Abstract
Pseudomonas aeruginosa produces glycolipidic surface-active molecules (rhamnolipids) which have potential biotechnological applications. Rhamnolipids are produced by P. aeruginosa in a concerted manner with different virulence-associated traits. Here, we review the rhamnolipids biosynthetic pathway, showing that it has metabolic links with numerous bacterial products such as alginate, lipopolysaccharide, polyhydroxyalkanoates, and 4-hydroxy-2-alkylquinolines (HAQs). We also discuss the factors controlling the production of rhamnolipids and the proposed roles this biosurfactant plays in P. aeruginosa lifestyle.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Pseudomonas aeruginosa is an environmental bacterium that can be isolated from many different habitats, including water, soil, and plants, but it is also an opportunistic human pathogen causing serious nosocomial infections (Costerton 1980; Lyczak et al. 2000). This bacterium was shown by Jarvis and Johnson (1949) to produce the biosurfactant rhamnolipids, which are amphiphilic molecules composed of a hydrophobic fatty acid moiety and a hydrophilic portion composed of one or two rhamnose. Rhamnolipid anabolic precursors without the sugar moiety, 3-(3-hydroxyalkanoyloxy)alkanoic acids (HAAs), are also released by the bacteria and display tensio-active properties (Déziel et al. 2003). While the production of rhamnolipids is characteristic of P. aeruginosa, some isolates of the nonpathogenic pseudomonads P. putida and P. chlororaphis as well as the pathogen Burkholderia pseudomallei were also recently shown to produce a variety of rhamnolipids (Häussler et al. 1998, 2003; Tuleva et al. 2002; Gunther et al. 2005). Rhamnolipids have several potential industrial and environmental applications due to their tensio-active properties (Lang and Wullbrandt 1999; Maier and Soberón-Chávez 2000). These uses include the production of fine chemicals, the characterization of surfaces and surface coatings, and usage as additives for environmental remediation, and they have even been reported to be useful as a biological control agent (Stanghellini and Miller 1997).
Here, we review the production of rhamnolipids, showing that their biosynthesis is dependent on central metabolic pathways, such as fatty acid and deoxythymidine diphosphate dTDP-activated sugars synthesis (also reviewed in the study of Soberón-Chávez 2004). We also describe that the production of this biosurfactant is very tightly regulated at the transcriptional level by the quorum-sensing (QS) response and by environmental conditions and that the production of polyhydroxyalkanoates (PHAs), other biotechnologically important compounds (Madison and Huisman 1999) made by P. aeruginosa, have some biosynthetic steps in common (Soberón-Chávez et al. 2005b). The rhamnolipids biosynthetic pathway has also steps in common with lipopolysaccharides (LPS; Rahim et al. 2000), alginate (Olvera et al. 1999), and 4-hydroxy-2-alkylquinolines (HAQs; Bredenbruch et al. 2005). The role that rhamnolipids play is not yet understood. They have been regarded as virulence factors (Kownatzki et al. 1987) and antimicrobials (Abalos et al. 2001), implicated in the development of biofilms (Davey et al. 2003) and, along with HAAs, shown to be indispensable for P. aeruginosa swarming motility (Köhler et al. 2000; Déziel et al. 2003).
Characterization of rhamnolipids
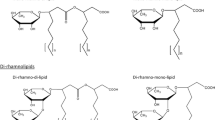
Rhamnolipids are typically constituted of a dimer of 3-hydroxyfatty acids linked through a beta glycosidic bond to a mono- or di-rhamnose moiety (Fig. 1). However, with resting cells or using naphthalene as carbon source, rhamnolipids containing only one fatty acid chain are also detected (Syldatk et al. 1985a,b; Déziel et al. 1999). It is unknown whether monomers of 3-hydroxyfatty acids can act as substrates for RhlB or if these smaller rhamnolipids result from the degradation of rhamnolipids containing two fatty acids.
Liquid chromatography coupled to mass spectrometry (LC/MS) allows the detection of more than 28 different rhamnolipid congeners in liquid cultures (Déziel et al. 1999). The alkyl chains of these congeners vary from C8 to C12, and some of these chains also contain one unsaturation. In liquid culture and under usual growth conditions, the two most abundant rhamnolipids observed are rhamnosyl-β-hydroxydecanoyl-β-hydroxydecanoate (Rha-C10-C10), a mono-rhamnolipid, and rhamnosyl-rhamnosyl-β-hydroxydecanoyl-β-hydroxydecanoate (Rha-Rha-C10-C10), a di-rhamnolipid. MS also revealed that for an isomeric rhamnolipid pair in which each compound contains two alkyl chains of different length (for instance Rha-C10-C8 and Rha-C8-C10), the congener with the shortest chain adjacent to the sugar is always more abundant than the other one by at least a factor of three. If the longest chain contains an unsaturation, the rhamnolipid with the shorter chain adjacent to the sugar is more than 20 times more abundant than the other congener. Free HAAs are also detected in the culture medium by LC/MS (Lépine et al. 2002; Déziel et al. 2003). These compounds are not degradation products of rhamnolipids, and the most abundant free HAA congener is C10-C10 (Lépine et al. 2002). As with rhamnolipids, for an isomeric HAA pair in which each compound contains two alkyl chains of different length, the congener with the shortest chain at the hydroxyl end is always more abundant than the other one. In fact, the relative proportion of each of these two free HAA congeners exactly matches the proportion of the two corresponding rhamnolipid congener containing the same two chains (Lépine et al. 2002). However, within the pool of free HAAs, those with a longer alkyl chain are proportionally less abundant than in the rhamnolipid congener pool, suggesting that the free HAAs in the culture medium are leftovers of the initial HAA pool used for rhamnolipid synthesis.
