Abstract
One of the major extracellular enzymes of the white-rot fungus Coriolus versicolor is laccase, which is involved in the degradation of lignin. We constructed a homologous system for the expression of a gene for laccase III (cvl3) in C. versicolor, using a chimeric laccase gene driven by the promoter of a gene for glyceraldehyde-3-phosphate dehydrogenase (gpd) from this fungus. We transformed C. versicolor successfully by introducing both a gene for hygromycin B phosphotransferase (hph) and the chimeric laccase gene. In three independent experiments, we recovered 47 hygromycin-resistant transformants at a transformation frequency of 13 transformants μg−1 of plasmid DNA. We confirmed the introduction of the chimeric laccase gene into the mycelia of transformants by a polymerase chain reaction in nine randomly selected transformants. Overproduction of extracellular laccase by the transformants was revealed by a colorimetric assay for laccase activity. We examined the transformant (T2) that had the highest laccase activity and found that its activity was significantly higher than that of the wild type, particularly in the presence of copper (II). Our transformation system should contribute to the efficient production of the extracellular proteins of C. versicolor for the accelerated degradation of lignin and aromatic pollutants.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Coriolus (Trametes) versicolor, a white-rot basidiomycete, has been studied extensively because of its ability to degrade lignin (Higuchi 1993; Kirk and Farrell 1987). The fungus secretes enzymes that catalyze cleavage of the side-chains and aromatic rings of lignin; and it can also degrade various persistent organic pollutants, such as chlorophenols (Itoh et al. 2000; Ullah et al. 2000). Laccase, a multi-copper oxidase, is one of the major extracellular components of the lignin-degradation system of C. versicolor (Kawai et al. 1988, 1999). Laccases are also exploited industrially in various oxidative processes (Jönsson et al. 1998). The genome of C. versicolor encodes several laccases and laccase III (CVL3) is the isoenzyme that is secreted at the highest level (Iimura et al. 1995). This isozyme is highly glycosylated and its four N-linked carbohydrate chains play important roles in the resistance of the enzyme to proteolysis and high temperatures (Yoshitake et al. 1993).
To study the mechanism of degradation of lignin and other phenolics by laccase III and to examine the relationship between the structure and function of this enzyme, we tried to develop a system for the efficient expression of a recombinant enzyme. A system for the production of such a recombinant protein would allow us to modify the properties of the enzyme, such as its redox potential and thermostability. The heterologous expression of fungal genes for laccases has been achieved in yeast and filamentous fungi (Cassland and Jönsson 1999; Liu et al. 2003; Saloheimo et al. 1991; Yaver et al. 1996). High productivity and efficient post-translational modification of the recombinant proteins have been demonstrated in these heterologous systems. Such systems are generally useful for appropriate secretion of the protein of interest and for time-saving production. However, in some cases, the properties of the resulting protein, such as molecular weight, rate of glycosylation rate and optimal temperature, differ from those of the native protein (Conesa et al. 2002; Liu et al. 2003). As a consequence of the requirements for complex post-transcriptional modification of extracellular proteins, the best systems in basidiomycetes involve homologous expression. In such systems, the recombinant protein is likely to be very similar to the wild-type protein (Ma et al. 2003).
Homologous expression systems for extracellular proteins of basidiomycetes have been reported for several enzymes (Irie et al. 2001; Li et al. 2000; Ma et al. 2003; Mayfield et al. 1994; Sollewijn Gelpke et al. 1999). The recombinant manganese peroxidase protein produced by transformed Phanerochaete chrysosporium had physical, kinetic, and spectral characteristics that were identical to those of the native protein from the wild-type fungus (Mayfield et al. 1994). By contrast to the major advantages in the development of homologous expression systems in other fungi, progress in the development of systems for the transformation of C. versicolor has been limited. Bartholomew et al. (2001) and Kim et al. (2002) succeeded in transforming this basidiomycete by introducing genes for resistance to phleomycin and hygromycin B (hph), respectively. In a recent study, we transformed C. versicolor using the hph gene under the control of a homologous promoter sequence from the gene for glyceraldehyde-3-phosphate dehydrogenase (gpd) of this fungus (Nitta et al. 2004). However, to our best knowledge, no system for the homologous expression of a secreted protein in C. versicolor has been reported to date.
