Abstract
The impact of orotate accumulation in the medically important bacterium Pseudomonas aeruginosa was studied by deleting pyrE, the gene encoding orotate phosphoribosyltransferase and responsible for converting orotate into orotate monophosphate within the de novo pyrimidine synthesis pathway. The pyrE mutant accumulated orotate and exhibited decreased production of hemolysin, casein protease, and elastase. Feeding orotate at a concentration of 51.25 μM to the wild type, PAO1, likewise decreased production of these factors except for hemolysin, which was not affected. A significant increase in the pigments pyocyanin and pyoverdin was also observed. Pyocyanin increase in the pyrE mutant was heightened when the mutant was supplemented with orotate. Although pyoverdin production in the wild-type PAO1 was unaffected by orotate supplementation, a decrease in the mutant’s production was observed when supplemented with orotate. These results indicate a significant reduction in virulence factor production in the pyrE mutant and reduction in some virulence factors in the wild type when supplemented with orotate.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Pseudomonas aeruginosa is an opportunistic pathogen that can cause severe infections in immune-compromised individuals such as patients with cystic fibrosis [24] and is also an important cause of nosocomial infections [4]. This bacterium is also notorious for its ability to resist multiple antibiotic families, making it a challenging infection to treat [12]. The ability to resist multiple types of antibiotics contributes to prolonged hospital stays and increased treatment costs [11]. P. aeruginosa also produces a plethora of virulence factors that contribute to the severity of infections [20].
Given the medical importance of P. aeruginosa, further study is necessary to understand the mechanism by which this organism causes disease. This includes understanding the role that pyrimidine synthesis, important in providing nucleotides for nucleic acid synthesis, plays in virulence factor production. Pyrimidine synthesis in bacteria has been extensively studied [1, 6, 13, 14]. The enzymes of the first three dedicated steps of the pyrimidine pathway (aspartate transcarbamylase encoded by pyrB, dihydroorotase encoded by pyrC, and dihydroorotate dehydrogenase encoded by pyrD) catalyze the production of orotate from carbamoyl phosphate and aspartate. The next step in the pathway is the production of orotate monophosphate (OMP) from orotate and phosphoribosyl pyrophosphate (PRPP) by orotate phosphoribosyltransferase. The gene encoding this enzyme, pyrE, is the subject of this work. The final step of the pyrimidine de novo pathway is the decarboxylation of OMP to uracil monophosphate by the product of the pyrF gene, OMP decarboxylase.
Mutations in the pyrE and pyrF genes lead to accumulation and secretion of orotate in bacteria [26]. A similar phenotype is seen in humans exhibiting orotic aciduria [25]. Brichta [2] first described a novel connection between pyrimidine synthesis and virulence factor production in 2003. A closer look at the effect of orotate on virulence factor production was undertaken here by comparing the impact of orotate accumulation in a pyrE − mutant to the wild-type PAO1.
Materials and Methods
Strains
The strains utilized in this work are listed in Table 1.
Construction of the pyrE Mutant
A pyrE mutant was constructed utilizing the technique described by Choi and Schweizer [3] using the Gateway® technology from Life Technologies™. A 74-bp segment of the 642 bp pyrE, extending from bp336 to bp411, was replaced by a gentamicin resistance cassette. The gentamicin resistance cassette was utilized for final screening of the mutant. The modified pyrE was amplified from the mutant with PCR and sequenced for confirmation.
