Abstract
The study of the draft genome of an Antarctic marine ciliate, Euplotes petzi, revealed foreign sequences of bacterial origin belonging to the γ-proteobacterium Francisella that includes pathogenic and environmental species. TEM and FISH analyses confirmed the presence of a Francisella endocytobiont in E. petzi. This endocytobiont was isolated and found to be a new species, named F. adeliensis sp. nov.. F. adeliensis grows well at wide ranges of temperature, salinity, and carbon dioxide concentrations implying that it may colonize new organisms living in deeply diversified habitats. The F. adeliensis genome includes the igl and pdp gene sets (pdpC and pdpE excepted) of the Francisella pathogenicity island needed for intracellular growth. Consistently with an F. adeliensis ancient symbiotic lifestyle, it also contains a single insertion-sequence element. Instead, it lacks genes for the biosynthesis of essential amino acids such as cysteine, lysine, methionine, and tyrosine. In a genome-based phylogenetic tree, F. adeliensis forms a new early branching clade, basal to the evolution of pathogenic species. The correlations of this clade with the other clades raise doubts about a genuine free-living nature of the environmental Francisella species isolated from natural and man-made environments, and suggest to look at F. adeliensis as a pioneer in the Francisella colonization of eukaryotic organisms.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Like their multicellular descendants, also single-celled eukaryotes host a huge variety of bacteria. Ciliates in particular are a preferential and stable home to bacteria, which may be carried either attached as epibionts to the cell body surface, as is the case of the association between a group (designated as “epixenosomes”) of Verrucomicrobia and Euplotidium itoi [1], or enclosed as endocytobionts inside the cell body. Being a principal component of the diet of ciliates, which are mostly phagotrophic and filter-feeding, bacteria can easily escape digestion and adopt a new intracellular lifestyle [2]. Roughly 250 ciliate species, among the nearly 10,000 that are in total known, have been detected to be hosts of endocytobiont bacteria. Large-size species of Paramecium, Euplotes, and Spirostomum may be home also of mixed populations of unrelated species of bacteria [3, 4].
The knowledge of the biology and life cycle of endocytobiont bacteria in ciliates is essentially limited to species of Holospora and Caedibacter, which are colonizers of the nuclear apparatus of freshwater species of Paramecium [5, 6]. These symbionts have been successfully isolated from host-cell homogenates, but any attempt of cultivation outside their hosts has failed as in the case of any other bacterial symbiont of aerobic ciliates [7].
A substantial contribution to improve this knowledge may now be provided by the isolation and cultivation of Francisella bacteria living as endocytobionts in marine species of Euplotes, a genus which is quite rich also in freshwater species extensively studied for their symbiotic associations with polymorphic populations of Polynucleobacter [8]. F. endociliophora, earlier described as a novel subspecies of F. noatunensis [9], is the first Francisella that has been isolated and genome-sequenced from a marine species of Euplotes, E. raikovi, dwelling in temperate waters [10]. Here, we report the isolation and genome sequencing of another new species of Francisella, F. adeliensis sp. nov., living as endocytobiont in a bipolar (Antarctic and Arctic) species of Euplotes, E. petzi.
The genus Francisella comprises species classified as facultative intracellular γ-proteobacteria potentially noxious to their hosts [11, 12]. F. tularensis, with its three subspecies, is a specialized intracellular pathogen of both invertebrate and vertebrate hosts, human beings included [13, 14]. F. noatunensis, with its two subspecies adapted to different hosts’ temperatures, is the etiological agent of the fish disease known as francisellosis [15, 16]. The endosymbiotic F. persica (ex Wolbachia persica) [17], together with the generalists F. philomiragia and F. novicida, may harm human beings with a compromised immune system [18,19,20].
The position that F. adeliensis takes in the genome-based phylogenetic tree provides new insights on Francisella diversity and helps to decipher the emergence of symbiosis and the evolution of pathogenicity in this genus.
Materials and Methods
Host E. petzi Cells
The E. petzi cells were isolated from a sample of seawater and sandy bottom collected by means of a sediment trap from Adelie Cove in Antarctica, at a depth of 27 m, a temperature of − 1.2 °C and a salinity of 34‰. Cultures were maintained in the laboratory in cold rooms, at 4 °C, under a cycle of 12 h of very low light and 12 h of dark, as previously described [21]. The green alga Dunaliella tertiolecta was used as food source.
