Abstract
The aim of this work is to describe the diversity and phylogeny of rhizobial bacteria associated to nodules of Ononis tridentata L. in different geographical regions of Spain. Twenty-two bacterial isolates were characterized using several molecular techniques (16S amplified ribosomal deoxyribonucleic acid restriction analysis, fingerprinting, and sequencing) and phylogenies were inferred from their 16S and nodC gene sequences. Phylogenetically, the isolates grouped with the genera Rhizobium, Mesorhizobium, Phylobacterium, and Bosea. The nodC gene, essential for nodulation, was detected for the first time in isolates close to the genera Bosea and Phyllobacterium. The bacteria isolated showed a high diversity at the genus, species, and strain level regardless of the geographical origin of the host plant. This is the first report describing bacteria associated to nodules of O. tridentata. This shrub legume is highly prized for the revegetation of gypsum soils in semiarid Mediterranean areas. Our molecular description of bacteria associated to this legume improves the current understanding of the ecology of this plant species. Our findings have implications for formulating suitable bacterial inocula to recover gypsum ecosystems.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Legumes form a large plant family, widespread all over the world. Many legume species establish symbiotic relationships with soil bacteria that are capable of fixing atmospheric nitrogen in specific symbiotic organs, the nodules. Most of the legume-nodulating bacteria described so far, and known as “rhizobia,” have been classified as belonging to the alpha subgroup of the Proteobacteria and to one of the six genera: Rhizobium, Sinorhizobium, Bradyrhizobium, Azorhizobium, Mesorhizobium, and Allorhizobium [9]. However, recent studies have ascribed bacteria able to nodulate legumes to other genera within α-and β-Proteobacteria [6, 26, 38, 41, 52]. Gamma-proteobacteria were found associated with legume nodules, although their presence and role is yet to be defined [4].
As a result of their nitrogen-fixing capacity, legumes can colonize nitrogen-deficient soils enhancing their fertility, which makes them optimal candidates for revegetation programs. However, despite their economic and environmental importance, only a small proportion of existing legume species and their rhizobial associates have been investigated. So far, most of these studies have been focused on herbaceous species of agronomic interest. Notwithstanding, the past few years have witnessed a surge in rhizobial biodiversity studies performed on wild shrubby and woody legumes in different ecosystems worldwide [6, 17, 21, 26, 46, 48] including semiarid Mediterranean regions [11, 27, 29, 30, 52, 53, 54]. In the Mediterranean ecosystems, shrubby and woody legumes are an essential part of many revegetation projects owing to their ecological benefits such as improving soil fertility, preventing soil erosion, and contributing to graze animal nutrition [11, 24, 28].
Within the Mediterranean basin, gypsum steppes are valuable ecosystems (protected by the EU Habitats Directive [2]), which are highly represented in Spain by some of the largest gypsum deposits in Europe [7]. Ononis tridentata L. is one of the shrub legumes commonly found in the gypsum soils of Spain. Characteristics of gypsum soils, such as low water retention, formation of hard surface crusts, poor structure, high infiltration and ion washing, and low availability of nutrients, are usually constrictive for the development of plants [22]. The successful revegetation of this type of soils using well-adapted wild legumes depends on the use of suitable indigenous rhizobial inocula to ensure seedling performance after out-planting [5, 28, 39]. To the best of our knowledge, no study has examined the bacterial species associated to nodules of O. tridentata. Characterizing such species will be essential both for preserving biodiversity and for revegetation projects targeted at gypsum ecosystems.
In the past 20 years, advances in molecular methods have served to revise the taxonomic classification of many bacterial groups. Definition of new bacterial genera and species and the description of their evolutionary relationships have been mainly inferred from 16S ribosomal ribonucleic acid (rRNA) gene sequences [44, 51]. Particularly in the case of rhizobia, nodulation-specific genes (nif and nod) have been widely used to perform evolutionary analysis [15, 42]. The nodC gene has often been chosen as a nodulation marker in different studies because it is essential for nodulation in most of the rhizobial species examined so far [14, 20, 33, 42]. This gene encodes the enzyme involved in the first step of the Nod factor assembly, and it has been also described as a determinant of host range [23]. However, it has been recently reported that some photosynthetic Bradyrhizobial strains can nodulate in the absence of conventional nod genes [10].
The aim of our study was to gain basic knowledge on the diversity and taxonomy of the rhizobial bacteria associated to nodules of O. tridentata in different regions of Spain. Different molecular techniques for characterization of isolates and phylogenies based on the 16S rRNA and nodC genes are discussed.
