Abstract
Flooded rice fields are important sources of atmospheric methane. Aerobic methanotrophs living in the vicinity of rice roots oxidize methane and act as environmental filters. Here, we present genome characteristics of a gammaproteobacterial methanotroph, isolate Sn10-6, which was isolated from a rice rhizosphere of a flooded field in India. Sn10-6 has been identified as a member of a putative novel genus and species within the family Methylococcaceae (Type I methanotrophs). The draft genome of Sn10-6 showed pathways for the following: methane oxidation, formaldehyde assimilation (RuMP), nitrogen fixation, conversion of nitrite to nitrous oxide, and other interesting genes including the ones responsible for survival in the rhizosphere environment. The majority of genes found in this genome were most similar to Methylovulum miyakonese which is a forest isolate. This draft genome provided insight into the physiology, ecology, and phylogeny of this gammaproteobacterial methanotroph.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
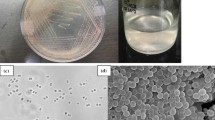
Flooded rice fields are important sources of methane, which is the second most important greenhouse gas responsible for global warming [1]. Aerobic methanotrophs dwelling in the vicinity of rice roots oxidize methane and thus play an important role in methane mitigation [2]. Classical methanotrophs are classified into two groups (Type I and Type II) methanotrophs and belong to Gammaproteobacteria and Alphaproteobacteria, respectively [3]. Recently, methanotrophs have also been reported from Verrucomicrobial phylum; however, these species are mostly present in acidic habitats [4, 5]. Though Type I methanotrophs play an important role in methane oxidation in the rhizospheric environments of flooded rice fields, very few members have been isolated and characterized and described from such habitats. To the best of our knowledge, only two species of gammaproteobacterial methanotrophs (Type I) have been formally described from rice field environments [6, 7]. These are Methylogaea oryzae which was isolated from rice fields in Uruguay [8] and Methylomonas koyamae which was isolated from Japanese rice fields [9]. Genome information for either of these species is not yet available. In our recent work, we enriched, isolated and characterized methanotrophs from a rice rhizosphere environment in India (Pandit, P., Rahalkar, M. and others, accepted). Isolate Sn10-6 was isolated from a dilution series liquid enrichment (10−6 dilution) of a rice rhizospheric soil sample incubated in presence of methane, air, and nitrogen. The sample was collected from a flooded rice field in Western India. Blast analysis of the 16S rRNA gene of Sn10-6 (KP793700.1) showed only 90–93.5 % similarity with Type I methanotrophic genera and hence represented a member of a putative novel genus and species within Methylococcaceae. Using EzTaxon server (http://www.ezbiocloud.net/eztaxon/), the closest neighbor was found to be Methylosarcina lacus LW14T (93.5 %). After phylogenetic analysis of 16S rRNA genes, Sn10-6 belonged to a distinct branch next to Methylovulum-Methylosoma clade.
We sequenced the genome of Sn10-6 in order to get a clearer idea of its taxonomy, physiology, and ecology. Moreover, Sn10-6 is one of the first gammaproteobacterial methanotroph recovered from rice rhizospheres of flooded paddy fields in a tropical country like India. Any information about the metabolism of strain Sn10-6 that can be obtained from analysis of the genome will also help us understand its role in biogeochemical cycling in flooded rice field ecosystems. We additionally investigated the genome for genes which might be helping the organism to survive in the rhizosphere environment. Thus, the main aim of this paper was to study the salient characteristics of the genome of a novel gammaproteobacterial (Type I) methanotroph which was isolated from a flooded rice field in India.
After DNA extraction and confirmation of the 16S rRNA gene sequence, whole genome sequencing of Sn10-6 was done using the Ion Torrent PGM (200 bp) with the 316TM sequencing chip. We performed de novo assembly using the MIRA 4.0.5 assembler [10] which generated 389 contigs. It was revealed that genome size of Sn10-6 is 4.58 Mb with a G + C content of 43.9 % (Table 1). The longest contig was 91,582 bp with N50 of 19,087 bp and N90 4856 bp. This data was then submitted to the Rapid Annotation using Subsystem Technology (RAST server) [11, 12] and NCBI for annotation. Annotation by NCBI revealed presence of 3413 proteins. We additionally used BlastKOALA (http://www.kegg.jp/blastkoala/) for prediction of KEGG pathways. Single copy of SSU rRNA and LSU rRNA were found in the genome along with 43 tRNAs (Table 1). Using the RAST server, the closest genome match was Methylobacter tundripaludum (genome id: 697282.3) with a score of 524.