Fluorescent In Situ Hybridization (FISH)
E. petzi cells were collected from severely starved cultures, transferred onto glass slides, fixed with 4% formaldehyde in phosphate-buffered saline (PBS) for 10 min at room temperature, and permeabilized by ethanol gradient (50%, 80%, and 100% of ethanol in water, for 10 min each). The fluorescein-labeled probe EUB338 (5′-GCTGCCTCCCGTAGGAT-3′) for eubacteria and the Cy3-labeled probe Bwall1448 (5′-CAACCATTCGCCGGGCCT-3′) for Francisella were synthesized by Integrated DNA Technologies (Coralville, IA, USA). Hybridization was performed following the method described by Hugenholtz et al. [22]. Briefly, 2 μl of each probe solution (50 ng/μl) in 20 μl hybridization buffer (0.9 M NaCl, 20 mM Tris-HCl pH 7.0, 15% formamide, 0.1% SDS) were added directly to the cells on slides. Hybridization was performed in a humid chamber at 46 °C for 3 h. Slides were then washed 20 min with washing buffer (318 mM NaCl, 20 mM Tris-HCl pH 7.0, 0.1% SDS) at 48 °C and air dried. Slides were embedded with anti-fading mounting medium and then inspected with a Nikon confocal microscope (Nikon, Amsterdam, The Netherlands).
Light and Transmission Electron Microscopy (TEM)
In light microscopy, bacteria were observed in vivo, as well as after staining with the fluorescent dye Syto-9 (used as recommended by the manufacturer Molecular Probes, Eugene, OR, USA), on an Axiophot microscope equipped with an Axio Cam MRC color video camera and an AxioVision computer-assisted image analysis system (Zeiss, Jena, Germany).
For TEM analysis, bacteria were fixed with 2.5% glutaraldehyde and 6% sucrose in 0.1 M cacodylate buffer, pH 7.2, for 2 h at 4 °C. After three washings at 4 °C in the same buffer, samples were post-fixed with 1% osmium tetroxide in 0.1 M cacodylate buffer, pH 7.2, for 1 h at 4 °C, washed in the same buffer, and dehydrated in a gradient ethanol series. Samples were then infiltrated with mixtures of LRWhite resin/ethanol in different percentages, embedded in pure LRWhite resin, and left to polymerize for 2 days at 50 °C. Resin blocks were cut with a Reichert Ultracut ultramicrotome using a diamond knife. Ultrathin sections (60–80 nm) were collected on copper grids, stained with uranyl acetate and lead citrate, and observed with a JEOL 1200EXII electron microscope (JEOL, Peabody, MA, USA). Micrographs were captured using a SIS VELETA CCD camera (Olympus, Muenster, Germany) equipped with iTEM software.
Isolation, Identification, and Culturing
The isolation of F. adeliensis was carried out from E. petzi cell lysates following the procedure previously used by Sjödin et al. to isolate F. endociliophora [10], taking care to incubate plates at 4 °C. Briefly, E. petzi cell samples were bead beaten and acid treated according to Humrighouse et al. [23], before being diluted in PBS and spread on Cysteine Heart Agar Blood (CHAB) culture plates, supplemented with 105 U/l penicillin and 40 mg/l vancomycin as described [24]. The culture plates were incubated at 4 °C for 1 to 2 weeks and monitored for bacterial growth. Colonies were then isolated and maintained in CHAB plates at 4 °C. To identify F. adeliensis from other contaminating bacteria, isolated colonies were picked, resuspended in 20 μl of water and immediately lysed by boiling for 3 min. Five microliters of each lysed cell suspension were used as template in PCR, run using two sets of primers: fw1 (5′-GCGTTTACCACGGAGTGATT-3′) and rv1 (5′-TGGAGCCTAGCGGGATC-3′); fw2 (5′-AGTCAGGGAGGAAGTTTATTTGGTT-3′) and rv2 (5′-CACCTTCCTCCGCCTTGT-3′). Positive clones were maintained in CHAB plates for subsequent analysis.
For dot-plate analysis, a Francisella colony was suspended in 1 ml PBS buffer and 5 microliters of serial dilutions were spotted on CHAB plates and incubated at 4, 10, 20, 30, and 37 °C. Plates were checked for bacterial growth every 3 days. For growth assays in liquid medium, an overnight culture was used to inoculate aliquots of 50 ml of T medium [25] to reach an OD600 ranging from 0.01 to 0.05. Flasks were then incubated at different temperatures, salinity, and CO2 concentrations. Bacterial growth was monitored by measuring the OD600 every day during the first week, then every 3 days. The number of generation/day was calculated using the Origin 8 software.
The presence of the enzymes catalase and oxidase was tested on agar plates using a 3% H2O2 solution and an oxidase-strip (OXOID, Monza, Italy), respectively. Motility was determined with the hanging drop technique.
Genome Sequencing and Assembly
Isolated DNA was sequenced using the Nextera XT library protocol on an Illumina MiSeq instrument in addition to a Pacific Biosciences RSII system (10-kb library, 2-h movie length), generating a total of 57,926 PacBio reads with an average read length of 11,653 bp, using single-molecular real-time (SMRT) cells. The initial draft of the genome was generated by assembling PacBio reads using the SMRT analysis system version 2.3.0. Polishing of the draft genome was performed using Illumina reads [26].
Phylogenetic Analysis
The phylogenetic analysis was inferred using the neighbor-joining method [27]. The evolutionary distances among Francisella genomes were computed using the number of differences method [28] and are in the units of the number of base differences per sequence. The analysis involved 139 nucleotide sequences, with a total of 213,734 positions in the final dataset. Positions containing gaps and missing data were eliminated. Evolutionary analyses were conducted in MEGA7 [29]. Fangia hongkongensis was included as outgroup to generate the genome-based phylogenetic tree.
