Abstract
Both diet and host phylogeny shape the gut microbial community, and separating out the effects of these variables can be challenging. In this study, high-throughput sequencing was used to evaluate the impact of diet and phylogeny on the gut microbiota of nine colobine monkey species (N = 64 individuals). Colobines are leaf-eating monkeys that fare poorly in captivity—often exhibiting gastrointestinal (GI) problems. This study included eight Asian colobines (Rhinopithecus brelichi, Rhinopithecus roxellana, Rhinopithecus bieti, Pygathrix nemaeus, Nasalis larvatus, Trachypithecus francoisi, Trachypithecus auratus, and Trachypithecus vetulus) and one African colobine (Colobus guereza). Monkeys were housed at five different captive institutes: Panxi Wildlife Rescue Center (Guizhou, China), Beijing Zoo, Beijing Zoo Breeding Center, Singapore Zoo, and Singapore Zoo Primate Conservation Breeding Center. Captive diets varied widely between institutions, but within an institution, all colobine monkey species were fed nearly identical or identical diets. In addition, four monkey species were present at multiple captive institutes. This allowed us to parse the effects of diet and phylogeny in these captive colobines. Gut microbial communities clustered weakly by host species and strongly by diet, and overall, colobine phylogenetic relationships were not reflected in gut microbiota analyses. Core microbiota analyses also identified several key taxa—including microbes within the Ruminococcaceae and Lachnospiraceae families—that were shared by over 90% of the monkeys in this study. Microbial species within these families include many butyrate producers that are important for GI health. These results highlight the importance of diet in captive colobines.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The gut microbiota—a community of microbes living within the gastrointestinal tract—play an important role in both health and evolution of host species. On a broad scale, Ley et al. (2008a) examined the gut microbiota of 60 mammalian species and found more similar gut microbial composition in animals that were closely related taxonomically [1]. This study also found that animals with similar diets (i.e., herbivores, carnivores, omnivores) had more similar gut microbial communities. Host phylogeny influences the microbial community through immune-related genes that shape host-microbiota interactions and through vertical transmission of the microbiota [2,3,4,5]. Diet shapes the gut microbial community by providing substrates that differentially support or enhance the growth of specific microbes [6,7,8]. The gut microbiota can enable adaptation to new dietary niches [1]; thus, the effects of diet and host phylogeny are often intertwined, as dietary changes—which are mediated through gut microbes—can play a role in host speciation [9]. The effects of both host phylogeny and diet on the gut microbiota have been examined in multiple studies: In a few of these studies, similar diets appear to drive convergence of gut microbial communities between host species [10, 11]. In other studies (e.g., turtle ants, zebrafish, Oriental river prawn, great apes, New World leaf-nosed bats), host phylogeny appears to have a stronger impact on gut microbial community composition [12,13,14,15,16]. One complicating factor in many of these studies is that differing host species often have different diets and environments—either in captivity or in the wild—making it challenging to separate out the effects of phylogeny from diet.
In this study, we had the unique opportunity to parse effects of diet and phylogeny by examining gut microbiota of several closely related leaf-eating monkey species (subfamily Colobinae) that shared the same captive diets. Colobine monkeys are Old World monkeys that occur in Asia and Africa. They have evolved several specialized anatomical and physiological features that facilitate consumption of a cellulose- and lignin-rich diet. These foregut fermenting primates have significantly longer gastrointestinal (GI) tracts and larger stomach surface areas than other mammals—including other herbivorous primates [17]. Expansion of the gut increased retention time, and the highly sacculated forestomachs, also known as the saccus and presaccus, became the site for microbial fermentation [17, 18]. It has been suggested that these adaptations allowed colobines to shift their diet gradually from frugivory to folivory, reducing their competition with apes [17]. While colobine digestive anatomy has been well described [17, 19,20,21], less is known about the composition and diversity of the colobine gut microbial community and how it potentially evolved from handling a high sugar, low fiber diet (ripe fruit) to a low sugar, high fiber diet (mature leaves) [22,23,24,25]. Studying the gut microbiota of colobines can provide insights into the evolution of folivory in these primates. This type of study is also important for another reason: colobines fare poorly in captivity and commonly exhibit GI problems (e.g., diarrhea, vomiting, bloat, etc.) [21, 25,26,27,28,29]. In the wild, colobines consume a variety of plants daily and seasonally. Captivity represents a rapid and dramatic change in diet and environment, which can significantly alter the gut microbiota and potentially have negative consequences on host health [30, 31].
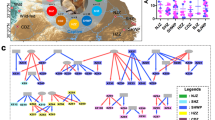
For colobine phylogeny, we referred to the phylogenies constructed by Wang et al. [32] and Liedigk et al. [33]. These phylogenies are based on both mitochondrial and nuclear DNA sequences. While there is some controversy about the phylogenetic tree at the species level, there is a general consensus at the genus level: Rhinopithicus is most closely related to Pygathrix and Nasalis followed by Trachypithecus. Colobus, the African colobine, is the most distantly related (Fig. 1). We predicted that if diet plays a stronger role in gut microbiota composition than phylogeny does, or if diet overwhelms subtle microbial community differences between closely related host species, then gut microbial communities should be more similar in all colobine species consuming the same diet. Alternately, if primate phylogeny plays a stronger role in gut microbiota composition than diet, then gut microbial communities should be more similar in conspecific or congeneric monkeys that consume different diets.
Methods
Study Design
Our study examined gut microbiota in nine colobine species: eight Asian colobines (Rhinopithecus brelichi, Rhinopithecus roxellana, Rhinopithecus bieti, Nasalis larvatus, Trachypithecus francoisi, Trachypithecus auratus, Trachypithecus vetulus, Pygathrix nemaeus) and one African colobine (Colobus guereza). These colobines were housed at five captive institutes: Beijing Zoo (China), Beijing Zoo Breeding Center (China), Panxi Wildlife Rescue Center (China), Singapore Zoo (Singapore), and Singapore Zoo Primate Conservation Breeding Center (Singapore). Six species (R. brelichi, R. roxellana, R. bieti, N. larvatus, P. nemaeus, T. auratus) were sampled at multiple captive institutes. We also sampled multiple monkey species within each captive institute. Diets varied by captive institute; however, within an institute, diets for all colobine species were identical or nearly so, as assessed through observation and review of diet sheets (Table 1). Therefore, “captive institute” is used in this study as a proxy for diet.
