Abstract
The ability to produce vanillin and/or vanillic acid from isoeugenol was screened using resting cells of various bacteria. The vanillin- and/or vanillic-acid-producing activities were observed in strains belonging to the genera Achromobacter, Aeromonas, Agrobacerium, Alcaligenes, Arthrobacter, Bacillus, Micrococcus, Pseudomonas, Rhodobacter, and Rhodococcus. Strain IE27, a soil isolate showing the highest vanillin-producing activity, was identified as Pseudomonas putida. We optimized the culture and reaction conditions for vanillin production from isoeugenol using P. putida IE27 cells. The vanillin-producing activity was induced by adding isoeugenol to the culture medium but not vanillin or eugenol. Under the optimized reaction conditions, P. putida IE27 cells produced 16.1 g/l vanillin from 150 mM isoeugenol, with a molar conversion yield of 71% at 20 °C after a 24-h incubation in the presence of 10% (v/v) dimethyl sulfoxide.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
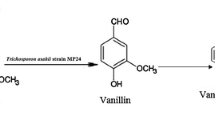
Lignin-related phenylpropanoides such as eugenol and isoeugenol have attracted attention as natural renewable sources for the production of useful chemicals (Rabenhorst 1996). Eugenol has great potential as a starting material for the synthesis of aromatic flavorings and aromas. Pseudomonas and Corynebacterium strains degrade eugenol to vanillin via coniferyl alcohol, coniferyl aldehyde, and ferulic acid as shown in Fig. 1 (Takada 1977; Takada and Kayahara 1983). The initial step of eugenol degradation was confirmed to be the double-bond-transferring hydroxylation catalyzed by eugenol dehydrogenase (Furukawa et al. 1998; Wieser et al. 1999). The gene loci ehyA and ehyB of Pseudomonas sp. HR199 were identified as the structural genes of eugenol hydroxylase (dehydrogenase) (Priefert et al. 1999). The biotransformation of eugenol to ferulic acid was demonstrated, in which Ralstonia eutropha H16 carrying ehyAB, calA, and calB genes encoding eugenol hydroxylase, coniferyl alcohol dehydrogenase, and coniferyl aldehyde dehydrogenase, respectively, was used (Overhage et al. 2002). The biotransformation of isoeugenol using isoeugenol-degrading bacteria has recently been reported. Bacillus sp. cells degraded isoeugenol to vanillic acid via vanillin, and its cell-free extract transformed isoeugenol to vanillin (Shimoni et al. 2000). The resting cells of P. putida I58 produced vanillic acid from isoeugenol with a high molar yield, without the accumulation of vanillin (Furukawa et al. 2003). The initial step of isoeugenol degradation is of interest because of oxidative cleavage of the double bond of isoeugenol. However, the enzyme(s) catalyzing the oxidative cleavage of isoeugenol have not been purified from isoeugenol-degrading microorganisms yet.
In the present study, to find microorganisms showing high vanillin-producing activity, we carried out an extensive screening of isoeugenol-degrading microorganisms and attempted the conversion of isoeugenol to vanillin using resting cells of Pseudomonas putida IE27, a novel isoeugenol-degrading bacterium.
Materials and methods
Chemicals
Isoeugenol, vanillin, and NZ amine were obtained from Wako Pure Chemical Industries (Japan). Peptone and casamino acids were purchased from Nihon Pharmacy (Japan). Meat extract was obtained from Kyokuto Seiyaku (Japan). Yeast extract was purchased from Oriental Yeast (Japan). Malt extract was obtained from Bacto (France). Soy bean hydrolyzate was provided by Ajinomoto (Japan). Corn steep liquor was obtained from Shono Denpun (Japan). All of the other chemicals used were of analytical grade and commercially available.