Materials and Methods
Sampling Zones
Naturally occurring root nodules of the gypsophyte O. tridentata were sampled at several locations within Spanish steppe regions (rainfall not exceeding 200 mm): (a) Almería (ALM), (b) Madrid–Guadalajara (MAD), and (c) Huesca (HUE; see Table 1 for UTM coordinates). Other companion legume species were found at each location: (a) ALM: Anthyllis cytisoides L., Coronilla juncea L., Dorycnium rectum (L.) Ser. in DC., Dorycnium pentaphyllum Scop., Genista ramosissima (Desf.) Poir. in Lam. Onobrychis stenorhiza DC., and Spartium junceum L., (b) MAD: Astragalus incanus L., Medicago minima L., Dorycnium pentaphyllum Scop., and (c) HUE: Astragalus incanus L., Coronilla scorpioides L., Genista scorpius L., and Vicia amphicarpa L. Chemical analysis of soil samples from the different regions revealed the following general characteristics: pH 7.8–7.9; EC 2,070–2,300 μS/cm; OM 0.7–1.1%; N 0.04–0.08%; C 0.4–0.6%; P2O5 5.5–18 mg/100 g, and SO4 135–152 mg/100 g.
Bacterial Isolation and Culture Conditions
Fragments of roots with attached nodules were transported in their natural substrate and stored at 4°C until further processing (within a week). Nodules were surface sterilized with 1% HgCl2 for 1 min and exhaustively washed in ten changes of autoclaved distilled water. Intact surface-sterilized nodules were rolled and placed on yeast extract–mannitol (YEM) plates as controls to detect possible contaminations. Each nodule was crushed on a sterile plate, and the bacteria were isolated on YEM agar plates [45]. Isolates were grown at 28°C, and the purity of the colonies was checked by repeated streaking of single colonies on YEM plates and by microscopy examination. Pure cultures were preserved in 20% glycerol at −80°C and deposited in the culture collection of the IRN-CCMA-Consejo Superior de Investigaciones Cientificas.
Approximately 20% of the nodules processed yielded bacterial colonies (one per nodule). Among them, 22 bacterial isolates of O. tridentata showing characteristics corresponding to rhizobia were selected for subsequent molecular analysis.
Several reference strains of rhizobial species in the genera Rhizobium (R. gallicum bv. gallicum strain R602 and R. gallicum bv. phaseoli strain P4I21), Sinorhizobium (S. meliloti strains Rm41 and Rm2011), Mesorhizobium (M. loti strain NZP2037), Sinorhizobium (S. fredii strain USDA205), Ochrobactrum (O. lupini strain LUP21) and Bradyrhizobium (B. sp. (Lupinus) strain ISLU16, and B. japonicum strain USDA110) were used for performing the amplified ribosomal deoxyribonucleic (rDNA) acid restriction analysis (ARDRA) analyses.
Isolation of Genomic DNA
The bacterial isolates were grown on trypton–yeast [32] agar plates at 28°C to minimize polysaccharide production. Cells were washed twice in phosphate-buffered saline (140 mM NaCl, 2.6 mM KCl, 10 mM Na2HPO4, 1.8 mM KH2PO4), pelleted by centrifugation (13,000 rpm for 3 min), and resuspended in 567 μL Tris–edta (TE) buffer (10 mM Tris HCl, 1 mM ethylenediamine tetraacetic acid [EDTA], pH 8.0). Each bacterial cell suspension was mixed with 30 μL 10% sodium dodecyl sulfate and 3 μL proteinase K (20 mg/mL), followed by incubation for 1 h at 37°C. Next, 100 μL of 5 M NaCl and 80 μL of 10% cetyl trimethylammonium bromide (in 0.7 M NaCl) were added, and the mixture was incubated for 20 min at 65°C. Samples were extracted twice with an equal volume of phenol/chloroform/isoamyl alcohol 25:24:1 and once with chloroform/isoamyl alcohol 24:1. Finally, the DNA was precipitated with isopropanol at 4°C, washed in 70% ethanol, and resuspended in 50 μL of autoclaved distilled water.
Amplification and Sequencing of the 16S rRNA Gene
The 16S rRNA gene was amplified using the primers fD1 and rD1, which are able to amplify almost the full-length 16S rDNA sequence in many bacterial genera [49]. Primers were synthesized by MWG-BiotechAG (Spain). Polymerase chain reaction (PCR) amplification was carried out in a 50-μL volume mixing the template DNA (5 ng/μL) with polymerase reaction buffer (100 mM Tris–HCl, 15 mM MgCl2, 500 mM KCl, pH 8.3), 25 μM (each) deoxyadenosine triphosphate, deoxycytidine triphosphate, deoxythymidine triphosphate, and deoxyguanosine triphosphate, 0.1 μM of each primer fD1 and rD1, and 2.5 U of Taq DNA polymerase (Roche, Germany). The following temperature profile was used for DNA amplification: an initial denaturation step of 95°C for 5 min, followed by 35 cycles of 95°C for 1 min, 55°C for 1 min, and 72°C for 2 min, and a final extension step of 72°C for 10 min. A negative control containing 1 μL of water instead of DNA was included in every PCR run to avoid contaminations. PCR amplifications were performed in a thermal cycler PCR Express instrument (Hybaid, UK). PCR products were examined by electrophoresis on a Tris–acetate–EDTA (TAE) 1% agarose gel (90 V) and purified using the GFX™ PCR Gel Band Purification Kit (Amersham Pharmacia Biotech, USA) before sequencing. Sequencing reactions were performed using the ABI PRISM™ Dye Terminator Cycle Sequencing Kit (Applied Biosystems, USA) with the primers IRF1, 1050R, 800F, and 800R [16] and analyzed in an automatic sequencer ABI PRISM 3730 sequencer (Applied Biosystems).
