillin production data corresponding to these bioassays are listed in Table 2 (supplementary material)
Secretion of IPN depends on phenylacetyl Co-A intracellular levels: phenylacetic acid directs the pathway to benzylpenicillin precluding IPN secretion
To study the correlation between intracellular phenylacetyl Co-A levels and IPN secretion, increasing concentrations (0, 0.6, 1.2, and 2.4 mg/ml) of phenylacetic acid, which is activated to phenylacetyl Co-A before being used as substrate by the IAT, were added to cultures of the parental Wis54-1255, phlAR+ [a strain overexpressing eightfold the phl gene encoding the phenylacetyl Co-A ligase (Lamas-Maceiras et al. 2006)], DPhl1 [a Δphl strain disrupted in the phl gene but keeping some phenylacetyl: Co-A ligase activity from alternative acyl-CoA ligase genes (Lamas-Maceiras et al. 2006)], and npe10-AB·C strains. Culture filtrates were taken at 72 h, and the amounts of secreted IPN were quantified by HPLC (Fig. 6a). Results indicated that IPN secretion drastically decreased in all strains (Wis54-1255, phlAR+, DPhl1, and npe10-AB·C) when increasing amounts of phenylacetic acid were added (Fig. 6b). In the Wis54-1255, phlAR+, and DPhl1 strains, this decrease is very likely due to intracellular conversion of IPN into benzylpenicillin. The finding that the disrupted Dphl1 mutant secretes more IPN than the Wis54-1255 or the phlAR+ strains (even when 1.2 mg/ml of phenylacetic acid is added to the culture medium) appears to be due to a lower formation of phenylacetyl Co-A in that strain and therefore to a lower consumption of IPN for benzylpenicillin synthesis. IPN secretion is, therefore, suppressed by the higher affinity of the IAT for IPN when sufficient intracellular phenylacetic-CoA (the second substrate of IAT) is available. Remarkably, the secretion of IPN also decreased when phenylacetic acid was added to cultures of the npe10-AB·C strain, though the IPN intracellular levels in this strain remained above those formed by the other strains (Fig. 6b). Because the npe10-AB·C strain lacks the IAT encoded by penDE and therefore there is no conversion of IPN into benzylpenicillin, a mechanism other than depletion of IPN by the IAT must also be controlling the secretion of IPN to the culture medium (see “Discussion” section).
Effects of internal phenylacetyl Co-A levels on IPN secretion. a Representative chromatograms of filtrates taken at 72 h from the Wis54-1255, phlAR+ (overexpressing the phl gene encoding the phenylacetyl Co-A ligase), DPhl1 (disrupted in the phl gene), and npe10-AB·C strains after addition of 0.6 mg/ml of phenylacetic acid (PA+) to the culture medium or without addition of this compound (PA−). IPN was added to these samples as internal control. b Relative IPN specific secretion (%) in filtrates taken at 72 h from cultures of the Wis54-1255, phlAR+, DPhl1, and npe10-AB·C strains when increasing amounts of phenylacetic acid were added to the culture medium. The inset plot shows the percentage of IPN secreted in these strains (mean ± SD from three independent cultures performed in duplicate). These values are relative to the IPN concentrations secreted by the Wis54-1255, phlAR+, DPhl1, and npe10-AB·C strains at 72 h without the addition of phenylacetic acid, which were set to 100%
Permeability of the P. chrysogenum membrane to benzylpenicillin
A possible uptake of exogenous penicillin in P. chrysogenum was proposed several years ago to explain an apparent feedback regulation of penicillin biosynthesis (Gordee and Day 1972; Nestaas and Demain 1981). The permeability to exogenous benzylpenicillin was tested in vivo by adding 100 μg/ml (0.279 mM) of benzylpenicillin to cultures of P. chrysogenum npe10, which cannot synthesize endogenous penicillin. Samples were taken at 24, 48, and 72 h, and extracellular and intracellular penicillin levels were quantified by HPLC. As shown in Fig. 7a, a compound coincident with the retention time of authentic benzylpenicillin was found in intracellular extracts at 24 h and increased until 72 h, suggesting that the npe10 is able to internalize some penicillin. Results of the quantification experiments (Fig. 7b) indicated that the P. chrysogenum npe10 strain internalizes external benzylpenicillin up to intracelullar levels similar to those shown by the Wis54-1255 strain.