Role of RhlA, RhlB, and RhlC in rhamnolipid production
P. aeruginosa produces rhamnolipids by three sequential reactions (Fig. 2). RhlA is involved in the synthesis of the fatty acid dimer moiety of rhamnolipids and free HAAs (Déziel et al. 2003; Cabrera et al., unpublished data), as discussed further below, and seems to be loosely bound to the inner membrane (Rahim et al. 2001). The next reaction is catalyzed by the membrane-bound RhlB rhamnosyltransferase and uses dTDP-l-rhamnose and an HAA as precursors, yielding mono-rhamnolipids (Ochsner et al. 1994; Rahim et al. 2001). These compounds are in turn the substrates, together with dTDP-l-rhamnose, of RhlC to produce di-rhamnolipids (Rahim et al. 2001). RhlC also seems to be loosely bound to the inner membrane (Rahim et al. 2001) and has sequence homology with rhamnosyltransferases involved in LPS synthesis but is specific for di-rhamnolipid synthesis (Rahim et al. 2001).
Regulation of rhamnolipid production at the transcriptional level
The rhlA and rhlB genes are arranged as an operon and are clustered with rhlR and rhlI, which encode proteins involved in their transcriptional regulation through the QS response (Lazdunski et al. 2004; Soberón-Chávez et al. 2005a), as described below. The rhlC gene is not linked in the chromosome to other rhl genes and forms an operon with a gene whose function is not known. This operon is regulated at the transcriptional level in a similar manner as rhlAB (Rahim et al. 2001).
QS response regulates at the transcriptional level the production of several virulence-associated traits, including rhamnolipids (Van Delden and Iglewski 1998), as well as hundreds of additional genes (Hentzer et al. 2003; Schuster et al. 2003; Wagner et al. 2003). The QS response depends on the production of two autoinducers, butanoyl-homoserine lactone (C4-HSL) and 3-oxo-dodecanoyl-homoserine lactone (3-oxo-C12-HSL), that bind to RhlR and LasR, respectively, to activate gene expression. C4-HSL and 3-oxo-C12-HSL are synthesized by RhlI and LasI, respectively (reviewed by Lazdunski et al. 2004; Soberón-Chávez et al. 2005a). The transcriptional activator LasR, once bound to 3-oxo-C12-HSL, promotes the expression of several genes (Whiteley et al. 1999), including the one coding for the transcription regulator RhlR (Latifi et al. 1996; Pesci et al. 1997; Medina et al. 2003a). The second QS genetic circuit responds to RhlR that, once bound to C4-HSL (Ochsner and Reiser 1995), promotes the expression, among others, of rhlAB (Ochsner et al. 1994) and rhlC (Rahim et al. 2001).
The transcriptional regulation of the rhlAB promoter not only depends on RhlR and C4-HSL. This operon is not expressed in the exponential phase of growth even in the presence of this protein and its autoinducer when P. aeruginosa is cultured in rich medium (Medina et al. 2003b), presumably due to its partial transcriptional dependence on the RpoS sigma factor (σS) (Medina et al. 2003b). Furthermore, RhlR activates rhlAB transcription when coupled with C4-HSL but represses its transcription when not coupled with its autoinducer (Medina et al. 2003c). In addition, the transcriptional regulator MvfR, which directs the synthesis of HAQs, influences the expression of multiple RhlR-dependent genes, including rhlAB (Déziel et al. 2005).
Regulation of production of dTDP-l-rhamnose and its role in rhamnolipids production
In P. aeruginosa, AlgC plays a central role in the biosynthetic pathway of dTDP-d-glucose, guanosine diphosphate (GDP)-d-rhamnose, GDP-mannose, and dTDP-l-rhamnose; it transforms mannose-6-phosphate to mannose-1-phosphate, a precursor of GDP-mannose and, thus, of LPS (Lam 2004) and the exopolysaccharide alginate, but it also catalyzes the conversion of glucose-6-phosphate to glucose-1-phosphate, a precursor of dTDP-d-glucose and dTDP-l-rhamnose (Coyne et al. 1994). We described that AlgC through its phospho-gluco-mutase activity is directly involved in rhamnolipids biosynthesis (Fig. 2; Olvera et al. 1999).
The dTDP-l-rhamnose biosynthetic pathway has been reported in different bacteria to consist of the conversion of glucose-1-phosphate via dTDP-glucose, dTDP-6-deoxy-d-xylo-4-hexulose, and dTDP-6-deoxy-l-lyxo-4-hexulose (Fig. 2). In P. aeruginosa, the enzymes catalyzing these conversions are encoded by rmlA, rmlB, rmlC, and rmlD, respectively, and form the rmlBCAD operon (Rahim et al. 2000). Mutations in the rml operon of P. aeruginosa serotypes containing l-rhamnose in their LPS, like PAO1, produce truncated LPS molecules (Rahim et al. 2000) and in all cases inhibit rhamnolipid production (Olvera and Soberón-Chávez, unpublished data). TDP-l-rhamnose is the limiting RhlB substrate for rhamnolipid production in recombinant Escherichia coli expressing the rhlAB operon (Cabrera et al., unpublished data). The limited availability of this activated sugar might be the cause of the reduced rhamnolipid production by other recombinant bacteria expressing the rhlAB operon (Ochsner et al. 1995b).