ARDRA of the 16S rDNA Region and Phenetic Analysis
The amplification products of the 16S rDNA region were independently digested using four restriction endonucleases HinfI, AluI, MspI, and TaqI, according to the manufacturer’s instructions (New England Biolabs, UK). Restricted DNA was analyzed by electrophoresis on 2.5% MS4-agarose gels (Pronadisa, Spain), performing 1.5-h runs at 100 V visualized by staining with ethidium bromide. Bacterial isolates were scored for the presence or absence of each restriction fragment generated by the four restriction endonucleases. TIFF files of the electrophoretograms were compared by image analysis generating a binary 1/0 matrix used to estimate the genetic relationships among the rhizobial isolates. Jaccard similarity indices were calculated for each pair of isolates using the Bionumerics software (Applied Maths, Kortrijk, Belgium). A dendrogram was generated from the similarity matrix using the unweighted pair group method with arithmetic mean method of analysis implemented in this software [35].
tDNA-PCR/IGS-PCR Fingerprinting and Phenetic Analysis
The primer pairs T5A/T3B and L1/G1 were used for low (transfer DNA [tDNA]-PCR) and high (intergenic spacer [IGS]-PCR) resolution fingerprinting, respectively. PCR reactions were performed according to the conditions described by Hirsch and Sigmund [13]. A 7-μL sample of the PCR product of each bacterial isolate was loaded on a 4–20% acrylamide gradient minigel (Bio-Rad) and run electrophoretically for 2 h at 100 V. Each isolate was assigned to its genome haplotype, defined by the combined band patterns obtained using both fingerprinting techniques. TIFF files of the electrophoretograms were compared by image analysis generating a binary data matrix (presence or absence of bands), which was used to estimate genetic relationships among the rhizobial isolates. Phenetic analyses and dendrogram construction were conducted as described above for the ARDRA technique.
nodC Amplification and Sequencing
The nodC gene was partially amplified using the primers pair nodCF–nodCI, according to the PCR procedure described by Laguerre et al. [15]. PCR products were examined by electrophoresis on TAE 1% agarose gel (90 V) and purified using the GFX™ PCR Gel Band Purification Kit (Amersham Pharmacia Biotech). Sequencing reactions were performed using the primers nodCF–nodCI and the same conditions as those described in the section above.
Phylogenetic Analysis of the 16S rRNA and nodC Genes
As a first approach to identifying the 22 selected isolates, we performed a BLAST search on the GenBank database. Nucleotide alignments of partial 16S and nodC sequences were generated using the multiple alignment program ClustalX 1.81 [40], and the ends of the alignments were trimmed using the Se-Al v2.0a11 Carbon software [25]. The TuneClustalX software [12] was used to obtain a mean Q score, as an estimate of the accuracy of the entire alignment. Phylogenetic analysis was performed by using the neighbor-joining algorithm [31] implemented in PAUP* ver. 4.0 b10 [37]. The trees were rooted with Bacillus licheniformis (DQ351939) or Ralstonia taiwanensis (AJ505303) as the out-groups for the 16S and nodC phylogenetic analyses, respectively. The confidence of the branches was measured by bootstrapping with 1,000 replicates.
Results
Bacterial Isolation and Amplification of the 16S rDNA and nodC Gene Regions
For the present work, we selected 22 bacterial isolates representative of the three steppe regions sampled that showed phenotypic characteristics corresponding to rhizobia (Table 1). The 22 isolates formed colonies that were visible after 3–5 days. All the pure bacterial cultures were Gram negative, produced colorless to whitish and round, convex colonies, with regular margins, and abundant extracellular gum on the surface.
Nearly the entire length of the 16S rRNA gene was amplified, and all the isolates yielded a single-band amplification product of approximately 1,490 bp. Partial amplification of the nodC gene rendered an amplification product of approximately 930 bp.
Molecular Characterization of Isolates
The 16S rDNA ARDRA analysis revealed a remarkable heterogeneity in the size of the restriction products (from 50 to 920 bp) obtained for the different isolates. As a rule, all the restriction enzymes processed the amplification products into two (TaqI) to eight fragments (AluI), and, as expected, the band patterns showed correlation (±50 pb) with the size of the amplification products previously observed. The isolates examined grouped as five different clusters in the dendrogram, with low similarity (50–60%) within each cluster (Fig. 1). Isolate Hue-6 was separated from the entire group. Clustering of the reference strains gave rise to two groups: group 1 comprised of O. lupini and R. gallicum bv. phaseoli with less than 60% similarity and group 2 comprising both strains of S. meliloti (>80% similarity) and S. fredii (75% similarity). The remaining reference strains: R. gallicum bv. gallicum, M. loti, B. sp. (Lupinus) and B. japonicum were separated from the rest.