In the present study, we report the overproduction of laccase III by transformed C. versicolor that harbored both a hph gene and a chimeric gene for laccase III (cvl3), driven by the ras promoter of Lentinus edodes and the gpd promoter of C. versicolor, respectively. The characteristics of the transformants and their possible utility are discussed.
Materials and methods
Strain and culture conditions
A dikaryotic strain of C. versicolor (IFO 1030) was used as the recipient in transformation experiments. Mycelia of C. versicolor were grown on potato/dextrose agar medium (Kyokuto, Tokyo) or in glucose/peptone (GP) liquid medium (30 g glucose, 10 g peptone, 1.5 g potassium dihydrogenphosphate, 500 mg magnesium sulfate, 2 mg thiamine hydrochloride) with and without copper (II) sulfate (0.2 mM) at 28°C.
Plasmid vector
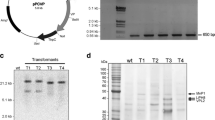
Two types of vector were used for the co-transformation of C. versicolor. We used LC1-hph (7.5 kbp), which was found to be useful in L. edodes (Sato et al. 1998). It includes both the promoter sequence of the ras gene from L. edodes and the terminator sequence of the priA gene from L. edodes for the transcription of hph (Schuren and Wessels 1994; Irie et al. 2001). We also used the plasmid shown in Fig. 1, pT7GPTLac (5.5 kbp), which contains full-length cDNA for laccase III of C. versicolor (1,563 bp; accession no. D13372) under the control of the gpd promoter of C. versicolor (844 bp; accession no. AB075243; Nitta et al. 2004), and a nopaline synthase (nos) terminator from pBI121 (BD Bioscience Clonetech, Tokyo, Japan).
Transformation procedure
We prepared protoplasts of C. versicolor as described by Sato et al. (1998), with slight modifications. After mycelia were cultured at 28°C for 2 days in SM medium (1% sucrose, 1% malt extract), they were collected, washed twice with M buffer (2 g l−1 sodium malate, pH 5.5, 0.6 M mannitol), and treated with a solution of enzymes [10 mg ml−1 cellulase Onozuka R-10 (Yakult Honsha, Tokyo, Japan), 1 mg ml−1 chitinase (Sigma Chemical, St Louis, Mo., USA), in M buffer] at 30°C for 2–3 h, with shaking at 50–60 rpm. After incubation, the protoplasts in the suspension were collected by filtration on Miracloth. They were suspended in STC buffer (0.01 M Tris-HCl, pH 7.5, 0.01 M calcium chloride, 1.2 M sorbitol) and recovered by centrifugation. Transformation was performed using polyethylene glycol (PEG4000; Wako Pure Chemical, Osaka, Japan) as described by Yanai et al. (1996)), with slight modifications. The protoplasts were suspended in STC buffer at 100 protoplasts ml−1 and then 100 μl of the suspension were mixed with both 4 μg of pLC1-hph and 16 μg of pT7GPTLac. The mixture was kept on ice for 5 min and was then supplemented with 25 μl of PEG solution (40% PEG4000, 10 mM Tris-HCl, pH 7.5, 50 mM calcium chloride). The mixture was kept on ice for 20 min. Then it was mixed gently with an additional 1 ml of PEG solution and allowed to stand at room temperature for 5 min. After the further addition of 2 ml of STC buffer, the mixture was centrifuged at 1,000 g for 5 min. The pelleted protoplasts were suspended in 4 ml of SM liquid medium and incubated at 28°C for 48 h. The mixture was then spread on an SM agar plate that had been prepared with 100 mg l−1 hygromycin B (Wako Pure Chemical).
Analysis of DNA
Total DNA was extracted by the cetyltrimethylammonium bromide method (Möller et al. 1992) from wild-type and hygromycin-resistant mycelia, which were cultured in GP-Cu liquid medium (containing 0.2 mM copper (II) sulfate, 100 μM 4-chloro-1-naphthol; Wako Pure Chemical) at 28°C for 7 days. For the polymerase chain reaction (PCR), we used primers that allowed specific amplification of a portion of the chimeric laccase III cDNA fragment (98 bp), namely cvl3-2F (5′-ggatcgcttccagctcaatg-3′) and cvl3-2R (5′-tcgtgcccttctggaagaaa-3′). This pair of primers also allowed amplification of a fragment of the endogenous cvl3 gene (146 bp) in C. versicolor, which includes 48 bp of the sequence of the first intron. As a control experiment, we also performed PCR with pT7GPTLac as template and the same primers.
