Abstract
The purpose of this study was to examine the variations in the microbial community structure of soil actinomycetes in fields with continuous cropping of cotton in Xinjiang Autonomous Region, China. Soil samples were collected from four depths in fields with 7-year continuous cotton cropping. The community structure of soil actinomycetes was examined using the 16S rRNA-based polymerase chain reaction–density gradient gel electrophoresis (PCR-DGGE) techniques. The microbial diversity indices of the soil samples from different depths generally decreased along with the period of continuous cotton cropping. When the period of continuous cropping of cotton reached 5 years, the diversity indices rose again and gradually stabilized at a level slightly lower than that of soils with original ecology (i.e., 0-year cotton cropping). Cluster analysis showed that at the 1–20-cm depth, the actinomycete community structure of the soil subjected to 1-year cotton cropping was similar to that of soil subjected to 0-year cotton cropping, whereas that of soils after 3-year continuous cotton cropping showed high similarity. At the 21–40-cm depth, the actinomycete community structure showed various changes but generally recovered to its original pattern after repeated fluctuations. Principal component analysis showed that at the 1–30-cm depth, the actinomycete community structure varied similarly regardless of the period of continuous cotton cropping. In contrast, there were no clear actinomycete community structure variation trends at the 31–40-cm soil depth. Homology comparison of sequences recovered from the DGGE bands showed that the obtained sequences shared similarities >88 %. Alignment with the known homologous sequences indicated a lack of microorganisms related to soil-borne cotton diseases. Continuous cotton cropping exerted significant influences on the community structure of soil actinomycetes in Xinjiang Autonomous Region, which were largely determined by the soil depth and the period of continuous cotton cropping. The microbial diversity of soil actinomycete communities gradually recovered after 5-year continuous cropping. Thereafter, a new actinomycete community structure that was beneficial for continuous cropping of cotton was formed and stabilized each year.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Cotton, a major economic crop in China, has an important position in the national economy. Xinjiang Autonomous Region is suitable for cotton cultivation for a variety of reasons, e.g., little rainfall, abundant sunshine, and substantial differences between day and night temperatures (annual rainfall of 110–200 mm, 1,000–1,500 mm evaporation, average temperature of 6–6.6 °C, 2,900 h of sunshine, altitude of 300–500 m, and temperature of 15–20 °C in August). Due to continuous cropping obstacles, cotton cultivation areas have substantially decreased in the middle and eastern parts of China. Consequently, a vast area of uncultivated land in Xinjiang has been reclaimed for cotton cultivation since the 1980s. For the 20 years up until 2011, the Xinjiang region raised China in cotton cultivation area, total yield, and yield per unit area. Due to the single crop species in Xinjiang (e.g., >90 % coverage of cotton in local major cultivation districts), crop rotation and/or intercropping is unlikely, leading to serious continuous cropping obstacles [1]. Similar to other cultivation areas, the continuous cropping of cotton in Xinjiang initially caused a large-scale outbreak of soil-borne diseases that resulted in substantial crop yield and quality losses. However, along with the increasing period of continuous cotton cropping, there are no clear trends in the continuous cropping obstacles while the crop yields remain high and stable. This has raised great interests from agriculturists.
Continuous cropping is common in agricultural production and can cause obstacles such as soil quality degradation, crop yield reduction, and aggravated crop pest-induced plant diseases [2, 3]. The fundamental reason for continuous cropping obstacles is related to rhizosphere soil microstructure disorders or deterioration. Crop rotation, fallow, and manure application have commonly been used to solve continuous cropping obstacles in agriculture [4–6]. These methods attempt to improve the soil microstructure, but their effects are temporary [7, 8]. A healthy and stable soil microstructure is essential to maintaining long-term continuous cropping and stable high crop yields. This study aimed to examine the shifts in soil microstructure in fields subjected to continuous cotton cropping (from the occurrence to the disappearance of continuous cropping obstacles, further to the stable long-term high cotton yield) in Xinjiang, China. The results showed that continuous cropping of a single species both exerted strong influences on soil microstructure and affected the community structure of soil microorganisms at different depths [9]. Although soil microbial communities at different depths responded to environmental changes at different rates, the associated soil microstructure was capable of spontaneous adjustment according to the consistent long-term environmental changes, thereby forming a new stable and healthy structural pattern that is beneficial for continuous cropping.