The techniques tDNA-PCR and IGS-PCR rendered polymorphism patterns consisting of 4 to 14 or 2 to 9 amplification products of less than 1,000 bp, respectively. The results of both fingerprinting techniques were combined to reflect the genotype diversity of the different isolates (Fig. 2). Overall, the isolates were genotypically dissimilar (>60%). Isolates Mad-3 and Mad-5 displayed the highest similarity (69%), whereas isolate Mad-1 showed 100% dissimilarity with respect to the remaining isolates (Fig. 2).
16S Sequencing and Phylogenetic Analyses
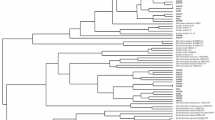
When compared with sequences of related species in the GenBank database, as a first approach to the identification of the selected isolates, 14 of them could be ascribed to the genus Rhizobium, three to Mesorhizobium, three to Phyllobacterium, and the remaining two were close to the genus Bosea, showing 97% of similarity (Table 1). The accession numbers for the 16S rRNA gene sequences obtained in this work are provided in Table 1. The data matrix constructed by aligning the 16S rRNA gene sequences belonged to 67 different isolates. The resulting phylogenetic tree showed two major clades (Fig. 3). A main clade, with total bootstrap support, grouped all isolates of the genera Rhizobium, Sinorhizobium, Mesorhizobium, and Phyllobacterium. A second main clade grouped different genera within the family Bradyrhizobiaceae (100% bootstrap).
Within the first main clade, two clusters could be differentiated: one formed by species belonging to the genera Rhizobium and Sinorhizobium and the other one grouping species of the genera Mesorhizobium and Phyllobacterium. The cluster containing the species in the genus Rhizobium grouped 14 O. tridentata isolates. Isolate Mad-3 grouped with Rhizobium leguminosarum bv. viceae, and three other (Alm-4, Mad-5, Mad-7) were close to this species (Fig. 3). Isolate Alm-2 was close to Rhizobium rhizogenes and Rhizobium tropici, whereas isolates Alm-3 and Mad-6 were close to Rhizobium etli (Fig. 3). Isolate Mad-4 was close to Rhizobium sullae. Isolate Hue-4 grouped with Rhizobium galegae, whereas isolate Hue-6 appeared separated but close to the cluster formed by Rhizobium huautlense and R. galegae. Remarkably, a separated cluster included isolates Alm-1, Alm-5, Alm-6, and Mad-2 close to Rhizobium rubi and Rhizobium giardinii. None of the O. tridentata isolates were included within the cluster corresponding to species of the genus Sinorhizobium (Fig. 3).
The clade of the genera Mesorhizobium and Phyllobacterium grouped six O. tridentata isolates (three within each genus, respectively; Fig. 3). Isolate Hue-3 grouped with Mesorhizobium mediterraneum, and isolate Mad-8 was close to this species, whereas isolate Mad-1 formed a branch with Mesorhizobium chacoense (Fig. 3). Isolates Hue-1 and Hue-8 grouped together close to isolate Hue-5 and to Phyllobacterium leguminum (Fig. 3).
The second main clade containing the genera within the Bradyrhizobiaceae family grouped two O. tridentata isolates (Hue-2 and Hue-7), which were close to the genus Bosea (Fig. 3).
nodC Sequencing and Phylogenetic Analysis
Amplified PCR products of the expected size (~930 bp) were obtained for 14 isolates. Eight isolates (Mad-2, Mad-6, Mad-8, Hue-1, Hue-3, Hue-4, Hue-5, and Hue-6) did not amplify for the nodC gene (Table 1). The analysis of the partial nodC gene sequences gave an alignment data matrix including 31 sequences. Isolate Alm-1 was not included in the phylogenetic analysis because only 444 nucleotides could be correctly assembled to obtain the nodC sequence (GenBank accession number EF364394) limiting the nucleotide alignment analysis.
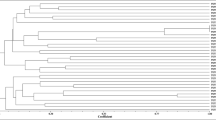
The resulting phylogenetic tree showed two major clades (Fig. 4). A main clade, with high bootstrap support (91%), grouped nodC sequences of R. leguminosarum and R. galegae (Fig. 4). Eight O. tridentata isolates grouped close to R. leguminosarum but formed an independent cluster with total bootstrap support (Fig. 4). The second main clade grouped species of the genera Rhizobium, Mesorhizobium and Bradyrhizobium (Fig. 4). Five O. tridentata isolates (Mad-1, Mad-7, Hue-2, Hue-7, and Hue-8) formed a separated cluster (with 91% bootstrap) close to species of the genus Mesorhizobium (Fig. 4). The isolates previously classified as close to the genera Phyllobacterium (Hue-8) and Bosea (Hue-2 and Hue-7) grouped together within this cluster.
Discussion
Molecular characterization of bacterial isolates associated to nodules of O. tridentata revealed Rhizobium, Mesorhizobium, Phyllobacterium, and Bosea as the predominant genera associated with this wild, gypsophyte shrub legume. As far as we know, this is the first description of bacteria isolated from the nodules of O. tridentata. Isolates described to be close to the genera Phyllobacterium and Bosea were only found in the geographic region of Fraga (HUE), where most taxonomic diversity at the genus level was observed.