Colorimetric assay of laccase activity
To assess the production of laccase by each transformant, we performed a colorimetric assay using GP-Cu liquid medium. Mycelia of each transformant that had been cultured on GP-agar medium with hygromycin B for 10 days were recovered as a disk (ca. 8 mm diam.) that was punched out with a cork borer. Each disk was transferred to a well in a 12-well micro-plate that contained 1 ml of GP-Cu liquid medium. The laccase activity of each transformant was easily detectable as a change in the color of the medium and the mycelial disk.
Assay of laccase activity during culture with and without copper (II)
We measured the activity of laccase secreted by the wild-type strain and transformant T2, using guaiacol as the substrate. Mycelial disks were recovered from cultures on GP-agar medium and incubated in 10 ml of GP medium with or without copper (II) sulfate (0.2 mM). No hygromycin was added to either medium. To measure laccase activity in the medium, we took 100-μl aliquots of the supernatant from each culture at 24-h intervals and combined them with sodium tartrate buffer (pH 4.0) that contained 5 mM guaiacol. The optical density of each mixture was monitored with a spectrophotometer at 436 nm at 28°C. One unit of laccase activity was defined as the amount of enzyme that oxidized 1 μmol min−1 guaiacol under the conditions of the reaction. Amounts of oxidized guaiacol were calculated from the molar extinction coefficient of 6,400 M−1 cm−1 reported by Eggert et al. (1996).
Analysis by reverse transcription-PCR
Using reverse transcription and PCR (RT-PCR), we examined transcripts from the chimeric and endogenous cvl3 genes, using total RNA isolated from wild-type and transformant T2 mycelia incubated with and without copper (II) sulfate (0.2 mM). The design of a pair of primers was based on the nucleotide sequence of the cvl3 gene. We also analyzed the expression of rRNA genes in each sample by RT-PCR, using commercial primers (Classic II 18S internal standards; Ambion, Austin, Tex., USA). In the analysis, first-strand cDNA was synthesized from total RNA using a random primer (for analysis of rRNA gene expression) or oligo-dT primer (for chimeric and endogenous cvl3 gene expression) and the cDNA was used as templates for PCR, using the BcaBEST RNA PCR kit ver. 1.1 (Takara Bio, Shiga, Japan). The primers used for detection of the transcripts derived from both endogenous and chimeric cvl3 genes were cvl3-2F (5′-ggatcgcttccagctcaatg-3′) and cvl3-3R (5′-tcgaagtggtctgcgtggta-3′). In addition, we also used a primer pair for specific amplification of the transcript from the chimeric gene: gpd-f1 (5′-ctcatcccatccaccacatccac-3′) and cvl3-2R (5′-tcgtgcccttctggaagaaa-3′). To distinguish amplified DNA fragments derived from first-strand cDNA from those derived from contaminating genomic DNA in total RNA, we designed pairs of primers for the amplification of intron-containing fragments. In addition, to check that the amplification by PCR was linear, each of the first-strand cDNAs derived from the total RNA (1 μg for cvl3, 0.15 μg for rRNA) was amplified by PCR, using at least four different numbers of amplification cycles (15, 20, 25, 30 cycles). The amplification program for both transcripts was: 94°C for 2 min, followed by 15, 20, 25 and 30 cycles at 94°C for 30 s, 55°C for 30 s, and 72°C for 90 s. After amplification, 10 μl of reaction mixture that reacted at 25 cycles was analyzed by agarose gel electrophoresis (Fig. 5).
