Abstract
The ocean is a natural habitat for antibiotic-producing bacteria, and marine aquaculture introduces antibiotics into the ocean to treat infections and improve aquaculture production. Studies have shown that the ocean is an important reservoir of antibiotic resistance genes. However, there is a lack of understanding and knowledge about the clinical importance of the ocean resistome. We investigated the relationship between the ocean bacterial resistome and pathogenic resistome. We applied high-throughput sequencing and metagenomic analyses to explore the resistance genes in bacterial plasmids from marine sediments. Numerous putative resistance determinants were detected among the resistance genes in the sediment bacteria. We also found that several contigs shared high identity with transposons or plasmids from human pathogens, indicating that the sediment bacteria recently contributed or acquired resistance genes from pathogens. Marine sediment bacteria could play an important role in the global exchange of antibiotic resistance.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The discovery of antimicrobial agents was one of the great triumphs of the twentieth century. However, we have moved from the age of antibiotics to the age of antibiotic resistance. The phenomenon of antibiotic resistance has snowballed into a serious public health concern. We are now at a crossroad where the antibiotics that we have relied on are becoming less effective and with the limited number of drugs under investigation. Fortunately, a few solutions are on the horizon. One of them is to understand the origin of antibiotic resistance genes and the gene pool of resistance genes. The concept of an “antibiotic resistome” was proposed in 2006 [1]. The resistome includes not only the resistance determinants of pathogenic bacteria but also non-pathogenic bacteria in environments such as soil. Numerous studies have shown that soil is a reservoir of antibiotic resistance, and a recent study provided evidence that soil has contributed to or acquired resistance determinants from pathogens [2]. Compared to soil, the resistome of marine bacteria has not been well documented. The ocean is a natural habitat of antibiotic-producing bacteria, and marine aquaculture introduces antibiotics into the ocean to treat infections and improve aquaculture production. However, the clinical importance of the ocean resistome remains poorly defined.
Horizontal gene transfer (HGT) is a common mechanism for the exchange of genetic information between bacterial species. Plasmids, as active mobile genetic elements, are transferred by transformation and conjugation. Plasmids carry various accessory genes that often confer an advantage to their host’s adaptation to different ecological niches [3]. Resistance genes are common accessory genes carried by plasmids, which facilitate their spread. To date, several studies have examined the distribution of resistance genes in plasmid communities of activated sludge using a metagenomic approach [4, 5]. These studies revealed the diversity and abundance of resistance genes and genetic elements in plasmids isolated from activated sludge. We also applied a metagenomic approach to explore the plasmids in sediment samples from a marine fish farm. The resistance genes on the plasmids were studied to decipher the relationship between the ocean bacterial resistome and pathogenic resistome.
In this study, we collected plasmids from cultured microbial communities of marine sediment. By high-throughput sequencing, we detected 58 resistance genes conferring resistance to 11 antibiotics. Our results demonstrated that marine sediment is a reservoir of antibiotic resistance genes. We also found that more than 40,000 reads were similar to genes from pathogenic isolates; 59 % of these reads were 100 % identical. Additionally, several contigs carrying antibiotic resistance genes shared >90 % identity with transposons or plasmids from human pathogens, encompassing coding and non-coding regions. High nucleotide sequence identity between sediment bacteria and pathogens implies that HGT occurred recently among these microorganisms.
Materials and Methods
Sample Collection and Plasmid Extraction
Samples were collected from fish farm sediments (Xiangshan Harbor, Zhejiang province, China) in November 2009. Sediments were suspended in seawater, and serially diluted samples were plated on seawater plates supplemented with 0.5 % (w/v) fish peptone and 0.1 % (w/v) yeast extract for 3 days at 30 °C. Growing bacteria were collected for plasmid extraction. Plasmids were extracted from mixed bacterial populations using the Large-Construct Kit (Qiagen, Valencia, CA, USA), which removes chromosomal DNA. The plasmid DNA was then digested using an ATP-dependent plasmid-safe DNase (Epicentre Biotechnologies, Madison, WI, USA) to remove any remaining chromosomal DNA according to the manufacturer’s instructions. Purified plasmid DNA samples were used for Solexa sequencing. The presence of chromosomal DNA was evaluated by PCR using the 16S rRNA gene universal primers 27 F and 1492R [6].