The genome of strain Sn10-6 showed all the necessary machinery required for methane metabolism (Fig. 1). Single copy of the pmo CAB gene cluster encoding particulate methane monooxygenase enzyme (pMMO), enzyme responsible for conversion of methane to methanol, the first step in the methane oxidation pathway, was found. Additionally, many of the methanotrophs also show a homologue of pmo CAB, denoted as pmx ABC operon [13]. Genes for pmx ABC and the soluble methane monooxygenase (sMMO) were both missing from the genome. sMMO is present only in a few genera within Type I methanotrophs and is mostly present in Type X and Type II methanotrophs [3]. pMMO genes in Sn10-6 were most similar to genes in M. miyakonese and M. tundripaludum as revealed by blastx analysis (Table 2). The second step in the pathway is conversion of methanol to formaldehyde by the enzyme methanol dehydrogenase. Genes coding for all subunits of methanol dehydrogenase showed highest homology to those from M. miyakonese (Table 2). Next step in the pathway is oxidation of formaldehyde to formate. In Sn10-6, the formaldehyde oxidation occurs probably by the tetrahydromethanopterin (H4MPT) pathway. Various enzymes for the H4MPT pathway were detected which included 5,10-methylenetetrahydrofolate reductase (EC 1.5.1.20), methylene tetrahydromethanopterin dehydrogenase (EC 1.5.99.9), formate-tetrahydrofolate ligase (EC 6.3.4.3), formiminotetrahydrofolate cyclodeaminase (EC 4.3.1.4) and N(5), N(10)-methenyltetrahydromethanopterin cyclohydrolase (EC 3.5.4.27). Tetrahydromethanopterin pathway has been discovered as one of the ancient and major pathways for formaldehyde oxidation [14] and is found to be present in some of the methanotrophic and methylotrophic genome sequences described recently [15]. The last step in the methane oxidation pathway is encoded by formate dehydrogenase enzyme. Formate dehydrogenase in Sn10-6 was most similar to that of Methylovulum miyakonese. Formaldehyde acts as the central intermediate in methanotrophs and is used for assimilation. In Type I methanotrophs, formaldehyde assimilation normally takes place via ribulose mono phosphate (RuMP) pathway [3]. Genes for RuMP pathway were found in Sn10-6 genome confirming that it was a typical Type I methanotroph (Fig. 1). Even though methanotrophs can use only C1 compounds (methane and in some cases methanol) as their carbon and energy source, genes for glycolysis, tricarboxylic acid (TCA) cycle are present in many species [13]. Similarly, in Sn10-6, genes for glycolysis, pentose phosphate pathway, TCA cycle were present. Similarly, genes encoding acetate kinase and acetyl Co-A synthetase were also detected. Many of the genes in the carbon metabolism were most related to the genes of M. miyakonese.
Genes related to methane metabolism as detected in Sn10-6 (shown in green). This figure was generated by submitting the assembled contigs (amino acids) to BlastKOALA (http://www.kegg.jp/blastkoala/) for prediction of KEGG pathways
We detected genes that coded for proteins involved in nitrogen metabolism which included genes for assimilatory nitrate reduction, nitrogen fixation, nitrite to nitrous oxide conversion, and urea or ammonium utilization (Fig. 2). Although dissimilatory nitrate reduction genes were absent in the genome, Sn10-6 shows genes required for conversion of nitrite to nitrous oxide. Genes of this pathway have been detected in M. miyakonese, Methylomicrobium alcaliphilum [15] and Methylomonas methanica MC09 [16] (Table 2). Nitrogen fixation can be an important process for bacteria living in rhizospheres with respect to their interactions with plants. The metabolic pathway for the conversion of nitrite to nitrous oxide in Sn10-6 was also noteworthy. Most of the nitrogen metabolism genes were also most related to M. miyakonese (nifH genes, nitric oxide reductase, and urease) and a few to M. tundripaludum (assimilatory nitrate reductase).