At the species level, the ARDRA analysis indicated intense genotypic heterogeneity of the isolates, even among those sharing a common geographical origin, with homologies ranging from 10 to 68%. There have been increasing reports of the wide genotypic diversity of rhizobial nodulating bacteria from wild woody legume species even within a given geographic region [6, 11, 17, 29, 30, 52, 53, 54]. Contrary to legume crops, the higher environmental pressure to which wild legume shrubs and trees are subjected leads to the greater taxonomic diversity of their rhizobial populations [17, 28].
The fingerprinting techniques used in our study (tDNA-PCR and IGS-PCR) proved to be efficient tools for the rapid and accurate differentiation of the genetic diversity of the O. trientata nodule isolates at the species level. These procedures regrouped the isolates into well-supported clusters, most of which were highly consistent with phylogenetic clades inferred from the 16S rRNA gene sequence analysis. Both tDNA-PCR and IGS-PCR have been extensively used to differentiate bacterial strains in taxonomic groups other than Rhizobiales [13, 36]. The wide genotypic diversity indicated by these fingerprinting techniques was consistent with the results obtained through ARDRA analysis of the 16S rDNA region. O. tridentata showed a remarkably wide array of possible symbionts and a high level of promiscuity, as it has been previously described for other shrubby legumes [52, 53].
From a phylogenetic perspective, legume root bacterial symbionts do not form a single monophyletic group, and different lineages within the α-proteobacteria are today recognized according to the systematics of Garrity et al. [9]. The phylogenetic analyses performed here confirmed the wide diversity of bacterial isolates associated to O. tridentata. The phylogeny of the 16S rRNA gene yielded two main clades. The first one grouped members of the Rhizobiaceae (Rhizobium and Sinorhizobium) and Phyllobacteriaceae (Mesorhizobium and Phyllobacterium), with 100% bootstrap support, and the second clade brought together members of the Bradyrhizobiaceae, such as Bosea, Bradyrhizobium, or Afipia, with full bootstrap support. The genus Rhizobium was grouped in a single cluster in which most of the sequences of O. tridentata isolates were placed. Some of these sequences clearly grouped with those of type strain species. This was the case for Alm-4, Mad-3, Mad-5, and Mad-7 with R. leguminosarum, Alm-2 with R. tropicii and R. rhizogenes, or Hue-4 with R. galegae, clearly defining the phylogenetic position of these O. tridentata isolates. Relationships among Rhizobium species have been widely described and indicate that R. rhizogenes is much closer to R. leguminosarum than to R. galegae in phylogenetic terms [17, 34, 47]. At first, the phylogenetic divergence of R. galegae suggested the possibility of a new genus in the Rhizobiaceae, but subsequent analyses including other Rhizobium-related species have revealed R. galegae to be a strict Rhizobium species [47, 50]. Our data confirm also the consideration of R. galegae as a Rhizobium sensu stricto.
Our analysis detected another separated Rhizobium cluster with high bootstrap support (91%) containing isolates Alm-1, Alm-5, Alm-6, and Mad-2 close to R. rubi and R. giardinii. Given the tree topology and the high bootstrap value for this group, we could speculate that our isolates could belong to a new rhizobial species. Further studies including more related species and complementary techniques such as DNA–DNA hybridization probes are needed to confirm this point.
A second cluster with a bootstrap support of 90% grouped isolates initially identified as Mesorhizobium and Phyllobacterium. Although the taxonomy of the genus Phyllobacterium is still poorly known, several molecular phylogenetic analyses have designated the genus Phyllobacterium as a sister group of Mesorhizobium [19, 34], as indicated by our results. The genus Phyllobacterium contains seven recognized species, three of them, P. leguminum, P. ifriqiyense, and P. trifolii, obtained from root nodules of legumes [19, 43]. In fact, an unexpected immuno-relationship between the lipopolysaccharides (key molecules in nodulation) of several rhizobial species and Phyllobacterium was previously reported [18]. Many Phyllobacterium strains have indeed been isolated from root nodules, but their nitrogen-fixing ability, the presence of nif-like genes, and their capacity to induce nodulation have not been clearly demonstrated [19]. Recently, a novel species designated Phyllobacterium trifolii sp. nov., which is able to nodulate Trifolium repens and Lupinus albus, has been proposed [43]. Our three putative Phyllobacterium isolates (Hue-1, Hue-5, and Hue-8) were grouped close to P. leguminum but clearly separated, indicating that they could belong to a new Phyllobacterium species, although further work is needed to support this hypothesis. The partial sequence of the nodC gene was obtained for isolate Hue-8, indicating its probable nodulation ability.