PyrE and PyrF Assays
The enzymatic activities of both PyrE and PyrF were measured using methods described by Beckwith et al. [1] and Schwartz and Neuhard [21, 22]. Cells were grown in Pseudomonas minimal media (PsMM) [15] to late log phase, reaching optical density (OD600 nm) of 0.9 A. Cells were lysed by sonication using a Branson 200 Cell Disruptor for 1 min intervals to 6 min total with power amplitude at setting 5, and a cell-free extract was produced by centrifugation at 10 °C for 25 min at 1876.9×g using a Sorvall RT600 with an H1000B rotor. The extract was dialyzed against 2 mM β-mercaptoethanol, 20 μM ZnSO4, 50 mM Tris–HCl, pH 8.0 at 4 °C for 18 h, and then kept at 4 °C until use, typically within 24 h. Reaction buffers contained 100 mM Tris/HCl buffer at pH 8.6, 6 mM MgCl2 for both enzymes. In addition, 0.25 mM orotate and 0.6 mM PRPP were added as substrates for PyrE or 0.2 mM OMP included for PyrF assays. 1 mg/ml protein from cell extracts was used for both reactions using a Bradford assay standard curve. Incubation was at 30 °C for 5 min before the addition of substrates. Reactions were monitored every 5 min for 15 min. PyrE activity was measured at λ 295 nm and PyrF at λ 290 nm. A change equal to 3.67 absorbance units is equal to 1 mM increase in OMP for PyrE, while a change equal to 1.38 absorbance units is equal to 1 mM decrease in OMP for PyrF.
Casein Protease
Casein protease activity was assessed for pyrE −, and wild-type PAO1 cells were grown in 5 ml peptone tryptic soy broth (PTSB) for 24 h. Cells were collected by centrifugation at 1876.9×g using a Sorvall RT600 centrifuge with a H1000B rotor. The supernatant was filter-sterilized using a 0.45-μM filter. Reaction solution contained 0.05 M Tris, pH 7.5, and 0.5 mM CaCl2 containing 0.3 % azocasein from Sigma-Aldrich™ [7]. The reaction contained 50 μl filtered supernatant in a 1-ml reaction solution. The reaction was incubated at 37 °C for 15 min and terminated with 0.5 ml of 10 % trichloroacetic acid. Samples were then centrifuged, and absorbance of the supernatant was determined using a spectrophotometer at λ 400 nm.
Elastase Production
Elastase production was determined using the elastin congo red assay [16]. Supernatants were prepared as for casein protease measurements. The reaction solution contained 0.1 M Tris, pH 7.2, and 1 mM CaCl2. The reaction contained 50 μl of supernatant in 1 ml reaction solution with 20 mg of elastin congo red (ECR) from Sigma-Aldrich™. The reaction mixture was incubated for 18 h at 37 °C. The mixture was placed on ice, and EDTA was added to 0.12 mM. After a 10 min incubation, the sample was centrifuged to remove remaining ECR, and the absorbance was measured at λ 495 nm.
Hemolysin Production
TSA plates supplemented with 5 % sheep blood were used to test for the production of hemolysin. Using a sterile toothpick, a culture samples from 24 h colonies were patched onto blood agar plates, which were then incubated for 48 h at 37 °C. Orotate-supplemented plates contained 51.25 μM orotate.
Orotate Accumulation and Secretion
Cells were grown at 37 °C in PsMM broth to OD600 nm 0.6 A. Cells were then removed by centrifugation to yield final cell pellets and supernatants. For orotate secretion, uninoculated medium with known concentration of orotate was used as reference to create an orotate standard curve at λ 278 nm. Uninoculated medium was used as blank and media for both mutant and wild-type P. aeruginosa samples. Cells were sonicated in breaking buffer (2 mM β-mercaptoethanol, 20 μM ZnSO4, 50 mM Tris–HCl, pH 8.0, and 20 % glycerol) without orotate supplement, and cell debris was removed by centrifugation. Orotate was measured in the supernatant at λ 278 nm [17].
Pyocyanin Production Assay
The method adapted from Essar et al. [5] was used for this assay. 30 ml of King’s A [8] broth was used in a 125-ml flask. Bacteria were grown in King’s A broth for 20 h in triplicate samples at 37 °C with shaking. To determine cell density, absorbance reading was measured at λ 600 nm for each flask. Five ml of the sample was transferred to a 15-ml conical centrifuge tube and centrifuged for 25 min at 1876.9×g using a H1000B rotor on a Sorval TR 6000B centrifuge. The supernatant was then filtered through a 0.45-μm filter into another 15 ml conical centrifuge tube. To precipitate organic matter, 3 ml of chloroform was added, and the mixture was vortexed 10 times for 2 s. The mixture was then centrifuged for 7 min at 1876.9×g using the same conditions mentioned above. The top layer was removed using a 5-ml pipet, 1.5 ml of 0.2 N HCL was added, and the mixture was vortexed 10 times for 2 s. The sample was centrifuged again at 1876.9×g for 7 min. The top pink layer was collected, and the absorbance was measured at λ 520.
