Abstract
Luminous bacteria are isolated from both Hydrozoa and Bryozoa with chitinous structures on their surfaces. All the specimens of the examined hydroid species (Aglaophenia kirchenpaueri, Aglaophenia octodonta, Aglaophenia tubiformis, Halopteris diaphana, Plumularia setacea, Ventromma halecioides), observed under blue light excitation, showed a clear fluorescence on the external side of the perisarc (chitinous exoskeleton) around hydrocladia. In the bryozoan Myriapora truncata, luminous bacteria are present on the chitinous opercula. All the isolated luminous bacteria were identified on the basis of both phenotypic and genotypic analysis. The isolates from A. tubiformis and H. diaphana were unambiguously assigned to the species Vibrio fischeri. In contrast, the isolates from the other hydroids, phenotypically assigned to the species Vibrio harveyi, were then split into two distinct species by phylogenetic analysis of 16S rRNA gene sequences and DNA–DNA hybridization experiments. Scanning electron microscopy analysis and results of culture-based and culture-independent approaches enabled us to establish that luminous vibrios represent major constituents of the bacterial community inhabiting the A. octodonta surface suggesting that the interactions between luminous bacteria and the examined hydrozoan and bryozoan species are highly specific. These interactions might have epidemiological as well as ecological implications because of the opportunistic pathogenicity of luminous Vibrio species for marine organisms and the wide-distribution of the hydrozoan and bryozoan functioning as carriers.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In the marine environment, only certain biological surfaces resist bacterial colonization for more or less extended periods, whereas, as a rule, bacteria rapidly colonize both living and nonliving submerged surfaces [20]. Despite their potentially important role in marine ecology, reports of interactions of epibiotic bacteria with marine macroorganisms are scarce and often circumstantial. A wide range of epibacterial densities (0 to 108 cells per square centimeter) is reported from the surfaces of algae [27, 44], sponges [49], Cnidaria [18, 50], and bryozoans [60]. Epibionts are able to survive in the natural environment longer than free-living forms and, by means of adhesive strategies, they can adapt to adverse conditions, e.g., organic matter limitation [15, 45]. The symbiosis of micro with macroorganisms is such a widespread phenomenon that it should have a profound impact on the physiology, ecology, and evolution of both hosts and symbiotic partners [38]. Thus, it is not surprising that the evolutionary patterns of closely related hosts and symbionts are often congruent. However, the mechanisms underlying the processes of cospeciation and host–symbiont specificity are poorly understood because most of animal–bacterial interactions cannot be experimentally initiated.
Some Vibrio species elaborate an extracellular chitinase and can feed on the chitinous structures of supporting organisms [6]. The colonization of the integument of copepods by Vibrio species is a well-described phenomenon especially in fecal-polluted vs. nonpolluted coastal zones [14, 28, 34, 51, 56]. Luminous bacteria, the simplest light-emitting organisms, can be either free-living in the water column with densities ranging from 102 to 104 cells per liter [37] or can be associated with other organisms [5]. The best-characterized species among luminous bacteria are Photobacterium phosphoreum, Vibrio fischeri, and Vibrio harveyi. Pujalte et al. [41] suggested that the association with guest organisms might contribute to survival and distribution of luminous bacteria in the marine environment. In contrast, the nature of the relationships between marine Metazoa and their associated luminous bacteria is not well known.
In a previous study, we described a novel association between V. harveyi and a benthic hydrozoan, Aglaophenia octodonta (Hydrozoa, Cnidaria) [50], suggesting that the association could be explained by the feeding activity of this microorganism on the hydroid chitinous structures. In the present study, we investigated the presence of luminous bacteria in some species of Hydrozoa and one species of Bryozoa, all characterized by the presence of chitinous structures on their surfaces. The selected species are commonly found at shallow depths on rocky coast of the Mediterranean Sea [12, 43].
Methods
Sampling
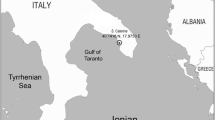
Batches of 50 colonies of the hydrozoan species Aglaophenia kirchenpaueri, Aglaophenia octodonta, Aglaophenia tubiformis, Halopteris diaphana, Plumularia setacea, Ventromma halecioides and 20 colonies of the bryozoan Myriapora truncata were collected by SCUBA diving along the Ionian coast of Apulia, Italy (Otranto Channel; 40° 08′ 39.8″ N, 18° 30′ 23.3″ E; Fig. 1) at 0–2 m of depth from May until November 2007. These species were selected because they do not emit their own light and avoid any interference with the epifluorescence microscope observations [11]. Colonies were transported in the laboratory under controlled temperature (5°C) and processed for isolation of luminous bacteria within 4 h from collection.
Epifluorescence Microscope Observation
Taking into account that luminous bacteria are able to emit fluorescence (excitation wavelength = 450 nm; emission wavelength = 540 nm), we utilized an epifluorescence microscope for our observations. Both the hydrozoans and the bryozoan colonies, in particular the hydrothecae and hydrocauli for hydrozoans and the opercula for bryozoan, were observed using a Zeiss Standard Axioplan microscope equipped with a halogen lamp (Hg 100) light. Blue light excitation with a BP 485/20 excitation filter, a FT 510 chromatic beam splitter and a LP 520 barrier filter were used to observe the slides prepared from each sample for light-emitting bacteria.