Permeability of the P. chrysogenum plasma membrane to benzylpenicillin. a Chromatograms showing the benzylpenicillin levels at 24, 48, and 72 h in the intracellular extracts of both the Wis54-1255 strain (left panel) and the npe10 strain cultured in presence of 0.1 mg/ml of exogenous benzylpenicillin (right panel). b Intracellular levels of benzylpenicillin (μg/mg dry weight) detected in the Wis54-1255 strain (continuous line) and in the npe10 strain cultured in presence of 0.1 mg/ml exogenous of benzylpenicillin (dotted line)
Calculation of the external/internal benzylpenicillin ratio indicated that the Wis54-1255 strain secretes the antibiotic very efficiently, accumulating it outside the cell and keeping an outside/inside ratio of approximately 70 along the culture time.
Discussion
Genome research has allowed us to identify the genes responsible for the biosynthesis of penicillin in P. chrysogenum and other fungi. These genes are clustered in the 56.8-kbp amplifiable region of P. chrysogenum together with other 13 ORFs (Fierro et al. 2006). However, a putative enhancing role of some of the protein encoded by these ORFs on penicillin biosynthesis or transport of the intermediates in and out of the peroxisomes or the cytoplasmic membrane is not clear to date and requires further research.
As shown in this article, IPN secretion was observed in the npe10 strain (a mutant lacking the whole 56.8-kbp region) when the pcbAB and pcbC genes were introduced in this strain (Fig. 1), indicating that this hydrophilic penicillin (independently of its conversion to the hydrophobic benzylpenicillin) is secreted by a transport system. In other words, the penDE gene (encoding the IAT) is not strictly required for β-lactam antibiotic biosynthesis and secretion and may be a late addition to the basic penicillin gene cluster (pcbAB–pcbC) (Liras and Martín 2006). This hypothesis is supported by the presence of a gene with high similarity to penDE in sequenced genomes of several ascomycetes that lack, however, the pen gene cluster, like Aspergillus clavatus (GenBank accession number XP_001271254), Neosartorya fischeri (GenBank accession number XP_001263202), or Aspergillus fumigatus (GenBank accession number XP_754359). Because IPN and benzylpenicillin production is partially restored in the npe10 strain after transformation with the pcbAB, pcbC, and penDE genes (Fig. 2), it is concluded that the rest of the ORFs present in the amplified region are not strictly essential for the transport of precursors or intermediates of penicillin or for proper targeting and processing of the biosynthetic enzymes. Nevertheless, we have found a fourfold decrease in both IPN and benzylpenicillin specific production in both the npe10-AB·C and npe10-AB·C·DE strains, as compared to the Wis54-1255 parental strain. Because the npe10-AB·C and npe10-AB·C·DE strains contain at least one copy of the pcbAB and pcbC genes (as it occurs in the Wis54-1255 strain) and because these genes are expressed from their own wild-type promoter, differences in IPN and benzylpenicillin levels are probably due to the absence of the other genes in the 56.8-kbp DNA region. This indicates that some of the genes of the 56.8-kbp region may be necessary for enhanced penicillin biosynthesis.
Another interesting finding is the fact that the secretion of IPN depends on phenylacetyl Co-A intracellular levels. It is clear that the decrease in the extracellular IPN when phenylacetic acid is added is a consequence of the conversion of IPN into benzylpenicillin. However, the npe10-AB·C strain lacks the penDE-encoded IAT. A decrease in the secretion of IPN was observed when increasing concentrations of phenylacetic acid were added to this strain (Fig. 6b), indicating that other mechanisms related to the intracellular levels of phenylacetyl-CoA control the secretion of IPN to the culture medium. Phenylacetic acid is toxic and it is transported very efficiently into P. chrysogenum (Hillenga et al. 1995), and this fungus has probably developed a detoxification mechanism to remove it (Lamas-Maceiras et al. 2006).
Secretion of ACV, IPN, and 6-APA might be a waste of intermediates for benzylpenicillin production. Important differences were found in the uptake of the penicillin biosynthesis intermediates. ACV is not taken up into P. chrysogenum cells, even when high concentrations of this compound were added to the culture medium (Fig. 3d). ACV can be found either in a reduced monomeric form or as an oxidized dimer (disulfide bisACV), which is formed in oxic (aerobic) culture medium. BisACV may be reduced back to LLD-ACV by the thioredoxin–thioredoxin reductase system in the cytoplasm. The fact that the oxidized dimer is the predominant ACV form in culture broths may explain the lack of internalization. Even when ACV is added as a monomer, no internalization was observed, probably because of the rapid oxidation under the conditions of the culture medium.