Primate Fecal Collection and Processing
A total of 70 fecal samples were collected from 64 individuals during the summers of 2010, 2012, and 2013. All individuals at each captive institute had access to both indoor and outdoor enclosures that were cleaned daily. Monkeys at the Beijing Zoo, Panxi Wildlife Rescue Center, and Singapore Zoo additionally had access to natural space including trees and soil. Average temperature and humidity across locations and collections dates were relatively similar (Table S1). As available, age, sex, grouping, and wild/captive born status were recorded (Table S2). Dietary data was also recorded at each captive institute (Table 1).
All feces were processed on Flinders Technology Associates (FTA) cards as previously described [34, 35]. Briefly, a sterile cotton swab (Dynarex, Orangeburg, NY, USA) was inserted into the center of the feces, rotated, and withdrawn. The swab was then applied to an FTA card (Whatman Inc., Florham Park, NJ), which was air dried and then stored in a Ziploc bag at room temperature with MiniPax desiccant packets (Multisorb Technologies Inc., Buffalo, NY, USA). Samples were stored for up to 38 months prior to DNA extraction.
Earth Microbiome Project (EMP) protocols were followed for DNA extraction, amplification, and library preparation [36] with one alteration: Twenty punches were made in each FTA sample circle using a 2-mm Harris Uni-Core biopsy punch (TedPella, Redding, CA, USA). These 20 punches were then used as the starting material for DNA extraction with a PowerSoil DNA isolation kit (MoBio Laboratories, Carlsbad, CA, USA). Extraction was performed at Purdue University (West Lafayette, IN, USA) or the Zhejiang Institute of Microbiology in Hangzhou, China. The remainder of sample processing, sequencing, and core amplicon data analysis were performed by the Earth Microbiome Project (www.earthmicrobiome.org) at the BioFrontiers Institute Next-Generation Genomics Facility at University of Colorado, Boulder, USA as previously described [34, 35]. All amplicon and metadata have been made public through the data portal (https://qiita.ucsd.edu/) [36]. Samples collected in 2010 and 2012 were paired-end sequenced on an Illumina HiSeq 2000. Samples collected in 2013 were paired-end sequenced on an Illumina MiSeq. MiSeq reads (150-base pair long) were trimmed to 100 base pairs to match HiSeq reads.
Microbial Taxonomic Assignment
16S amplicons were de-multiplexed, quality-filtered, and clustered using default parameters in Quantitative Insights Into Microbial Ecology (QIIME version 1.8.0), a software that facilitates microbial community analysis [37]. A total of 11.2 million reads (mean = 36,000; standard deviation = 9600) resulted after filtering. Sequences were grouped into operational taxonomic units (OTUs) at a sequence similarity of 97%. Sequences were then assigned to OTUs via open-reference picking. This was done through UCLUST in QIIME, which used the Greengenes reference dataset (version 13_8, release date August 2013 [38]) (http://greengenes.secondgenome.com). Representative OTU sequences were aligned to a Greengenes reference alignment [39]. De novo OTUs were classified using RDP classifier and the Greengenes reference set (version 13_8, release date August 2013) with a minimum 80% confidence threshold [40]. Samples were rarified at 15,029 reads.
Statistical Analyses
Observed OTUs, Shannon, Simpson, Chao1, and PD Whole Tree diversity indices were calculated in QIIME to compare gut microbial species richness between colobine species and genera. These values were then compared using ANOVA or Kruskal-Wallis tests with Tukey’s HSD or Dunn’s post hoc pairwise comparisons respectively in RStudio (version 0.99.465). In all diversity index analyses, two outliers were noted. One outlier, a nursing 3-month-old male infant R. roxellana, C008, had a very low gut microbial diversity. Human studies indicate that gut microbial diversity of nursing infants is low compared to adults but expands rapidly when solid food is added to the diet [41, 42]. C008 was removed from diversity index analyses because he was the only infant sampled in our study. Thus, his age and diet (primarily milk) were distinct from that of all other monkeys. The other outlier was a sub-adult female R. brelichi, G032, who had the lowest overall gut microbial diversity (4.1 standard deviations below the average) of any monkey we sampled. She was co-housed with two other R. brelichi that did not exhibit low gut microbial diversity. No other information is available regarding the health or history of G032 at the time of sampling. Because of the significant effect G032 had on some results, analyses are reported with and without this individual included.
Microbial composition of each sample was assessed at various taxonomic levels (phylum through genus) using QIIME. A core microbiota analysis was also run in QIIME to identify OTUs (bacterial taxa) present in >90% of the samples included in this study. The goal of this analysis is to identify taxa that may be shared across host species potentially due to the importance of these taxa in association with host evolution and dietary choices.
Beta diversity was assessed using UniFrac distances within QIIME [43]. UniFrac distances are measured as the distances between microbial communities accounting for phylogenetic relationships between microbes. Principal coordinate analysis (PCoA) and Unweighted Pair Group Method with Arithmetic mean (UPGMA) allowed visualization of UniFrac distance patterns. Unweighted PCoA, which only accounts for the microbial species present, not their abundance, was used to determine if monkeys cluster by primate species or diet (i.e., captive institute).
Supervised learning analyses—also known as Random Forests—were performed in QIIME to determine if host phylogeny or diet could be used to differentiate samples based on microbial composition (OTUs) [44, 45]. This analysis was performed after filtering out all OTUs that were present in fewer than two samples. Twenty percent of the data were used as a test set, while 80% of the data were used as a training set. One thousand decision trees were generated based on groups (genera, diet) and microbial composition (OTUs.) Results from this analysis produced an error ratio. The error ratio is the error of random guessing over the error in the test sets summed. Finally, a PERMANOVA to test the effects of both host genera and diet was run in R Studio (version 0.99.465; vegan package, adonis function) [46].
Results
Differences in Microbial Diversity Associated with Phylogeny but Not Diet
Diversity indices were calculated for the microbial communities within each individual. Shannon, Simpson, PD Whole Tree, and observed OTU metrics all produced similar results (Figs. 2 and S1); thus, we chose to focus our analyses on the Shannon Diversity Index that accounts for both microbial richness and abundance. Chao1 results differed slightly (Fig. S1). When analyzing Shannon diversity by host species alone, N. larvatus had a significantly lower microbial diversity than all other species except T. vetulus, R. bieti, and P. nemaeus (Table S3). At a genus level, Nasalis monkeys had significantly lower microbial diversity than all other genera except Pygathrix (Shannon without G032 ANOVA: F 4, 63 = 8.53, p < 0.0001—Fig. 2; Shannon with G032 Kruskal-Wallis: p < 0.0001; post hoc pairwise comparisons: Nasalis-Colobus without G032: p = 0.008; with G032: Benjamini-Hochberg adjusted (BH): p = 0.007; Nasalis-Rhinopithecus without G032: p < 0.0001; with G032 BH: p = 0.0003; Nasalis-Trachypithecus without G032: p = 0.0001; with G032 BH adjusted: p = 0.0003).