Phenotypic Characterization of Bacterial Isolates
In the laboratory, hydroids and bryozoans were washed in sterile seawater (0.2 μm pore-filtered). Hydrocauli-bearing hydrocladia of the hydroid species examined to obtain about 3,000–3,500 polyps (approximately 1 g) and calcified stolons of M. truncata (approximately 1 g) were diluted with an equal volume of phosphate-buffered peptone water (Beckton Dickinson and Company) and homogenized for 90 s in a sterile Waring blender at 5°C. Serial dilutions of each homogenate were plated onto Marine Agar 2216 (Beckton Dickinson and Company; seeding with 0.1 ml). The plates were incubated for 24–48 h at 30°C. At the end of the incubating period, bioluminescence was detected in a dark room from the emission of visible light. The luminous bacterial colony types were picked out and streaked onto Marine Agar to obtain pure cultures. Phenotypic identification followed the schemes of Baumann and Schubert [7], Alsina and Blanch [3], and Garrity et al. [22]. Tests performed were: Gram stain, growth in thiosulfate–citrate–bile salts–sucrose agar (TCBS; Beckton Dickinson and Company), bioluminescence on luminescence agar [62], oxidase and catalase assays, sensitivity to the vibriostatic agent O/129, OF test, amino acids decarboxylase reaction. Growth at 0%, 3%, and 8.0% NaCl, growth at 4°C, 30°C, and 35°C, indole production, gelatinase production, lipase hydrolysis, Voges–Proskauer test, utilization of citrate, and acid production from inositol, arabinose, and sucrose were also determined. The assay of Berger and Reynolds [8] was employed to determine the chitinolytic activity of the isolates. A further characterization was done by screening the utilization of different carbon sources in a concentration of 1% (l-arabinose, d-glucosamine, xylose, mannose, arabinose, cellobiose, glucose, galactose, trehalose, melibiose, lactose, mannitol, sorbitol, inositol, and sucrose) added in Bacto Phenol Red Agar Base (Beckton Dickinson and Company) with the addition of NaCl to give a final concentration of 2.5%.
16S rRNA Gene Sequence Analysis of Bacterial Isolates
High-molecular-weight genomic DNA from the different bacterial isolates was prepared according to standard procedures. Strains were grown in 100-ml nutrient broth (Beckton Dickinson and Company) containing 3% NaCl with shaking at 30°C to an optical density at 550 nm of 0.8. After centrifugation, pellets were washed with 50 ml of sodium chloride–Tris–ethylenediaminetetra-acetic acid (EDTA) buffer (500 mM NaCl, 50 mM Tris–Cl pH 8, 5 mM EDTA) and then resuspended in 4 ml of a solution containing 50 mM Tris–Cl pH 8, 25% sucrose, 1 mM EDTA. Lysozyme (1 mg ml−1) treatment was carried out at 0°C for 10 min, then EDTA was added to a final concentration of 40 mM, and samples were incubated at 0°C for 10 min. Proteinase K (100 μg ml−1) treatment was performed for 2 h at 65°C after addition of sodium dodecyl sulfate (SDS) to a final concentration of 1%. Nucleic acids were extracted by phenol-chloroform and isoamylic alcohol (24:1) extraction according to standard procedures [48], and 15 μg ml−1 ribonuclease A were used to remove RNA. After phenol-chloroform and isoamylic alcohol (24:1) extraction and ethanol precipitation, high-molecular-weight DNA was collected by spooling using Shepherd's crooks [48].
The 16S-rRNA-encoding genes were amplified using the bacterial-specific primers 16SEB20-42-F (5′-TGGCTCAGGAYGAACGCTGGCGG-3′) and 16SEB1488-R (5′-TACCTTGTTACGACTTCACC-3′) [58]. The two primers were designed to amplify a 1,488-bp-long DNA fragment (from nucleotide 20 to nucleotide 1507 in the Escherichia coli 16S rRNA gene). Polymerase chain reactions (PCR) were performed as follows: initial denaturation at 95°C for 5 min, followed by 30 cycles of denaturation at 95°C for 45 s, annealing for 30 s at 58°C and extension at 72°C for 2 min. They were carried out in a Perkin-Elmer Cetus DNA Thermal Cycler 2400. PCR products were isolated through 1% agarose gels in 1X Tris–acetate-EDTA buffer (40 mM Tris–acetate, 1 mM EDTA, pH 8.0), recovered by using the Quiaex II Gel extraction kit (Quiagen), cloned into pGEM® -T Easy Vector by using the TA Cloning kit (Promega), and finally sequenced using primers M13F and M13R as a service by MWG Biotech Custom Sequencing Service (Germany) or by Primm s.r.l (Italy).
DNA similarity searches were carried out using Basic Local Alignment Search Tool [4] at National Center for Biotechnology Information (http://www.ncbi.nlm.nih.gov/). Sequence alignments were performed with Clustal W [53] at European Bioinformatics Institute (http://www.ebi.ac.uk/). Phylo_win software [21] was used to infer evolutionary trees according to neighbor-joining methods [47] using the Kimura two-parameter model [29]. Tree robustness was assessed by bootstrap resampling (1,000 replicates each) [13].