Synthesis of the rhamnolipid fatty acid moiety and of HAAs
Synthesis of the fatty acid moiety of rhamnolipids diverges from the P. aeruginosa general fatty acid biosynthetic pathway at the level of the ketoacyl reduction (Campos-García et al. 1998). The enzyme responsible for draining the fatty acid precursors of rhamnolipids away from the general biosynthetic pathway is called RhlG and shows significant sequence homology with numerous nicotinamide adenine dinucleotide phosphate dependent ketoacyl reductases. RhlG is specifically involved in rhamnolipids production and also affects PHA synthesis (Fig. 2). General fatty acid content and autoinducer production in a rhlG mutant remain unaffected (Campos-García et al. 1998). However, it was recently reported that RhlG is involved in providing acyl carrier protein (ACP) fatty acid precursors for the synthesis of HAQs (Bredenbruch et al. 2005; Fig. 2), which include the QS-related Pseudomonas quinolone signal (PQS) (Déziel et al. 2004). The chain length of the fatty acid portion of rhamnolipids seems less affected by the culture medium or carbon source than PHAs are (Lépine, unpublished data).
Burger et al. (1966) observed that adding 3-hydroxydecanoic acid or C10-C10 HAA to a partially purified P. aeruginosa extract leads to the production of rhamnolipids. They thus hypothesized that 3-hydroxyfatty acids and HAAs are the precursors of these biosurfactants. The stereochemistry at the chiral center of the 3-hydroxyfatty acids included in rhamnolipids is the same as the one found in PHAs. The most abundant 3-hydroxyfatty acid found in PHAs is C10, which, as already mentioned, is also the most abundant in rhamnolipids and HAAs. These elements point to a common origin between PHA and rhamnolipids, and recently, we reported direct experimental evidence in support of this metabolic relation (Soberón-Chávez et al. 2005b).
Although, as already mentioned, some rhamnolipids contain only one 3-hydroxyfatty acid linked to one or two rhamnose moiety, the most abundant rhamnolipids produced by P. aeruginosa contain a 3-hydroxyfatty acid dimer. This brings to question how these fatty acid dimer precursors are made. Rehm et al. (2001) suggested they might arise from partial degradation of PHAs. A variety of PHA depolymerases are known to produce mono- and di- and even trilipids (Schirmer et al. 1993; Jendrossek et al. 1996). However, this hypothesis is difficult to reconcile with the observation that for two isomeric HAAs containing two alkyl chains of different length, the most abundant of these congeners is always the one with the short alkyl chain at the hydroxy terminal end (Lépine et al. 2002). Since PHAs are random copolymers (Barbuzzi et al. 2004), it is unlikely that such relative abundances could be maintained for HAA synthesis. In fact, it was recently shown that PHA synthesis is not required for the production of rhamnolipids (Pham et al. 2004).
The observation that an rhlB mutant produces free HAAs, while a rhlA mutant does not (Déziel et al. 2003), indicates that RhlA is probably responsible for the synthesis of these compounds. The reason why rhamnolipids contain preferentially a shorter alkyl chain adjacent to the sugar remains to be elucidated. This could be due to a preferential synthesis by RhlA of HAAs with the shorter alkyl chain at the hydroxyl end of the molecule, or it could be due to preferential coupling of such HAAs, from the HAA pool, to the sugar by the RhlB rhamnosyltransferase. The fact that the relative abundances of free HAAs with the short alkyl chain at the hydroxyl terminal end of the molecule match almost exactly the relative abundances of mono- or di-rhamnolipid with the short alkyl chain adjacent to the sugar indicates that it is RhlA and not RhlB that is responsible for this preferred regioselectivity. In addition, the fact that rhamnolipids with only one hydroxyfatty acid have been observed rather indicates that RhlB is not highly specific for the fatty acid portion it couples to the sugar.
We recently showed that expressing rhlAB in E. coli leads to the production of the same rhamnolipids and HAA congeners as observed in P. aeruginosa, indicating that RhlA is the enzyme responsible for the synthesis of HAAs and the fatty acid moiety of rhamnolipids (Cabrera et al., unpublished data). That the same spectrum of HAAs is produced in a different host also indicates that it is RhlA that dictates the type of fatty acid that is incorporated into HAAs and not the fatty acid relative abundance in the cell. What remains to be determined is the RhlA substrate: it might be the free 3-hydroxy acid linked either to ACP or to coenzyme A or all these species as suggested recently (Cabrera et al., unpublished data).
Environmental and growth conditions influencing the production of rhamnolipids
Rhamnolipids are so-called “secondary metabolites”, and as such, their production coincides with the onset of the stationary phase (Venkata Ramana and Karanth 1989; Déziel et al. 1996). This is in agreement with the fact that, as discussed above, transcription from the rhlAB promoter is primarily regulated in a cell density-dependent manner by QS. However, rhamnolipid production also requires appropriate growth conditions. Mostly because of their commercial/biotechnological interest as alternatives to synthetic surfactants, these cultivation factors have been extensively investigated (Ochsner et al. 1995a).
Rhamnolipid production seems possible from most carbon sources supporting bacterial growth. Nevertheless, oil of vegetable origin, such as soybean (Lang and Wullbrandt 1999), corn (Linhardt et al. 1989), canola (Sim et al. 1997), and olive (Robert et al. 1989), provides the highest productivity. Among water-soluble substrates, mannitol is especially effective (Robert et al. 1989). In contrast to PHAs, the carbon source does not generally affect the composition of rhamnolipids produced presumably because their fatty acid is synthesized de novo (Fig. 2). A noticeable exception was observed when P. aeruginosa 57RP was grown on the aromatic hydrocarbon naphthalene: 80% of the total rhamnolipids contained only one fatty acid moiety instead of HAAs (Déziel et al. 1999).