Sequencing and Assembly of the Plasmid Metagenome
Plasmid DNA from fish farms was sequenced by the Beijing Institute of Genomics (Chinese Academy of Sciences) using a Solexa GAII sequencer (Illumina, San Diego, CA, USA). Mate pair was used for sequencing with 1.5-kb inserts. More than 5 Gb of raw data were generated for plasmid DNA. Platform Galaxy was used to process the raw data [7], including removing adaptor sequences, low-quality trimming sequences (reads of less than 60 bp or with more than 20 bases with Sanger quality scores less than 30), and duplicate sequences. All clean reads were assembled using the program Velvet [8]. Different k-mer size ranges, from 29 to 61, were used to assemble the clean reads. The best assembly was produced using a k-mer size of 41; the outputs included 2,789 contigs with N50 of 18,690 and a maximum contig size of 233,555. The contigs were blasted against the non-redundant nucleotide database of GenBank using the blastn program (E-value cutoff of 10−10).
Mapping Reads to the NCBI Plasmid Genome Database and the ARDB
BLAT was used to map the plasmid reads to the NCBI Plasmid Genome Database (NCBI RefSeq database, 3,085 sequences, ftp://ftp.ncbi.nlm.nih.gov/refseq/release/plasmid/). Reads with alignment lengths above 70 bp were maintained for further analysis.
The replication proteins of plasmids were extracted from the NCBI plasmid database according to their annotated names [9]. All reads were aligned against the replication protein using blastx. The BLAST hits were filtered with an identity above 80 % and alignment lengths of more than 60 bp; only the best hits were taken as annotations for the corresponding reads. Predicted proteins present in the plasmids were grouped by their functional domains according to the Pfam database and corresponding alignment was executed using blastp [10].
Blastx was also used to align the plasmid metagenome data to protein sequences in the Antibiotic Resistance Genes Database (ARDB) [11], which includes more than 7,000 proteins that confer resistance to 249 different antibiotics. The threshold was set to an alignment length ≥60 bp and an identity ≥80 %, and only antibiotic genes with more than three mapped reads were kept for further research.
Identification of Plasmid pNM5k
Blastn was used to align the contigs to the NCBI Plasmid Genome Database. Contig231 was similar to the reported plasmid. Contig231 was amplified by overlap extension using primers 231 F and 231R (231 F: AGGTCCATAGGTAGCAACTTCTGATTCAGCAAGCGTACAG, 231R: GAAGTTGCTACCTATGGACCTTGATCAATCCACTTACTTGT), and the PCR product was transformed into Escherichia coli DH10B. Transformants were selected on Luria–Bertani plates containing tetracycline (8 mg L−1). The plasmid was extracted from the transformants and sequenced. The MIC for transformants was determined using broth microdilution assays, as recommended by the Clinical and Laboratory Standards Institute (http://www.clsi.org/).
Results
Diversity of Plasmids in Marine Sediment
A plasmid purification kit and Plasmid-Safe™ DNase (Epicentre Biotechnologies) were used to obtain pure plasmid DNA. However, amplification of the 16S rRNA gene indicated that chromosomal DNA was not completely removed. Plasmid DNA from the fish farms was sequenced using a Solexa GAII sequencer, and more than 5 Gb of raw data were generated. Galaxy was used to obtain clean reads [7]. In total, 27,409,674 clean reads of 2.19 Gb were obtained for further analysis.
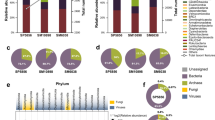
BLAT was used to map the reads to the NCBI Plasmid Genome Database. A total of 2,907,002 reads (10.6 %) aligned to 652 plasmids. Plasmids that were mapped to more than 10 reads were selected for further analysis. The distribution of plasmid hosts were Actinobacteria (3 %), Firmicutes (14 %), Proteobacteria (82 %), and others (1 %; Fig. 1). Proteobacteria was the most abundant in the host of plasmid. Proteobacteria include Betaproteobacteria (3 %), Alphaproteobacteria (25 %), and Gammaproteobacteria (54 %; Fig. 1). Our study provides new data regarding the genetic diversity of plasmids in marine sediment.
An analysis of replication proteins matched by reads showed that these replication proteins were assigned to 10 replication protein-related domain families, RP-C_C, RepB, SeqA, RPA, Rep_trans, Rep_3, Rep_1, Phage_GPA, ParBc, and DnaB_C (Table 1). Rep-1 was the most abundant. Our results revealed substantial divergence of different plasmids in sediment.