Shannon diversity indices for primate species that had two or more individuals housed at more than one captive institute were also analyzed by diet. One model was run for each host species with diet as the sole regressor. There were no significant differences in gut microbial diversity by diet (N. larvatus—Singapore Zoo versus Singapore Zoo Primate Conservation Breeding Center diet: F 1, 14 = 2.23; p = 0.16; R. roxellana—Panxi Wildlife Rescue Center versus Beijing Zoo diet: F 1, 7 = 0.001; p = 0.97; R. bieti—Beijing Zoo versus Beijing Zoo Breeding Center diet: F 1, 5 = 1.51; p = 0.27; R. brelichi—Panxi Wildlife Rescue Center versus Beijing Zoo diet: F 1, 8 = 2.24; p = 0.17). When primate species and diet were combined in an ANOVA with collection year included as a random effect, species but not diet was a significant predictor of Shannon diversity (species: F 8, 53 = 5.25, p < 0.0001; diet/captive institute: F 3, 53 = 2.67, p = 0.058). The interaction term between species and diet was not significant (F 3, 53 = 0.71, p = 0.55).
Microbial Composition Varied by Phylogeny and Diet
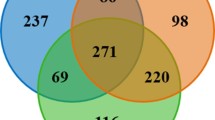
A core microbiota analysis found that the colobine monkeys we sampled share many of the same gut bacterial taxa (Table 2). One hundred percent of the monkeys sampled shared two OTUs (roughly equivalent to bacterial species) in the phylum Firmicutes, order Clostridiales. Ninety percent of the monkeys sampled shared a total of 38 OTUs, 27 of which were in the phylum Firmicutes. Common families of these shared Firmicutes OTUs include Lachnospiraceae and Ruminococcaceae.
A PCoA based on unweighted beta diversity values indicated that gut microbial communities cluster weakly by primate species (Fig. 3a), moderately by primate genus (Fig. 3b), and strongly by diet (Fig. 3c). For all Principal Coordinate Analyses, PC1 accounted for 8% of the variation and samples along this axis separated roughly by country of sampling, with samples collected in China clustering separately from those collected in Singapore. PC2 accounted for 6% of the variation and samples along this axis separated roughly by captive institute within China (Beijing Zoo, Beijing Zoo Breeding Center, Panxi Wildlife Rescue Center) or within Singapore (Singapore Zoo, Singapore Zoo Primate Conservation Breeding Center). A UPGMA based on unweighted UniFrac distances also indicated clustering of gut microbial communities by primate genus and diet (Fig. 4). The left side of the figure, labeled by primate genera, shows that while there is clustering evident by genera, there were also some clusters (e.g., the Trachypithecus, Colobus, Rhinopithecus cluster in the center) that could only be explained by diet. All five of the monkeys in this central cluster were housed at the Beijing Zoo Breeding Center and received the same diet.
Principal Coordinate Analysis (PCoA) on unweighted UniFrac distances between a colobine species, b colobine genera, and c diets based on captive institute. Unweighted UniFrac distances account for microbial species richness but not evenness. In a, arrow indicates a 3-month-old R. roxellana infant (C008) that was still nursing at the time of sample collection. The dashed square indicates a juvenile female R. roxellana, C005, who was housed singly. This sample was collected in 2010. The dashed circle indicates the same individual (C005) sampled in 2013, at which time she was co-housed with a large family group
Unweighted Pair Group Method with Arithmetic mean (UPGMA) using unweighted UniFrac distances. On the left, samples are identified by primate genera. On the right, the same samples are identified by diet. Arrow indicates R. roxellana individuals C008 and C005. (Note: Captive institute is used a proxy for diet). SZPCBC Singapore Zoo Primate Conservation Breeding Center
Two R. roxellana individuals separated out from the R. roxellana cluster in both PCoA and UPGMA analyses (Figs. 3a–c and 4). One of these individuals was C008 (the nursing infant). The second individual, C005, was a singly housed sub-adult female. No information is known about the health of C005; although, it is possible that stress (from being housed singly) could have had altered her gut microbiota [47, 48].
Supervised learning and PERMANOVA analyses indicated that diet had a stronger effect on microbial composition than primate phylogeny. Supervised learning analysis by primate species yielded an error ratio (error of random guessing over the sum of the error in the test sets) of 3.7. In contrast, analysis by diet produced an error ratio of 6.2. Error ratios greater than 2 indicate a significant difference among groups; higher error ratios indicate greater differences. When host species and diet were combined in a PERMANOVA analysis, diet had a significant effect while species had only a marginally significant effect on microbial composition (diet: pseudo-F = 1.17, R 2 = 0.067, p = 0.016; host species: pseudo-F = 1.09, R 2 = 0.124, p = 0.053). The diet/species interaction was not significant (diet: pseudo-F = 0.926, R 2 = 0.04, p = 0.865).
Discussion
Our results revealed that both diet and host phylogeny shape the gut microbiota in colobine monkeys. This finding is not unique to colobines (ants [49]; Antarctic seals, [50]; myrmecophages [10]; woodrats [51]; fish [52]; hominids [12]; bees, wasps, beetles, termites [53]). In our study, clear patterns of microbial composition emerged based on diet and primate genus. Microbial diversity played a lesser role in these patterns. Additionally, diet had a stronger influence on the gut microbial community than host phylogeny did—potentially because hosts were so closely related phylogenetically that there may have been few differences in their gut microbiota.