Genes related to nitrogen metabolism as detected in Sn10-6 (shown in green). This figure was generated by submitting the assembled contigs (amino acids) to BlastKOALA (http://www.kegg.jp/blastkoala/) for prediction of KEGG pathways
In addition, genes coding for type IV pili, flagella, and chemotaxis were also detected. These genes could especially be important for the motility of the organism according to the changing methane and oxygen gradients within the rhizosphere microenvironment. We also found another interesting genes, i.e., MazF/MazE genes which are responsible for toxin-antitoxin genes involved in programmed cell death occurring after nutrient starvation [17, 18]. After blast analysis, MazF (toxin) genes in Sn10-6 were found most related to that of M. miyakonese. None of the other gammaproteobacterial methanotrophs (other than M. miyakonese and Sn10-6) have been reported to have MazE/MazF, and hence, their function in these two methanotrophs warrants further investigation. Nutrient starvation might be a common situation for the rice field methanotrophs during the dry periods of the year [19], and therefore, these genes would be worth of further exploration. Apart from these genes, we also detected genes for carotenoid pigment, flagellar apparatus, and rod shape which explain the morphology of this pale pink pigmented motile methanotroph.
Thus, studying the draft genome of Sn10-6 gave us important insights into the metabolism and helped us to understand its lifestyle as a rice rhizosphere associated gammaproteobacterial methanotroph living in paddy fields of a tropical country.
References
Bachelet D, Neue H-U (1993) Methane emissions from wetland rice areas of Asia. Chemosphere 26:219–237
Conrad R (2009) The global methane cycle: recent advances in understanding the microbial processes involved. Environ Microbiol Rep 1:285–292
Hanson RS, Hanson TE (1996) Methanotrophic bacteria. Microbiol Mol Biol Rev 60:439–471
Islam T, Jensen S, Reigstad LJ, Larsen Ø, Birkeland N-K (2008) Methane oxidation at 55°C and pH 2 by a thermoacidophilic bacterium belonging to the Verrucomicrobia phylum. Proc Natl Acad Sci U S A 105:300–304
Pol A, Heijmans K, Harhangi HR, Tedesco D, Jetten MSM, Camp HJMOd (2007) Methanotrophy below pH 1 by a new Verrucomicrobia species. Nature 450:874–878
Dianou D, Ueno C, Ogiso T, Kimura M, Asakawa S (2012) Diversity of cultivable methane-oxidizing bacteria in microsites of a rice paddy field: investigation by cultivation method and fluorescence in situ hybridization (FISH). Microbes Environ 27:278–287
Ferrando L, Tarlera S (2009) Activity and diversity of methanotrophs in the soil-water interface and rhizospheric soil from a flooded temperate rice field. J Appl Microbiol 106:306–316
Geymonat E, Ferrando L, Tarlera SE (2011) Methylogaea oryzae gen. nov., sp. nov., a mesophilic methanotroph isolated from a rice paddy field. Int J Syst Evol Microbiol 61:2568–2572
Ogiso T, Ueno C, Dianou D, Huy TV, Katayama A, Kimura M, Asakawa S (2012) Methylomonas koyamae sp. nov., a type I methane-oxidizing bacterium from floodwater of a rice paddy field. Int J Syst Evol Microbiol 62:1832–1837
Chevreux B, Wetter T, Suhai S (1999) Genome sequence assembly using trace signals and additional sequence information. Computer Science and Biology Proceedings of the German Conference on Bioinformatics (GCB), vol. 99, Hanover, pp. 45–56.