Isolation of isoeugenol-degrading microorganisms
For the isolation of isoeugenol-degrading microorganisms, a conventional enrichment culture was carried out at 28 °C. Soils were used as the source for the isolation of isoeugenol-degrading microorganisms. The soils were put in 50 ml of the following medium I containing 0.005–0.5% (v/v) isoeugenol. Medium I comprised 3 g (NH4)2HPO4, 2 g KH2PO4, 1 g NaCl, 0.05 g MgSO4·7H2O, 0.01 g FeSO4·7H2O, 0.005 g MnSO4·xH2O, 5 ml metal solution (Uchida et al. 2003), and 5 ml vitamin mixture (Yoshida et al. 2004) in 1 l of tap water, pH 7.0. The microorganisms isolated on agar plates were stored on slants of nutrient medium: 5 g peptone, 5 g meat extract, 0.5 g yeast extract, and 2 g NaCl in 1 l of tap water, pH 7.0.
Culture conditions and preparation of resting cells
Subculture was done at 28 °C for 12 h with reciprocal shaking at 120 rpm in a test tube containing 5 ml of the above nutrient medium. The subculture was transferred to 50 ml of the same medium in a 500-ml shaking flask and incubated at 28 °C. After a 12-h cultivation, 0.5% (v/v) isoeugenol was added to the culture broth, after which cultivation was carried out for another 12 h. Cells were harvested by centrifugation at 9,000×g at 4 °C for 20 min, followed by washing twice with 50 mM potassium phosphate buffer (pH 7.0) after which they were suspended in the same solution.
The cell growth of P. putida IE27 was estimated turbidimetrically at 610 nm, and 0.374 mg dry cell weight ml−1 was equivalent to 1.0 unit of optical density at 610 nm.
Resting cells reaction
The reaction mixture contained 300 μmol isoeugenol, 1 mmol potassium phosphate buffer (pH 7.0), and resting cells derived from 75 ml culture broth at a final volume of 10 ml. The reaction was carried out in a 50-ml conical flask with reciprocal shaking (160 strokes min 1) at 30 °C, started by the addition of isoeugenol and stopped by adding 10 ml ethanol. The reaction mixture was centrifuged at 13,000×g for 5 min and the resulting supernatant was analyzed by high-performance liquid chromatography (HPLC). Vanillin-producing activity was defined as the amount of vanillin formed per minute by resting cells derived from 1 ml of culture broth.
The effect of pH was investigated using 100 mM of the following buffers: citrate–sodium citrate (pH 3.0–6.0), acetate–sodium acetate (pH 4.0–5.5), potassium phosphate (pH 6.0–8.0), Tris–HCl (pH 7.5–9.0), and glycine–NaOH (pH 9.0–11.0). To optimize the reaction conditions, the following organic solvents were tested: methanol, ethanol, 1-propanol, 2-butanol, 1-heptanol, 1-octanol, acetone, acetonitrile, diisopropyl ether, t-butylethyl ether N,N-dimethylformamide, dimethyl sulfoxide (DMSO), ethyl acetate, butyl acetate, chloroform, n-heptane, n-octane, and cyclohexane.
Optimization of culture conditions for P. putida IE27
To optimize the culture medium, the following carbon and/or nitrogen sources were used: glucose, maltose, sucrose, fructose, lactose, glycerol, succinic acid, sodium citrate, sodium fumarate, (NH4)2HPO4, NaNO3, NH4Cl, sodium l-glutamine, NZ amine, casamino acids, yeast extract, peptone, meat extract, malt extract, corn steep liquor, and soy bean hydrolyzate.
Identification of reaction products
Vanillin produced by P. putida IE27 cells was extracted from the reaction mixture with chloroform and concentrated by evaporation. Vanillin was extracted from the concentrate with hot water and kept at 4 °C; it was obtained as a white needle crystalline solid in water. The product was identified by 1H NMR and 13C NMR with authentic vanillin as a reference: (a) 1H NMR (500 MHz, CDCl3): δ 3.97 (3 H, s), 6.31 (1 H, s), 7.05 (1 H, d, J=8.6 Hz), 7.26 to 7.44 (2 H, m) and 9.83 (1 H, s) and (b) 13C NMR (125 MHz, CDCl3): δ 56.1, 108.7, 114.4, 127.5, 129.8, 147.1, 151.7, and 191.0.