Pyoverdin Production Assay
50 ml of King’s B broth [8] in a 125-ml flask was inoculated with freshly grown cells. The broth was incubated for 12 h with shaking at 37 °C. After 12 h, cell density was measured at λ 600 nm. 1 ml of culture was centrifuged at 10053×g in a Speed Fuge® HSC10 K microcentrifuge using a HSR-24 rotor. The absorbance of the supernatant was measured at λ 405 nm. The results were recorded as 12-h λ 405 nm/λ 600 nm [22].
Results
PyrE/PyrF Activities and Orotate Accumulation
A complete loss of PyrE activity was observed in the mutant. As expected, the resultant absence of pyrF inducer (OMP) eliminated expression of this gene, too. It was found that the PAO1 had 1.96 μM orotate in cell extracts, while the pyrE − had 3.36 μM. This represented a 58 % increase in the mutant strain. Wild-type PsMM cultures had no detectable orotate in the medium, while medium from pyrE − cultures contained 0.403 μM of orotate.
Elastase and Casein Protease Assays
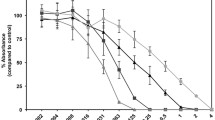
pyrE − cultures exhibited a sharp decrease in elastase production as compared to PAO1, with an average reading of 0.035 A at λ 495 nm/λ 600 nm, which compared to 0.102 A for the wild type. Elastase production in the wild-type fed orotate decreased to an average reading of 0.037 A, which is a similar level to that of the mutant (Fig. 1). No casein protease was measurable for the pyrE − cells. Casein protease production by the PAO1 showed a 64 % reduction when cells were supplemented with orotate, falling from an average reading at λ 495/λ 600 for non-supplemented wild type of 0.0797 to 0.0512 A for orotate-supplemented (at 51.25 μM) wild type (Fig. 2).
Hemolysis
The wild type showed β-hemolysis both with and without orotate supplementation. The pyrE − exhibited α-hemolysis. However, when fed orotate, the pyrE − lost its hemolysis completely (γ-hemolysis) (Fig. 3).
Pyocyanin Production
Pyocyanin production increased significantly by the pyrE − compared to PAO1. PAO1 produced 4.2 μM, while there was an increase in pyocyanin production to 12.34 μM. When feeding the pyrE − orotate at 51.25 μM, it was observed that there was an additional increase in pyocyanin production to 15.1 μM, while PAO1 increased to 4.97 μM (Fig. 4).
Pyoverdin Production
The pyrE − produced almost double the amount of pyoverdin produced by PAO1 when measured at 12 h of incubation. When adding orotate to the media at a concentration of 51.25 μM, PAO1 did not show any increase in pyoverdin production, while the pyrE − dropped to levels similar to that of PAO1 (Fig. 5).
Discussion
Previous studies have shown that uracil increases quorum sensing activity in P. aeruginosa and, as a result, increases virulence factor production [23]. Because uracil increases quorum sensing activity, it was expected that orotate would act likewise given the similarity in structure. Surprisingly, the opposite was found as orotate supplementation or overproduction decreased the production of the virulence factors casein protease, elastase, and hemolysin. Brichta [2] found a connection between pyrimidine synthesis and virulence factor production through construction of a double knockout on pyrC that led to decreased virulence factor production. In comparison, we find that knocking out pyrC led to decreased production of hemolysin, casein protease, elastase, pyoverdin, and pyocyanin. Casein protease was decreased in the pyrC − unlike the pyrE − where casein protease activity was completely absent. Pigment production in the pyrC − showed decreased pyoverdin and pyocyanin production, while the pyrE − showed an increase in these pigments. The pyoverdin increase is possibly due to the accumulation of dihydroorotate which is proposed to be a part of the pyoverdin structure [10]. What is unclear is why the pyrE − showed a sharp increase in pyocyanin production. When feeding wild-type PAO1 orotate at 51.25 μM, pyocyanin increased slightly as also seen when the mutant is fed orotate. With dihydroorotate being structurally a part of pyoverdin, it is possible that orotate may also be involved in the structure of pyocyanin or it may be a stimulator of the pyocyanin synthesis pathway. There is a similar pattern of increase in both pyocyanin and pyoverdin levels, indicating that orotate may be involved in the production of both. Further studies could explore these relationships, including utilizing transcriptomics or 14C labeled orotate.