Materials and Methods
Sample Collection
In August 2010, soil samples (gray desert soil: pH 8.59; total organic matter, 13.2 g kg−1; clay, 243.5 g kg−1; total N, 0.611 g kg−1; total P, 0.987 g kg−1; total K, 44.9 g kg−1; exchangeable N, 39.4 mg kg−1; exchangeable P, 20.9 mg kg−1; exchangeable K, 212.0 mg kg−1) were collected from fields subjected to 0- (uncultivated), 1-, 3-, 5-, 10-, 15-, and 20-year continuous cotton cropping in Shihezi, Xinjiang Autonomous Region, China (E 85°21′81″–25′55″, N 44°25′47″–26′96″). For each period of continuous cotton cropping, 40-cm-deep soil cores were collected from five plots by vertical drilling, and the samples were taken from four depths, i.e., 1–10, 11–20, 21–30, and 31–40 cm. Soils from the same depth with the same continuous cotton cropping period were mixed in equal amounts to form a composite sample (n = 28). The samples were transported to the laboratory immediately after collection and stored at 4 °C prior to experimental analyses conducted within 7 days.
DNA Extraction and 16S rRNA Gene Polymerase Chain Reaction Amplification
DNA was extracted from 0.5 g of soil from each sample using a MoBio Ultra-Clean™ soil DNA kit (MoBio Laboratories, USA). Using the Moreira method [10], we analyzed the total DNA in the next step. During the first polymerase chain reaction (PCR) cycle, the 16S rRNA fragment was amplified using the actinomycete-specific primers 27F/16Sact1114r (5′-AGAPTTTGATCCTGGCTCAG-3′/5′ -GAGTTGACCCCGGCRGT-3′) [11, 12]. PCR was performed in a total volume of 25 μL in 200-μL microtubes, which contained 0.15 μL each primer (60 pmol each), 2.8 μL 2.5 mmol/L dNTP mixture, 2.5 μL 10× buffer, 0.5 μL (2.0 U/μL) Taq enzymes, 0.2 μL (about 10 ng) DNA template, and 18.85 μL Milli-Q water. The PCR conditions were as follows: 5 min at 94 °C; 60 s at 94 °C, 45 s at 65 °C (0.5 °C decrease per cycle for the first 20 cycles, finally down to 55 °C), 120 s at 72 °C for a total of 30 cycles; and 72 °C for 7 min. During the second PCR cycle, the actinomycete 16S rRNA V3 fragment was amplified using P338fGC/P518r primers [13]. PCR was performed in a total volume of 50 μL in 200-μL microtubes, which contained 1 μL of the product of the first PCR cycle (1:100 diluted), 0.25 μL each primer (100 pmol each), 5.6 μL 2.5 mmol/L dNTP mixture, 5 μL 10× buffer, 1 μL (2.0 U/ μL) Taq enzymes, and 37.15 μL Milli-Q water. The reaction protocol was as follows: 5 min at 94 °C; 60 s at 94 °C, 45 s at 55 °C, 60 s at 72 °C for 30 cycles; and 72 °C for 5 min.
Polymerase Chain Reaction–Density Gradient Gel Electrophoresis Analysis and Band Identification
PCR samples were loaded onto 6–8 % (w/v) polyacrylamide gels in 0.5× TAE buffer (20 mM Tris–acetate, 1 mM ethylenediaminetetraacetic acid; pH 8.0). The polyacrylamide gels were made with denaturing gradients of 30–70 % (where 100 % denaturant contained 7 M urea and 40 % formamide). The gels (length, 18 cm) were run for 4 h at 150 V and 60 °C, after which time they were soaked for 1 h in SYBR Green I nucleic acid stain solution (1:10,000 dilution; Molecular Probes, Leiden, the Netherlands) and immediately photographed using an FR-200 UV–visible analysis system (Furi Company, Japan).