Microbial Diversity
All colobine genera except Pygathrix had significantly greater gut microbial diversity than Nasalis. Interestingly, Pygathrix and Nasalis are the most closely related genera phylogenetically and in the wild, Colobus, Trachypithecus, and Rhinopithecus, all tend to consume higher proportions of leaves—and particularly mature leaves—than Nasalis and Pygathrix (Table 3). In general, mature leaves contain higher amounts of cellulose, lignin, and tannins [66, 67] and lower amounts of protein than young leaves [68]. As a result, mature leaves are more difficult to digest than young leaves, and microbial fermentation may play an even more important role in primate species that consume higher proportions of mature leaves in their diet. Perhaps historically, the Colobus, Trachypithecus, and Rhinopithecus gut microbial communities expanded in diversity to increase their fermentative capacity, while Nasalis and Pygathrix, with their lower consumption of mature leaves, did not require a “fermentation boost.” Higher fiber consumption [69], and notably in wild colobine monkeys as compared captive colobine monkeys [30, 31], has been linked with increased microbial diversity in multiple recent studies. Unique behavioral physiology of Nasalis monkeys may also contribute to their microbial dynamics: Nasalis monkeys are the only colobines in which facultative merycism (rumination) has been reported [70, 71]. Regurgitation and remastication could potentially impact Nasalis gut microbial diversity by exposing forestomach ingesta and microbes to varying pH, oxygen, and nutrient conditions that reduce microbial diversity [72].
There were no significant differences in Shannon diversity indices as a function of diet (captive institute) within a primate species. This may have been due to small sample size. A power analysis (G*Power, version 3.1.5.1) indicated that the following sample sizes would have been necessary to detect significant differences based on diet: N. larvatus—41 monkeys at each captive institute; R. bieti—43 monkeys at each institute; R. brelichi—22 monkeys at each institute; and R. roxellana—130 monkeys at each institute. Our actual sample sizes were as follows: N. larvatus—16 monkeys total; R. bieti—7 monkeys total; R. brelichi—8 monkeys total; and R. roxellana—7 monkeys. Alternatively, diet may affect composition but not diversity of the gut microbiota. In a study on obesity, humans exhibited no change in microbial diversity but a dramatic change in gut microbial species composition before and after diet therapy [73]. Similarly, horses subjected to two different diets (high-energy forage versus forage-concentrate) differed in gut microbial species composition but not diversity [74]. Different but not fewer microbial species may thrive in the presence of different dietary substrates, and even if we had had larger sample sizes, we may still have made the same observation that, within a host species, microbial diversity did not differ across diets.
Microbial Composition
Our study identified a colobine “core microbiota” or a group of bacteria shared by captive colobine hosts. Because colobines are leaf-eating monkeys with similar digestive physiology, they might be expected to share microbes associated with the digestion of cellulose. That is, in fact, what we found. Most of the shared taxa were in the Lachnospiraceae and Ruminococcaceae families, including several OTUs in the genus Oscillospira (family Ruminococcaceae). Oscillospira species are found on leaf and grass cuticles within the gut of ruminants [75]. Notably, these bacteria are not found on leaves or grass in nature [75, 76]; rather, they are gut microbes adapted to living on and degrading leaves in the GI tract. Microbes within the Ruminococcaceae and Lachnospiraceae families have been reported in wild folivorous primates, and one study by Amato et al. [77] suggests that the changing abundances of these microbes seasonally are linked to increases and decreases in dietary energy consumption. Bacterial species within the Ruminococcaceae and Lachnospiraceae families also contain enzymes, transport mechanisms, and metabolic pathways that enable degradation of complex plant material, such as cellulose, hemicellulose, and lignin [78]. Many of these species also produce butyrate as a metabolic product of fiber degradation [79]. Butyrate is a short chain fatty acid (SCFA) critical to colonocyte health and immune defense [80, 81]. Butyrate also has anti-inflammatory properties and has been reported to alleviate obesity and the risk metabolic disease [82, 83]—both of which are growing concerns in captive populations. Inflammatory GI diseases have an inverse relationship with the abundance of bacterial species in the Lachnospiraceae [84] and Ruminococcaceae families [85]. The core microbiota emphasized the broad dietary similarities between leaf-eating monkeys and the co-evolution of gut microbes within colobines. The core also highlighted taxa (Ruminococcaceae and Lachnospiraceae) that may play an important role in colobine health. Altered abundances or reductions in these taxa could be contributing to captive colobine GI problems. Given the strong effect we observed of diet on the colobine gut microbial community, dietary modifications may be a promising way to improve captive colobine GI health. Future work comparing the gut microbial communities of wild and captive colobines as well as healthy and unhealthy colobines is needed to further assess the potential role of the gut microbiota and the core taxa in colobine health.
Microbial diversity and core microbiota analyses provided some support for our prediction based on primate phylogeny. Notably, the fact that primate phylogeny produced a signal—albeit not strong—across different regions and climates and despite different diets indicates that host evolution does indeed play a role in shaping the microbial community. However, overall, host phylogeny was a weaker predictor of gut microbial composition. Diet, on the other hand, was a stronger predictor of gut microbiota. PCoA, UPGMA, PERMANOVA, and supervised learning analyses supported our prediction based on diet: gut microbial communities were more similar in colobine species consuming the same diet. PCoA and UPGMA analyses demonstrated moderate clustering of samples by primate species and genera but strong clustering of samples by diet. Muegge et al. (2011) came to a similar conclusion while examining the effect of diet and host phylogeny on the gut microbiota of 33 mammalian species. Animals clustered by diet (herbivores, omnivores, and carnivores), but the gut microbiota did not reflect mammalian phylogeny. For example, colobus monkeys clustered with other herbivores like sheep and kangaroos, rather than with other primates such as chimpanzees or baboons, which are omnivores [11].
Similarly, in our study, colobine phylogeny was not reflected in our gut microbial analyses. The African colobine (Colobus) was the outgroup relative to the Asian colobines (Rhinopithecus, Trachypithecus, Pygathrix, Nasalis). African and Asian colobines diverged approximately 10 million years ago, and all subsequent divergences within the Asian colobine clade occurred more recently [86]. However, the Colobus samples did not diverge from the Asian colobines in PCoA clusters or in UPGMA branching. There are several possible explanations for this: (1) a long-term captive diet overwhelmed subtle or previous differences in Colobus gut microbiota or (2) folivory in primates is thought to have evolved from frugivory [17, 20]. If the common ancestor to both African and Asian colobines was frugivorous, then folivory may have evolved independently in both sets of colobines. Convergence would then explain similarities in gut microbial composition between African and Asian colobines. A recent study on distantly related myrmecophages (ant and termite eating mammals; e.g., anteaters, aardvarks, aardwolves) found convergence of the gut microbiota due to highly similar diets [10]. A similar result was found in a study of 283 ant species: distantly related herbivorous ant species shared more similar gut microbial communities with each other than they did with more closely related carnivorous ants [87]. In these cases, gradual diet change due to competition or changing resources for the host probably altered the gut microbial environment. In turn, the gut microbial community changed. Similarly, both African and Asian colobines may have begun eating unripe fruits, seeds, and young leaves in order to avoid competition with frugivores [20]. Colobine gut microbiota may have then adapted to this dietary change, and microbes that processed sugar were outcompeted by microbes that processed secondary compounds found in seeds and leaves. This, in turn, may have primed the colobine digestive system to degrade cellulose, hemicellulose, and lignin; thus, colobines could begin consuming mature leaves [20].