Results
We co-transformed protoplasts of C. versicolor with pLC1-hph and pT7GPTLac and recovered 47 independent hygromycin-resistant mycelial colonies on a selection medium that contained 100 mg l−1 hygromycin B. The transformation efficiency, calculated from the amount of pLC1-hph used in each experiment was approximately 13 transformants μg−1 of plasmid DNA. All of the transformed mycelia tested were able to grow rapidly on GP medium that contained more than 150 mg l−1 hygromycin B, on which wild-type mycelia were unable to grow rapidly (Fig. 2a). To confirm the introduction of the chimeric cvl3 gene, we prepared total DNA from nine randomly selected transformants and amplified the gene by PCR, using the total DNA as template (Fig. 3). To our surprise, we detected a specific DNA fragment (98 bp) derived from the chimeric gene in all of the nine transformants that we tested. Thus, all nine transformants tested harbored the introduced gene for laccase. As expected, a fragment derived from the native cvl3 gene (146 bp) was also amplified in the analysis of all DNA samples from wild-type mycelia, from mycelia transformed with only pLC1-hph, and from mycelia transformed with both pLC1-hph and pT7GPTLac (Fig. 3). These results indicate that our co-transformation procedure with two plasmid DNAs was effective for the introduction of foreign genes into C. versicolor.
a,b Growth and laccase activity. a Growth of the wild type (WT), two transformants generated with pLC1-hph (hph-1, hph-2) and nine transformants generated with both pLC1-hph and pT7GPTLac (T1–T49) on GP medium plus copper (II) sulfate (0.2 mM) and hygromycin B (150 mg l−1). b Colorimetric assay of the laccase activity of the wild type and various transformants. Mycelia were cultured in liquid medium that contained 100 μM 4-chloro-1-naphthol as substrate. Changes in the color of the medium resulted from oxidation of the substrate by laccase produced by the mycelial disks
PCR analysis of total DNA prepared from the wild type and transformants. PCR was performed with plasmid pT7GPTLac (PL) and total DNA as template and primers specific for the amplification of both the endogenous and introduced chimeric cvl3 genes. The arrowhead indicates amplified DNA derived from the cvl3 cDNA (98 bp) that was introduced into the transformants. The arrow indicates amplified DNA (146 bp) derived from the endogenous cvl3 gene in the wild type and transformants. Total DNA from mycelia transformed with pLC1-hph (hph-1) alone was included in the analysis
We performed a colorimetric assay to assess the laccase activities of the transformants, using mycelial disks prepared from each transformant. In addition to the wild type and mycelia transformed with pLC1-hph alone, we tested nine transformants that harbored both pLC1-hph and pT7GPTLac in GP liquid medium that contained 4-chloro-1-naphthol and copper (II) sulfate. As indicated in Fig. 2b, some of the substrate-containing medium in wells that contained transformed mycelia turned dark blue during an overnight incubation at 28°C. Indeed, a change in the color of the medium in the well that contained transformant T2 was observed within a few hours after transfer of the mycelia to the medium. The results of this analysis suggested that transformants T2, T3, T10, T14, and T20 secreted laccase extracellularly at greater levels than the wild type.
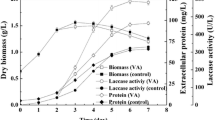
To analyze the transformants in greater detail, we examined the effects of copper on the laccase activities of the wild type and the T2 transformant, since laccase is a multi-copper oxidase. Figure 4 shows the time-courses of the activities in culture medium with and without 0.2 mM copper (II) sulfate. The activity in copper-containing medium increased significantly in the case of both the wild type and the transformant, indicating that copper stimulates the extracellular laccase activity of C. versicolor. The laccase activity of the transformant in medium to which copper (II) sulfate was added was significantly higher than that of the copper-added wild type during the analysis (until 144 h). The activity of the transformant in the absence of copper (II) sulfate was also stronger than that of the wild type with and without copper (II) sulfate, but the activity of the wild type in the presence of copper (II) exceeded that of the transformant without copper (II) at 120 h. The mycelial dry weights of the transformant with copper (II) sulfate measured at 48 h (15.7 mg) and 144 h (46.0 mg) showed levels similar to those of the wild type (18.2 mg and 50.1 mg, respectively), indicating that the higher activity of the transformant does not derive from a higher mycelial growth rate. Even though the time-course of laccase activity of the transformant remains to be confirmed, a peak in the activity of the wild-type mycelium of C. versicolor was observed after 10 days of incubation (Collins and Dobson 1997). Thus, the activity of the transformant should elevate continuously after 144 h. Our present data indicate that the introduced chimeric cvl3 gene was appropriately expressed in C. versicolor and that the inclusion of copper (II) in the medium was essential for the efficient overproduction of active laccase by the transformants.