DNA–DNA Hybridization Analyses
To perform DNA–DNA hybridization experiments, the genomic DNAs (5 μg) from the different bacterial isolates were restricted with BamHI, serially diluted in a buffer containing 2 × sodium citrate–sodium chloride (SSC buffer) [48] and 50% (v/v) formamide, heated at 95°C for 5 min, and immobilized onto nylon filters by slow filtration in a slot-blot apparatus (Schleicher and Schull) in duplicate. A prehybridization step was performed by adding a solution containing 5 × SSC (1 × SSC: 0.015 M NaCl, 15 mM Na–citrate [pH7]), 5 × Denhardt's solution [48], 0.1% SDS, 50 mM sodium phosphate buffer (pH 6.5), 50% (v/v) formamide, 500 μg denatured salmon sperm DNA per milliliter for 6 h at 36°C. This temperature represents stringent conditions for V. harveyi, for which the optimal renaturation temperature (34.05°C) is calculated as [(0.51 × G + C content) + 47] – 36 [23], where 36°C is the correction for the presence of 50% formamide [36]. A G + C content of 45.2 mol% was assumed for V. harveyi according to previous study [54]. Hybridization was carried out for 12 h at 36°C, a hybridization buffer similar to the prehybridization solution but containing 32P-labeled BamHI-restricted genomic DNAs from the different Vibrio strains in place of salmon sperm DNA. The genomic DNA from the type strain V. harveyi LMG 4044T was used as a reference. The BamHI-restricted genomic DNAs were labeled with 32P by random priming according to standard procedures [48]. After hybridization, filters were washed three times with a solution containing 2 × SSC, 0.2% SDS at 36°C, and then subjected to autoradiography. The semiquantitative analysis of the hybridization signals was performed by densitometry using a Scanmaster 3 (Howtek, Inc., Hudson, NH, USA), a high-performance desktop flatbed color scanner equipped with RFLPrintTM (Pdi, Huntington Station, NY, USA) software package or by directly counting the radioactivity bands by a PhosphoImager SI (Molecular Dynamics, Inc., Sunnyvale, CA, USA).
GenBank Accession
EU031643: V. fischeri HD-1, partial 16S rRNA gene; EU031644: V. fischeri AT-1, partial 16S rRNA gene; EU031646: V. harveyi AK-1, partial 16S rRNA gene; EU031649: Vibrio sp. MT-1, partial 16S rRNA gene; EU031648: Vibrio sp. PS-1, partial 16S rRNA gene; EU031647: Vibrio sp. VH-1, partial 16S rRNA gene.
Scanning Electron Microscopy on Aglaophenia octodonta
Twenty colonies of A. octodonta were washed several times with sterile seawater (0.2 μm pore-filtered) to assure the elimination of any bacteria settled on the surfaces and then fixed in 2.5% glutaraldehyde in 0.1 M cacodylate buffer (CB), pH 7.5, overnight. The colonies were then washed three times in CB and postfixed for 1 h in 2% osmium tetroxide in CB. After washing, the colonies were dehydrated in a graded acetone series, dried in a critical-point drier, and coated with gold. Colonies were observed with a Philips 515 scanning electron microscope (SEM) operated at 20 kV.
Quantitative Analysis of Bacteria Living on Aglaophenia octodonta Surface
In the laboratory, A. octodonta colonies (approximately 1 g) were gently washed in sterile seawater (0.2 μm pore-filtered) to assure the elimination of any bacteria settled on the surfaces, suspended in sterile seawater and sonicated three times (Branson Sonifier 2200, 60 W, 47 kHz for 1 min in an ice bath) to optimize surface bacteria detachment. The sonication was interrupted for 30 s every minute, during which time the samples were shaken manually. To enumerate surface bacteria including luminous bacteria, 1 or 5 ml of the sonicated sample and appropriate decimal dilutions were plated in parallel onto Marine Agar 2216 or TCBS Agar (Beckton Dickinson and Company) and after incubation for 2 days at 22°C and at 30°C, respectively, the culturable bacteria were counted according to the colony-forming units (CFU) method.
PCR–Single-Strand Conformational Polymorphism Analysis
Bacteria were recovered from A. octodonta colonies (about 1 g) surface and high-molecular-weight genomic DNA was extracted as described [17, 58]. Due to limited amount of the microbial sample (about 3 × 104 CFU/g), the protocol for DNA extraction was adapted to smaller (20-fold) volumes. After the extraction, the DNA was amplified using the bacteria-specific primers Com1-F (5′-CAGCAGCCGCGGTAATAC-3′) and Com2-R (5′-CCGTCAATTCCTTTGAGTTT-3′) [30]. These primers were designed to amplify 409-bp-long DNA fragments (from nucleotide 519 to nucleotide 927 in the E. coli 16S rRNA gene) that could be resolved by single-strand conformational polymorphism (SSCP) as described [17, 58]. To this purpose, PCR products were purified by High Pure PCR product Purification Kit (Boehringer Mannheim, Germany), denatured and resolved on 10% polyacrylamide gel (acrylamide–N,N-methylenebisacrylamide 49:1) in 0.8X Tris–borate–EDTA (72 mM Tris–borate, 1.6 mM EDTA) containing 5% glycerol. Bands identified after silver staining were excised with razorblades and single-strand DNAs were eluted from the gel by using the Quiaex II DNA purification kit (Quiagen). The resulting DNAs were reamplified, cloned into pGEM® -T Easy Vector by using the TA Cloning kit (Promega), and finally sequenced using primers M13F and M13R as a service by MWG Biotech.