Results
Identification
Total DNA preparations of E. petzi subjected to high-throughput sequencing generated 24,800 assembled contigs (Villalobo and Vallesi, unpublished), of which approximately 800 (equivalent to a total of 1.6 Mb) revealed a close similarity to bacterial sequences available from public databases, with the highest matching value of each contig systematically resulting against gene sequences of Francisella species.
Among the 800 contigs, one of 5091 bp included the 16S and 23S rRNA gene sequences plus the sequences of the tRNAIle and tRNAAla genes (Fig. 1a). Therefore, it revealed to be a typical bacterial rDNA operon. Using the SILVA INcremental Aligner bioinformatics tool [30], the 16S rRNA gene sequence of this operon was classified as belonging to Francisella with 94.39% identity and 97 score along 1480 bp. Given that the 3% cutoff rule [31] for a 16S divergence among species was fulfilled, the new 16S rRNA gene sequence was assumed to belong to a new Francisella species for which the proposed name is Francisella adeliensis nov. sp.. The species name is after that of the Antarctic cove, Adelie, from which E. petzi, the F. adeliensis host, was collected.
F. adeliensis identification. a Schematic representation of the F. adeliensis rDNA operon. The relative positions of fluorescent FISH probes and primers used in colony-PCR are indicated. The 33-bp sequence exclusive of F. adeliensis rDNA operon is shown. b Fluorescent in situ hybridization of E. petzi cells: a, signal from fluorescein-labeled probe EUB338 for all eubacteria; b, signal from Cy3-labeled probe Bwall1448 specific for Francisella; c, co-localization of signals of the two labeled probes. Scale bar = 20 μm
Analyzed in the BLASTN 2.6.1 database [32] for its closest identity, the F. adeliensis 16S rRNA gene sequence showed the best alignment (only seven nucleotide variations along 1376 bp) with the 16S gene sequence of an unnamed and uncultured γ-proteobacterium reported to be a chemoautotrophic symbiont on gills of deep-sea clams and mussels collected at a 10-m depth from the fjord of Saanich Inlet, British Columbia [33]. The other two closest counterparts were the 16S sequences of F. endociliophora [10] and F. salina [24], with 96% of sequence identity along the 1481-bp gene length.
Intracellular Localization
To verify whether F. adeliensis resides as endosymbiont inside E. petzi, or it coexists as environmental bacteria with E. petzi in culture, E. petzi cells were starved for 10 days to avoid any possible bacterial contamination from undigested food, and analyzed by fluorescent in situ hybridization (FISH) with two distinct probes: the first (“EUB338,” see “Materials and Methods”) specific to a 16S rRNA-sequence conserved in most bacterial species, and the second (“Bwall1448”) specific to a 23S rRNA region unique to Francisella [34]. Both probes generated fluorescent signals within the cytoplasm of E. petzi cells (Fig. 1b), and their co-localization provided evidence that F. adeliensis was the only guest.
TEM analysis of E. petzi cells, suspended without food for not less than 1 week before being used, confirmed the presence of numerous bacteria (Fig. 2a, b). Only occasionally were bacteria observed to be individually dispersed in the cytoplasm, each confined inside a membranous-bound vesicle (Fig. 2e), or apparently free in the cytosol (Fig. 2f, g). Much more often, they appeared clustered together in larger fusogenic membrane-bound structures (Fig. 2c, d). These structures were quite heterogeneous in size and number of enclosed bacteria, and usually located in close proximity of the host’s somatic and transcriptionally active nucleus (macronucleus).
TEM of E. petzi cells containing F. adeliensis. a, b Micrographs showing bacteria individually dispersed in the host cytoplasm, or associated together in groups enclosed in membrane-bound compartments. c–g Magnifications of the boxed areas in a and b. MAC macronucleus, AZM adoral zone ciliary membranelles
Phenotypic Traits
Individual F. adeliensis colonies were screened by PCR using two sets of specific primers (Fig. 1a). Primers (“fw1” and “rv1,” see “Materials and Methods”) of one set were designed to amplify a 360-bp fragment containing a 33-bp sequence lying between the two tRNA coding regions and without counterparts in the rDNA operons of other Francisella species. Primers (“fw2” and “rv2”) of the second set were designed to amplify a 660-bp fragment of the 16S rRNA coding region shared among other Francisella species. Products sequenced from both amplifications showed to fully match the genomic data, confirming the taxonomic identity of the isolated colonies with F. adeliensis.