Analytical methods
Isoeugenol, vanillin, and vanillic acid were determined by HPLC (Shimadzu LC-10AS and SPD-10A system) with a Wakosil-II 5C18 HG column (4.6×150 mm, Wako Pure Chemical Industries, Japan). Elution was done at 40 °C with a 40:57:3 (v/v/v) mixture of methanol, water, and acetic acid. The flow rate was maintained at 1.0 ml/min, and the absorbance of eluates was monitored at 280 nm.
Results
Screening of isoeugenol-degrading microorganisms
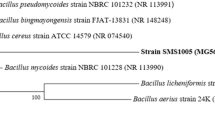
Twenty isoeugenol-degrading microorganisms were isolated through the enrichment culture using isoeugenol as a sole carbon source. Using the soil isolates and 245 bacteria stocked in our laboratory, vanillin-producing activity was examined by the resting cells reaction. Strain IE27, a soil isolate, exhibited the highest vanillin-producing activity. In our bacterial collection, the resting cells of 17 strains belonging to the genera Achromobacter, Aeromonas, Agrobacterium, Alcaligenes, Arthrobacter, Bacillus, Micrococcus, Pseudomonas, Rhodobacter, and Rhodococcus formed vanillin and/or vanillic acid from isoeugenol (Table 1). Vanillin- and/or vanillic-acid-forming activity was widely distributed among not only gram-negative but also gram-positive bacteria. In our previous studies on vanillic acid production from isoeugenol using P. putida I58 (Furukawa et al. 2003), the resting cells of the bacterium converted 10 mM isoeugenol to 9.8 mM vanillic acid by a 40-min incubation, during which 0.3 mM vanillin accumulated transiently in the reaction mixture under the optimized conditions. When the resting cells of strain IE27 were incubated with 10 mM isoeugenol, 4.2 mM vanillin accumulated predominantly after a 24-h incubation, with the formation of 2.1 mM vanillic acid. This indicates that strain IE27 is a promising vanillin producer. The 16S ribosomal DNA sequence of strain IE27 showed 99.9% and 99.7% identities to P. putida KT2440 and P. putida ATCC17453, respectively. From these results, strain IE27 was identified to be P. putida.
Optimization of culture conditions for P. putida IE27
The vanillin-producing activity of P. putida IE27 was calculated to be 2.01 nmol min−1 ml−1 by resting cells reaction for 30 min. To enhance the activity of P. putida IE27 cells, culture conditions for the bacteria were optimized using 300 ml of medium I containing 0.05% (v/v) isoeugenol in 2-l flask. Activity was measured by reaction using the resting cells. The addition of 0.1% (v/v) glycerol and 1% (w/v) yeast extract was effective in promoting the growth of P. putida IE27, resulting in an enhancement of vanillin-producing activity. The optimum concentration of isoeugenol was 0.025% (v/v), while bacterial growth was partially inhibited at the higher concentration of 0.05% (v/v). The highest activity of 210 nmol min−1 ml−1 was obtained after a 24-h cultivation. The vanillin-producing activity was induced by adding isoeugenol, p-hydroxycinnamic acid, ferulic acid, and trans-anethole to the medium, the activities being 87.5, 26.5, 14.7, and 3.0 nmol min−1 (mg dry cell weight)−1, respectively. Vanillin and eugenol induced no activity at all.
Substrate specificity
Substrate specificity was examined by the resting cells reaction (Table 2). Isoeugenol was the most suitable substrate. The resting cells of P. putida IE27 acted on 2-methoxy-4-vinylphenol and trans-anethole to produce vanillin and anisaldehyde, respectively, although their activities were much lower than that of isoeugenol. The following compounds were inert as substrates: eugenol, vanillic acid, vanillyl alcohol, anisaldehyde, ferulic acid, coniferyl alcohol, 3,4-dihydroxycinnamic acid, 3,4-dimethoxycinnamic acid, p-methoxycinnamic acid, p-hydroxycinnamic acid, 3-pyridinepropionic acid, α-methoxycinnamic acid, styrene, trans-β-methylstyrene, trans-stilbene, trans-stilbene oxide, benzoic acid, acrylic acid, and trans-2-hexenoic acid.