Regarding IPN, it was found that this compound was taken up very poorly inside P. chrysogenum. In fact, no IPN was detected in the cells when it was added to cultures of the npe10 strain. This is in contrast to what was found with 6-APA, which was detected at relatively high concentrations in intracellular extracts of the npe10 strain when it was added to the culture medium (Fig. 4), indicating that exogenous 6-APA is taken up efficiently. The disparity observed between the uptake of IPN and 6-APA may be due to differences in the polarity of these two intermediates. Thus, the higher polarity of IPN may decrease the passive transport of this compound through the P. chrysogenum membrane. In turn, 6-APA, which is more hydrophobic, is transported more efficiently inside this microorganism.
The penicillin biosynthetic pathway occurs in different cellular compartments. ACV synthetase and IPN synthase are cytosolic enzymes, whereas IAT and phenylacetyl Co-A ligase are targeted into peroxisomes (Müller et al. 1991, 1992, 1995; Gledhill et al. 1997; Van der Lende et al. 2002; Lamas-Maceiras et al. 2006). Results obtained with the npe10-DE strain proves that exogenous 6-APA not only enters the cell membrane but also is transported across the peroxisomal membrane because benzylpenicillin was synthesized when this intermediate was added to cultures of this strain (Fig. 5). The fact that IPN has to be transported through the peroxisomal membrane to synthesize penicillin, together with the poor IPN uptake observed through the plasma membrane, indicates that an active IPN transport system must be present in the peroxisomal membrane to assure an adequate pool of IPN inside the peroxisomal matrix.
The secretion of antibiotics is a useful tool in nature for the microorganisms that produce them (Martín et al. 2005). In some producer organisms, these compounds are also toxic to them in high concentrations (Cundliffe 1991). Therefore, the producer strains have developed mechanisms for protection against their own antibiotics. One of these mechanisms is the membrane permeability barriers coupled to efficient efflux mechanisms for removal and secretion of drug molecules from the cells (Cundliffe 1991; Martín et al. 2005). It is known that very high concentrations of benzylpenicillin and other hydrophobic penicillins exert toxic effects on eucaryotic cells due to lipophilic interactions with the plasma membrane and intracellular membranes (Sikkema et al. 1995). Indeed, benzylpenicillin is actively pumped out of the cells.
In summary, most of the endogenously synthesized penicillin is efficiently secreted and only minor amounts of the final product are reintroduced into the cells, explaining the well-known accumulation of extracellular penicillin in the culture broths.
References
Aharonowitz Y, Cohen G, Martín JF (1992) Penicillin and cephalosporin biosynthetic genes: structure, regulation, and evolution. Annu Rev Microbiol 46:461–495
Álvarez E, Cantoral JM, Barredo JL, Díez B, Martín JF (1987) Purification to homogeneity and characterization of the acyl-CoA: 6-APA acyltransferase of Penicillium chrysogenum. Antimicrob Agents Chemother 31:1675–1682
Andrade AC, Van Nistelrooy JG, Peery RB, Skatrud PL, De Waard MA (2000) The role of ABC transporters from Aspergillus nidulans in protection against cytotoxic agents and in antibiotic production. Mol Gen Genet 263:966–977
Barredo JL, van Solingen P, Díez B, Álvarez E, Cantoral JM, Kattevilder A, Smaal EB, Groenen MAM, Veenstra AE, Martín JF (1989) Cloning and characterization of the acyl-coenzyme A: 6-aminopenicillanic-acid-acyltransferase gene of Penicillium chrysogenum. Gene 83:291–300
Byford MF, Baldwin JE, Shiau CY, Schofield CJ (1997) The mechanisms of ACV synthetase. Chem Rev 97:2631–2650
Cantoral JM, Díez B, Barredo JL, Álvarez E, Martín JF (1987) High frequency transformation of Penicillium chrysogenum. Biotechnology 5:494–497