On CHAB plates, F. adeliensis colonies look spherical, white and slightly mucoidal, formed by roundish (diameter, 1–1.2 μm) bacteria (Fig. 3a), which are non-motile, catalase-positive, and oxidase-negative. In solid medium, colonies are visible after 3 days of incubation at temperatures ranging from 20 to 30 °C, and require 6–12 days to grow when incubated at 4 and 10 °C (Fig. 3b). In liquid medium, the highest growth rate was measured at 20 and 30 °C and the lowest at 4 °C (Fig. 3c). The mean numbers of generations/day were counted to be 0.11, 0.29, 0.53, and 0.47 at 4, 10, 20, and 30 °C, respectively, and no growth was observed at 37 °C. Roughly one half of bacteria inoculated on plates at 37 °C died after 16 h of incubation and none survived after 48 h.
F. adeliensis colonies and growth. a Colonies grown on a CHAB plate and bacteria as they appear after staining with Syto-9 fluorescent dye (left picture), and in vivo in differential interference contrast microscopy (right picture). Scale bar, 10 μm. b Dot-plate analysis. Serial dilutions of a F. adeliensis suspension were spotted on plates and plates incubated at the indicated temperatures for the indicated times (days). c Growth curves in liquid medium incubated at the indicated temperatures. Data from a representative experiment are shown; experiments were repeated three times with equivalent results
In the presence of 5% CO2, F. adeliensis cultures grew with OD600 values approximately 60% lower than those measured in ambient atmosphere (0.04%). Instead, no significant variation in the growth rate was observed in cultures left to grow in liquid medium containing salt concentrations ranging from 0 to 35‰, implying that F. adeliensis is a strongly euryhaline bacterium (data not shown).
Genomic Features
The F. adeliensis genome extends for 2,054,094 bp, a length matching the mean genome size of other Francisella species (1.96 ± 0.14 Mbp, Table 1) much more closely than the size of any other bacterial genome (3.82 ± 1.8 Mbp) [35]. It contains 1880 predicted protein coding sequences, 38 tRNA genes, ten rRNA genes (four 5S rRNA, three 16S rRNA, and three 23S rRNA) and one tmRNA gene (Table 1). Its average nucleotide identity (ANI) with the closest Francisella genomes is in the range of 77–78.8% (Table 2), which is distant from the 95–96% range usually taken as the minimum threshold value to consider two genome sequences as belonging to the same species [36]. Consistently with an intracellular lifestyle, the average 32.6% G+C content of the F. adeliensis genome closely reflects the 32.38 ± 0.24% content of the other endosymbiotic Francisella, and is significantly lower than the average G+C content (49.1 ± 12.4%) shown by free-living bacteria [35].
Based on a search for transposable elements and phages carried out with PHASTER and ISFinder softwares [37, 38], the F. adeliensis genome contains prophage sequences like other Francisella. However, it includes only one ISFtu4 insertion sequence element (E-value 1e−15).
Analysis of the F. adeliensis genome for the presence of the ten igl (intracellular growth locus) and five pdp (pathogenicity determinant proteins) genes, which composethe so-called Francisella pathogenicity island responsible for the virulence of F. tularensiss [39, 40], indicated that all igl genes are present, while the pdp gene set lacks the pdpC and pdpE genes.
The observation that F. adeliensis requires complex media to grow in culture suggested a loss of genes responsible for the synthesis of essential amino acids. This hypothesis was verified by screening the F. adeliensis genome for the presence of genes responsible for the synthesis of arginine, cysteine, histidine, lysine, methionine and tyrosine for which the pathogenic F. tularensis is known to be auxotrophic [41]. Only the histidine and arginine biosynthesis appeared to be genetically supported: the histidine biosynthesis by the complete set of relevant genes, and the arginine biosynthesis by the argJ gene that likely replaces the lack of argA, argD, and argE genes [42]. Instead, the biosynthesis of the other four amino acids appeared genetically not supported. The F. adeliensis genome lacks the genes dapD, dapC, and dapE encoding enzymes responsible for the lysine biosynthesis [43], as well as the gene encoding cystathionine γ-synthase responsible for the methionine and cysteine biosynthesis [44]. With regard to the tyrosine biosynthesis, the genome contains the complete gene set for the shikimate pathway, but it lacks the gene encoding prephenate dehydrogenase which converts prephenic acid to 4-hydroxyphenyl-pyruvic acid [45]. In conclusion, F. adeliensis shows to be prototrophic for arginine and histidine, and auxotrophic for cysteine, lysine, methionine, and tyrosine.
Phylogenetic Relationships
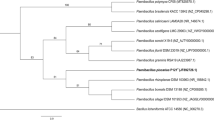
To assess the F. adeliensis interspecific relationships, the F. adeliensis genome was compared with the other Francisella genomes available from NCBI using 139 gene sequences for a total of 213,734 nucleotide positions. As shown in Fig. 4, F. adeliensis forms its own clade with a high statistical support. Together with the clade formed by F. frigiditurris, a species recently isolated from the water of a cooling tower [46], it precedes the split of four other major clades in which all the other Francisella species are subdivided in full accord with the recently proposed genome-based Francisella phylogeny [46, 47]. One of the four clades is specific to species, such as F. tularensis and F. novicida, that are pathogenic to terrestrial hosts, and F. persica (formerly Wolbachia persica) isolated from ticks [12, 17]. The second one includes species such as F. noatunensis that are pathogenic to fish, as well as F. salina isolated from a seawater sample [46]. The third one includes species such as F. endociliophora and F. halioticida isolated from marine hosts, together with F. uliginis isolated from a seawater sample [9, 12, 46]. And the fourth one is specific to Francisella species that have been isolated from waters of cooling systems, and are usually described as “environmental” species and regarded as belonging to the genus Allofrancisella [48, 49].