Conditions for vanillin production by resting cells of P. putida IE27
The reaction conditions for vanillin production from isoeugenol by resting cells of P. putida IE27 were optimized. We studied the effect of isoeugenol concentration on vanillin production and found that it was high when the reaction mixture contained 30–200 mM isoeugenol (Fig. 2a). Although the resting cells of P. putida IE27 contained the oxidation activity of vanillin to vanillic acid, the formation of the latter type was significantly inhibited in the presence of high concentrations of isoeugenol. The thermal stability of vanillin-producing activity was also investigated. After the resting cells had been incubated at temperatures up to 20 °C for 30 min, no loss of activity was observed. Treatments at 25, 30, 40, and 50 °C caused 15, 28, 30, and 100% losses of initial activity, respectively. The effect of reaction temperature on vanillin production was examined at 20 and 30 °C (Fig. 2b). Although vanillin was slowly produced by incubation at 20 °C, 62 mM vanillin accumulated in the reaction mixture. When the reaction was carried out at 30 °C, the formation of vanillin was decelerated after a 6-h incubation, probably owing to the thermal stability of isoeugenol-degrading enzyme(s). The effect of various buffers at different pHs was examined. Glycine–NaOH buffer (pH 10.5) was the most effective; its optimum concentration was 100 mM. To enhance the productivity of vanillin, the effect of organic solvents was investigated. The addition of N,N-dimethylformamide or DMSO at 10% (v/v) was effective in promoting vanillin-producing activities of 106 and 114%, respectively, of that in the case of no solvent added. The optimum concentration of DMSO was found to be 10% (v/v) (Fig. 2c). The addition of DMSO also inhibited the formation of vanillic acid. The addition of 2 ml of ethyl acetate, butyl acetate, 1-octanol, and 1-heptanol in 10 ml of reaction mixture did not affect the activity. The other organic solvents tested considerably inhibited the vanillin-producing activity of P. putida IE27 cells.
Effect of isoeugenol concentration, reaction temperature, and DMSO concentration on vanillin production by P. putida IE27 cells. The reaction mixture contained 2 mmol isoeugenol, 95 mg cells as dry weight, and 100 mM potassium phosphate buffer (pH 7.0) in a total volume of 10 ml. Reaction was carried out at 30°C for 24 h with shaking at 160 strokes min−1. Isoeugenol concentration (a), reaction temperature (b), and DMSO concentration (c) were varied as indicated. Closed circles, vanillin formation at 30°C; open circles, vanillin formation at 20°C; triangles, vanillic acid
Vanillin production by resting cells of P. putida IE27
The time course of vanillin production was examined under the optimized reaction conditions (Fig. 3). After a 24-h incubation at 20 °C in the presence of 10% (v/v) DMSO, 106 mM (16.1 g/l) vanillin had accumulated from 150 mM isoeugenol with a molar conversion yield of 71%. The formation of vanillic acid was severely constrained to 2.8 mM. As the remaining isoeugenol was 23 mM in the reaction mixture, 18 mM of the added isoeugenol might have vaporized from the mixture during the reaction. The vanillin produced was partly solidified because of its low solubility in the reaction mixture.