The second main clade supported by a 100% bootstrap value included sequences from other phylogenetically distant α-proteobacteria belonging to the family Bradyrhizobiaceae. Two O. tridentata isolates (Hue-2 and Hue-7) were grouped within this clade close to the genus Bosea but independently placed indicating a putative new species. Several new α- and β-rhizobia genera, including Burkholderia, Cupriavidus, and Ochrobactrum, have recently been associated with the root nodules of various legumes [3, 6, 8, 41, 53]. Furthermore, recently, a strain belonging to the genus Bosea was isolated from the root nodules of Ononis vaginalis, although its nodulation capacity has not yet been tested [53]. In our study, partial sequence of the nodC gene has been obtained for Hue-2 and Hue-7, indicating the nodulation ability of these isolates. The goals of a future study will be to establish the phenotypic characters, taxonomic identification (DNA–DNA hybridization probes), host range, nodulation, and nitrogen-fixation efficiency, in particular of isolates close to the genera Phyllobacterium and Bosea.
Results from the phylogenetic analysis of some of the nodC gene sequences were not totally congruent with those based on the 16S rRNA gene, as previously pointed out [1, 15, 19]. The existence of diverse nodC lineages has been often explained as a consequence of the different rates of evolution of these sequences, the coevolution of the nodC sequences in symbiont and host, or the lateral transfer of Sym genes [15, 47].
If it is taxonomically confirmed that our isolates Hue-2, Hue-7, and Hue-8 belong to these genera, this would be the first report of a nodC-like gene in Bosea and Phyllobacterium.
In summary, our findings represent a first approach to the molecular characterization of bacteria isolated from a representative sample of the wild shrub O. tridentata growing across Spain. Increasing our knowledge about these nodulating rhizobacteria will have applications in formulating appropriate inocula for reclaiming gypsum ecosystems.
References
Aguilar OM, López MV, Donato M, Morón B, Soria-Diaz ME, Mateos C, Gil-Serrano A, Sousa C, Megías M (2006) Phylogeny and nodulation signal molecule of rhizobial populations able to nodulate common beans other than the predominant species Rhizobium etli present in soils from the northwest of Argentina. Soil Biol Biochem 38:573–586
Council of the European Community (1992) Directive 92/43 of the Council of the European Community on the conservation of habitats and wild fauna and flora. European Community, Brussels, Belgium
Barret CF, Parker MA (2006) Coexistence of Burkholderia, Cupriavidus, and Rhizobium sp nodule bacteria on two Mimosa spp. in Costa Rica. Appl Environm Microbiol 72:1198–1206
Benhizia Y, Benhizia H, Benguedouar A, Muresu R, Giacomini A, Squartini A (2004) Gamma proteobacteria can nodulate legumes of the genus Hedysarum. Syst Appl Microbiol 27:462–468
Brockwell J, Bottomley PJ, Thies JE (1995) Manipulation of rhizobia microflora for improving legume productivity and soil fertility. Plant and Soil 174:143–180
Chen W-M, Moulin L, Bontemps C, Vandamme P, Béna G, Boivin-Masson C (2003) Legume symbiotic nitrogen fixation by β-Proteobacteria is widespread in nature. J Bacteriol 185:7266–7272
Dana ED, Mota JF (2006) Vegetation and soil recovery on gypsum outcrops in semi-arid Spain. J Arid Environm 65:444–459
Elliot GN, Chen W-M, Chou J-H, Wang H-C, Sheu S-Y, Perin L, Reis VM, Moulin L, Simon MF, Bontemps C, Sutherland JM, Bessi R, de Faria SM, Trinick MJ, Prescott AL, Sprent JI, James EK (2007) Burkholderia phymatum is a highly effective nitrogen-fixing symbiont of Mimosa spp. and fixes nitrogen ex planta. New Phytol 173:168–180
Garrity GM, Bell JA, Lilburn TG (2003) Taxonomic outline of Prokaryotes. In: Garrity GM (ed) Bergey’s manual of systematic bacteriology. 2nd edn. Springer, New York
Giraud E, Moulin L, Vallenet D, Barbe V, Cytryn E, Avarre JC, Jaubert M, Simon D, Cartieaux F, Prin Y, Bena G, Hannibal L, Fardoux J, Kojadinovic M, Vuillet L, Lajus A, Cruveiller S, Rouy Z, Mangenot S, Segurens B, Dossat C, Franck WL, Chang WS, Saunders E, Bruce D, Richardson P, Normand P, Dreyfus B, Pignol D, Stacey G, Emerich D, Vermeglio A, Medigue C, Sadowsky M (2007) Legumes symbioses: absence of Nod genes in photosynthetic bradyrhizobia. Science 316 (5829):1307–1312