Cantoral JM, Gutiérrez S, Fierro F, Gil-Espinosa S, van Liempt H, Martín JF (1993) Biochemical characterization and molecular genetics of nine mutants of Penicillium chrysogenum impaired in penicillin biosynthesis. J Biol Chem 268:737–744
Cardoza RE, Moralejo FJ, Gutiérrez S, Casqueiro J, Fierro F, Martín JF (1998) Characterization and nitrogen-source regulation at the transcriptional level of the gdhA gene of Aspergillus awamori encoding an NADP-dependent glutamate dehydrogenase. Curr Genet 34:50–59
Casqueiro J, Bañuelos O, Gutiérrez S, Hijarrubia MJ, Martín JF (1999) Intrachromosomal recombination between direct repeats in Penicillium chrysogenum: gene conversion and deletion events. Mol Gen Genet 261:994–1000
Cundliffe E (1991) Office of Naval Research lecture. Antibiotics and the search for new principles. J Ind Microbiol 7:157–161
Demain AL (1983) Biosynthesis of β-lactam antibiotics. In: Demain AL, Solomon NA (eds) Antibiotics containing the β-lactam structure. Springer, Berlin Heidelberg New York, pp 189–228
Díez B, Álvarez E, Cantoral JM, Barredo JL, Martín JF (1987) Isolation and characterization of pyrG mutants of Penicillium chrysogenum by resistance to 5′-fluorotic acid. Curr Genet 12:277–282
Díez B, Gutiérrez S, Barredo JL, van Solingen P, van der Voort LHM, Martín JF (1990) The cluster of penicillin biosynthetic genes. J Biol Chem 265:16358–16365
Elander RP (2003) Industrial production of beta-lactam antibiotics. Appl Microbiol Biotechnol 61:385–392
Fernández FJ, Gutiérrez S, Velasco J, Montenegro E, Marcos AT, Martín JF (1994) Molecular characterization of three loss-of-function mutations in the isopenicillin N-acyltransferase gene (penDE) of Penicillium chrysogenum. J Bacteriol 176:4941–4948
Fernández FJ, Cardoza RE, Montenegro E, Velasco J, Gutiérrez S, Martín JF (2003) The isopenicillin N acyltransferases of Aspergillus nidulans and Penicillium chrysogenum differ in their ability to maintain the 40-kDa alphabeta heterodimer in an undissociated form. Eur J Biochem 270:1958–1968
Fierro F, Barredo JL, Díez B, Gutiérrez S, Fernández FJ, Martín JF (1995) The penicillin gene cluster is amplified in tandem repeats linked by conserved hexanucleotide sequences. Proc Natl Acad Sci USA 92:6200–6204
Fierro F, Montenegro E, Gutiérrez S, Martín JF (1996) Mutants blocked in penicillin biosynthesis show a deletion of the entire penicillin gene cluster at a specific site within a conserved hexanucleotide sequence. Appl Microbiol Biotechnol 44:597–604
Fierro F, García-Estrada C, Castillo NI, Rodríguez R, Velasco-Conde T, Martín JF (2006) Transcriptional and bioinformatic analysis of the 56.8 kb DNA region amplified in tandem repeats containing the penicillin gene cluster in Penicillium chrysogenum. Fungal Genet Biol 43:618–629
Gledhill L, Greaves PA, Griffin JP (1997) Phenylacetyl-CoA ligase from Penicillium chrysogenum. Patent IPN WO97/02349, Smithkline Beecham, UK
Gordee EZ, Day LE (1972) Effect of exogenous penicillin on penicillin biosynthesis. Antimicrob Agents Chemother 1:315–321
Gutiérrez S, Diez B, Alvarez E, Barredo JL, Martin JF (1991) Expression of the penDE gene of Penicillium chrysogenum encoding isopenicillin N acyltransferase in Cephalosporium acremonium: production of benzylpenicillin by the transformants. Mol Gen Genet 225:56–64
Hillenga DJ, Versantvoort H, van der Molen S, Driessen A, Konings WN (1995) Penicillium chrysogenum takes up the penicillin G precursor phenylacetic acid by passive diffusion. Appl Environ Microbiol 61:2589–2595
Kirschbaum J (1986) Penicillin G, potassium. In: Florey K (ed) Analytical profiles of drug substances, vol 15. Academic Press, New York, pp 427–509
Lamas-Maceiras M, Vaca I, Rodríguez E, Casqueiro J, Martín JF (2006) Amplification and disruption of the phenylacetyl-CoA ligase gene of Penicillium chrysogenum encoding an aryl-capping enzyme that supplies phenylacetic acid to the isopenicillin N acyltransferase. Biochem J 395:147–155
Liras P, Martín JF (2006) Gene clusters for beta-lactam antibiotics and control of their expression: why have clusters evolved, and from where did they originate? Int Microbiol 9:9–19