Extracellular laccase activity of the wild type and transformant T2 during culture in GP medium with and without copper sulfate (0.2 mM). Activity was measured with guaiacol as the substrate. The oxidation of guaiacol was monitored in terms of the increase in absorbance at 436 nm. Activity was defined in terms of the amount of substrate oxidized per milliliter of medium per minute at 28°C. Squares T2 transformant with CuSO4, diamonds T2 transformant without CuSO4, circles wild type with CuSO4, triangles wild type without CuSO4
To examine the effect of the introduced chimeric cvl3 gene on the level of cvl3 transcripts, we used RT-PCR with total RNA prepared from 6-day-old wild-type and transformant T2 mycelia cultured in GP medium with and without copper (II) sulfate. We could detect transcripts from the chimeric cvl3 gene only in the transformant and not in wild-type mycelia (Fig. 5b). Judging from the result shown in Fig. 5b, the expression of the chimeric gene in the transformant should not be affected by the concentration of copper (II) sulfate. By contrast, copper (II) ions availability should influence the expression of the endogenous cvl3 gene in the transformant, because a larger amount of the transcripts was detected in the presence of copper (II) sulfate (Fig. 5a). These results indicate that the enhanced laccase activity in the transformant could be derived, in part at least, from expression of the chimeric cvl3 gene.
a–c RT-PCR analysis of cvl3 transcripts in wild-type and transformant T2 mycelia cultured with and without copper sulfate. The analysis was carried out using total RNA isolated from individual 6-day-old mycelia. a Transcript levels monitored with a non-specific primer pair for both the endogenous (endo) and introduced chimeric (chi) genes. b Transcript levels monitored with a specific primer pair for the chimeric genes. c Transcript level of rRNA genes in each mycelium
Discussion
We succeeded in co-transforming C. versicolor using an hph gene and a chimeric gene for laccase III. The resultant transformants had apparently elevated extracellular laccase activity, indicating that the product of the chimeric gene functioned appropriately. We confirmed the introduction of the laccase gene by PCR, using isolated total DNA from individual transformants as template. We did not determine whether the introduced plasmid DNAs were integrated into the chromosome or whether they replicated autonomously and extrachromosomally in C. versicolor. However, we confirmed the integration of the hph gene, driven by a gpd promoter, into the genome of C. versicolor by Southern blotting analysis in a previous study (Nitta et al. 2004). Thus, it is possible to think that the chimeric cvl3 gene was also integrated into the genome of each transformant in the present study.
The efficiency of transformation in the present study (13 transformants μg−1 DNA) was low, compared with results achieved in other basidiomycetes. A high frequency (more than 100 transformants μg−1 DNA) was obtained in P. chrysosporium with a bialaphos-resistance gene or the ade or ura gene as the selectable marker (Akileswaran et al. 1993; Alic et al. 1989; Ma et al. 2003). In C. versicolor, Kim et al. (2002) reported that they obtained 25–50 transformants μg−1 plasmid DNA with a hygromycin-resistance gene. Although the reason for the low efficiency of our transformation is unclear, data shown in Figs. 2, 3, 5 suggest that the introduced gene was appropriately expressed in the transformants.
Copper ions in the culture medium are important for extracellular laccase activity because the enzyme is a multi-copper oxidase. The addition of copper (II) sulfate (0.4–400 μM) to the growth medium resulted in increased levels of both the expression of the laccase gene and the activity of its product in C. versicolor (Collins and Dobson 1997). The optimal concentration of copper ions in a heterologous system for expression laccases in Pichia pastoris was also reported to range between 200 μM and 400 μM (Liu et al. 2003; O’Callaghan et al. 2002). When we examined the effect of the availability of copper on activity in extracellular fractions of wild-type and T2 mycelia, we found (as indicated in Fig. 4) that the addition of copper (II) sulfate stimulated the extracellular laccase activity of both the wild type and the transformant. In particular, the activity of the transformant was significantly higher than that of the wild type when mycelia were cultured under the same conditions in the presence of copper sulfate. Furthermore, even though the activity of the wild type, in the presence of copper sulfate, increased dramatically after 96 h, the activity of transformant T2 under the same conditions was still very high. The results from RT-PCR analysis suggest that the elevated activity in the transformant derives from activated transcription of the introduced cvl3 gene (Fig. 5b). These data also suggest that copper availability is still a limiting factor for laccase activity, even when the chimeric cvl3 gene is strongly expressed by a constitutive promoter in the transformant.