Elevated C/N (Guerra-Santos et al. 1984; Venkata Ramana and Karanth 1989) and C/P (Mulligan et al. 1989) ratios promote rhamnolipids production, while high concentrations of divalent cations, especially iron, are inhibitory (Guerra-Santos et al. 1986; Venkata Ramana and Karanth 1989). Actually, nitrogen-limiting conditions do not favor rhamnolipids production per se, but production starts with the exhaustion of nitrogen (Robert et al. 1989; Venkata Ramana and Karanth 1989; Manresa et al. 1991). Production of rhamnolipids is inhibited by the presence of NH4 +, glutamine, asparagine, and arginine as nitrogen source and promoted by NO3 −, glutamate, and aspartate (Mulligan and Gibbs 1989; Venkata Ramana and Karanth 1989; Köhler et al. 2000; Déziel, unpublished data). It has been repeatedly demonstrated that NO3 − is the best nitrogen source for rhamnolipid production (Venkata Ramana and Karanth 1989; Manresa et al. 1991; Arino et al. 1996), and we have seen that it indeed elicits higher rhlAB expression than NH4 + (Déziel et al. 2003). On the other hand, high levels of NH4 + or glutamine reduce rhamnolipid production, and this is correlated with a lower glutamine synthase activity (Mulligan and Gibbs 1989). The RpoN sigma factor (σ54) controls this enzyme, which is upregulated under nitrogen-limiting conditions (Totten et al. 1990). This sigma factor is also required for transcription of the rhlAB genes (Ochsner et al. 1994), one reason being that rhlR transcription is partially σ54-dependent (Medina et al. 2003a). The basis for the preference for nitrate is unknown. One suggestion was that P. aeruginosa, which is capable of denitrification, is also using NO3 − as an electron acceptor even in the presence of oxygen (Manresa et al. 1991). Interestingly, Sabra et al. (2002) recently proposed that P. aeruginosa is producing rhamnolipids to reduce oxygen transfer rate as a means to protect itself from oxidative stress, and it appears that this mechanism is activated by iron deficiency (Kim et al. 2003). However, excellent rhamnolipid production is also obtained in the absence of oxygen (Chayabutra et al. 2001).
Functions of rhamnolipids
Although rhamnolipids have been extensively studied, their natural function is still highly speculative. They actually seem to play multiple roles. First, since they display potent surface tension-reducing and emulsifying activities, these molecules are considered surfactants and, as a result, have been mostly studied for their ability to solubilize and promote the uptake of hydrophobic substrates, especially hydrocarbons such as n-alkanes (Itoh and Suzuki 1972; Koch et al. 1991; Zhang and Miller 1995; Beal and Betts 2000). Another mechanism through which rhamnolipids enhance the biodegradation of poorly soluble molecules is by causing the cell surface to become more hydrophobic (Zhang and Miller 1994; Al-Tahhan et al. 2000). Nevertheless, it is unlikely that the intended function of rhamnolipids is to facilitate the assimilation of insoluble substrates, as they are also efficiently produced when grown on soluble substrates.
An alternative ecological role for these surface-active molecules relates to their toxicity against a variety of microorganisms, which might confer a competitive advantage in niche colonization, P. aeruginosa being a notoriously successful and ubiquitous bacterium. Rhamnolipids display antibacterial activity mostly against Gram-positives and also a few Gram-negatives. Furthermore, antiviral, antifungal, mycoplasmacidal, algicidal, zoosporicidal, and antiamoebal activities have been reported (Itoh et al. 1971; Lang and Wagner 1993; Stanghellini and Miller 1997; Abalos et al. 2001; Cosson et al. 2002; Wang et al. 2005).
Since rhamnolipid synthesis is regulated by QS, a mechanism controlling the production of most virulence factors in P. aeruginosa (Smith and Iglewski 2003), they are regarded as virulence-associated exoproducts. However, rhamnolipids are certainly among the less well-understood virulence factors released by this bacterium. Indeed, they have been attributed with a plethora of biological activities, most of which can be ascribed to their detergent-like properties. Early on, rhamnolipids were identified as the heat-stable hemolysin of P. aeruginosa, and this hemolytic activity was their first suspected role in pathogenesis (Sierra 1960; Al-Dujaili 1976; Johnson and Allen 1978; Johnson and Boese-Marrazzo 1980; Fujita et al. 1988). They were also proposed to act by solubilizing the phospholipids of lung surfactant, making them more accessible to cleavage by the phospholipase C secreted by P. aeruginosa (Kurioka and Liu 1967). Further studies showed rhamnolipids to exhibit several effects on mammalian cells, such as disruption of the polymorphonuclear leukocyte chemotactic responses (Shryock et al. 1984), inhibition of the normal macrophage function (McClure and Schiller 1992, 1996), stimulation of the release of cytokines from airway epithelial cells (Bedard et al. 1993), interference with normal ciliary function, inhibition of the functional cilia of tracheal epithelium and slowing down of the human ciliary beat frequency (Hastie et al. 1986; Hingley et al. 1986; Read et al. 1992; Kanthakumar et al. 1996), and mucus glycoconjugate secretagogue activity (Somerville et al. 1992; Fung et al. 1995). P. aeruginosa cell-to-cell communications mechanisms, such as QS, rely on the exchange of lipidic intercellular signals (Juhas et al. 2005). Not surprising, one of these signals, PQS, was recently shown to be solubilized by rhamnolipids, hinting at an additional function for this biosurfactant (Calfee et al. 2005). PQS plays an important role in the transcriptional regulation of genes involved in P. aeruginosa virulence (Déziel et al. 2004; Wade et al. 2005). While all these reports were obtained from in vitro experiments, rhamnolipids have also been detected in sputum samples of cystic fibrosis patients colonized with P. aeruginosa (Kownatzki et al. 1987). Nevertheless, the importance of rhamnolipids' contribution to pathogenesis has yet to be demonstrated in vivo.