The banding patterns of the density gradient gel electrophoresis (DGGE) profile were analyzed using Quantity One v4.62 software (Bio-Rad Company, USA), Gel-Pro analyzer 4.0 (Media Cybernetics Company, USA), and Excel software (Microsoft Company, USA). The position and intensity of each band were determined automatically by the program. Each band's intensity was divided by the average band's intensity of the sample to minimize the influence of the different DNA concentrations between samples [14]. All data were the average of three trials.
Statistical Analysis
The diversity index of the soil actinomycetes communities was evaluated using the Shannon–Wiener index (H), richness (S), and evenness (E H ) based on the following equation [15]:
where p i is the ratio of the intensity of a single band to the total intensity of all bands within the same lane, and S is the total number of bands in each lane.
Cluster analysis was performed using the unweighted pair group method with arithmetic mean. The band data were subjected to principal component analysis (PCA) using the Statistical Package for the Social Sciences version 16.0 for Windows (SPSS Incorporated 2007). The principal components were then extracted from the correlation matrix. The standardized data were then projected onto the principal axes, plotted in two dimensions, and examined for clustering behavior [16].
DNA Cloning, Sequencing, Nucleotide Sequence Accession Numbers, and Phylogenetic Analysis
High-density DGGE bands were recovered from the uncultivated sample, and specific bands were recovered from other continuous cropping samples. DNA band recovery and the subsequent cloning procedures were performed according to Daniela [16]. Sequencing (conducted by Shenggong Company, China) of the colonies containing the correct target gene insert was conducted using vector primers (P338fGC/P518r). The obtained sequences were submitted to the GenBank database under accession numbers JN572602-JN572631 and JN572673. A BLAST search was used to explore the sequence similarity with others in the GenBank database. Sequence alignment was conducted using the Clustal X version 2.1 software. A phylogenetic tree of the 16S rRNA gene V3 fragments was built using the neighbor-joining method.
Results
Microbial Diversity Indices
With extended continuous cotton cropping periods, microbial diversity indices (H, S, and E H ) of soil actinomycetes at different depths showed various changes (Table 1). The three indices were highest for soil actinomycetes in uncultivated soils at various depths, and rapid decreases occurred within the first year of continuous cotton cropping. However, the decreasing rate gradually slowed down as the continuous cotton cropping periods increased and then rose again after 5 years of continuous cropping. Among the different depths, the diversity indices of soil actinomycete communities showed the largest reduction and rebounds at the 21–30-cm depth. The three diversity indices of soil actinomycetes gradually recovered at levels slightly lower than those of uncultivated soils after 10-year continuous cropping, except that the E H value was slightly higher than that of uncultivated soils at the 31–40-cm depth.
Similarity and Cluster Analysis
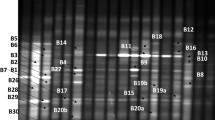
Cluster analysis showed that at the 1–20-cm depth, the uncultivated and 1-year continuously cropped samples were similar and clustered into a single small group, whereas those 3-, 5-, 10-, 15-, and 20-year continuously cropped samples were similar and clustered into one big group (Figs. 1, 2, 3, and 4, right). At the 21–30-cm depth, the uncultivated sample had low similarity with other samples and was individually clustered into a small group, whereas those subjected to 3- and 15-year continuous cropping had high similarity and were clustered into a separate small group. The third cluster included four samples from the fields subjected to the shortest and longest continuous cropping periods, i.e., 1, 5, 10, and 20 years. At the 31–40-cm depth, the 20-year continuously cropped sample had the least similarity with the other samples, while the latter were clustered into one big group.