González-Andrés J, Alegre J, Ceresuela JL (2005) The rhizobia nodulating shrubs for revegetation of arid lands: isolation of native strains and specificity of the plant–rhizobia interaction by cross inoculation tests. Arid Land Res Manag 19:307–326
Hall BG (2004) Phylogenetic trees made easy: A how-to manual, 2nd edn. Sinauer, Sunderland, MA
Hirsch CF, Sigmund JM (1995) Use of polymerase chain reaction (PCR) fingerprinting to differentiate bacteria for microbial products screening. J Indust Microbiol 15:85–93
Kalita M, Stepkowski T, Lotock B, Malek W (2006) Phylogeny of nodulation genes and symbiotic properties of Genista tinctoria bradyrhizobia. Arch Microbiol 186:87–97
Laguerre G, Nour SM, Macheret V, Sanjuan J, Drouin P, Amarger N (2001) Classification of rhizobia based on nodC and nifH gene analysis reveals a close phylogenetic relationship among Phaseolus vulgaris symbionts. Microbiology 147:981–993
Lane DJ (1991) 16S/23S rRNA sequencing. In: Strackebrandt E, Goodfellow M (eds) Nucleic acid techniques in bacterial systematics. Chischester, Wiley, pp 115–175
Liu J, Wang ET, Chen WX (2005) Diverse rhizobia associated with woody legumes Wisteria sinensis, Cercis racemosa and Amorpha fruticosa grown in the temperate zone of China. Syst Appl Microbiol 28:465–477
Lucas MM, Peart JL, Brewin NJ, Kannenberg EL (1996) Isolation of monoclonal antibodies reacting with the core component of lipopolysaccharide from Rhizobium leguminosarum strain 3841 and mutant derivatives. J Bacteriol 178:2727–2733
Mantelin S, Fisher-Le Saux M, Zakhia F, Béna G, Bonneau S, Jeder H, de Lajudie P, Cleyet-Marel JC (2006) Emended description of the genus Phyllobacterium and description of four novel species associated with plant roots: Phyllobacterium bourgognense sp. nov., Phyllobacterium ifriqiyense sp. nov., Phyllobacterium leguminum sp. nov. and Phyllobacterium brassiccearum sp. nov. Int J Syst Evol Microbiol 56:827–839
Moschetti G, Peluso AL, Protopapa A, Anastasio M, Pepe O, Defez R (2005) Use of nodulation pattern, stress tolerance, nodC gene amplification, RAPD-PCR and RFLP–16S rDNA analysis to discriminate genotypes of Rhizobium leguminosarum biovar viciae. Syst Appl Microbiol 28:619–631
Odee DW, Haukka K, McInroy SG, Sprent JI, Sutherland JM, Young JPW (2002) Genetic and symbiotic characterisation of rhizobia isolated from tree and herbaceous legumes grown in soils from ecologically diverse sites in Kenya. Soil Biol Biochem 34:801–811
Palacio S, Escudero A, Montserrat-Martí G, Maestro M, Milla R, Albert MJ (2007) Plants living on gypsum: beyond the specialist model. Ann Bot 99:333–343
Perret X, Staehelin C, Broughton WJ (2000) Molecular basis of symbiotic promiscuity. Microbiol Mol Biol Res 64:180–201
Pugnaire FI, Lázaro R (2000) Seed bank and understory species composition in a semi-arid environment: the effect of shrub age and rainfall. Ann Bot 86:807–813
Rambaut A (1996) Se-Al: sequence alignment editor. Available at http://evolve.zoo.ox.ac.uk/
Rasolomampianina R, Bailly X, Fetiarison R, Bena G, Ramaroson L, Raherimandimby M, Moulin L, de Lajudie P, Dreyfus B, Avarre JC (2005) Nitrogen-fixing nodules from rose wood legume trees (Dalbergia spp.) endemic to Madagascar host seven different genera belonging to (α- and β-Proteobacteria). Mol Ecol 14:4135–4146
Requena N, Jiménez I, Toro M, Barea JM (1997) Interactions between plant-growth promoting rhizobacteria (PGPR), arbuscular mycorrhizal fungi and Rhizobium spp. in the rhizosphere of Anthyllis cytisoides, a model legume for revegetation in the Mediterranean semi-arid ecosystems. New Phytol 136:667–677
Requena N, Pérez-Solís E, Azcón-Aguilar C, Jeffries P, Barea JM (2001) Management of indigenous plant-microbe symbioses aids restoration of desertified ecosystems. Appl Environ Microbiol 67:495–498
Rodriguez-Echeverría S, Pérez-Fernández MA (2005) Potential use of Iberian shrubby legumes and rhizobia inoculation in revegetation projects under acidic soil conditions. Appl Soil Ecol 29:203–208
Romdhane SB, Nasr H, Samba-Mbaye R, Neyra M, Ghorbal MH, de Lajudie P (2006) Genetic diversity of Acacia tortilis ssp. raddiana rhizobia in Tunisia assessed by 16S and 16S-23S rDNA genes analysis. J Appl Microbiol 100:436–445
Saitou N, Nei M (1987) The neighbour-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sambrook J, Fritsch EF, Manniatis T (1989) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory, Cold Spring Harbor, NY