Time course of vanillin production by P. putida IE27 cells. The reaction was aerobically agitated using a magnetic stirrer at 20°C in a total volume of 10 ml containing 1.5 mmol isoeugenol, 319 mg cells as dry weight, 100 mM glycine–NaOH buffer (pH 10.5), and 10% (v/v) DMSO. Circles, vanillin; triangles, vanillic acid; square, isoeugenol
Vanillin production using isoeugenol as substrate and solvent
Zhao et al. (2005) recently reported a biotransformation by Bacillus fusiformis CGMCC1347 cells using 60% (v/v) isoeugenol as both substrate and solvent. We also examined vanillin production by P. putida IE27 in the presence of a large amount of isoeugenol. In the optimized reaction mixture, 60% (v/v) isoeugenol (corresponding to 3.97 M) was substituted for 10% (v/v) DMSO and 150 mM isoeugenol. After a 24-h incubation at 20 °C, 136 mM (20.7 g/l) vanillin was produced in 10 ml of the mixture. During the reaction, the viscosity of the reaction mixture increased owing to the formation of emulsion. The addition of 10 ml ethyl acetate was effective in decreasing the viscosity, resulting in an enhancement of vanillin productivity. In the presence of ethyl acetate, 3.00 mmol of vanillin was produced in 24 h in 20 ml of the reaction mixture. In 13.7 ml of the ethyl acetate phase, 87% of the vanillin produced was dissolved at 191 mM (29.1 g/l). The product concentration was about two times higher than that in the presence of 10% (v/v) DMSO. However, 92% of the added isoeugenol remained in the reaction mixture.
Discussion
As the microbial degradation of isoeugenol had been reported previously only with Pseudomonas and Bacillus strains (Priefert et al. 2001), we extensively surveyed isoeugenol-degrading microorganisms, especially the formation of vanillin and/or vanillic acid from isoeugenol using a variety of bacterial strains. P. putida IE27 was found to show the highest vanillin-producing activity among isoeugenol-degrading bacteria tested. In previous studies on the bioconversion of isoeugenol, the cell-free extract of Bacillus sp. transformed isoeugenol into vanillin with a molar yield of 14% by a 48-h reaction, the product concentration being 0.9 g/l of the reaction mixture (Shimoni et al. 2000). In our previous studies on biotransformation of isoeugenol using P. putida I58, the resting cells of the bacterium produced 1.6 g/l vanillic acid with a molar yield of 98% by a 40-min incubation, without the accumulation of vanillin (Furukawa et al. 2003). In contrast with P. putida I58, the present soil isolate, strain IE27, produced predominantly vanillin from isoeugenol. Although P. putida IE27 cells contained the oxidation activity of vanillin to vanillic acid, a high concentration of isoeugenol inhibited the activity, leading to efficient vanillin production. The vanillinoxidizing activity of P. putida I58 is probably less sensitive to isoeugenol. In the present study, we have attempted to optimize the culture and reaction conditions to enhance the productivity of vanillin from isoeugenol using P. putida IE27. Although isoeugenol shows antimicrobial activity against most bacteria, P. putida IE27 grew well in the presence of 0.025% (v/v) isoeugenol under optimized culture conditions. The vanillin-producing activity of P. putida IE27 was induced effectively by isoeugenol, which was also the best substrate, suggesting that the physiological substrate of isoeugenoldegrading enzyme(s) is isoeugenol. The low solubility of isoeugenol in water (at most 6 mM) was disadvantageous to the efficient biotransformation of isoeugenol to vanillin. The addition of 10% (v/v) DMSO, however, enhanced its solubility, resulting in higher vanillin productivity. The higher concentration of isoeugenol solubilized by DMSO resulted in the inhibition of vanillic acid formation. Thus, the resting cells of P. putida IE27 attained 16.1 g/l vanillin (106 mM) from 150 mM isoeugenol with a molar conversion yield of 71% after a 24-h incubation in the presence of 10% (v/v) DMSO (Fig. 3).