Marine Sediment as a Reservoir of Antibiotic Resistance Genes
We analyzed the reads using blastx against the ARDB. A stringent cutoff (alignment length ≥60 bp; number of mapped reads ≥3) was adopted. The reads were assigned to 58 genes with high homology (≥80 % amino acid identity), 96 genes with identities of 60–80 %, and 37 genes with identities of 40–60 % to previously known resistance genes. Genes with low homology suggest the presence of previously unknown resistance genes in the bacteria from marine sediments, which could be infrequently exchanged with human pathogens. We selected 58 genes with high homology for further analysis. These genes conferred resistance to 11 classes of antibiotics, including 12 resistance genes for aminoglycoside, 10 for tetracycline, 7 for β-lactam, 7 for chloramphenicol, 4 for trimethoprim, 3 for glycopeptide, 2 for bacitracin and fluoroquinolone, 1 for macrolides, sulfonamide, and streptogramin, and 8 for multidrug efflux (Fig. 2). Our results indicated that marine sediment is a reservoir of antibiotic resistance genes.
According to our result, sequences coding for tetracycline resistance genes were highly rich in the plasmid data. Ten tet resistance genes were detected in the sediment samples, including tetC, tet33, tetK, tet41, tetB, tetL, tet35, tet32, and tetB(P). Resistance genes coding for the tetL efflux pump were the most abundant. Recent studies have demonstrated that tetracycline genes are prevalent in the aquaculture environment [12, 13]. However, previous studies have showed that tetA, tetB, tetD, tetE, and tetM are the most frequent tetracycline resistance genes in marine fish farms [14–16].
Sediment Bacteria Shared Common Antibiotic Resistance Genes with Human Pathogens
More than 40,000 reads were similar to genes in pathogenic isolates (≥80 % amino acid identity). A total of these reads were 100 % identical. Also, 4,565 reads mapped to the gene aph33Ib which encodes aminoglycoside O-phosphotransferase (YP_002112964) in Salmonella enterica serovar Schwarzengrund. S. enterica serovar Schwarzengrund is a common cause of human gastroenteritis which is spreading internationally [17]. A total of 233 reads mapped to the gene qnrA which encodes a pentapeptide family protein (ACE95087) in Klebsiella pneumoniae with 100 % identity. QnrA proteins protect DNA from quinolone binding [18]. The aquatic bacterium Shewanella algae has been identified as a reservoir of qnrA [19].
We identified six contigs carrying antibiotic resistance genes with >90 % identity to transposon or plasmid of human pathogens (Table 2). Contig891 contained four functional genes: tnpA which encodes a putative transposase, tnpR which encodes a putative resolvase, and strA and strB genes which encode two different phosphotransferases, which are both required for streptomycin resistance. Contig891 with a length of 5,396 bp shared 99 % identity with the mobile element sequence of human pathogenic bacteria (Erwinia amylovora, K. pneumoniae, Bordetella bronchiseptica, Alcaligenes faecalis, Pseudomonas syringae, S. enterica, and Yersinia ruckeri) (Table 2). Additionally, the contig was 99 % identical to a transposon from the fish pathogenic bacterium Aeromonas salmonicida subsp. salmonicida 1682/92 [20]. Contig178 with a length of 3,139 bp included two open reading frames (ORFs). One ORF showed 95 % identity with the macrolide-resistant mefE-encoded efflux pump, while the other encodes a product that shares 95 % identity with an ATP-binding cassette transporter (mel) involved in macrolide resistance [21]. Interestingly, the contig was well aligned to a mega element which was prevalent in Streptococcus pneumoniae (Table 2). Contig218 was 99 % identical to transposon Tn558 with 43 % coverage including florfenicol–chloramphenicol exporter protein FexA, a putative transposase and a hypothetical protein (Table 2). Tn558 was detected in different staphylococcal species [22].
Contig231 was 5,109 bp in length and had three main functional genes: rep, mob, and tetL. rep belongs to the Rep_1 superfamily (PF01446) replication protein; mob is a DNA relaxase that nicks supercoiled DNA in a site- and strand-specific fashion at oriT sites in preparation for conjugation [23], and tetL encodes a tetracycline efflux pump. tetL showed 93 % identity with tetracycline resistance genes from Staphylococcus aureus and Enterococcus faecalis. In addition, contig231 was 99 % identical to plasmid pBSDMV9, which was detected on farms with chicken waste [24]. We used PCR to show that the contig derived from circular molecules are likely a complete circular plasmid. PCR products were transformed into E. coli DH10B by electroporation. Transformant’s tetracycline resistance increased eightfold than that of control. The results demonstrated that the plasmid could replicate and the tetracycline resistance gene functionally expressed in different hosts, indicating dissemination of resistance determents by the plasmid. The contig was identified as a plasmid named pNM5. Mapping reads to the plasmid showed that pNM5 was prevalent in marine sediment samples with high relative abundance (0.45 %).