Compared with the yield of laccase from the wild-type strain, the yield of laccase from transformants with a chimeric cvl3 gene was elevated and the enzyme was secreted effectively. Although we have not yet confirmed the properties of the recombinant protein, such as its molecular weight, pH stability, and thermostability, our homologous expression system appears to have the potential to contribute to the exploitation of both the secreted proteins and the mycelia of C. versicolor in processes such as the bio-bleaching of wood pulp and bioremediation.
References
Akileswaran L, Alic M, Clark EK, Hornick JL, Gold MH (1993) Isolation and transformation of uracil auxotrophs of the lignin-degrading basidiomycete Phanerochaete chrysosporium. Curr Genet 23:351–356
Alic M, Kornegay JR, Pribnow D, Gold MH (1989) Transformation by complementation of an adenine auxotroph of the lignin-degrading basidomycete Phanerochaete chrysosporium. Appl Environ Microbiol 55:406–411
Bartholomew K, Dos Santos G, Dumonceaux T, Charles T, Archibald F (2001) Genetic transformation of Trametes versicolor to phleomycin resistance with the dominant selectable marker shble. Appl Microbiol Biotechnol 56:201–204
Cassland P, Jönssson LJ (1999) Characterization of a gene encoding Trametes versicolor laccase A and improved hetelogous expression in Saccharomyces cerevisae by decreased cultivation temperature. Appl Microbiol Biotechnol 52:393–400
Collins PJ, Dobson AWD (1997) Regulation of laccase gene transcription in Trametes versicolor. Appl Environ Microbiol 63:3444–3450
Conesa A, Jeenes D, Archer DB, Van Den Hondel C, Punt PJ (2002) Calnexin overexpression increases manganese peroxidase production in Aspergillus niger. Appl Environ Microbiol 68:846–851
Eggert C, Temp U, Eriksson K-E (1996) The ligninolytic system of the white rot fungus Pycnoporus cinnabarinus: purification and characterization of the laccase. Appl Environ Microbiol 62:1151–1158
Higuchi T (1993) Biodegradation mechanism of lignin by white-rot basidiomycetes. J Biotechnol 30:1–8
Iimura Y, Katayama Y, Kawai S, Morohoshi N (1995) Degradation and solubilization of C-13, C-14 side-chain labeled synthetic lignin (dehydrogenative polymerizate) by laccase-III of Coriolus versicolor. Biosci Biotechnol Biochem 59:903–905
Irie T, Honda Y, Watanabe T, Kuwahara M (2001) Homologous expression of recombinant manganese peroxidase genes in ligninolytic fungus Pleurotus ostreatus. Appl Microbiol Biotechnol 55:566–570
Itoh K, Fujita M, Kumano K, Suyama K, Yamamoto H (2000) Phenolic acid affect transformations of chlorophenols by a Coriolus versicolor laccase. Soil Biol Biochem 32:85–91
Jönsson LJ, Palmqvist E, Nilvebrant NO, Hahn-Hägerdal B (1998) Detoxification of wood hydrolysate with laccase and peroxidase from the white-rot fungus Trametes versicolor. Appl Microbiol Biotechnol 49:691–697
Kawai S, Umezawa T, Higuchi T (1988) Degradation mechanisms of phenolic beta-1 lignin substructure model compounds by laccase of Coriolus versicolor. Arch Biochem Biophys 262:99−110
Kawai S, Asukai M, Ohya N, Okita K, Ito T, Ohashi H (1999) Degradation of a non-phenolic beta-O-4 substructure and of polymeric lignin model compounds by laccase of Coriolus versicolor in the presence of 1-hydroxybenzotriazole. FEMS Microbiol Lett 170:51–57
Kim K, Leem Y, Kim K, Kim K, Choi HT (2002) Transformation of the medical basidiomycete Trametes versicolor to hygromycin B resistance by restriction enzyme mediated integration. FEMS Microbiol Lett 209:273–276