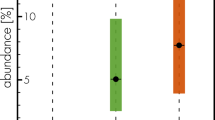
Evolutionary relationships of F. adeliensis. The optimal tree with the sum of branch length = 284,909.68814135 is shown. The tree is drawn to scale, with branch lengths in the same units as those of the evolutionary distances used to infer the phylogenetic tree. The scale bar corresponds to 5000 nucleotide differences. Growth style and environment of each species are indicated by colored dots on the right; the six major branches of the tree are enclosed in colored rectangles. The position of F. adeliensis is highlighted in bold
Discussion
The isolation reported here of F. adeliensis from an Antarctic strain of E. petzi follows the isolation of F. endociliophora from E. raikovi [10], which is a species distributed in the Caspian and Mediterranean Seas and Eastern Atlantic Ocean [50], and the identification of DNA sequences of a taxonomically undetermined Francisella in the genome of E. focardii [51], which is a species endemic to Antarctic coastal waters [52]. Altogether, these findings strongly suggest that Francisella/Euplotes associations are relatively common in the marine environment, and two additional considerations reinforce this hypothesis. The first consideration is related to the bipolar biogeographic distribution that characterizes the species structure of the F. adeliensis’s host, E. petzi [21, 53]. Embracing Arctic and White Sea populations in addition to Antarctic and peri-Antarctic ones, this distribution clearly implies that the F. adelinesis association with E. petzi is likely not restrained to the Antarctic waters where it has been detected. Being extended to the high latitudes of both the hemispheres, it appears to be virtually global and the analysis of other bipolar Euplotes species for their symbiotic associations with F. adeliensis and/or its close relatives may definitively establish these global dimensions. The second and more significant consideration is related to the psychrophilic and euryhaline behavior shown by F. adeliensis. Growing well at temperatures ranging from 4 to 30 °C and promptly adapting to 0–35 ‰ variations in the ambient salinity, F. adeliensis appears capable of colonizing other organisms independently of their adaptation to live in marine, brackish, or lacustrine habitats of either cold or temperate areas.
The 16S rRNA gene sequences are the molecules of choice for phylogenetic reconstructions, but their use in devising a Francisella phylogenetic tree has frequently been biased by branches supported by low bootstrap values due to the particularly high degree of conservation that these sequences show in Francisella. Only the recent availability of genomic data provided more solid grounds to trace the phylogenetic relationships among Francisella species, producing phylogenetic trees with more solid statistic support [46, 54]. In the genome-based tree updated with the inclusion of F. adeliensis (shown above in Fig. 4), F. adeliensis branches surprisingly distant from all the intracellular Francisella, including F. endociliophora endocytobiont in E. raikovi. It correlates much closer to the two earliest branching clades that are uniquely formed by Francisella species, namely F. frigiditurris, Allofrancisella frigidaquae, and A. guangzhouensis, isolated from cooling towers. As such, they are collectively regarded as environmental species.
Granted that these species are really free living—considering the strong acidic conditions used for their isolation, it cannot be excluded that they have actually been isolated from some eukaryotic microorganisms living inside the cooling towers—this correlation implies that F. adeliensis foreruns the Francisella adaptive evolution in replacing a free-living lifestyle with an intracellular/endosymbiotic style. And the F. adeliensis acquisition of the endosymbiotic lifestyle is likely to be quite ancient, considering that a single IS element is present in its genome. In effect, a low number of mobile genetic elements is widely accepted to be a distinctive trait of an ancient stage of intracellular life and an expansion of these elements is regarded to be distinctive of initial stages of host restriction [55, 56]. In addition, the finding that F. adeliensis is auxotrophic for cysteine, lysine, methionine, and threonine, and likely depends on the host for nutrient supply, establishes a close physiological analogy with pathogenic strains of F. tularensis, whose virulence depends on the activity of the Francisella pathogenicity island cluster of genes [40]. In a mouse model of tularaemia, it has been shown that among these genes F. tularensis and F. novicida particularly need the expression of the pdpC gene in order to escape from phagosomes and become free in the cytosol [57]. Neither the pdpE gene, which is not directly involved in F. tularensis virulence, nor the pdpC gene was identified in the F. adeliensis genome. However, evidence from TEM analysis indicated that F. adeliensis may produce cytosolic stages in addition to more common fusogenic membrane-bound structures closely recalling the “Francisella containing vacuoles” involved in the autophagy-mediated mechanism of F. tularensis re-entry into the endocytic compartment [58]. Although these cytosolic stages suggest that Euplotes is a potential ecological reservoir for the evolution of pathogenic Francisella, the observation that F. adeliensis is unable to proliferate at 37 °C would exclude any ability to colonize and be harmful to homothermic, warm-blood organisms.