Even though it was shown in a previous study by Santiago and West [19] that pyoverdin also increased slightly in Pseudomonas putida pyrC mutants, for the pyrE mutant in the same study, a much higher increase in pyoverdin production was seen. This may indicate that dihydroorotate is not absolutely essential for pyoverdin production in P. putida. It is also observed in P. aeruginosa that pyrC double knockouts showed a sharp drop in pyoverdin production which differs from the results observed in P. putida [2]. Supplementation of media with orotate clearly decreased virulence factor production in the wild-type strain of P. aeruginosa, and the pyrE − strain also showed a significant decrease in virulence factors, which is likely due to the accumulation of orotate in these cells. The decreases in virulence factors seen in this study could also be due to disruption of the quorum sensing network by orotate.
The reduction in virulence factor production as a result of disrupting the pyrimidine synthesis pathway could provide a target with implications for the control of P. aeruginosa infections. Finding approaches which attack the P. aeruginosa pyrimidine synthesis pathway without harming eukaryotic cell pyrimidine synthesis would be a significant step in this direction. However, due to the similarities between the eukaryotic pyrimidine synthesis pathway and the bacterial pathway, it might be difficult to achieve that goal. Similar strategies, such as 5-flourouracil used as a treatment for skin cancer [9], would be a starting point for consideration. Other pyrimidine analog might be utilized to inhibit the pyrimidine pathway in bacteria in concert with antibiotics. This might especially be useful when bacteria form complex biofilms inside the body where they are protected from phagocytosis and traditional antibiotics. It would also be intriguing to take a look at the effect of external application of orotate to subjects infected with P. aeruginosa. Exogenous feeding of orotate could reduce disease symptoms and help to combat the infection.
Further study on the increased production of pyocyanin could also have commercial implications. In 2005, Rabaey et al. [18] showed that pyocyanin increased electron transfer in biofuel cells. They found that introduction of mutants that were deficient in pyocyanin production resulted in reduction of power output to 5 %. Output increased up to 50 % when pyocyanin was added. The addition of pyocyanin also enhanced power output by other bacteria within the biofuel cell. Additional studies on Pseudomonas strains that are known to produce high amounts of pyocyanin could determine if knocking out pyrE might further increase pyocyanin production. If so, higher pyocyanin production in P. aeruginosa could contribute to improved biofuel cell output.
References
Beckwith JR, Pardee AB, Austrian R, Jacob F (1962) Coordination of the synthesis of enzymes in the pyrimidine pathway of E. coli. J Mol Biol 5:618–634
Brichta DM (2003) Construction of a Pseudomonas aeruginosa dihydroorotase mutant and the discovery of a novel link between pyrimidine biosynthetic intermediates and the ability to produce virulence factors. Ph.D. Dissertation, University of North Texas
Choi KH, Schweizer HP (2005) An improved method for rapid generation of unmarked Pseudomonas aeruginosa deletion mutants. BMC Microbiol 5:30
Edmond MB, Wallace SE, McClish DK, Pfaller MA, Jones RN, Wenzel RP (1999) Nosocomial blood infection in United States hospitals: a three-year analysis. Clin Infect Dis 29:239–244
Essar DW, Eberly L, Hadero A, Crawford IP (1990) Identification and characterization of genes for a second anthranilate synthase in Pseudomonas aeruginosa: interchangeability of the two anthranilate synthases and evolutionary implications. J Bacteriol 172:884–900