López-Nieto MJ, Ramos FR, Luengo JM, Martín JF (1985) Characterization of the biosynthesis in vivo of a-aminoadipyl-cysteinyl-valine in Penicillium chrysogenum. Appl Microbiol Biotechnol 22:343–351
Martín JF (2000a) α-Aminoadipyl-cysteinyl-valine synthetases in β-lactam producing organisms. From Abraham’s discoveries to novel concepts of non-ribosomal peptide synthesis. J Antibiot 53:1008–1021
Martín JF (2000b) Molecular control of expression of penicillin biosynthesis genes in fungi: regulatory proteins interact with a bidirectional promoter region. J Bacteriol 182:2355–2362
Martín JF, Ingolia TD, Queener SW (1990) Molecular genetics of penicillin and cephalosporin antibiotic biosynthesis. In: Leong SA, Berka R (eds) Molecular industrial mycology. Marcel Dekker, New York, pp 149–195
Martín JF, Casqueiro J, Liras P (2005) Secretion systems for secondary metabolites: how producer cells send out messages of intercellular communication. Curr Opin Microbiol 8:282–293
Meesschaert B, Eyssen H (1981) Intercorversion of penicillins by mycelium of Penicillium chrysogenum. FEMS Microbiol Lett 10:115–118
Müller WH, van der Krift TP, Krouwer AJ, Woösten HA, van der Voort LH, Smaal EB, Verkleij AJ (1991) Localization of the pathway of the penicillin biosynthesis in Penicillium chrysogenum. EMBO J 10:489–495
Müller WH, Bovenberg RA, Groothuis MH, Kattevilder F, Smaal EB, Van der Voort LH, Verkleij AJ (1992) Involvement of microbodies in penicillin biosynthesis. Biochim Biophys Acta 1116:210–213
Müller WH, Essers J, Humbel BM, Verkleij AJ (1995) Enrichment of Penicillium chrysogenum microbodies by isopycnic centrifugation in nycodenz as visualized with immuno-electron microscopy. Biochim Biophys Acta 1245:215–220
Nestaas E, Demain AL (1981) Influence of penicillin instability on interpretation of feedback regulation experiments. Appl Microbiol Biotechnol 12:170–172
Newbert RW, Barton B, Greaves P, Harper J, Turner G (1997) Analysis of a commercially improved Penicillium chrysogenum strain series: involvement of recombinogenic regions in amplification and deletion of the penicillin biosynthesis gene cluster. J Ind Microbiol Biotechnol 19:18–27
Ramos FR, López-Nieto MJ, Martín JF (1985) Isopenicillin N synthetase of Penicillium chrysogenum, an enzyme that converts delta-(L-alpha-aminoadipyl)-L-cysteinyl-D-valine to isopenicillin N. Antimicrob Agents Chemother 27:380–387
Sikkema J, de Bont JAM, Poolman B (1995) Mechanisms of membrane toxicity of hydrocarbons. Microbiol Rev 59:210–222
Van der Lende TR, van de Kamp M, Berg M, Sjollema K, Bovenberg RA, Veenhuis M, Konings WN, Driessen AJ (2002) delta-(L-alpha-Aminoadipyl)-L-cysteinyl-D-valine synthetase, that mediates the first committed step in penicillin biosynthesis, is a cytosolic enzyme. Fungal Genet Biol 37:49–55
Acknowledgements
This work was supported by grants of the European Union (Eurofung II) and the Junta de Castilla and León (Project LE13/04). C. García-Estrada is supported by the Torres Quevedo Program (PTQ04-3-0411). I. Vaca received a fellowship of the Diputación de León. We thank Jorge Martín for providing the Acremonium chrysogenum (Cephalosporium acremonium) N2 ACV preparations and B. Martín, J. Merino, A. Casenave, and B. Aguado for their excellent technical assistance.
Author information
Authors and Affiliations
Corresponding author
Electronic Supplementary Material
Below is the link to the electronic supplentary material.
Table 2
Production of benzylpenicillin (ng/mg dry weight) by the npe10-DE strain after addition of 0,0278 mM or 0,278 mM exogenous IPN or exogenuous 6-APA. (DOC 34 kb)
Rights and permissions
About this article
Cite this article
García-Estrada, C., Vaca, I., Lamas-Maceiras, M. et al. In vivo transport of the intermediates of the penicillin biosynthetic pathway in tailored strains of Penicillium chrysogenum . Appl Microbiol Biotechnol 76, 169–182 (2007). https://doi.org/10.1007/s00253-007-0999-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-007-0999-4