More recently, swarming motility was explicitly demonstrated to require HAAs and rhamnolipids (Köhler et al. 2000; Déziel et al. 2003), and it was proposed that this multicellular behavior is related to biofilm development (Déziel et al. 2003). Indeed, a notion of rhamnolipids playing a central role in the normal formation of biofilm architecture is emerging (Davey et al. 2003; Schooling et al. 2004; Lequette and Greenberg 2005).
Finally, it is noteworthy that, since HAAs are concurrently produced and often coextracted with rhamnolipids (Lépine et al. 2002), it is likely that many reports about rhamnolipids actually included HAAs in the preparations. Therefore, the role HAAs play in P. aeruginosa activities besides a contribution to swarming motility will require further investigations.
References
Abalos A, Pinazo A, Infante MR, Casals M, Garcia F, Manresa A (2001) Physicochemical and antimicrobial properties of new rhamnolipids produced by Pseudomonas aeruginosa AT10 from soybean oil refinery wastes. Langmuir 17:1367–1371
Al-Dujaili AH (1976) Toxic activity against alveolar macrophages of products of Pseudomonas aeruginosa isolated from respiratory and non-respiratory sites. J Hyg (Lond) 77:211–220
Al-Tahhan RA, Sandrin TR, Bodour AA, Maier RM (2000) Rhamnolipid-induced removal of lipopolysaccharide from Pseudomonas aeruginosa: effect on cell surface properties and interaction with hydrophobic substrates. Appl Environ Microbiol 66:3262–3268
Arino S, Marchal R, Vandecasteele J-P (1996) Identification and production of a rhamnolipidic biosurfactant by a Pseudomonas species. Appl Microbiol Biotechnol 45:162–168
Barbuzzi T et al (2004) Microbial synthesis of poly(3-hydroxyalkanoates) by Pseudomonas aeruginosa from fatty acids: identification of higher monomer units and structural characterization. Biomacromolecules 5:2469–2478
Beal R, Betts WB (2000) Role of rhamnolipid biosurfactants in the uptake and mineralization of hexadecane in Pseudomonas aeruginosa. J Appl Microbiol 89:158–168
Bedard M, McClure CD, Schiller NL, Francoeur C, Cantin A, Denis M (1993) Release of interleukin-8, interleukin-6, and colony-stimulating factors by upper airway epithelial cells: implication for cystic fibrosis. Am J Respir Cell Mol Biol 9:455–462
Bredenbruch F, Nimtz M, Wray V, Morr M, Müller R, Häussler S (2005) Biosynthetic pathway of Pseudomonas aeruginosa 4-hydroxy-2-alkylquinolines. J Bacteriol 187:3630–3635
Burger MM, Glaser L, Burton RM (1966) Formation of rhamnolipids of Pseudomonas aeruginosa. Methods Enzymol 8:441–445
Calfee MW, Shelton JG, McCubrey JA, Pesci EC (2005) Solubility and bioactivity of the Pseudomonas quinolone signal are increased by a Pseudomonas aeruginosa-produced surfactant. Infect Immun 73:878–882
Campos-García J, Caro AD, Nájera R, Miller-Maier RM, Al-Tahhan RA, Soberón-Chávez G (1998) The Pseudomonas aeruginosa rhlG gene encodes an NADPH-dependent β-ketoacyl reductase which is specifically involved in rhamnolipid synthesis. J Bacteriol 180:4442–4451
Chayabutra C, Wu J, Ju LK (2001) Rhamnolipid production by Pseudomonas aeruginosa under denitrification: effects of limiting nutrients and carbon substrates. Biotechnol Bioeng 72:25–33
Cosson P et al (2002) Pseudomonas aeruginosa virulence analyzed in a Dictyostelium discoideum host system. J Bacteriol 184:3027–3033
Costerton JW (1980) Pseudomonas aeruginosa in nature and disease. In: Sabath CD (ed) Pseudomonas aeruginosa: the organism, diseases it causes and their treatment. Hans Huber Publishers, Bern, Switzerland pp 15–24
Coyne MJ Jr, Russell KS, Coyle CL, Goldberg JB (1994) The Pseudomonas aeruginosa algC gene encodes phosphoglucomutase, required for the synthesis of a complete lipopolysaccharide core. J Bacteriol 176:3500–3507
Davey ME, Caiazza NC, O'Toole GA (2003) Rhamnolipid surfactant production affects biofilm architecture in Pseudomonas aeruginosa PAO1. J Bacteriol 185:1027–1036
Déziel É, Paquette G, Villemur R, Lépine F, Bisaillon J-G (1996) Biosurfactant production by a soil Pseudomonas strain growing on polycyclic aromatic hydrocarbons. Appl Environ Microbiol 62:1908–1912
Déziel E, Lépine F, Dennie D, Boismenu D, Mamer OA, Villemur R (1999) Liquid chromatography/mass spectrometry analysis of mixtures of rhamnolipids produced by Pseudomonas aeruginosa strain 57RP grown on mannitol or naphthalene. Biochim Biophys Acta 1440:244–252
Déziel E, Lépine F, Milot S, Villemur R (2003) rhlA is required for the production of a novel biosurfactant promoting swarming motility in Pseudomonas aeruginosa: 3-(3-hydroxyalkanoyloxy)alkanoic acids (HAAs), the precursors of rhamnolipids. Microbiology 149:2005–2013