Description of Francisella adeliensis Sp. Nov.
Francisella adeliensis (a.de.lien’sis. L. adj. of Adelie) is named after Adelie Cove, the location in Antarctica where the host, the ciliate Euplotes petzi, was collected in 2005 [21]. The type strain is deposited at the Swedish Defence Research Institute (FOI), Francisella strain collection # FSC1327. Within its host, F. adeliensis resides in the cytoplasm, as determined by TEM and FISH analysis carried out with the Francisella-specific probe Bwall1448 [34]. Cells are Gram-negative, roundish in shape, non-motile, catalase-positive, and oxidase-negative. They grow at a wide range of temperature (4–30 °C), salinity (0–35‰), and carbon dioxide concentrations (0.04–5%). The F. adeliensis complete genome sequence is available at GenBank, with the accession number CP021781, and supporting sequencing data are deposited in Bioproject PRJNA389235.
References
Petroni G, Spring S, Schleifer KH, Verni F, Rosati G (2000) Defensive extrusive ectosymbionts of Euplotidium (Ciliophora) that contain microtubule-like structures are bacteria related to Verrucomicrobia. Proc. Natl. Acad. Sci. U. S. A. 97:1813–1817
Görtz HD (2006) Symbiotic associations between ciliates and prokaryotes. In: Dworkin M, Falkow S, Rosenberg E, Schleifer K-H, Stackebrandt E (eds) The prokaryotes. Springer, New York, pp 364–402
Fokin SI (2012) Frequency and biodiversity of symbionts in representatives of the main classes of Ciliophora. Eur. J. Protistol. 48:138–148
Dziallas C, Allgaier M, Monaghan MT, Grossart HP (2012) Act together—implications of symbioses in aquatic ciliates. Front. Microbiol. 3:288
Fujishima M, Görtz H-D (1983) Infection of macronuclear anlagen of Paramecium caudatum with the macronucleus-specific symbiont Holospora obtusa. J. Cell Sci. 64:137–146
Fujishima M (2009) Infection and maintenance of Holospora species in Paramecium caudatum. In: Fujishima M (ed) Endosymbionts in Paramecium. Springer, Dordrecht Heidelberg London New York, pp 201–226
Schweikert M, Fujishima M, Görtz HD (2013) Symbiotic associations between ciliates and prokaryotes. In: Rosenberg E, DeLong EF, Lory S, Stackebrandt E, Thompson F (eds) The prokaryotes: prokaryotic biology and symbiotic associations. Springer, Berlin, Heidelberg, pp 427–463
Vannini C, Ferrantini F, Ristori A, Verni F, Petroni G (2012) Betaproteobacterial symbionts of the ciliate Euplotes: origin and tangled evolutionary path of an obligate microbial association. Environ. Microbiol. 14:2553–2563
Schrallhammer M, Schweikert M, Vallesi A, Verni F, Petroni G (2011) Detection of a novel subspecies of Francisella noatunensis as endosymbiont of the ciliate Euplotes raikovi. Microbial Ecol 61:455–464
Sjödin A, Öhrman C, Bäckman S, Lärkeryd A, Granberg M, Lundmark E et al (2014) Complete genome sequence of Francisella endociliophora strain FSC1006 isolated from a laboratory culture of the marine ciliate Euplotes raikovi. GenomeA 2:e01227–e01214
Sjöstedt A (2005) Family III Francisellaceae fam nov. In: Garrity GM (ed) Bergeys manual of systematic bacteriology. Springer, Baltimore, pp 199–210
Colquhoun DJ, Larsson P, Duodu S, Forsman M (2014) The family Francisellaceae. In: Rosenberg E, EF DL, Lory S, Stackebrandt E, Thompson F (eds) The prokaryotes. Springer, Berlin Heidelberg, pp 287–314
Dorofe'ev KA (1947) Classification of the causative agent of tularemia. Symposium Research Works Institute Epidemiology and Microbiology Chita 1:170–180
Olsufjev NG, Meshcheryakova IS (1983) Subspecific taxonomy of Francisella tularensis McCoy and Chapin 1912. Int. J. Syst. Bacteriol. 33:872–874
Ottem KF, Nylund A, Karlsbakk E, Friis-Moller A, Kamaishi T (2009) Elevation of Francisella philomiragia subsp noatunensis Mikalsen et al (2007) to Francisella noatunensis comb nov [syn Francisella piscicida Ottem et al (2008) syn nov] and characterization of Francisella noatunensis subsp orientalis subsp nov two important fish pathogens. J. Appl. Microbiol. 106:1231–1243
Colquhoun DJ, Duodu S (2011) Francisella infections in farmed and wild aquatic organisms. Vet. Res. 8(42):47
Larson MA, Nalbantoglu U, Sayood K, Zentz EB, Cer RZ, Iwen PC, Francesconi SC, Bishop-Lilly KA, Mokashi VP, Sjöstedt A, Hinrichs SH (2016) Reclassification of Wolbachia persica as Francisella persica comb nov and emended description of the family Francisellaceae. Int. J. Syst. Evol. Microbiol. 66:1200–1205