Discussion
Plasmids, as mobile genetic elements, are involved in the dissemination of antibiotic resistance. Many resistance genes located on plasmids have been isolated from activated sludge [4, 5]. In this study, we applied high-throughput sequencing and metagenomic analyses to study the contribution of resistance genes carried by plasmid in marine sediment to global resistance exchange.
Studies have been done to detect resistance genes in marine aquaculture environment. Tetracycline resistance genes have been identified in fish farms throughout the world [14, 25]. However, reports of resistance genes conferring to other types of antibiotic resistance are rare [26]. In our study, resistance genes on plasmids are found to confer resistance to 11 types of antibiotics through diverse mechanisms. Twelve resistance genes for aminoglycoside were detected in marine sediment, which encode phosphotransferase and nucleotidylyltransferase separately. Genes of aminoglycoside N-acetyltransferase and nucleotidylyltransferase (aadA1, aadA2, aac61b) were found in isolates from ornamental fish [27]. Our study is the first report about the occurrence of genes encoding aminoglycoside phosphotransferase in the marine environment. Also, to the best of our knowledge, genes conferring resistance to streptogramins, glycopeptides, and bacitracin were first reported in an aquaculture environment. Overall, our study identified numerous putative resistance determinants, which profiled the reservoir of resistance genes in sediment bacteria.
To understand the exchange of resistance genes between sediment bacteria and human pathogens, we aligned our data against the pathogen resistome. More than 40,000 reads were assigned to genes in pathogenic isolates with high homology ≥80 % (amino acid identity), 59 % of which were 100 % identical. Additionally, six contigs carrying antibiotic resistance genes were well aligned (>90 % identity) to transposon or plasmid from human pathogens, encompassing coding and non-coding regions (from 1.5 to 5.4 kb, Table 2). The high nucleotide identity detected between sediment bacteria and pathogens implies that HGT occurred recently among these microorganisms.
Contig891 with a length of 5,393 bp shared 99 % identity with the conserved backbone of transposon Tn5393 [28]. The insertion of different mobile elements into the Tn5393 backbone indicates that recombination events occurred frequently within the transposon (Fig. 3). The transposon backbone could embed into plasmids such as broad-host-range IncP-1 [29]. Therefore, the transposon can be mobilized into unrelated bacteria [30]. Contig178 was well aligned to the mega element with 58 % coverage, including two macrolide resistance genes (Table 2). The mega element, which is 5.5 kb in size, can be found inserted at different chromosomal sites. [31]. The presence of a mega element is the most prevalent mechanism of S. pneumoniae macrolide resistance in the USA, Canada, and the UK [32]. Two resistance genes carried by the mega element, mefE and mel, are prevalent in S. pneumoniae and have been detected in other clinical streptococcal species [33]. However, our results demonstrate the occurrence of mefE and mel in marine sediment.
In this study, we applied high-throughput sequencing and metagenomic analyses to examine the resistance genes of the plasmids in a fish farm. Numerous putative resistance determinants were found in sediment bacteria indicating that marine sediment serves as a pool of resistance genes. We also found several contigs that shared high identity with transposons or plasmids indicating that the sediment bacteria recently contributed or acquired resistance genes from human pathogens. Thus, the sediment bacteria may play an important role in the global exchange of antibiotic resistance.