PERMANOVA and supervised learning analyses provided additional support for our prediction based on diet. Although error ratios were greater than 2 for analysis by diet and primate species, samples classified by diet had a higher error ratio (indicating lower classification error) than samples classified by primate species. Additionally, only diet was significant by PERMANOVA. These results, taken together, illustrate that the colobine gut microbial community is shaped by multiple factors. Diet played a large role in gut microbial composition, but host phylogeny could not be discounted. Another example of the interplay between diet and phylogeny comes from a controlled study on rodents: two closely related species of woodrats were captured from the wild and brought into captivity [51]. Both species were maintained under identical conditions and fed an identical diet. After 6 months, both species still had distinct gut microbial communities, and both retained over 50% of the OTUs they had as wild rodents [51]. In other words, ~50% of their gut microbiota was altered by captivity and a captive diet. The other ~50% was unaffected by the new environment and retained a likeness to their wild conspecifics, potentially due to host phylogeny or host genetics.
In our study, we used captive institute as a proxy for diet. We report the diets offered at each institute, but notably, diet offered may vary from the diet consumed and we did not track this information. Additionally, there are other features of captive institutes that could potentially contribute to the differences we observed in our study. For example, soil, air, or water microbes may differ between China and Singapore or between locations within China and Singapore. There may be different husbandry or sanitation practices at each institute that result in varying microbial exposures to the monkeys. And frequent visitors or tourists at some of the captive institutes may create stressors or increased potential for bacterial transmission from humans. Some studies suggest that free living or environmental sources of bacteria contribute little to the overall gut microbial community [50, 51], but we cannot rule this out as a possibility.
Conclusions
Our study highlights the importance of diet in shaping captive colobine gut microbiota. Differences in microbial diversity between Nasalis monkeys and other colobine genera and the presence of a core microbiota suggest the influence of host phylogeny on the gut microbiota. UPGMA, PCoA, PERMANOVA, and supervised learning analyses indicate the strong effect of diet on colobine gut flora. While we cannot change colobine phylogeny, we can certainly alter captive colobine diet in attempt to improve the GI health of these endangered primate species. This study provides a foundation for further work on colobine gut microbiota and for improving captive diets with both host and microbiota in mind.
Change history
11 September 2017
An erratum to this article has been published.
References
Ley R, Hamady M, Lozupone C, Turnbaugh PJ, Ramey RR, Bircher JS, Schlegel ML, Tucker TA, Schrenzel MD, Knight R, Gordon JI (2008) Evolution of mammals and their gut microbes Science 320:1647–1651
Zhang C, Zhang M, Wang S, Han R, Cao Y, Hua W, Mao Y, Zhang X, Pang X, Wei C, Zhao G, Chen Y, Zhao L (2010) Interactions between gut microbiota, host genetics and diet relevant to development of metabolic syndromes in mice ISME J 4:232–241. doi:10.1038/ismej.2009.112
Zhao L, Wang G, Siegel P, He C, Wang H, Zhao W, Zhai Z, Tian F, Zhao J, Zhang H, Sun Z, Chen W, Zhang Y, Meng H (2013) Quantitative genetic background of the host influences gut microbiomes in chickens Sci Rep 3. doi:10.1038/srep01163
McKnite AM, Perez-Munoz ME, Lu L, Williams EG, Brewer S, Andreux PA, Bastiaansen JWM, Wang X, Kachman SD, Auwerx J, Williams RW, Benson AK, Peterson DA, Ciobanu DC (2012) Murine gut microbiota is defined by host genetics and modulates variation of metabolic traits PLoS One 7:e39191. doi:10.1371/journal.pone.0039191
Blekhman R, Goodrich JK, Huang K, Sun Q, Bukowski R, Bell JT, Spector TD, Keinan A, Ley RE, Gevers D, Clark AG (2015) Host genetic variation impacts microbiome composition across human body sites Genome Biol 16:191. doi:10.1186/s13059-015-0759-1
Wang M, Radlowski EC, Monaco MH, Fahey GC, Gaskins HR, Donovan SM (2013) Mode of delivery and early nutrition modulate microbial colonization and fermentation products in neonatal piglets J Nutr 143:795–803. doi:10.3945/jn.112.173096
De Filippo C, Cavalieri D, Di Paola M, Ramazzotti M, Poullet J, Massart S, Collini S, Pieraccini G, Lionetti P (2010) Impact of diet in shaping gut microbiota revealed by a comparative study in children from Europe and rural Africa Proc Natl Acad Sci U S A 107:14691–14696. doi:10.1073/pnas.1005963107
Scott KP, Gratz SW, Sheridan PO, Flint HJ, Duncan SH (2013) The influence of diet on the gut microbiota Pharmacol Res 69:52–60. doi:10.1016/j.phrs.2012.10.020
Burin G, Kissling WD, Guimarães PR, Şekercioğlu ÇH, Quental TB (2016) Omnivory in birds is a macroevolutionary sink Nat Commun 7:11250. doi:10.1038/ncomms11250
Delsuc F, Metcalf JL, Parfrey LW, Song SJ, González A, Knight R (2013) Convergence of gut microbiomes in myrmecophagous mammals Mol Ecol 23:1301–1317. doi:10.1111/mec.12501
Muegge BD, Kuczynski J, Knights D, Clemente JC, González A, Fontana L, Henrissat B, Knight R, Gordon JI (2011) Diet drives convergence in gut microbiome functions across mammalian phylogeny and within humans Science 332:970–974. doi:10.1126/science.1198719