Hollis DG, Weaver RE, Steigerwalt AG, Wenger JD, Moss CW, Brenner DJ (1989) Francisella philomiragia comb nov (formerly Yersinia philomiragia) and Francisella tularensis biogroup novicida (formerly Francisella novicida) associated with human disease. J. Clin. Microbiol. 27:1601–1608
Mailman TL, Schmidt MH (2005) Francisella philomiragia adenitis and pulmonary nodules in a child with chronic granulomatous disease. Can J Infect Dis Med Microbiol 16:245–248
Sjöstedt A (2007) Tularemia: history epidemiology pathogen physiology and clinical manifestations. Ann. N. Y. Acad. Sci. 1105:1–29
Di Giuseppe G, Erra F, Frontini F, Dini F, Vallesi A, Luporini P (2014) Improved description of the bipolar ciliate Euplotes petzi and definition of its basal position in the Euplotes phylogenetic tree. Eur. J. Protistol. 50:402–411
Hugenholtz P, Tyson GW, Blackall LL (2002) Design and evaluation of 16S rRNA-targeted oligonucleotide probes for fluorescence in situ hybridization. Methods Mol. Biol. 179:29–42
Humrighouse BW, Adcock NJ, Rice EW (2011) Use of acid treatment and a selective medium to enhance the recovery of Francisella tularensis from water. Appl. Environ. Microbiol. 77:6729–6732
Petersen JM, Carlson J, Yockey B, Pillai S, Kuske C, Garbalena G et al (2009) Direct isolation of Francisella spp from environmental samples. Lett. Appl. Microbiol. 48:663–667
Pavlovich NV, Mishan'kin BN (1987) Transparent nutrient medium for culturing Francisella tularensis. Antibiot Med Biotekhnol 32:133–137
Walker BJ, Abeel T, Shea T, Priest M, Abouelliel A, Sakthikumar S, Cuomo CA, Zeng Q, Wortman J, Young SK, Earl AM (2014) Pilon: an integrated tool for comprehensive microbial variant detection and genome assembly improvement. PLoS One 9:e112963
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 4:406–425
Nei M, Kumar S (2000) Molecular evolution and phylogenetics. Oxford University Press, New York
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 70 for bigger datasets. Mol. Biol. Evol. 33:1870–1874
Pruesse E, Peplies J, Glöckne FO (2012) SINA: accurate high-throughput multiple sequence alignment of ribosomal RNA genes. Bioinformatics 28:1823–1829
Kim M, Oh HS, Park SC, Chun J (2014) Towards a taxonomic coherence between average nucleotide identity and 16S rRNA gene sequence similarity for species demarcation of prokaryotes. Int. J. Syst. Evol. Microbiol. 64:346–351
Morgulis A, Coulouris G, Raytselis Y, Madden TL, Agarwala R, Schäffer AA (2008) Database indexing for production MegaBLAST searches. Bioinformatics 24:1757–1764
Walsh DA, Zaikova E, Howes CG, Song YC, Wright JJ, Tringe SG, Tortell PD, Hallam SJ (2009) Metagenome of a versatile chemolithoautotroph from expanding oceanic dead zones. Science 326:578–582
Splettstoesser WD, Seibold E, Zeman E, Trebesius K, Podbielski A (2010) Rapid differentiation of Francisella species and subspecies by fluorescent in situ hybridization targeting the 23S rRNA. BMC Microbiol. 10(72):72
Merhej V, Royer-Carenzi M, Pontarotti P, Raoult D (2009) Massive comparative genomic analysis reveals convergent evolution of specialized bacteria. Biol. Direct 4(13):13
Stackebrandt E, Ebers J (2006) Taxonomic parameters revisited: tarnished gold standards. Microbiol. Today 33:152–155
Arndt D, Grant JR, Marcu A, Sajed T, Pon A, Liang Y, Wishart DS (2016) PHASTER: a better faster version of the PHAST phage search tool. Nucl Acids Res 44:W16–W21
Siguier P, Pérochon J, Lestrade L, Mahillon J, Chandler M (2006) ISfinder: the reference centre for bacterial insertion sequences. Nucl Acids Res 34:D32–D36
Gray CG, Cowley SC, Cheung KK, Nano FE (2002) The identification of five genetic loci of Francisella novicida associated with intracellular growth. FEMS Microbiol. Lett. 215:53–56
Nano FE, Zhang N, Cowley SC, Klose KE, Cheung KK, Roberts MJ, Ludu JS, Letendre GW, Meierovics AI, Stephens G, Elkins KL (2004) A Francisella tularensis pathogenicity island required for intramacrophage growth. J. Bacteriol. 186:430–6436
Santic M, Abu Kwaik Y (2013) Nutritional virulence of Francisella tularensis. Front. Cell. Infect. Microbiol. 3:112