Déziel E et al (2004) Analysis of Pseudomonas aeruginosa 4-hydroxy-2-alkylquinolines (HAQs) reveals a role for 4-hydroxy-2-heptylquinoline in cell-to-cell communication. Proc Natl Acad Sci U S A 101:1339–1344
Déziel E et al (2005) The contribution of MvfR to Pseudomonas aeruginosa pathogenesis and quorum sensing circuitry regulation: multiple quorum sensing-regulated genes are modulated without affecting lasRI, rhlRI or the production of N-acyl-L-homoserine lactones. Mol Microbiol 55:998–1014
Fujita K, Akino T, Yoshioka H (1988) Characteristics of the heat-stable extracellular hemolysin from Pseudomonas aeruginosa. Infect Immun 56:1385–1387
Fung DC, Somerville M, Richardson PS, Sheehan JK (1995) Mucus glycoconjugate complexes released from feline trachea by bacterial toxin. Am J Respir Cell Mol Biol 12:296–306
Guerra-Santos LH, Käppeli O, Fiechter A (1984) Pseudomonas aeruginosa biosurfactant production in continuous culture with glucose as carbon source. Appl Environ Microbiol 48:301–305
Guerra-Santos LH, Käppeli O, Fiechter A (1986) Dependence of Pseudomonas aeruginosa continuous culture biosurfactant production on nutritional and environmental factors. Appl Microbiol Biotechnol 24:443–448
Gunther NW IV, Nunez A, Fett W, Solaiman DK (2005) Production of rhamnolipids by Pseudomonas chlororaphis, a nonpathogenic bacterium. Appl Environ Microbiol 71:2288–2293
Hastie AT, Hingley ST, Higgins ML, Kueppers F, Shryock T (1986) Rhamnolipid from Pseudomonas aeruginosa inactivates mammalian tracheal ciliary axonemes. Cell Motil Cytoskeleton 6:502–509
Häussler S, Nimtz M, Domke T, Wray V, Steinmetz I (1998) Purification and characterization of a cytotoxic exolipid of Burkholderia pseudomallei. Infect Immun 66:1588–1593
Häussler S, Rohde M, von Neuhoff N, Nimtz M, Steinmetz I (2003) Structural and functional cellular changes induced by Burkholderia pseudomallei rhamnolipid. Infect Immun 71:2970–2975
Hentzer M et al (2003) Attenuation of Pseudomonas aeruginosa virulence by quorum sensing inhibitors. EMBO J 22:3803–3815
Hingley ST, Hastie A, Kueppers F, Higgins ML, Weinbaum G, Shryock T (1986) Effect of ciliostatic factors from Pseudomonas aeruginosa on rabbit respiratory cilia. Infect Immun 51:254–262
Itoh S, Suzuki T (1972) Effect of rhamnolipids on growth of Pseudomonas aeruginosa mutant deficient in n-paraffin-utilizing ability. Agric Biol Chem 36:2233–2235
Itoh S, Honda H, Tomita F, Suzuki T (1971) Rhamnolipids produced by Pseudomonas aeruginosa grown on n-paraffin (mixture of C12, C13 and C14 fractions). J Antibiot 24:855–859
Jarvis FG, Johnson MJ (1949) A glycolipide produced by Pseudomonas aeruginosa. J Am Chem Soc 71:4124–4126
Jendrossek D, Schirmer A, Schlegel HG (1996) Biodegradation of polyhydroxyalkanoic acids. Appl Microbiol Biotechnol 46:451–463
Johnson MK, Allen JH (1978) The role of hemolysin in corneal infections with Pseudomonas aeruginosa. Invest Ophthalmol Vis Sci 17:480–483
Johnson MK, Boese-Marrazzo D (1980) Production and properties of heat-stable extracellular hemolysin from Pseudomonas aeruginosa. Infect Immun 29:1028–1033
Juhas M, Eberl L, Tummler B (2005) Quorum sensing: the power of cooperation in the world of Pseudomonas. Environ Microbiol 7:459–471
Kanthakumar K et al (1996) The effect of bacterial toxins on levels of intracellular adenosine nucleotides and human ciliary beat frequency. Pulm Pharmacol 9:223–230
Kim EJ, Sabra W, Zeng AP (2003) Iron deficiency leads to inhibition of oxygen transfer and enhanced formation of virulence factors in cultures of Pseudomonas aeruginosa PAO1. Microbiology 149:2627–2634
Koch AK, Käppeli O, Fiechter A, Reiser J (1991) Hydrocarbon assimilation and biosurfactant production in Pseudomonas aeruginosa mutants. J Bacteriol 173:4212–4219
Köhler T, Curty LK, Barja F, Van Delden C, Pechère J-C (2000) Swarming of Pseudomonas aeruginosa is dependent on cell-to-cell signaling and requires flagella and pili. J Bacteriol 182:5990–5996
Kownatzki R, Tummler B, Doring G (1987) Rhamnolipid of Pseudomonas aeruginosa in sputum of cystic fibrosis patients. Lancet 1:1026–1027
Kurioka S, Liu PV (1967) Effect of the hemolysin of Pseudomonas aeruginosa on phosphatides and on phospholipase c activity. J Bacteriol 93:670–674
Lam JS (2004) Lipopolysaccharides of Pseudomonas aeruginosa. In: Ramos JL (ed) The pseudomonads. Biosynthesis of macromolecules and molecular metabolism. Kluwer/Plenum, New York, pp 3–52
Lang S, Wagner F (1993) Biological activities of biosurfactants. In: Kosaric N (ed) Biosurfactants: production, properties, applications. Dekker, New York, pp 251–268
Lang S, Wullbrandt D (1999) Rhamnose lipids—biosynthesis, microbial production and application potential. Appl Microbiol Biotechnol 51:22–32