Ochman H, Worobey M, Kuo C-H, Ndjango J-BN, Peeters M, Hahn BH, Hugenholtz P (2010) Evolutionary relationships of wild hominids recapitulated by gut microbial communities PLoS Biol 8:e1000546. doi:10.1371/journal.pbio.1000546
Roeselers G, Mittge EK, Stephens WZ, Parichy DM, Cavanaugh CM, Guillemin K, Rawls JF (2011) Evidence for a core gut microbiota in the zebrafish ISME J 5:1595–1608. doi:10.1038/ismej.2011.38
Sanders JG, Powell S, Kronauer DJC, Vasconcelos HL, Frederickson ME, Pierce NE (2014) Stability and phylogenetic correlation in gut microbiota: lessons from ants and apes Mol Ecol 23:1268–1283. doi:10.1111/mec.12611
Tzeng T-D, Pao Y-Y, Chen P-C, Weng FC-H, Jean WD, Wang D (2015) Effects of host phylogeny and habitats on gut microbiomes of oriental river prawn (Macrobrachium nipponense) PLoS One 10:e0132860. doi:10.1371/journal.pone.0132860
Carrillo-Araujo M, Taş N, Alcántara-Hernández RJ, Gaona O, Schondube JE, Medellín RA, Jansson JK, Falcón LI (2015) Phyllostomid bat microbiome composition is associated to host phylogeny and feeding strategies Front Microbiol 6:447. doi:10.3389/fmicb.2015.00447
Chivers D (1994) Functional anatomy of the gastrointestinal tract. In: Davies AG, Oates JF (eds) Colobine monkeys: their ecology, behaviour and evolution. Cambridge University Press, Cambridge, pp. 205–227
Kay RNB, Davies AG (1994) Digestive physiology. In: Davies AG, Oates J (eds) Colobine monkeys: their ecology, behavior and evolution. Cambridge University Press, Cambridge, pp. 229–249
Caton JM (1998) The morphology of the gastrointestinal tract of Pygathrix nemaeus (Linneaus, 1771). In: Jablonski NG (ed) The natural history of the doucs and snub-nosed monkeys. World Scientific Publishing Co., Singapore, pp. 129–154
Lambert JE (1998) Primate digestion: interactions among anatomy, physiology, and feeding ecology Evol Anthropol:8–20
Nijboer J, Clauss M (2006) The digestive physiology of colobine primates. In: Nijboer J (ed) Fibre intake and faeces quality in leaf-eating primates. Utrecht Publishing and Archiving Service, The Netherlands, pp. 9–28
Bauchop T, Martucci RW (1968) Ruminant-like digestion of the langur monkey Science 161:698–700
Yildirim S, Yeoman C, Sipos M, Torralba M, Wilson B, Goldberg T, Stumpf R, Leigh S, White B, Nelson K (2010) Characterization of the fecal microbiome from non-human wild primates reveals species specific microbial communities PLoS One 5:e13963
Wu C, Yang F, Gao R, Huang Z, Xu B, Dong Y, Hong T, Tang X (2010) Study of fecal bacterial diversity in Yunnan snub-nosed monkey (Rhinopithecus bieti) using phylogenetic analysis of cloned 16S rRNA gene sequences Af J Biotech 9:6278–6289
Amato KR, Metcalf JL, Song SJ, Hale VL, Clayton J, Ackermann G, Humphrey G, Niu K, Cui D, Zhao H, Schrenzel MD, Tan CL, Knight R, Braun J (2016) Using the gut microbiota as a novel tool for examining colobine primate GI health Glob Ecol Conserv 7:225–237. doi:10.1016/j.gecco.2016.06.004
Edwards M (1997) Leaf-eating primates: nutrition and dietary husbandry. Nutrition Advisory Group Handbook
Agoramoorthy G, Alagappasamy C, Hsu MJ (2004) Can proboscis monkeys be successfully maintained in captivity? A case of swings and roundabouts Zoo Biol 23:433–544. doi:10.1002/zoo.20018
Davies AG, Oates J (1994) Colobine monkeys: their ecology, behavior, and evolution. Cambridge University Press, New York,
Sutherland-Smith M, Janssen D, Lowenstine L (1998) Gastric analyses of colobine primates. AAZV Conference Proceedings: 136–139
Clayton JB, Vangay P, Huang H, Ward T, Hillmann BM, Al-Ghalith GA, Travis DA, Long HT, Tuan BV, Minh VV, Cabana F, Nadler T, Toddes B, Murphy T, Glander KE, Johnson TJ, Knights D (2016) Captivity humanizes the primate microbiome Proc Natl Acad Sci 113:10376–10381. doi:10.1073/pnas.1521835113
Amato K, Yeoman C, Kent A, Righini N, Carbonero F, Estrada A, Gaskins H, Stumpf R, Yildirim S, Torralba M, Gillis M, Wilson B, Nelson K, White B, Leigh S (2013) Habitat degradation impacts black howler monkey (Alouatta pigra) gastrointestinal microbiomes ISME J 7:1344–1353. doi:10.1038/ismej.2013.16
Wang XP, Yu L, Roos C, Ting N, Chen CP, Wang J, Zhang Y (2012) Phylogenetic relationships among the colobine monkeys revisited: new insights from analyses of complete mt genomes and 44 nuclear non-coding markers PLoS One 7:e36274
Liedigk R, Yang M, Jablonski NG, Momberg F, Geissmann T, Lwin N, Hla TH, Liu Z, Wong B, Ming L, Yongcheng L, Zhang Y-PP, Nadler T, Zinner D, Roos C (2012) Evolutionary history of the odd-nosed monkeys and the phylogenetic position of the newly described Myanmar snub-nosed monkey Rhinopithecus strykeri PLoS One 7:e37418. doi:10.1371/journal.pone.0037418
Hale VL, Tan CL, Knight R, Amato KR (2015) Effect of preservation method on spider monkey (Ateles geoffroyi) fecal microbiota over 8 weeks J Microbiol Methods 113:16–26. doi:10.1016/j.mimet.2015.03.021
Hale VL, Tan CL, Niu K, Yang Y, Cui D, Zhao H, Knight R, Amato KR (2016) Effects of field conditions on fecal microbiota J Microbiol Methods 130:180–188. doi:10.1016/j.mimet.2016.09.017
Gilbert JA, Folker M, Dion A, Pavan B, Brown CT, Christopher TB, Narayan D, Jonathan AE, Dirk E, Dawn F, Wu F, Daniel H, Janet J, Rob K, James K, Eugene K, Kostas K, Joel K, Nikos K, Rachel M, Alice M, Christopher Q, Jeroen R, Alexander S, Ashley S, Rick S (2010) Meeting report: the terabase metagenomics workshop and the vision of an Earth Microbiome Project Stand Genomic Sci 3:243–248. doi:10.4056/sigs.1433550