References
D'Costa VM, McGrann KM, Hughes DW, Wright GD (2006) Sampling the antibiotic resistome. Sci 311(5759):374–377. doi:10.1126/science.1120800
Forsberg KJ, Reyes A, Wang B, Selleck EM, Sommer MO, Dantas G (2012) The shared antibiotic resistome of soil bacteria and human pathogens. Sci 337(6098):1107–1111. doi:10.1126/science.1220761
Brown Kav A, Sasson G, Jami E, Doron-Faigenboim A, Benhar I, Mizrahi I (2012) Insights into the bovine rumen plasmidome. Proc Natl Acad Sci USA 14(109):7
Szczepanowski R, Bekel T, Goesmann A, Krause L, Krömeke H, Kaiser O, Eichler W, Pühler A, Schlüter A (2008) Insight into the plasmid metagenome of wastewater treatment plant bacteria showing reduced susceptibility to antimicrobial drugs analysed by the 454-pyrosequencing technology. J Biotechnol 136:11
Zhang T, Zhang XX, Ye L (2011) Plasmid metagenome reveals high levels of antibiotic resistance genes and mobile genetic elements in activated sludge. PLoS One 6(10):e26041. doi:10.1371/journal.pone.0026041
Weisburg WG, Barns SM, Pelletier DA, Lane DJ (1991) 16S ribosomal DNA amplification for phylogenetic study. J Bacteriol 173(2):697–703
Blankenberg D, Gordon A, Von Kuster G, Coraor N, Taylor J, Nekrutenko A (2010) Manipulation of FASTQ data with Galaxy. Bioinforma 26(14):1783–1785. doi:10.1093/bioinformatics/btq281
Zerbino DR, Birney E (2008) Velvet: algorithms for de novo short read assembly using de Bruijn graphs. Genome Res 18(5):821–829. doi:10.1101/gr.074492.107
Ma Y, Paulsen IT, Palenik B (2012) Analysis of two marine metagenomes reveals the diversity of plasmids in oceanic environments. Environ Microbiol 14(2):453–466. doi:10.1111/j.1462-2920.2011.02633.x
Sonnhammer EL, Eddy SR, Birney E, Bateman A, Durbin R (1998) Pfam: multiple sequence alignments and HMM-profiles of protein domains. Nucleic Acids Res 26(1):320–322
Liu B, Pop M (2009) ARDB—Antibiotic Resistance Genes Database. Nucleic Acids Res 37:D443–447. doi:10.1093/nar/gkn656
Seyfried EE, Newton RJ, Rubert KF, Pedersen JA, McMahon KD (2010) Occurrence of tetracycline resistance genes in aquaculture facilities with varying use of oxytetracycline. Microb Ecol 59(4):799–807. doi:10.1007/s00248-009-9624-7
Gao P, Mao D, Luo Y, Wang L, Xu B, Xu L (2012) Occurrence of sulfonamide and tetracycline-resistant bacteria and resistance genes in aquaculture environment. Water Res 46(7):2355–2364. doi:10.1016/j.watres.2012.02.004
Di Cesare A, Vignaroli C, Luna GM, Pasquaroli S, Biavasco F (2012) Antibiotic-resistant enterococci in seawater and sediments from a coastal fish farm. Microb Drug Resist. doi:10.1089/mdr.2011.0204
Tamminen M, Karkman A, Lohmus A, Muziasari WI, Takasu H, Wada S, Suzuki S, Virta M (2011) Tetracycline resistance genes persist at aquaculture farms in the absence of selection pressure. Environ Sci Technol 45(2):386–391. doi:10.1021/es102725n
Dang H, Ren J, Song L, Sun S, An L (2008) Diverse tetracycline resistant bacteria and resistance genes from coastal waters of Jiaozhou Bay. Microb Ecol 55(2):237–246. doi:10.1007/s00248-007-9271-9
Akiyama T, Khan AA (2012) Molecular characterization of strains of fluoroquinolone-resistant Salmonella enterica serovar Schwarzengrund carrying multidrug resistance isolated from imported foods. J Antimicrob Chemother 67(1):101–110. doi:10.1093/jac/dkr414
Nordmann P, Poirel L (2005) Emergence of plasmid-mediated resistance to quinolones in Enterobacteriaceae. J Antimicrob Chemother 56(3):463–469. doi:10.1093/jac/dki245
Poirel L, Rodriguez-Martinez JM, Mammeri H, Liard A, Nordmann P (2005) Origin of plasmid-mediated quinolone resistance determinant QnrA. Antimicrob Agents Chemother 49(8):3523–3525. doi:10.1128/AAC.49.8.3523-3525.2005
L'Abee-Lund TM, Sorum H (2000) Functional Tn5393-like transposon in the R plasmid pRAS2 from the fish pathogen Aeromonas salmonicida subspecies salmonicida isolated in Norway. Appl Environ Microbiol 66(12):5533–5535