Zhao et al. (2005) recently reported on vanillin production in the presence of 60% (v/v) isoeugenol using B. fusiformis CGMCC1347 cells, in which isoeugenol was used as both the substrate and the organic solvent of a two-phase system. A high level of vanillin accumulation was attained after a 72-h incubation (32.5 g/l, probably in the isoeugenol phase). When we carried out the conversion of isoeugenol into vanillin in the presence of 60% (v/v) isoeugenol, P. putida IE27 cells likewise exhibited a similar level of vanillin accumulation; however, the reaction mixture became so viscous and sticky that it was difficult to separate vanillin from the reaction mixture. We then carried out the reaction by adding ethyl acetate to reduce the viscosity of the reaction mixture, which resulted in the accumulation of 191 mM (29.1 g/l) vanillin in the ethyl acetate layer after a 24-h incubation. However, we ultimately realized that the conversion of isoeugenol into vanillin in the two-phase system using ethyl acetate was neither a practical nor an efficient procedure. The conversion yield of isoeugenol to vanillin proved to be extremely low, and the isolation of vanillin was laborious. Therefore, it became clear that, from a practical standpoint, for efficient vanillin production, the one-phase system adding 10% (v/v) DMSO to dissolve isoeugenol in water was superior to the two-phase system.
References
Furukawa H, Wieser M, Morita H, Sugio T, Nagasawa T (1998) Purification and characterization of eugenol dehydrogenase from Pseudomonas fluorescens E118. Arch Microbiol 171:37–43
Furukawa H, Morita H, Yoshida T, Nagasawa T (2003) Conversion of isoeugenol into vanillic acid by Pseudomonas putida I58 cells exhibiting high isoeugenol-degrading activity. J Biosci Bioeng 96:401–403
Overhage J, Steinbüchel A, Priefert H (2002) Biotransformation of eugenol to ferulic acid by a recombinant strain of Ralstonia eutropha H16. Appl Environ Microbiol 68:4315–4321
Priefert H, Overhage J, Steinbüchel A (1999) Identification and molecular characterization of the eugenol hydroxylase genes (ehyA/ehyB) of Pseudomonas sp. strain HR199. Arch Microbiol 172:354–363
Priefert H, Rabenhorst J, Steinbüchel A (2001) Biotechnological production of vanillin. Appl Microbiol Biotechnol 56:294–314
Rabenhorst J (1996) Production of methoxyphenol-type natural aroma chemicals by biotransformation of eugenol with a new Pseudomonas sp. Appl Microbiol Biotechnol 46:470–474
Shimoni E, Ravid U, Shoham Y (2000) Isolation of a Bacillus sp. capable of transforming isoeugenol to vanillin. J Biotechnol 78:1–9
Takada K (1977) Degradation of eugenol by a microorganism. Agric Biol Chem 41:925–929
Takada K, Kayahara H (1983) Initial steps of eugenol degradation pathway of a microorganism. Agric Biol Chem 47:2639–2640
Uchida A, Ogawa M, Yoshida T, Nagasawa T (2003) Regioselective hydroxylation of quinolinic acid, lutidinic acid and isocinchomeronic acid by resting cells of pyridine dicarboxylic acid-degrading microorganisms. Appl Microbiol Biotechnol 62:337–341
Wieser M, Furukawa H, Morita H, Yoshida T, Nagasawa T (1999) Synthesis of (S)-1-(4-hydroxyphenyl)alcohols by eugenol dehydrogenase from Pseudomonas fluorescens E118. Tetrahedron Asymmetry 10:1627–1630
Yoshida T, Hayakawa Y, Matsui T, Nagasawa T (2004) Purification and characterization of 2,6-dihydroxybenzoate decarboxylase reversibly catalyzing nonoxidative decarboxylation. Arch Microbiol 181:391–397
Zhao LQ, Sun ZH, Zheng P, Zhu LL (2005) Biotransformation of isoeugenol to vanillin by a novel strain of Bacillus fusiformis. Biotechnol Lett 27:1505–1509
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Yamada, M., Okada, Y., Yoshida, T. et al. Biotransformation of isoeugenol to vanillin by Pseudomonas putida IE27 cells. Appl Microbiol Biotechnol 73, 1025–1030 (2007). https://doi.org/10.1007/s00253-006-0569-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-006-0569-1