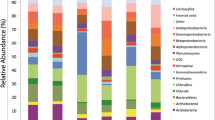
Density gradient gel electrophoresis profiles (left) of the polymerase chain reaction products from actinomycetes in soils at the 21–30-cm depth and their cluster analysis (right) with 0-, 1-, 3-, 5-, 10-, 15-, and 20-year successive monoculture cotton cropping (The 13–17 and e–f bands were excised)
Density gradient gel electrophoresis profiles (left) of the polymerase chain reaction products from actinomycetes in soils at the 31–40-cm depth and their cluster analysis (right) with 0-, 1-, 3-, 5-, 10-, 15-, and 20-year successive monoculture cotton cropping (The 18–24 and g–h bands were excised)
PCA Analysis
As shown in Fig. 5, the first principal component was the most important at various soil depths, with a variance contribution rate >40 %. The first principal component was positively correlated with the soil samples except for the 1–10- and 21–30-cm uncultivated soil, 11–20-cm 1-year continuously cropped soil, and 31–40-cm 20-year continuously cropped soil samples. In particular, except for the uncultivated soil, the soils had the highest correlations at the 21- to 30-cm depth, and all of them fluctuated between the positive and negative axes of the second principal component.
Homologous Sequence Similarity Analysis
DGGE bands 1–24 shown in Fig. 1, 2, 3, and 4 (left) represent the high-density bands from the uncultivated soil, whereas bands a–h represent the high-density bands from soils subjected to different continuous cotton cropping periods. A phylogenetic tree was constructed based on the selected DNA sequences recovered from the DGGE gel as well as their known relatives retrieved from GenBank (Fig. 6). The 32 DNA sequences recovered from the DGGE gel shared 88–100 % sequence homology, of which several sequences shared 100 % homology with known microorganisms such as Rhodococcus sp. Pc2 Fnocardiaceae, Bacterium Ellin 5004, and unclassified Actinobacteridae of different genera. Further inspection of the known microorganisms that are mostly related to DNA sequences recovered from the DGGE gel indicated a lack of microorganisms related to soil-borne diseases such as cotton blight and verticillium wilt in the tested soils. This finding differed from those results reported by other studies in terms of continuous cropping obstacles [17, 18].
Discussion
Community Structure of Soil Actinomycetes Shifted Due to Long-Term Continuous Cotton Cropping
The community structure of soil microorganisms is mainly determined by environmental factors such as soil properties [19], whereas crop type and tillage management measures also cause shifts in the community structure of soil microorganisms [20, 21]. In this study, 16S rRNA-based PCR-DGGE fingerprinting demonstrated that long-term continuous cotton cropping resulted in continuous changes in the community structure of soil actinomycetes, which shifted from the original ecological structure to a relatively stable and healthy pattern suitable for long-term continuous cotton cropping. In this process, the community structure of soil actinomycetes underwent the most rapid changes during the first year of cotton cultivation, mainly showing reduced microbial diversity that possibly benefited cotton growth and the appearance of some species that could induce the obstacle of cotton continuous cropping [18]. This finding suggested that cotton cultivation had a strong influence on the community structure of soil actinomycetes.
With extension of the continuous cropping from 1 to 3 years, three microbial diversity indices started to rebound in the major distribution area of the cotton root at the 11–31-cm depth. When the field was subjected to 5-year continuous cotton cropping, microbial diversity indices rebounded at both 1–10-cm depth (surface layer) and 30–40-cm depth (plow pan). These findings indicated that continuous cotton cropping had different influences on the community structure of soil actinomycetes regarding soil depth. It was likely that the interaction between cotton roots in the major distribution zone and the soil microorganisms was intense. The microbial diversity indices gradually stabilized in the field subjected to 10-year continuous cotton cropping, while cluster analysis showed that the soils subjected to multi-year continuous cropping were largely grouped together. These findings demonstrated that the soil microbial communities spontaneously formed a new and relatively stable community structure under the stress of a long-term external environment.