Ambrose KD, Nisbet R, Stephens DS (2005) Macrolide efflux in Streptococcus pneumoniae is mediated by a dual efflux pump (mel and mef) and is erythromycin inducible. Antimicrob Agents Chemother 49(10):4203–4209. doi:10.1128/AAC.49.10.4203-4209.2005
Kehrenberg C, Schwarz S (2006) Distribution of florfenicol resistance genes fexA and cfr among chloramphenicol-resistant Staphylococcus isolates. Antimicrob Agents Chemother 50(4):1156–1163. doi:10.1128/AAC.50.4.1156-1163.2006
Murray KD, Aronstein KA, de Leon JH (2007) Analysis of pMA67, a predicted rolling-circle replicating, mobilizable, tetracycline-resistance plasmid from the honey bee pathogen, Paenibacillus larvae. Plasmid 58(2):89–100. doi:10.1016/j.plasmid.2007.02.001
You Y, Hilpert M, Ward MJ (2012) Detection of a common and persistent tet(L)-carrying plasmid in chicken-waste-impacted farm soil. Appl Environ Microbiol 78(9):3203–3213. doi:10.1128/AEM.07763-11
Akinbowale OL, Peng H, Barton MD (2007) Diversity of tetracycline resistance genes in bacteria from aquaculture sources in Australia. J Appl Microbiol 103(5):2016–2025. doi:10.1111/j.1365-2672.2007.03445.x
Zhao JY, Dang H (2012) Coastal seawater bacteria harbor a large reservoir of plasmid-mediated quinolone resistance determinants in Jiaozhou Bay, China. Microb Ecol 64(1):187–199. doi:10.1007/s00248-012-0008-z
Verner-Jeffreys DW, Welch TJ, Schwarz T, Pond MJ, Woodward MJ, Haig SJ, Rimmer GS, Roberts E, Morrison V, Baker-Austin C (2009) High prevalence of multidrug-tolerant bacteria and associated antimicrobial resistance genes isolated from ornamental fish and their carriage water. PLoS One 4(12):e8388. doi:10.1371/journal.pone.0008388
Mantengoli E, Rossolini GM (2005) Tn5393d, a complex Tn5393 derivative carrying the PER-1 extended-spectrum beta-lactamase gene and other resistance determinants. Antimicrob Agents Chemother 49(8):3289–3296. doi:10.1128/AAC.49.8.3289-3296.2005
Petrovski S, Stanisich VA (2011) Embedded elements in the IncPbeta plasmids R772 and R906 can be mobilized and can serve as a source of diverse and novel elements. Microbiol 157(Pt 6):1714–1725. doi:10.1099/mic.0.047761-0
Miriagou V, Papagiannitsis CC, Kotsakis SD, Loli A, Tzelepi E, Legakis NJ, Tzouvelekis LS (2010) Sequence of pNL194, a 79.3-kilobase IncN plasmid carrying the blaVIM-1 metallo-beta-lactamase gene in Klebsiella pneumoniae. Antimicrob Agents Chemother 54(10):4497–4502. doi:10.1128/AAC.00665-10
Del Grosso M, Camilli R, Iannelli F, Pozzi G, Pantosti A (2006) The mef(E)-carrying genetic element (mega) of Streptococcus pneumoniae: insertion sites and association with other genetic elements. Antimicrob Agents Chemother 50(10):3361–3366. doi:10.1128/AAC.00277-06
Zhanel GG, Wang X, Nichol K, Nikulin A, Wierzbowski AK, Mulvey M, Hoban DJ (2006) Molecular characterisation of Canadian paediatric multidrug-resistant Streptococcus pneumoniae from 1998–2004. Int J Antimicrob Agents 28(5):465–471. doi:10.1016/j.ijantimicag.2006.08.005
Jonsson M, Swedberg G (2006) Macrolide resistance can be transferred by conjugation from viridans streptococci to Streptococcus pyogenes. Int J Antimicrob Agents 28(2):101–103. doi:10.1016/j.ijantimicag.2006.02.023
Acknowledgments
This work was supported by grants from the Knowledge Innovation Program of the Chinese Academy of Sciences (grant no. KSCX2-EW-J-6) and the National High-Technology Research and Development Program (“863” Program) of China (grant no. 2012AA092001).
Author information
Authors and Affiliations
Corresponding author
Additional information
Jing Yang, Chao Wang, and Chang Shu contributed to this work equally.
Rights and permissions
About this article
Cite this article
Yang, J., Wang, C., Shu, C. et al. Marine Sediment Bacteria Harbor Antibiotic Resistance Genes Highly Similar to Those Found in Human Pathogens. Microb Ecol 65, 975–981 (2013). https://doi.org/10.1007/s00248-013-0187-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-013-0187-2