Latifi A, Foglino M, Tanaka K, Williams P, Lazdunski A (1996) A hierarchical quorum-sensing cascade in Pseudomonas aeruginosa links the transcriptional activators LasR and RhlR (VsmR) to expression of the stationary-phase sigma factor RpoS. Mol Microbiol 21:1137–1146
Lazdunski AM, Ventre I, Sturgis JN (2004) Regulatory circuits and communication in Gram-negative bacteria. Nat Rev Microbiol 2:581–592
Lépine F, Déziel E, Milot S, Villemur R (2002) Liquid chromatographic/mass spectrometric detection of the 3-(3-hydroxyalkanoyloxy)alkanoic acid precursors of rhamnolipids in Pseudomonas aeruginosa cultures. J Mass Spectrom 37:41–46
Lequette Y, Greenberg EP (2005) Timing and localization of rhamnolipid synthesis gene expression in Pseudomonas aeruginosa biofilms. J Bacteriol 187:37–44
Linhardt RJ, Bakhit R, Daniels L, Mayerl F (1989) Microbially produced rhamnolipid as a source of rhamnose. Biotechnol Bioeng 33:365–368
Lyczak JB, Cannon CL, Pier GB (2000) Establishment of Pseudomonas aeruginosa infection: lessons from a versatile opportunist. Microbes Infect 2:1051–1060
Madison LL, Huisman GW (1999) Metabolic engineering of poly(3-hydroxyalkanoates): from DNA to plastic. Microbiol Mol Biol Rev 63:21–53
Maier RM, Soberón-Chávez G (2000) Pseudomonas aeruginosa rhamnolipids: biosynthesis and potential applications. Appl Microbiol Biotechnol 54:625–633
Manresa A et al (1991) Kinetic studies on surfactant production by Pseudomonas aeruginosa 44T1. J Ind Microbiol 8:133–136
McClure CD, Schiller NL (1992) Effects of Pseudomonas aeruginosa rhamnolipids on monocyte-derived macrophages. J Leukoc Biol 51:97–102
McClure CD, Schiller NL (1996) Inhibition of macrophage phagocytosis by Pseudomonas aeruginosa rhamnolipids in vitro and in vivo. Curr Microbiol 33:109–117
Medina G, Juarez K, Diaz R, Soberón-Chávez G (2003a) Transcriptional regulation of Pseudomonas aeruginosa rhlR, encoding a quorum-sensing regulatory protein. Microbiology 149:3073–3081
Medina G, Juarez K, Soberón-Chávez G (2003b) The Pseudomonas aeruginosa rhlAB operon is not expressed during the logarithmic phase of growth even in the presence of its activator RhlR and the autoinducer N-butyryl-homoserine lactone. J Bacteriol 185:377–380
Medina G, Juarez K, Valderrama B, Soberón-Chávez G (2003c) Mechanism of Pseudomonas aeruginosa RhlR transcriptional regulation of the rhlAB promoter. J Bacteriol 185:5976–5983
Mulligan CN, Gibbs BF (1989) Correlation of nitrogen metabolism with biosurfactant production by Pseudomonas aeruginosa. Appl Environ Microbiol 55:3016–3019
Mulligan CN, Mahmourides G, Gibbs BF (1989) The influence of phosphate metabolism on biosurfactant production by Pseudomonas aeruginosa. J Bacteriol 12:199–210
Ochsner UA, Reiser J (1995) Autoinducer-mediated regulation of rhamnolipid biosurfactant synthesis in Pseudomonas aeruginosa. Proc Natl Acad Sci U S A 92:6424–6428
Ochsner UA, Fiechter A, Reiser J (1994) Isolation, characterization, and expression in Escherichia coli of the Pseudomonas aeruginosa rhlAB genes encoding a rhamnosyltransferase involved in rhamnolipid biosurfactant synthesis. J Biol Chem 269:19787–19795
Ochsner UA, Hembach T, Fiechter A (1995a) Production of rhamnolipid biosurfactants. Adv Biochem Eng Biotechnol 53:89–118
Ochsner UA, Reiser J, Fiechter A, Witholt B (1995b) Production of Pseudomonas aeruginosa rhamnolipid biosurfactants in heterologous hosts. Appl Environ Microbiol 61:3503–3506
Olvera C, Goldberg JB, Sánchez R, Soberón-Chávez G (1999) The Pseudomonas aeruginosa algC gene product participates in rhamnolipid biosynthesis. FEMS Microbiol Lett 179:85–90
Pesci EC, Pearson JP, Seed PC, Iglewski BH (1997) Regulation of las and rhl quorum sensing in Pseudomonas aeruginosa. J Bacteriol 179:3127–3132
Pham TH, Webb JS, Rehm BH (2004) The role of polyhydroxyalkanoate biosynthesis by Pseudomonas aeruginosa in rhamnolipid and alginate production as well as stress tolerance and biofilm formation. Microbiology 150:3405–3413
Rahim R, Burrows LL, Monteiro MA, Perry MB, Lam JS (2000) Involvement of the rml locus in core oligosaccharide and O polysaccharide assembly in Pseudomonas aeruginosa. Microbiology 146(Pt 11):2803–2814
Rahim R et al (2001) Cloning and functional characterization of the Pseudomonas aeruginosa rhlC gene that encodes rhamnosyltransferase 2, an enzyme responsible for di-rhamnolipid biosynthesis. Mol Microbiol 40:708–718
Read RC et al (1992) Effect of Pseudomonas aeruginosa rhamnolipids on mucociliary transport and ciliary beating. J Appl Physiol 72:2271–2277
Rehm BH, Mitsky TA, Steinbuchel A (2001) Role of fatty acid de novo biosynthesis in polyhydroxyalkanoic acid (PHA) and rhamnolipid synthesis by pseudomonads: establishment of the transacylase (PhaG)-mediated pathway for PHA biosynthesis in Escherichia coli. Appl Environ Microbiol 67:3102–3109