Similar Variations in Actinomycete Community Structure and Cotton Yield During Continuous Cotton Cropping
Similar to other cotton cultivation districts (probably due to the large amount of endosulfan, monocrotophos, cypermethrin, lyphosate, foliar fertilizer, urea and diamine), newly reclaimed land in Xinjiang commonly shows serious continuous cropping obstacles such as reduced yields and pest-induced disease outbreaks. However, after extension of the continuous cropping period, not only are pest diseases spontaneously alleviated in cotton fields but crop yields are also recovered to their original high and stable state. Results from the present study showed that during the process of continuous cotton cropping, the actinomycete community structural variations could be related to the occurrence and alleviation processes of continuous cropping obstacles. In particular, the reduction of soil actinomycete diversity and its rebound period, the occurrence of cotton pest diseases [8, 22, 23], the loss of crop yields, and the subsequent rebound period were nearly consistent at the 11–30-cm depth (arable layer), suggesting a close relationship between soil microbial community structure and function and crop yield [24]. This finding also implies that long-term continuous cropping may not cause continued destruction of soil microbial community structures since the microbial communities can spontaneously adjust their composition to adapt to external environmental changes, thereby maintaining a new stable and healthy long-term community structure.
The Choice of Research Methods and Other Possible Factors May Cause Soil Actinomycete Community Structure Shift
Using the 16S rRNA-based PCR-DGGE technique [25], this study examined the entire shifting process of actinomycete community structures at different soil depths during continuous cotton cropping in Xinjiang for up to 20 years. Soil actinomycetes were targeted at different depths since previous research has shown that long-term cotton cropping can lead to the shifting of bacterial soil types to fungal soil types [26]. Despite the distribution of different actinomycete communities at different depths, variations in composition occurred during the entire continuous cropping period. In addition, the number of bands on the DGGE gel was between those of bacteria and fungus, thereby facilitating the data collection process. Soil sample collection was carried out at 15-cm line intervals (line spacing, 60 cm) since the soil microbes were affected by the crop root at this position, reflecting the composition of the microbial community structure of the entire soil environment.
Our results showed that the shift in the actinomycete community composition was most sensitive in the major distribution zone of the cotton roots, i.e., 11–31-cm depth, where they first showed the rebound and stabilization tendencies. Regardless of soil depth, the actinomycete community structure variations were quite similar, ultimately forming a relatively stable community composition. With regard to the spontaneous formation of the healthy microbial community structure in cotton fields subjected to long-term continuous cotton cropping, we speculated that it was primarily related to the local saline–alkali soil type, dry climate, and cropping patterns of half-year plough and half-year fallow in Xinjiang.
References
Han CL, Liu J, Zhang WF, Liu M, Huang WJ, Gao XM, Zhang HZ (2010) Biocycling of nine mineral elements of soil-cotton system in Xinjiang oasis. Acta Ecologica Sinica 30(22):6234–6241
Vargas GS, Meriles J, Conforto C, Figoni G, Basanta M, Lovera E, March GJ (2009) Field assessment of soil biological and chemical quality in response to crop management practices. World J Microb Biot 25(3):439–448
He JZ, Zheng Y, Chen CR (2010) Microbial composition and diversity of an upland red soil under long-term fertilization treatments as revealed by culture-dependent and culture-independent approaches. J Soil Sediment 8:349–358
Fu QL, Liu C, Ding NF, Lin YC, Guo B, Luo JF, Wang HL (2012) Soil microbial communities and enzyme activities in a reclaimed coastal soil chromosequence under rice-barley cropping. J Soil Sediment 12:1134–1144
Alvey S, Yang CH, Buerkert A (2003) Cereal legume rotation effects on rhizosphere bacterial community structure in west African soils. Biol Fert Soils 37:73–82
Martin I, Mun LM, Yunta F (2007) Tillage and croprotation effects on barley yield and soil nutrients on a Calciortidic Haploxeralf. Soil Till Res 92:1–9