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Pena AG, Goodrich JK, Gordon JI, Huttley GA, Kelley ST, Knights D, Koenig JE, Ley RE, Lozupone CA, McDonald D, Muegge BD, Pirrung M, Reeder J, Sevinsky JR, Turnbaugh PJ, Walters WA, Widmann J, Yatsunenko T, Zaneveld J, Knight R (2010) QIIME allows analysis of high-throughput community sequencing data Nat Methods 7:335–336
McDonald D, Price MN, Goodrich J, Nawrocki EP, DeSantis TZ, Probst A, Andersen GL, Knight R, Hugenholtz P (2011) An improved Greengenes taxonomy with explicit ranks for ecological and evolutionary analyses of bacteria and archaea ISME J 6:610–618. doi:10.1038/ismej.2011.139
Caporaso JG, Bittinger K, Bushman FD, DeSantis TZ, Andersen GL, Knight R (2010) PyNAST: a flexible tool for aligning sequences to a template alignment Bioinformatics 26:266–267. doi:10.1093/bioinformatics/btp636
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy Appl Environ Microbiol 73:5261–5267. doi:10.1128/aem.00062-07
Mackie RI, Sghir A, Gaskins HR (1999) Developmental microbial ecology of the neonatal gastrointestinal tract Am J Clin Nutr 69:1035S–1045S
Spor A, Koren O, Ley R (2011) Unravelling the effects of the environment and host genotype on the gut microbiome Nat Rev Microbiol 9:279–290. doi:10.1038/nrmicro2540
Lozupone C, Knight R (2005) UniFrac: a new phylogenetic method for comparing microbial communities Appl Environ Microbiol 71:8228–8235. doi:10.1128/aem.71.12.8228-8235.2005
Knights D, Costello E, Knight R (2011) Supervised classification of human microbiota FEMS Microbiol Rev 35:343–359. doi:10.1111/j.1574-6976.2010.00251.x
Breiman L (2001) Random forests Mach Learn 45:5–32. doi:10.1023/a:1010933404324
Oksanen J, Blanchet FG, Kindt R, Legendre P, Minchin PR, O'Hara RB, Simpson GL, Solymos P, Stevens MHH, Wagner H (2013) vegan: community ecology package. R package version 20-10
Soderholm JD, Perdue MH (2001) Stress and intestinal barrier function Am J Phys 280:G7–G13
O'Mahony SM, Marchesi JR, Scully P, Codling C, Ceolho A-M, Quigley EMM, Cryan JF, Dinan TG (2009) Early life stress alters behavior, immunity, and microbiota in rats: implications for irritable bowel syndrome and psychiatric illnesses Biol Psychiatry 65:263–267
Anderson K, Russell J, Moreau C, Kautz S, Sullam K, Hu Y, Basinger U, Mott B, Buck N, Wheeler D (2012) Highly similar microbial communities are shared among related and trophically similar ant species Mol Ecol 21:2282–2296. doi:10.1111/j.1365-294X.2011.05464.x
Nelson T, Rogers T, Carlini A, Brown M (2013) Diet and phylogeny shape the gut microbiota of Antarctic seals: a comparison of wild and captive animals Environ Microbiol 15:1132–1145. doi:10.1111/1462-2920.12022
Kohl KD, Skopec MM, Dearing MD (2014) Captivity results in disparate loss of gut microbial diversity in closely related hosts Conserv Physiol 2:cou009. doi:10.1093/conphys/cou009
Sullam K, Essinger S, Lozupone C, O'Connor M, Rosen G, Knight R, Kilham S, Russell J (2012) Environmental and ecological factors that shape the gut bacterial communities of fish: a meta-analysis Mol Ecol 21:3363–3378. doi:10.1111/j.1365-294X.2012.05552.x
Colman D, Toolson E, Takacs-Vesbach C (2012) Do diet and taxonomy influence insect gut bacterial communities? Mol Ecol 21:5124–5137. doi:10.1111/j.1365-294X.2012.05752.x
Xiang Z-F, Liang W-B, Nie S-G, Li M (2012) Diet and feeding behavior of Rhinopithecus brelichi at Yangaoping, Guizhou Am J Primatol 74:551–560
Bleisch WV, Jiahua X (1998) Ecology and behavior of the Guizhou snub-nosed langur (Rhinopithecus [Rhinopithecus] brelichi), with a discussion of socioecology in the genus. In: Jablonski NG (ed) The natural history of the doucs and snub-nosed monkeys. World Scientific Publishing, Singapore,
Guo S, Li B, Watanabe K (2007) Diet and activity budget of Rhinopithecus roxellana in the Qinling Mountains, China Primates 48:268–276. doi:10.1007/s10329-007-0048-z
Ding W, Zhao Q-K (2004) Rhinopithecus bieti at Tacheng, Yunnan: diet and daytime activities Int J Primatol 25:583–598. doi:10.1023/B:IJOP.0000023576.60883.e5
Bennett EL, Davies AG (1994) The ecology of Asian colobines. In: Davies AG, Oates J (eds) Colobine monkeys: their ecology, behavior, and evolution. Cambridge University Press, Cambridge,
Oates J (1994) The natural history of African colobines. Cambridge University Press, Cambridge,
Yeager CP, Kool K (2000) The behavioral ecology of Asian colobines. Cambridge University Press, Cambridge,
Rawson BM (2006) Activity budgets in black-shanked douc langurs (Pygathrix nigripes) Int J Primatol 27(Suppl 1) Abstract 307
Duc HM, Baxter GS, Page MJ (2009) Diet of Pygathrix nigripes in southern Vietnam Int J Primatol 30:15–28. doi:10.1007/s10764-008-9325-y
Lippold LK (1998) Natural history of douc langurs. In: Jablonski NG (ed) The natural history of the doucs and snub-nosed monkeys. World Scientific Publishing, Singapore,
Tinh NT, Long HT, Tuan BV, Vy TH, NA T (2012) The feeding behaviour and phytochemical food content of grey-shanked douc langurs (Pygathrix cinerea) at Kon Ka Kinh National Park, Vietnam Vietnamese J Primatol 2:25–35
Matsuda I, Tuuga A, Higashi S (2009) The feeding ecology and activity budget of proboscis monkeys Am J Primatol 71:478–492. doi:10.1002/ajp.20677
Raven PH, Evert RF, Eichhorn SE (2005) Biology of plants. W.H. Freeman & Company, New York,
McKey D (1974) Adaptive patterns in alkaloid physiology Am Nat 108:305–320
Hladik CM (1978) Adaptive strategies of primates in relation to leaf eating. Smithsonian Institution Press, Washington,
Sonnenburg ED, Smits SA, Tikhonov M, Higginbottom SK, Wingreen NS, Sonnenburg JL (2016) Diet-induced extinction in the gut microbiota compounds over generations Nature 529:212–215. doi:10.1038/nature16504