Robert M et al (1989) Effect of the carbon source on biosurfactant production by Pseudomonas aeruginosa 44T1. Biotechnol Lett 11:871–874
Sabra W, Kim EJ, Zeng AP (2002) Physiological responses of Pseudomonas aeruginosa PAO1 to oxidative stress in controlled microaerobic and aerobic cultures. Microbiology 148:3195–3202
Schirmer A, Jendrossek D, Schlegel HG (1993) Degradation of poly(3-hydroxyoctanoic acid) [P(3HO)] by bacteria: purification and properties of a P(3HO) depolymerase from Pseudomonas fluorescens GK13. Appl Environ Microbiol 59:1220–1227
Schooling SR, Charaf UK, Allison DG, Gilbert P (2004) A role for rhamnolipid in biofilm dispersion. Biofilms 1:91–99
Schuster M, Lostroh CP, Ogi T, Greenberg EP (2003) Identification, timing, and signal specificity of Pseudomonas aeruginosa quorum-controlled genes: a transcriptome analysis. J Bacteriol 185:2066–2079
Shryock TR, Silver SA, Banschbach MW, Kramer JC (1984) Effect of Pseudomonas aeruginosa rhamnolipid on human neutrophil migration. Curr Microbiol 10:323–328
Sierra G (1960) Hemolytic effect of a glycolipid produced by Pseudomonas aeruginosa. Antonie Van Leeuwenhoek 26:189–192
Sim L, Ward OP, Li Z-Y (1997) Production and characterization of a biosurfactant isolated from Pseudomonas aeruginosa UW-1. J Ind Microbiol Biotechnol 19:232–238
Smith RS, Iglewski B (2003) P. aeruginosa quorum-sensing systems and virulence. Curr Opin Microbiol 6:56–60
Soberón-Chávez G (2004) Biosynthesis of rhamnolipids. In: Ramos J-L (ed) Pseudomonas. Biosynthesis of macromolecules and molecular metabolism. Kluwer/Plenum, New York, pp 173–189
Soberón-Chávez G, Aguirre-Ramírez M, Ordóñez L (2005a) Is Pseudomonas aeruginosa only sensing quorum? Crit Rev Microbiol 131:171–182
Soberón-Chávez G, Aguirre-Ramirez M, Sanchez R (2005b) The Pseudomonas aeruginosa RhlA enzyme is involved in rhamnolipid and polyhydroxyalkanoate production. J Ind Microbiol Biotechnol, in press, published on line in June 4th
Somerville M et al (1992) Release of mucus glycoconjugates by Pseudomonas aeruginosa rhamnolipid into feline trachea in vivo and human bronchus in vitro. Am J Respir Cell Mol Biol 6:116–122
Stanghellini ME, Miller RM (1997) Biosurfactants: their identity and potential efficacy in the biological control of zoosporic plant pathogens. Plant Dis 81:4–12
Syldatk C, Lang S, Matulovic U, Wagner F (1985a) Production of four interfacial active rhamnolipids from n-alkanes or glycerol by resting cells of Pseudomonas species DSM 2874. Z Naturforsch [C] 40:61–67
Syldatk C, Lang S, Wagner F, Wray V, Witte L (1985b) Chemical and physical characterization of four interfacial-active rhamnolipids from Pseudomonas spec. DSM 2874 grown on n-alkanes. Z Naturforsch [C] 40:51–60
Totten PA, Lara JC, Lory S (1990) The rpoN gene product of Pseudomonas aeruginosa is required for expression of diverse genes, including the flagellin gene. J Bacteriol 172:389–396
Tuleva BK, Ivanov GR, Christova NE (2002) Biosurfactant production by a new Pseudomonas putida strain. Z Naturforsch [C] 57:356–360
Van Delden C, Iglewski BH (1998) Cell-to-cell signaling and Pseudomonas aeruginosa infections. Emerg Infect Dis 4:551–560
Venkata Ramana K, Karanth NG (1989) Factors affecting biosurfactant production using Pseudomonas aeruginosa CFTR-6 under submerged conditions. J Chem Technol Biotechnol 45:249–257
Wade DS, Calfee W, Rocha ER, Ling EA, Engstrom E, Coleman JP, Pesci EC (2005) J Bacteriol 187:4372–4380
Wagner VE, Bushnell D, Passador L, Brooks AI, Iglewski BH (2003) Microarray analysis of Pseudomonas aeruginosa quorum-sensing regulons: effects of growth phase and environment. J Bacteriol 185:2080–2095
Wang X, Gong L, Liang S, Han X, Zhu C, Li Y (2005) Algicidal activity of rhamnolipid biosurfactants produced by Pseudomonas aeruginosa. Harmful Algae 4:433–443
Whiteley M, Lee KM, Greenberg EP (1999) Identification of genes controlled by quorum sensing in Pseudomonas aeruginosa. Proc Natl Acad Sci U S A 96:13904–18909
Zhang Y, Miller RM (1994) Effect of a Pseudomonas rhamnolipid biosurfactant on cell hydrophobicity and biodegradation of octadecane. Appl Environ Microbiol 60:2101–2106
Zhang Y, Miller RM (1995) Effect of rhamnolipid (biosurfactant) structure on solubilization and biodegradation on n-alkanes. Appl Environ Microbiol 61:2247–2251
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Soberón-Chávez, G., Lépine, F. & Déziel, E. Production of rhamnolipids by Pseudomonas aeruginosa . Appl Microbiol Biotechnol 68, 718–725 (2005). https://doi.org/10.1007/s00253-005-0150-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-005-0150-3