Meriles JM, Vargas Gil S, Haro R, March GJ, Guzman CA (2008) Selected soil-borne fungi under glyphosate application and crop residues from a long-term field experiment. Biol Agric Hortic 26(2):193–205
Garbeva P, van Veen JA, van Elsas JD (2004) Microbial diversity in soil: selection of microbial populations by plant and soil type and implications for disease suppressiveness. Annu Rev Phytopathol 42:243–270
Huang JW, Yang JK, Duan YQ (2010) Bacterial diversities on unaged and aging flue-cured tobacco leaves estimated by 16S rRNA sequence analysis. Applied Microbiol Biot 88(2):553–562
Moreira D (1998) Efficient removal of PCR inhibitors using agarose-embedded DNA preparations. Nucleic Acids Res 26:3309–3310
Bruce KD, Hiorns WD, Hobman JL, Osborn AM, Strike P, Ritchie DA (1992) Amplification of DNA from native populations of soil bacteria by using the polymerase chain reaction. Appl Environ Microb 58:3413–3416
Martina K, Jan K, Tamas F, Ladislav C, Marek O, Genevieve LG, Yvan ML, Marketa SM (2008) Development of a 16S rRNA gene-based prototype microarray for the detection of selected actinomycetes genera. Anto Leeuw Int J G 94:439–453
Ovreas L, Fomey L, Daae FL (1997) Distribution of bacterioplankton in meromictic lake saelevannet, as determined by denaturing gradient gel electrophoresis of PCR. Amplified gene fragments coding for 16S rRNA. Appl Environ Microb 63:3367–3373
Graham MH, Haynes RJ (2005) Catabolic diversity of soil microbial communities under sugarcane and other land uses estimated by biolog and substrate-induced respiration methods. Appl Soil Ecol 29:155–164
Hill TCJ, Walsh KA, Harris JA (2003) Using ecological diversity measures with bacterial communities. FEMS Microbiol Ecol 43:1–11
Daniela RDF, Raquel VF, Mario C, Teresa CDM, Marrio JP, Bruno BC, Antonio C (2012) Impact of water quality on bacterioplankton assemblage along certima river basin (central western Portugal) accessed by PCR-DGGE and multivariate analysis. Environ Monit Assess 184:471–485
Li C, Li XM, Kong WD (2010) Effect of monoculture soybean on soil microbial community in the northeast china. Plant Soil 330:423–433
Zhang XL, Li X, Zhan CG (2011) Ecological risk of long-term chlorimuron-ethyl application to soil microbial community:an in situ investigation in a continuously cropped soybean field in northeast china. Environ Sci Pollut R 18:407–415
Girvan MS, Bullimore J, Pretty JN, Osborn AM, Ball AS (2003) Soil type is the primary determinant of the composition of the total and active bacterial communities in arable soils. Appl Environ Microb 69:1800–1809
Acosta-Martinez V, Dowd S, Sun Y, Allen V (2008) Tag-encoded pyrosequencing analysis of bacterial diversity in a single soil type as affected by management and land use. Soil Biol Biochem 40:2762–2770
Zhang Y, Du BH, Jin ZG, Li ZH, Song HN, Ding YQ (2011) Analysis of bacterial communities in rhizosphere soil of healthy and diseased cotton (Gossypium sp.) at different plant growth stages. Plant Soil 339:447–455
Robert P, Larkin (2008) Relative effects of biological amendments and crop rotations on soil microbial communities and soilborne diseases of potato. Soil Biol Biochem 40:1341–1351
Berg G, Smalla K (2009) Plant species and soil type cooperativelyshape the structure and function of microbial communities in the rhizosphere FEMS. Microbiol Ecol 68:1–13
Wei XR, Hao MD, Shao MA (2006) Changes in soil properties and the availability of soil micronutrients after 18 years of cropping and fertilization. Soil Till Res 91:120–130
Muyzer G, De Waal EC, Uitterlinden AG (1993) Profiling of complex microbial populations by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes coding for 16S rRNA. Appl Environ Microb 59(3):695–700
Ge Y, Zhang JB, Zhang LM, Yang M, He JZ (2008) Long-term fertilization regimes and diversity of an agricultural affect bacterial community structure soil in northern china. J Soil Sediment 8:43–50
Acknowledgments
This study was supported by the National Natural Science Foundation of China (30860016) and the key disciplines of Xinjiang Normal University (Microbiology).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhang, W., Long, X., Huo, X. et al. 16S rRNA-Based PCR-DGGE Analysis of Actinomycete Communities in Fields with Continuous Cotton Cropping in Xinjiang, China. Microb Ecol 66, 385–393 (2013). https://doi.org/10.1007/s00248-012-0160-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-012-0160-5