Results
Microscope Evidence for Luminous Bacteria Biofilm in Hydrozoa and Bryozoa Species
In thecate hydroids, such as A. kirchenpaueri (Fig. 2a), A. octodonta (Fig. 2c), A. tubiformis (Fig. 3a), H. diaphana (Fig. 3c), P. setacea (Fig. 4a), V. halecioides (Fig. 4c), hydrocauli and hydrocladia are ectoendodermal tubular prolongations of hydranths' gastric cavities, enveloped by a protective chitinous layer, or perisarc. Perisarcal structures are complex, being mainly composed of chitin and proteins; in some Hydrozoa, they are associated with calcareous elements (coenosteum). The chitinous perisarc forms a solid theca around the hydranths (the hydrotheca), the reproductive organs (the gonotheca), and the protective polyps, or dactylozooids (the dactylotheca or nematotheca). All the specimens of the examined hydroid species, observed under blue light excitation, showed a clear fluorescence on the external side of the perisarc (chitinous exoskeleton) around hydrocladia. In particular, this fluorescence was concentrated in the folds along the hydrocaulus and at the base of the hydrothecae (Figs. 2b,d, 3b,d, and 4b,d). Localization of fluorescence in the chitinous structures of these hydroids suggested, as previously observed in A. octodonta [50], an involvement of luminous bacteria known for their capacity to elaborate an extracellular chitinase and feed on the chitinous structures of supporting organisms [6]. This hypothesis was supported by isolation of luminous bacteria from all hydrozoan homogenates. Moreover, an interesting issue is that all the bacterial isolates were chitinase positive. Taking into account that in the bryozoan M. truncata, the operculum is made up of chitin, we checked and found luminous bacteria on the skeletal structure near to the opercula. The halo of fluorescence observed in the invertebrates studied was due to the development of a bacterial biofilm. Indeed, when the examined species were treated with antibiotics, the fluorescence disappeared suggesting the involvement of bacteria (data not shown).
A. kirchenpaueri photomicrographs, living material: a hydrothecae at transmitted light, b hydrothecae at epifluorescence, the halo of fluorescence is due to bacterial biofilm; A. octodonta photomicrographs, living material c hydrothecae at transmitted light, d hydrothecae at epifluorescence, the halo of fluorescence is due to bacterial biofilm. Scales: (a–d) 500 μm
A. tubiformis photomicrographs, living material: a hydrotheca at transmitted light, b hydrotheca at epifluorescence, the halo of fluorescence is due to bacterial biofilm; H. diaphana photomicrographs, living material: c hydrotheca at transmitted light, d hydrotheca at epifluorescence, the halo of fluorescence is due to bacterial biofilm. Scales: (a and b) 200 μm, (c and d) 250 μm
P. setacea photomicrographs, living material: a particular of hydrocladia at transmitted light, b particular of hydrocladia at epifluorescence, the halo of fluorescence is due to bacterial biofilm; V. halecioides photomicrographs, living material: c particular of hydrocladia at transmitted light, d particular of hydrocladia at epifluorescence, the halo of fluorescence is due to bacterial biofilm. Scales: (a and b) 100 μm, (c) 500 μm, (d) 250 μm
Identification of Luminous Bacteria
All the isolated luminous bacteria were identified on the basis of both phenotypic and genotypic analysis. Five independent isolates were examined from each type of animal. The identity of the isolates was investigated by phylogenetic analysis using almost the entire sequences of the 16S-rRNA-encoding genes (Fig. 5). All the 16S rRNA sequences of luminescent vibrios from A. tubiformis and H. diaphana were identical and exhibited an identity of 99.8% to the 16S rRNA sequence of the V. fischeri type strain ATCC 7744T. These isolates were named, respectively, V. fischeri AT1 to AT5 and V. fischeri HD1 to HD5. The isolates from A. kirchenpaueri (AK1 to AK5) grouped with the V. harveyi type strains LMG 4404T and ATCC 35084T, while those from P. setacea (PS1 to PS5), V. halecioides (VH1 to VH5), and M. truncata (MT1 to MT5) clustered with V.-harveyi-related strains LMG 20370 and R-14913. These two strains were previously assigned to the species V. harveyi on the basis of phenotypic criteria but later suspected to belong to a new species on the basis of DNA–DNA similarity, amplified fragment length polymorphism, and multilocus sequence analyses [24, 52, 55]. 16S rRNA gene sequences identities between AK1–AK5, PS1–PS5, VH1–VH5, MT1–MT5, and the V. harveyi type strain ATCC 35084T were, respectively, 99.7%, 98.1%, 97.7%, and 97.9%. On the basis of these results, only the five isolates AK1–AK5 could be unambiguously assigned to the species V. harveyi and indicated as V. harveyi AK1 to AK5, while PS1–PS5, VH1–VH5 and MT1–MT5 were assigned to a new V.-harveyi-related species together with the luminous Vibrio species (Vibrio sp. AO1 in Fig. 5) from A. octodonta previously described [50].
Phylogenetic analysis of 16S rRNA gene sequences of luminous Vibrio species isolated from the hydroids: taxonomic position of V. fischeri HD1 and V. fischeri AT1, V. harveyi AK1, and Vibrio sp. MT1, VH1, and PS1 is indicated with respect to several type strains. AO1 is a Vibrio species previously isolated form A. octodonta [50]. Bootstrap values ≥60 are reported at the branch points
This conclusion was supported by DNA–DNA hybridization experiments under stringent conditions (see “Methods”) using the DNA from V. harveyi type strain ATCC 4404T as a reference. AK1–AK5 were confirmed to belong to the species V. harveyi because the DNA similarity value with V. harveyi ATCC 4404T was 91.2%, while values below the 70% proposed by Wayne et al. [61] to delimit a species were obtained with the other isolates (Table 1).