Kirk TK, Farrell RL (1987) Enzymatic “combustion”: the microbial degradation of lignin. Annu Rev Microbiol 41:465−505
Li B, Rotsaert FAJ, Gold MH, Renganathan V (2000) Homologous expression of recombinant cellobiose dehydrogenase in Phanerochaete chrysosporium. Biochem Biophys Res Commun 270:141–146
Liu W, Chao Y, Liu S, Qian S (2003) Molecular cloning and characterization of a laccase gene from the basidiomycete Fome lignosus and expression in Pichia pastoris. Appl Microbiol Biotechnol 63:174–181
Ma B, Mayfield MB, Gold MH (2003) Homologous expression system of Phanerochaete chrysosporium manganese peroxidase, using bialaphos resistance as a dominant selectable marker. Curr Genet 43:407–414
Mayfield MB, Kishi T, Alic M, Gold MH (1994) Homologous expression of recombinant manganese peroxidase in Phanerochaete chrysosporium. Appl Environ Microbiol 60:4303–4309
Möller E, Bahnweg G, Sandermann H, Geiger H (1992) A simple and efficient protocol for isolation of high molecular weight DNA from filamentous fingi, fruit bodies and infected plant tissues. Nucleic Acids Res 20:6115–6116
Nitta Y, Miyazaki Y, Nakamura M, Iimura Y, Shishido K, Kajita S, Morohoshi N (2004) Molecular cloning of the promoter region of the glyceraldehyde-3-phosphate dehydrogenase gene that contributes to the construction of a new transformation system in Coriolus versicolor. Mycoscience 45:131–136
O’Callaghan J, O’Brien MM, Dobson ADW (2002) Optimization of the expression of Tremetes versicolor laccase gene in Pichia pastoris. J Ind Microbiol Biotechnol 29:55–59
Saloheimo M, Leena M, Paavola N (1991) Heterologous production of a lignolytic enzyme: expression of the Phlebia radiate laccase gene in Trichoderma reesei. Bio/Technology 9:987–990
Sato T, Yaegashi K, Ishii S, Hirano T, Kajiwara S, Shishido K, Enei H (1998) Transformation of the edible basidiomycete Lentinus edodes by restriction enzyme-mediated integration of plasmid DNA. Biosci Biotechnol Biochem 62:2346−2350
Schuren FHJ, Wessels JGH (1994) Highly-efficient transformation of the homobasidiomycete Schizophyllum commune to phleomycin resistance. Curr Genet 26:179−183
Sollewijn Gelpke MD, Mayfield M, Cereghino GPL, Gold MH (1999) Homologous expression of recombinant lignin peroxidase in Phanerochaete chrysosporium. Appl Environ Microbiol 65:1670–1674
Ullah MA, Bedford CT, Evans CS (2000) Reactions of pentachlorophenol with laccase from Coriolus versicolor. Appl Microbiol Biotechnol 53:230−234
Yanai K, Yonekura K, Usami H, Hirakawa M, Kajiwara S, Yamazaki T, Shishido K, Adachi T (1996) The integrative transformation of Pleurotus ostreatus using bialaphos resistance as a dominant selectable marker. Biosci Biotechnol Biochem 60:472−475
Yaver DS, Xu F, Golightly EJ, Brown KM, Brown SH, Rey MW, Schneider P, Halkeir T, Mondore K, Dalbøge H (1996) Purification, characterization, molecular cloning, and expression of two laccase genes from the white rot basidiomycete Trametes villosa. Appl Environ Microbiol 62:834–841
Yoshitake A, Katayama Y, Nakamura M, Iimura Y, Kawai S, Morohoshi N (1993) N-linked carbohydrate chains protect laccase-III from proteolysis in Coriolus versicolor. J Gen Microbiol 139:179–185
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kajita, S., Sugawara, S., Miyazaki, Y. et al. Overproduction of recombinant laccase using a homologous expression system in Coriolus versicolor. Appl Microbiol Biotechnol 66, 194–199 (2004). https://doi.org/10.1007/s00253-004-1663-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-004-1663-x