Evans DR, Guy HI (2004) Mammalian pyrimidine biosynthesis: afresh insight into an ancient pathway. J Biol Chem 279:30335–33038
Kessler E, Safrin M, Olson JC, Ohman DE (1993) Secreted LasA of Pseudomonas aeruginosa is a staphylolytic protease. J Biol Chem 268:7503–7508
King EO, Ward MK, Raney DE (1954) Two simple media for the demonstration of pyocanin and fluorescin. J Lab Clin Med 44:301–307
Love WE, Bernhard JD, Bordeaux JS (2009) Topical imiquimod or fluorouracil therapy for basal and squamous cell carcinoma: asystematic review. Arch Dermatol 145:1431–1438
Maksimova NP, Blazhevich OV, Fomichev IuK (1993) The role of pyrimidines in the biosynthesis of the fluorescing pigment pyoverdin Pm in Pseudomonas putida M. Mol Gen Mikrobiol Virusol 5:22–26
Mauldin PD, Salgado CD, Hansen IS, Durup DT, Bosso JA (2009) Attributable hospital cost and length of stay associated with health care-associated infections caused by antibiotic-resistant gram-negative bacteria. Antimicrob Agents Chemother 54:109–115
Morita Y, Tomida J, Kawamura Y (2013) Responses of Pseudomonas aeruginosa to antimicrobials. Front Microbiol 4:422
Neuhard J, Kelln RA, Stauning E (1986) Cloning and structural characterization of the Salmonella typhimurium pyrC gene encoding dihydroorotase. Eur J Biochem 157:335–342
O’Donovan GA, Neuhard J (1970) Pyrimidine metabolism in microorganisms. Bacteriol Rev 34:278–343
Ornston LN, Stanier RY (1966) The conversion of catechol and protocatechuate to beta-ketoadipate by Pseudomonas aeruginosa. J Biol Chem 241:3776–3786
Pearson JP, Pesci EC, Iglewski BH (1997) Roles of Pseudomonas aeruginosa las and rhl quorum-sensing systems in control of elastase and rhamnolipid biosynthesis genes. J Bacteriol 179:5756–5767
Peters GJ, Laurensse E, Leyva A, Pinedo HM (1987) A sensitive, nonradiometric assay for dihydroorotic acid dehydrogenase using anion-exchange high-performance liquid chromatography. Anal Biochem 15:32–38
Rabaey K, Boon N, Höftee M, Verstraete W (2005) Microbial phenazine production enhances electron transfer in biofuel cells. Environ Sci Technol 39:3401–3408
Santiago MF, West TP (2002) Control of pyrimidine formation in Pseudomonas putida ATCC 17536. Can J Microbiol 48:1076–1081
Schuek KN, Breidenstein E, Hancock RE (2012) Pseudomonas aeruginosa: a persistent pathogen in cycstic fibrosis and hospital-associated infections. In: Dougherty TJ, Pucci MJ (eds) Antibiotic discovery and development, 1st edn. Springer, New York, pp 679–715
Schwartz M, Neuhard J (1975) Control of expression of the pyr genes in Salmonella typhimurium: effects of variations in uridine and cytidine nucleotide pools. J Bacteriol 121:814–822
Stintzi A, Barnes C, Xu J, Raymond KN (2000) Microbial iron transport via a siderophore shuttle: a membrane ion transport paradigm. Proc Natl Acad Sci USA 97:10691–10696
Ueda A, Attila C, Whiteley M, Wood KT (2009) Uracil influences quorum sensing and biofilm formation in Pseudomonas aeruginosa and fluorouracil is an antagonist. Microb Biotechnol 2:62–74
Wilson R, Dowling RB (1998) Lung infections. 3. Pseudomonas aeruginosa and other related species. Thorax 53:213–219
Winkler JK, Suttle DP (1988) Analysis of UMP synthase Gene and mRNA structure in hereditary orotic aciduria fibroblasts. Am J Hum Genet 43:86–94
Womack JE, O’Donovan GA (1978) Orotic acid excretion in some wild-type strains of Escherichia coli K-12. J Bacteriol 136:825–827
Acknowledgments
The authors would like to acknowledge the late Dr. Gerard A. O’Donovan for his contributions to the development of this work and Dr. Robert C. Benjamin for his insightful comments on the manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Niazy, A., Hughes, L.E. Accumulation of Pyrimidine Intermediate Orotate Decreases Virulence Factor Production in Pseudomonas aeruginosa . Curr Microbiol 71, 229–234 (2015). https://doi.org/10.1007/s00284-015-0826-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-015-0826-6