The assignment of the bacterial isolates to the above-mentioned species was consistent with the results of morphological, cultural, and biochemical tests (Table 2) according to the schemes proposed by Baumann and Schubert [7], Alsina and Blanch [3], Garrity et al. [22], and Farmer et al. [19]. However, due to lower resolution with respect to phylogenetic and DNA–DNA hybridization analyses, these tests were not able to discriminate between the V. harveyi and the V.-harveyi-related isolates.
Scanning Electron Microscopy and Quantitative Estimation of A. octodonta Surface Bacteria
We further investigated the specificity of the association between hydroids and epibiotic bacteria by both SEM inspection of A. octodonta surface and quantitative assessment of luminous bacteria on the hydroid perisarc by using culture-based and culture-independent approaches. We chose A. octodonta as a model because, among the hydroids employed in this study, it is the most common along rocky coasts of the Mediterranean Sea [50]. SEM analysis demonstrated the adhesion of several vibrios to the hydroid perisarc (magnification ×500; Fig. 6a). Further magnification (×25,000) showed that the bacteria observed had the typical morphology of vibrios (Fig. 6b).
The results of cultural analysis using in parallel Marine Agar 2216 and TCBS Agar demonstrated that luminous vibrios represented a conspicuous component (6 × 103 CFU/g, 20%) of the total culturable A. octodonta surface bacteria (3 × 104 CFU/g) that was dominated by Vibrio species (>80%). This finding was supported by culture-independent approach (Fig. 7). To this purpose, three distinct batches of A. octodonta (sampled in May, August, and November 2007) were analyzed for the presence of epibiotic bacteria by PCR–SSCP using bacteria-specific primers Com1-F and Com2-R. These primers target a 409-bp-long (in E. coli) central region of prokaryotic 16S rRNA gene and allow high taxonomic resolution, often at the species level [30]. The SSCP profile of the amplified 16S rRNA gene pool revealed a relatively low degree of complexity of the microbial community with predominance of few common bands in all samples (a–d; Fig. 7, lanes 1–3). These bands were excised from the gel, eluted, cloned, and subjected to nucleotide sequence analysis. To avoid any bias due to the reamplification step, only PCR products showing the same SSCP profile as the original bands were further processed. The sequence corresponding to bands a, when challenged in the databank, poorly matched (81% of identity) with that of Rubritalea marina strain Pol012 (accession number DQ302104.1) and other Verrucomicrobiaceae, suggesting that it might belong to an organism phylogenetically distant from any known bacterial division. Bands b and d represented to the two complementary DNA strands of the 16S rRNA gene whose sequence exhibited the highest similarity (91.2%) with that of an unknown microorganism from deep-sea octocoral phylogenetically close to as-yet-uncultivated Spirochaeta (accession number DQ395500.1). Band c comigrated with the major signal (band c') resulting from PCR amplification of luminous Vibrio sp. AO1 DNA (Fig. 7, lane 4). Indeed, when subjected to nucleotide sequencing, band c and c' gave the same result corresponding to 16S rRNA gene of the Vibrio core group including the species V. harveyi (and V.-harveyi-related), Vibrio alginolyticus, Vibrio pelagius, Vibrio natriegens, and Vibrio mytili.
PCR–SSCP analysis of the bacterial community colonizing A. octodonta surface. Three distinct batches of A. octodonta sampled in May, August, and November (lanes 1–3, respectively) were analyzed for the presence of epibiotic bacteria by PCR–SSCP using 16S rRNA gene-specific primers. The PCR–SSCP profiles were compared with that obtained by amplifying the DNA from Vibrio sp. AO1, the luminous strain isolated from the hydroid, with the same primers. Arrowheads mark the positions of specific DNA bands (a, b, c, d, c') that were eluted from the gel, cloned, and subjected to nucleotide sequence analysis
Discussion
Available information on the interaction of luminous bacteria with marine invertebrates is limited [42]. By contrast, a wide literature is available on the symbiotic colonization of V. fischeri and the Hawaiian bobtail squid, Euprymna scolopes [10, 16, 35, 46, 63]. In this association, newly hatched squids acquire V. fischeri from the surrounding seawater in which they are present at a few hundred CFU per milliliter [33] in a total background of about 106 other marine bacteria per milliliter. The relationship between V. fischeri and its squid host is complex and highly specific. The specificity of the association suggests the specialized colonization mechanisms in the bacterial symbiont have coevolved with cognate recognition mechanisms in the squid host [59].
As regards the interaction between the luminous bacteria (V. fischeri and V. harveyi) and hydrozoans or bryozoan observed in our study, the exact molecular mechanism involved is still unknown. All the investigated invertebrate species are characterized by the presence of chitinous surface on which the luminous bacteria identified could find a suitable substrate. Because the luminous bacteria isolated showed a chitinolytic activity, the results of the present study support the original hypothesis of Hood and Meyers [25] that a primary role of some vibrios could be the colonization and initiation of the degradation of chitinous material in aquatic ecosystems.