Matsuda I, Sha JCM, Ortmann S, Schwarm A, Grandl F, Caton J, Jens W, Kreuzer M, Marlena D, Hagen KB, Clauss M (2015) Excretion patterns of solute and different-sized particle passage markers in foregut-fermenting proboscis monkey (Nasalis larvatus) do not indicate an adaptation for rumination Physiol Behav 149:45–52. doi:10.1016/j.physbeh.2015.05.020
Matsuda I, Tuuga A, Hashimoto C, Bernard H, Yamagiwa J, Fritz J, Tsubokawa K, Yayota M, Murai T, Iwata Y, Clauss M (2014) Faecal particle size in free-ranging primates supports a ‘rumination’ strategy in the proboscis monkey (Nasalis larvatus) Oecologia 174:1127–1137. doi:10.1007/s00442-013-2863-9
Russell JB, Rychlik JL (2001) Factors that alter rumen microbial ecology Science 292:1119–1122. doi:10.1126/science.1058830
Ley RE, Turnbaugh P, Klein S, Gordon JI (2006) Microbial ecology: human gut microbes associated with obesity Nature 444:1022–1023. doi:10.1038/4441022a
Willing B, Voros A, Roos S, Jones C, Jansson A, Lindberg JE (2009) Changes in faecal bacteria associated with concentrate and forage-only diets fed to horses in training Equine Vet J 41:908914. doi:10.2746/042516409x447806
Mackie R, Aminov R, Hu W, Klieve A, Ouwerkerk D, Sundset M, Kamagata Y (2003) Ecology of uncultivated Oscillospira species in the rumen of cattle, sheep, and reindeer as assessed by microscopy and molecular approaches Appl Environ Microbiol 69:6808–6815
Clarke R (1979) Niche in pasture-fed ruminants for the large rumen bacteria Oscillospira, Lampropedia, and Quin's and Eadie's ovals Appl Environ Microbiol 37:654–657
Amato KR, Leigh SR, Kent A, Mackie RI, Yeoman CJ, Stumpf RM, Wilson BA, Nelson KE, White BA, Garber PA (2015) The gut microbiota appears to compensate for seasonal diet variation in the wild black howler monkey (Alouatta pigra) Microb Ecol 69:434–443. doi:10.1007/s00248-014-0554-7
Biddle A, Stewart L, Blanchard J, Leschine S (2013) Untangling the genetic basis of fibrolytic specialization by Lachnospiraceae and Ruminococcaceae in diverse gut communities Diversity 5:627–640. doi:10.3390/d5030627
Barcenilla A, Pryde SE, Martin JC, Duncan SH, Stewart CS, Henderson C, Flint HJ (2000) Phylogenetic relationships of butyrate-producing bacteria from the human gut Appl Environ Microbiol 66:1654–1661. doi:10.1128/aem.66.4.1654-1661.2000
Velazquez OC, Lederer HM, Rombeau JL (1997) Butyrate and the colonocyte: production, absorption, metabolism, and therapeutic implications. In: Kritchevsky D, Bonfield C (eds) Dietary fiber in health and disease. Plenum Press Div Plenum Publishing Corp, New York, pp. 123–134
Donohoe DR, Garge N, Zhang X, Sun W, O’Connell TM, Bunger MK, Bultman SJ (2011) The microbiome and butyrate regulate energy metabolism and autophagy in the mammalian colon Cell Metab 13:517–526. doi:10.1016/j.cmet.2011.02.018
Brahe LK, Astrup A, Larsen LH (2013) Is butyrate the link between diet, intestinal microbiota and obesity-related metabolic diseases? Obes Rev 14:950–959. doi:10.1111/obr.12068
Hong J, Jia Y, Pan S, Jia L, Li H, Han Z, Cai D, Zhao R (2016) Butyrate alleviates high fat diet-induced obesity through activation of adiponectin-mediated pathway and stimulation of mitochondrial function in the skeletal muscle of mice Oncotarget 7:56071–56082. doi:10.18632/oncotarget.11267
Frank D, St Amand A, Feldman R, Boedeker E, Harpaz N, Pace N (2007) Molecular-phylogenetic characterization of microbial community imbalances in human inflammatory bowel diseases Proc Natl Acad Sci U S A 104:13780–13785. doi:10.1073/pnas.0706625104
Fujimoto T, Imaeda H, Takahashi K, Kasumi E, Bamba S, Fujiyama Y, Andoh A (2013) Decreased abundance of Faecalibacterium prausnitzii in the gut microbiota of Crohn's disease J Gastroenterol Hepatol 28:613–619. doi:10.1111/jgh.12073
Stewart C-B, Disotell TR (1998) Primate evolution—in and out of Africa Curr Biol 8:R582–R588. doi:10.1016/S0960-9822(07)00367-3
Russell J, Moreau C, Goldman-Huertas B, Fujiwara M, Lohman D, Pierce NE (2009) Bacterial gut symbionts are tightly linked with the evolution of herbivory in ants Proc Natl Acad Sci U S A 106:21236–21241. doi:10.1073/pnas.0907926106
Acknowledgements
We would like to thank Richard D. Howard for his support of this project, thoughtful suggestions on analysis, and review of this manuscript. Krista Nichols kindly provided laboratory space for the molecular work. We also thank Bong Suk-Kim and Gaenna Rogers for their assistance in the laboratory. Finally, we thank the reviewers for the time and thought they put into providing helpful feedback on this manuscript. This project was funded by the Margot Marsh Biodiversity Foundation (VLH, CLT), San Diego Zoo Global, the Offield Family Foundation, the Earth Microbiome Project, the Howard Hughes Medical Institute (RK), Purdue College of Veterinary Medicine International Programs (VLH), and Fanjingshan National Nature Reserve. VLH was supported by a Purdue University Andrews Fellowship and a Purdue Research Foundation Research Grant.
Author information
Authors and Affiliations
Corresponding author
Additional information
An erratum to this article is available at https://doi.org/10.1007/s00248-017-1070-3.
Rights and permissions
About this article
Cite this article
Hale, V.L., Tan, C.L., Niu, K. et al. Diet Versus Phylogeny: a Comparison of Gut Microbiota in Captive Colobine Monkey Species. Microb Ecol 75, 515–527 (2018). https://doi.org/10.1007/s00248-017-1041-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-017-1041-8