Aziz RK, Bartels D, Best AA, DeJongh M, Disz T, Edwards RA, Formsma K, Gerdes S, Glass EM, Kubal M, Meyer F, Olsen GJ, Olson R, Osterman AL, Overbeek RA, McNeil LK, Paarmann D, Paczian T, Parrello B, Pusch GD, Reich C, Stevens R, Vassieva O, Vonstein V, Wilke A, Zagnitko O (2008) The RAST Server: rapid annotations using subsystems technology. BMC Genomics 9:75
Overbeek R, Olson R, Pusch GD, Olsen GJ, Davis JJ, Disz T, Edwards RA, Gerdes S, Parrello B, Shukla M, Vonstein V, Wattam AR, Xia F, Stevens R (2014) The SEED and the Rapid Annotation of microbial genomes using Subsystems Technology (RAST). Nucleic Acids Res 42:D206–214
Hamilton R, Kits KD, Ramonovskaya VA, Rozova ON, Yurimoto H, Iguchi H, Khmelenina VN, Sakai Y, Dunfield PF, Klotz MG, Knief C, Camp HJMO, Jetten MSM, Bringel F, Vuilleumier S, Svenning MM, Shapiro N, Woyke T, Trotsenko YA, Stein LY, Kalyuzhnayaa MG (2015) Draft genomes of gammaproteobacterial methanotrophs isolated from terrestrial ecosystems. Genome Announc 3:1–3
Kalyuzhnaya MG, Korotkova N, Crowther G, Marx CJ, Lidstrom ME, Chistoserdova L (2005) Analysis of gene islands involved in methanopterin-linked C1 transfer reactions reveals new functions and provides evolutionary insights. J Bacteriol 187:4607–4614
Vuilleumier S, Khmelenina VN, Bringel F, Reshetnikov AS, Lajus A, Mangenot S, Rouy Z, Op den Camp HJ, Jetten MS, Dispirito AA, Dunfield P, Klotz MG, Semrau JD, Stein LY, Barbe V, Medigue C, Trotsenko YA, Kalyuzhnaya MG (2012) Genome sequence of the haloalkaliphilic methanotrophic bacterium Methylomicrobium alcaliphilum 20Z. J Bacteriol 194:551–552
Boden R, Cunliffe M, Scanlan J, Moussard H, Kits KD, Klotz MG, Jetten MS, Vuilleumier S, Han J, Peters L, Mikhailova N, Teshima H, Tapia R, Kyrpides N, Ivanova N, Pagani I, Cheng JF, Goodwin L, Han C, Hauser L, Land ML, Lapidus A, Lucas S, Pitluck S, Woyke T, Stein L, Murrell JC (2011) Complete genome sequence of the aerobic marine methanotroph Methylomonas methanica MC09. J Bacteriol 193:7001–7002
Aizenman E, Engelberg-Kulkat H, Glaser G (1996) An Escherichia coli chromosomal "addiction module" regulated by 3′,5′-bispyrophosphate: a model for programmed bacterial cell death and differentiation. Proc Natl Acad Sci USA 93:6059–6063
Katsuhiko K, Hanaoka F, Burley S (2003) Crystal structure of the MazE/MazF complex: molecular bases of antidote-toxin recognition. Mol Cell 11:875–884
Roslev P, King GM (1995) Aerobic and anaerobic starvation metabolism in methanotrophic bacteria. Appl Environ Microbiol 61:1563–1570
Acknowledgments
This work was supported by Department of Biotechnology under the DBT BioCARe programme (BT/BioCARe/06/840/2012) sanctioned to M. C. R and fellowship sanctioned to P. S. P. is acknowledged. CSIR research fellowship to Preeti Arora and Soham Pore is sincerely acknowledged.
Nucleotide sequence accession number
The draft genome sequence of Methylococcaceae bacterium strain Sn10-6 is deposited in the GenBank database under the accession number LAJX00000000.1 and the bioproject is registered as PRJNA278928.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Rahalkar, M.C., Pandit, P.S., Dhakephalkar, P.K. et al. Genome Characteristics of a Novel Type I Methanotroph (Sn10-6) Isolated from a Flooded Indian Rice Field. Microb Ecol 71, 519–523 (2016). https://doi.org/10.1007/s00248-015-0699-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-015-0699-z