In the present study, V. fischeri, V. harveyi, and V.-harveyi-related species were the luminous bacteria isolated with the selected hydrozoans and bryozoan. V. harveyi is almost ecologically identical to V. fischeri. Almashanu et al. [2] observed that the β subunit of V. fischeri luciferase forms active enzyme with the α subunit of V. harveyi luciferase showing the affinity of these two bacterial species. This result supports the specificity of the interaction observed in our study between these luminous bacteria and the investigated invertebrates: in fact, among the strains of luminous bacteria, only V. fischeri, V. harveyi, and V.-harveyi-related species were found on the surfaces of the examined invertebrates. Moreover, quantitative analyses demonstrated that luminous vibrios represent a conspicuous component of the total culturable bacteria colonizing the surface of A. octodonta (about 20% of the total count). This conclusion was consistent with the result of the PCR–SSCP analysis of 16S rRNA gene pool of the local bacterial community. In addition to bacteria belonging to the Vibrio core group (including the luminous V.-harveyi-related strain AO1), this analysis revealed the presence of at least two additional uncultivated microorganisms phylogenetically distant from any known species (bands b or d) or bacterial division (band a; Fig. 5). It is noteworthy that PCR–SSCP patterns were very similar irrespective of the sampling time (May to November). Thus, as observed for V. fischeri and the Hawaiian bobtail squid E. scolopes, also in the case of A. octodonta, the relationship between luminous V.-harveyi-related species and this hydroid is highly specific. This issue was also supported by scanning electron microscopy images demonstrating the colonization of this hydroid perisarc by vibrios (Fig. 6a).
The ancestral position of the Hydrozoa with respect to the Metazoa [9] suggests that these invertebrates functioned from the past until now as habitat “islands” providing a unique set of environmental conditions for the survival of the luminous bacteria isolated over the time [50]. The possibility of using chitin as a feeding substrate evidently exploited other chitin-coated Metazoa, from the Crustacea (as is well known for copepods) to the Bryozoa (as shown here).
Further studies will be undertaken to investigate the molecular mechanisms involved in such host–bacteria relationships and to better understand whether the interaction observed is only related to chitin utilization by V. fischeri, V. harveyi, and the V.-harveyi-related species (parasitic association) or if it has also an ecological role for the examined hydrozoan and bryozoan species. None of the invertebrate species studied, in fact, is able to emit its own light. Thus, the presence of luminous bacteria, able to emit light continuously, probably provides a behaviorally useful light source (symbiotic association) as already suggested [50].
Furthermore, both luminous bacterial isolated are considered opportunistic pathogens, as suggested by the name of the disease commonly referred to as luminous vibriosis: the expression of luminescence has long been associated with the virulence in pathogenic strains of these bacteria. In particular, V. harveyi is now known to cause mass mortalities in several marine organisms including pearl oysters (Pinctada maxima) [39], fish (Solea senegalensis) [64], sea horses (Hippocampus sp.) [1], and lobsters (Panulirus homarus) [26]. Virulence in V. harveyi has been attributed the production of an extracellular protein referred to as toxin T1 [40]. V. harveyi and V. fischeri, together with other luminous vibrios, have been involved in disease outbreaks in shrimp larviculture facilities [31] and, to a lesser extent, in grow-out ponds [32] as well as penaeid prawn farms across the world [57]. In this framework, the interaction between luminous bacteria with A. kirchenpaueri, A. octodonta, A. tubiformis, H. diaphana, P. setacea, V. halecioides, and M. truncata might have epidemiological implications because these hydrozoan and bryozoan species are widely distributed in the Mediterranean Sea and might constitute natural reservoirs of the pathogens for marine organisms.
Moreover, the interaction between the above-mentioned invertebrates and luminous bacteria provides an unusually tractable system for studying the evolution of these relationships because the selected species have a wide biogeographic distribution.
References
Alcaide E, Gil Sanz C, Esteve D, Sanjuan D, Amaro C, Silveira L (2001) Vibrio harveyi disease in seahorse, Hippocampus sp. J Fish Dis 2:311–313
Almashanu S, Gendler I, Hadar R, Kuhn J (1996) Interspecific luciferase b subunit hybrids between Vibrio harveyi, Vibrio fischeri and Photobacterium leiognathi. Prot Eng 9:803–809
Alsina M, Blanch AR (1994) A set of keys for biochemical identification of environmental Vibrio species. J Appl Bacteriol 76:79–85
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Baumann P, Baumann L (1984) Genus II. Photobacterium Beijerinck 1889. In: Kreig NR, Holt JG (eds) Bergey's manual of systematic bacteriology, vol. 1. Williams & Wilkins, Baltimore