Xu Y, Labedan B, Glansdorff N (2007) Surprising arginine biosynthesis: a reappraisal of the enzymology and evolution of the pathway in microorganisms. Microbiol. Mol. Biol. Rev. 71:36–47
Velasco AM, Leguina JI, Lazcano A (2002) Molecular evolution of the lysine biosynthetic pathways. J. Mol. Evol. 55:445–449
Ferla MP, Patrick WM (2014) Bacterial methionine biosynthesis. Microbiology 160:1571–1584
Ahmad S, Jensen RA (1988) The phylogenetic origin of the bifunctional tyrosine-pathway protein in the enteric lineage of bacteria. Curr. Microbiol. 16:295–310
Challacombe JF, Petersen JM, Hodge D, Pillai S, Kuske CR (2017) Whole-genome relationships among Francisella bacteria of diverse origins define new species and provide specific regions for detection. Appl. Environ. Microbiol. 83:e02589–e02516
Sjödin A, Svensson K, Öhrman C, Ahlinder J, Lindgren P, Duodu S, Johansson A, Colquhoun DJ, Larsson P, Forsman M (2012) Genome characterisation of the genus Francisella reveals insight into similar evolutionary paths in pathogens of mammals and fish. BMC Genomics 13:268
Qu PH, Chen SY, Scholz HC, Busse HJ, Gu Q, Kämpfer P et al (2013) Francisella guangzhouensis sp nov isolated from air conditioning systems. Int. J. Syst. Evol. Microbiol. 63:3628–3635
Qu PH, Li Y, Salam N, Chen SY, Liu L, Gu Q et al (2016) Allofrancisella inopinata gen nov sp nov and Allofrancisella frigidaquae sp nov isolated from water-cooling systems and transfer of Francisella guangzhouensis Qu et al 2013 to the new genus as Allofrancisella guangzhouensis comb nov. Int. J. Syst. Evol. Microbiol. 66:4832–4838
Jiang J, Zhang Q, Warren A, Al-Rasheid KA, Song W (2010) Morphology and SSU rRNA gene-based phylogeny of two marine Euplotes species E orientalis spec nov and E raikovi (Ciliophora Euplotida). Eur. J. Protistol. 46:121–132
Pucciarelli S, Devaraj RR, Mancini A, Ballarini P, Castelli M, Schrallhammer M, Petroni G, Miceli C (2015) Microbial consortium associated with the Antarctic marine ciliate Euplotes focardii: an investigation from genomic sequences. Microbial Ecol 70:484–497
Valbonesi A, Luporini P (1993) Biology of Euplotes focardii an Antarctic ciliate. Polar Biol. 13:489–493
Di Giuseppe G, Dini F, Vallesi A, Luporini P (2015) Genetic relationships in bipolar species of the protist ciliate Euplotes. Hydrobiologia 761:71–83
Ahlinder J, Ohrman C, Svensson K, Lindgren P, Johansson A, Forsman M et al (2012) Increased knowledge of Francisella genus diversity highlights the benefits of optimised DNA-based assays. BMC Microbiol. 12:220
Moran NA, Plague GR (2004) Genomic changes following host restriction in bacteria. Curr. Opin. Genet. Dev. 14:627–633
Dutta C, Paul S (2012) Microbial lifestyle and genome signatures. Curr Genomics 13:153–162
Brodmann M, Dreier RF, Broz P, Basler M (2017) Francisella requires dynamic type VI secretion system and ClpB to deliver effectors for phagosomal escape. Nat. Commun. 8:15853
Checroun C, Wehrly TD, Fischer ER, Hayes SF, Celli J (2006) Autophagy-mediated reentry of Francisella tularensis into the endocytic compartment after cytoplasmic replication. Proc. Natl. Acad. Sci. U. S. A. 103:14578–14583
Acknowledgments
E.V. was supported by the Universidad de Sevilla (Movilidad Docente ERASMUS). We acknowledge the National Genomics Infrastructure (NGI)/Uppsala Genome Center and UPPMAX for providing assistance in massive parallel sequencing and computational infrastructure..
Funding
This work was financially supported by the PNRA (Programma Nazionale di Ricerca in Antartide) from the Italian Ministry of Research (MIUR), FAR (Fondo Ateneo Ricerca) from the University of Camerino, by the Swedish Ministry of Defence (A4040), and the Swedish Civil Contingencies Agency (B4010). The work performed at NGI/Uppsala Genome Center was funded by RFI/VR and Science for Life Laboratory, Sweden.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Vallesi, A., Sjödin, A., Petrelli, D. et al. A New Species of the γ-Proteobacterium Francisella, F. adeliensis Sp. Nov., Endocytobiont in an Antarctic Marine Ciliate and Potential Evolutionary Forerunner of Pathogenic Species. Microb Ecol 77, 587–596 (2019). https://doi.org/10.1007/s00248-018-1256-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-018-1256-3