Baumann P, Baumann L, Bang SS, Woolkalis MJ (1980) Revaluation of the taxonomy of Vibrio, Beneckea, and Photobacterium-abolition of the genus Beneckea. Curr Microbiol 4:127–133
Baumann PS, Schubert RHW (1984) The family II Vibrionaceae Veron. In: Krieg NR (ed) Bergey's manual of systematic bacteriology, vol. 1. Williams & Williams, Baltimore
Berger LR, Reynolds DM (1958) The chitinase system of a strain of Streptomyces griseus. Biochem Biophys Acta 29:522–534
Boero F, Gravili C, Pagliara P, Piraino S, Bouillon J, Schmid V (1998) The cnidarian premises of metazoan evolution: from triploblasty, to coelom formation, to metamere. Ital J Zool 65:5–9
Boettcher KJ, Ruby EG (1990) Depressed light emission by symbiotic Vibrio fischeri of the sepiolid squid Euprymna scolopes. J Bacteriol 172:3701–3706
Bouillon J (1995) Classe des Hydrozoaires (Hydrozoa Owen, 1843). In: Grassé PP, Doumenc D (eds) Traité de Zoologie. Masson, Paris
Bouillon J, Medel MD, Pagès F, Gili JM, Boero F, Gravili C (2004) Fauna of the Mediterranean Hydrozoa. Sci Mar 68:5–438
Brown JK (1994) Bootstrap hypothesis tests for evolutionary trees and other dendrograms. Proc Natl Acad Sci U S A 91:12293–12297
Carli A, Pane L, Casareto L, Bertone S, Pruzzo C (1993) Occurrence of Vibrio alginolyticus in Ligurian coast rock pools (Tyrrhenian Sea, Italy) and its association with the copepod Tigriopus fulvus (Fisher 1860). Appl Environ Microbiol 59:1960–1962
Carman KR, Dobbs FC (1997) Epibiotic microorganisms on copepods and other aquatic crustaceans. Micros Res Tech 37:116–135
DeLoney CR, Bartley TM, Visick KL (2002) Role for phosphoglucomutase in Vibrio fischeri–Euprymna scolopes symbiosis. J Bacteriol 184:5121–5129
Di Giacomo M, Paolino M, Silvestro D, Vigliotta G, Imperi F, Visca P, Alifano P, Parente D (2007) Microbial community structure and dynamics of dark fire-cured tobacco fermentation. Appl Environ Microbiol 73:825–837
Ducklow HW, Mitchell R (1979) Bacterial populations and adaptations in the mucus layers on living corals. Limnol Oceanogr 24:4715–4725
Farmer JJ III, Janda JM, Brenner FW, Cameron DN, Birkhead KM (2005) Genus 1. Vibrio Pacini 1854, 411AL. In: Brenner DJ, Krieg NR, Staley JT (eds) Bergey's manual of systematic bacteriology the proteobacteria part B the gammaproteobacteria vol. 2. 2nd edn. Springer, New York
Fletcher M, Marshall KC (1982) Are solid surfaces of ecological significance to aquatic bacteria? Adv Microbial Ecol 6:199–236
Galtier N, Gouy M, Gautier C (1996) SEAVIEW and PHYLO_WIN: two graphic tools for sequence alignment and molecular phylogeny. Comput Appl Biosci 12:543–548
Garrity G, Brenner DJ, Krieg NR, Staley JR (2005) Bergey's manual of systematic bacteriology, 2nd edn. Williams & Wilkins, Baltimore
Gillis M, De Ley J, De Cleene M (1970) The determination of molecular weight of bacterial genome DNA from renaturation rates. Eur J Biochem 12:143–153
Gomez-Gil B, Soto-Rodríguez S, García-Gasca A, Roque A, Vazquez-Juarez R, Thompson FL, Swings J (2004) Molecular identification of Vibrio harveyi-related isolates associated with diseased aquatic organisms. Microbiology 150:1769–1777
Hood MA, Meyers SP (1977) Microbiological and chitinoclastic activities associated with Penaeus setiferus. J Oceanogr Soc Jpn 33:235–241
Jawahar AT, Keleemur RM, Leema JMT (1996) Bacterial disease in cultured spiny lobster, Panulirus homarus (Linnaeus). J Aquacult Trop 11:187–192
Johnson CR, Muir DG, Reysenbach AL (1991) Characteristic bacteria associated with surfaces of coralline algae: a hypothesis for bacterial induction of marine invertebrate larvae. Mar Ecol Prog Ser 74:281–294
Kaneko T, Colwell RR (1975) Adsorption of Vibrio parahaemolyticus onto chitin and copepods. Appl Microbiol 29:269–274
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Lane DJ, Pace B, Olsen GJ, Stahl DA, Sogin ML, Pace NR (1985) Rapid determination of 16S ribosomal RNA sequences for phylogenetic analyses. Proc Natl Acad Sci U S A 82:6955–6959
Lavilla-Pitogo CR, Leano EM, Paner MG (1990) Occurrence of luminous bacterial disease of Penaeus monodon larvae in the Philippines. Aquaculture 91:1–13
Lavilla-Pitogo CR, Leano EM, Paner MG (1998) Mortalities of pond-cultured juvenile shrimp, Penaeus monodon, associated with dominance of luminescent vibrios in the rearing environment. Aquaculture 164:337–349
Lee K-H, Ruby EG (1995) Symbiotic role of the viable but nonculturable state of Vibrio fischeri in Hawaiian coastal seawater. Appl Environ Microbiol 61:278–283
Maugeri TL, Carbone M, Fera MT, Irrera GP, Gugliandolo C (2004) Distribution of potentially pathogenic bacteria as free living and plankton associated in a marine coastal zone. J Appl Microbiol 97:354–361
McCann J, Stabb EV, Millikan DS, Ruby EG (2003) Population dynamics of Vibrio fischeri during infection of Euprymna scolopes. Appl Environ Microbiol 69:5928–5934
McConaughy BL, Laird CD, McCarthy BJ (1969) Nucleic acid reassociation in formamide. Biochemistry 8:3289–3295
Nealson HK, Haygood GM, Tebo MB, Roman M, Miller E, McCosker EJ (1984) Contribution by symbiotically luminous fishes to the occurrence and bioluminescence of luminous bacteria in seawater. Microb Ecol 10:69–77
Nishiguchi MK, Ruby EG, McFall-Ngai MJ (1998) Competitive dominance among strains of luminous bacteria provides an unusual form of evidence for parallel evolution in sepiolid squid–Vibrio symbioses. Appl Environ Microbiol 64:3209–3213
Pass DA, Dybdahl R, Mannion MM (1987) Investigation into the causes of mortality of the pearl oyster, Pinctada maxima (Jamson), in Western Australia. Aquaculture 65:149–169
Pizzuto M, Hirst RG (1995) Classification of isolates of Vibrio harveyi virulent to Penaeus monodon larvae by protein profile analysis and M13 DNA fingerprinting. Dis Aquat Org 21:61–68
Pujalte MJ, Ortigosa M, Macian MC, Garay E (1999) Aerobic and facultative anaerobic heterotrophic bacteria associated to Mediterranean oysters and seawater. Int Microbiol 2:259–266
Ramesh A, Venugopalan VK (1984) Colloque International de bactériologie marine. Actes de colloques. IFREMER, CNRS, Brest, pp 1–5
Riedl R (1970) Fauna und Flora der Adria, 2nd edn. Verlag Paul Parey, Hamburg
Rosowski JR (1992) Specificity of bacterial attachment sites on the filamentous diatom Navicula confervacea (Bacillariophyceae). Can J Microbiol 38:676–686
Roszak DB, Colwell RR (1987) Survival strategies of bacteria in the natural environment. Microbiol Rev 51:365–379
Ruby EG, Lee K-H (1998) The Vibrio fischeri–Euprymna scolopes light organ association: current ecological paradigms. Appl Environ Microbiol 64:805–812
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sambrook J, Russel DW (2001) Molecular cloning in laboratory. Manual, 3rd edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Santavy DL, Colwell RR (1990) Comparison of bacterial communities associated with the Caribbean sclerosponge Ceratoporella nicholsoni and ambient seawater. Mar Ecol Prog Ser 67:73–82
Stabili L, Gravili C, Piraino S, Boero F, Alifano P (2006) Vibrio harveyi associated with Aglaophenia octodonta (Hydrozoa, Cnidaria). Microb Ecol 52:603–608
Tamplin ML, Gauzens AL, Huq AL, Sack DA, Colwell RR (1990) Attachment of Vibrio cholerae serogroup O1 to zooplankton and phytoplankton of Bangladesh waters. Appl Environ Microbiol 56:1977–1980
Thompson FL, Gevers D, Thompson CC, Dawyndt P, Naser S, Hoste B, Munn CB, Swings J (2005) Phylogeny and molecular identification of vibrios on the basis of multilocus sequence analysis. Appl Environ Microbiol 71:5107–5115
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Thompson FL, Hoste B, Vandemeulebroecke K, Engelbeen K, Denys R, Swings J (2002) Vibrio trachuri Iwamoto et al. 1995 is a junior synonym of Vibrio harveyi (Johnson and Shunk 1936) Baumann et al. 1981. Int J Syst Evol Microbiol 52:973–976
Thompson FL, Hoste B, Vandemeulebroecke K, Swings J (2001) Genomic diversity amongst Vibrio isolates from different sources determined by fluorescent amplified fragment length polymorphism. Syst Appl Microbiol 24:520–538
Venkateswaran K, Kim SW, Nakano H, Onbè T, Hashimoto H (1989) The association of Vibrio parahaemolyticus serotypes with zooplankton and its relationship with bacterial indicators of pollution. Syst Appl Microbiol 11:194–201
Vidgen M, Carson J, Higgins M, Owens L (2006) Changes to the phenotypic profile of Vibrio harveyi when infected with the Vibrio harveyi myovirus-like (VHML) bacteriophage. J Appl Microbiol 100:481–487
Vigliotta G, Nutricati E, Carata E, Tredici SM, De Stefano M, Pontieri P, Massardo DR, Prati MV, De Bellis L, Alifano P (2007) Clonothrix fusca Roze 1896, a filamentous, sheathed, methanotrophic gamma-proteobacterium. Appl Environ Microbiol 73:3556–3565
Visick KL, McFall-Ngai MJ (2000) An exclusive contract: specificity in the Vibrio fischeri–Euprymna scolopes partnership. J Bacteriol 182:1779–1787
Walls JT, Ritz DA, Blackman MJ (1993) Fouling, surface bacteria and antibacterial agents of four bryozoan species found in Tasmania, Australia. Exp Mar Biol Ecol 169:1–13
Wayne LG, Brenner DJ, Colwell RR, Grimont PAD, Kandler O, Krichevsky MI, Moore LH, Moore WEC, Murray RGE, Stackebrandt E, Starr MP, Trüper HG (1987) Report of the ad hoc committee on reconciliation of approaches to bacterial systematics. Int J Syst Bacteriol 37:463–464
West PA, Colwell RR (1984) Identification and classification of vibrionaceae: an overview. In: Colwell RR (ed) Vibrios in the environment. Wiley, New York
Whistler CA, Ruby EG (2003) GacA regulates symbiotic colonization traits of Vibrio fischeri and facilitates a beneficial association with an animal host. J Bacteriol 185:7202–7212
Zorrilla I, Arijo S, Chabrillon M, Diaz P, Martinez-Manzanares E, Balebona MC, Morinigo MA (2002) Vibrio species isolated from diseased farmed sole, Solea senegalensis. J Fish Dis 26:103–108
Acknowledgements
Financial support was provided by MURST (COFIN and FIRB projects) and the European Community (MARBEF and IASON networks). Christian Vaglio helped in the field. Thanks are due to Dr. Marcella Elia for the technical assistance in the scanning electron microscopy.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Stabili, L., Gravili, C., Tredici, S.M. et al. Epibiotic Vibrio Luminous Bacteria Isolated from Some Hydrozoa and Bryozoa Species. Microb Ecol 56, 625–636 (2008). https://doi.org/10.1007/s00248-008-9382-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-008-9382-y