Sarita S, Sharma PK, Priefer UB, Prell J (2005) Direct amplification of rhizobia nodC sequences from soil total DNA and comparison to nodC diversity of root nodule isolates. FEMS Microbiol Ecol 54:1–11
Sawada H, Kuykendall LD, Young JM (2003) Changing concepts in the systematics of bacterial nitrogen-fixing legume symbionts. J Gen Appl Microbiol 49:155–179
Sneath PHA, Sokal RR (1973) Numerical taxonomy. WH Freeman, San Francisco
Storms V, Van den Vreken N, Gillis M, Vandamme P (2002) Evaluation of tRNA intergenic length polymorphism (tDNA-PCR) for the differentiation of the members of the Burkholderia cepacia complex. Syst Appl Microbiol 25:376–385
Swofford DL (2002) PAUP*: Phylogenetic Analysis Using Parsimony (*and other methods), version 4. Sinauer, Sunderland, MA
Sy A, Giraud E, Jurad P, García N, Willems A, de Lajudie P, Prin Y, Neyra M, Gillis M, Bombin-Masson C, Dreyfus B (2001) Methylotropic Methylobacterium bacteria nodulate and fix nitrogen in symbiosis with legumes. J Bacteriol 183:214–220
Thrall PH, Millsom DA, Jeavons AC, Waayers M, Harvey GR, Bagnall DJ, Brockwell J (2005) Seed inoculation with effective root-nodule bacteria enhances revegetation success. J Appl Ecol 42:740–751
Thompsom JD, Gibson TJ, Plewniak F, Jeanmoungin F, Higgins DG (1997) The CUSTAL X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Trujillo ME, Willems A, Abril A, Planchuelo AM, Rivas R, Ludeña D, Mateos PF, Martínez-Molina E, Velázquez E (2005) Nodulation of Lupinus albus by strains of Ochrobactrum lupini sp. nov. Appl Environ Microbiol 71:1318–1327
Ueda T, Suga Y, Yahiro N, Matsuguchi T (1995) Phylogeny of Sym plasmids of Rhizobia by PCR-based sequencing of a nodC segment. J Bacteriol 177:48–472
Valverde A, Velázquez E, Fernández-Santos F, Vizcaino N, Rivas R, Mateos PF, Martínez-Molina E, Igual JM, Willems A (2005) Phyllobacterium trifolii sp. nov., nodulating Trifolium and Lupinus in Spanish soils. Int J Syst Evol Micr 55:1985–1989
van Berkum P, Eardly BD (1998) Molecular evolutionary systematics of the Rhizobiaceae. In: Spaink HP, Kondorosi A, Hooykaas PJJ (eds) The Rhizobiaceae: molecular biology of model plant-associated bacteria. Kluwer, Dordrecht, pp 1–24
Vincent JM (1970) A manual for the practical study of root-nodules bacteria. IBP Handbook. Blackwell, Oxford, UK
Vinuesa P, Silva C, Lorite MJ, Izaguirre-Mayoral ML, Bedmar EJ, Martínez-Romero E (2005) Molecular systematics of rhizobia based on maximum likelihood and Bayesian phylogenies inferred from rrs, atpD, recA and nifH sequences, and their use in the classification of Sesbania microsymbionts from Venezuelan wetlands. Syst Appl Microbiol 28:702–716
Wang ET, Martínez-Romero E (2000) Sesbania herbacea–Rhizobium huautlense nodulation in flooded soils and comparative characterization of S. herbacea-nodulating Rhizobia in different environments. Microb Ecol 41:25–32
Weir BS, Turner SJ, Silvester WB, Park D-C, Young JM (2004) Unexpectedly diverse Mesorhizobium strains and Rhizobium leguminosarum nodulate native legume genera of New Zealand, while introduced legume weeds are nodulated by Bradyrhizobium species. Appl Environ Microbiol 70:5980–5987
Weisburg WG, Barns SM, Pelletier DA, Lane DJ (1991) 16S ribosomal DNA amplification for phylogenetic study. J Bacteriol 173:697–703
Young JP, Haukka KE (1996) Diversity and phylogeny of rhizobia. New Phytol 133:87–94
Young JM, Park DC, Weir BS (2004) Diversity of 16S rRNA sequences of Rhizobium spp. implications for species determinations. FEMS Microbiol Lett 238:125–131
Zakhia F, Jeder H, Domergue O, Willems A, Cleyet-Marel JC, Gillis M, Dreyfus B, de Lajudie P (2004) Characterisation of wild legume nodulating bacteria (LNB) in the infra-arid zone of Tunisia. Syst Appl Microbiol 27:380–395
Zakhia F, Jeder H, Willems A, Gillis M, Dreyfus B, de Lajudie P (2006) Diverse bacteria associated with root nodules of spontaneous legumes in Tunisia and first report for nifH-like gene within the genera Microbacterium and Starkeya. Microb Ecol 51:375–393
Zerhari K, Aurag J, Khbaya B, Kharchaf D, Filali-Maltouf A (2000) Phenotypic characteristics of rhizobia isolates nodulating Acacia species in the arid and Saharan regions of Morocco. Lett Appl Microbiol 30:351–357
Acknowledgments
The authors thank Dr. Noelle Amarger, Dr. Herman Spaink, and Dr. Encarna Velázquez for providing some of the reference strains. The authors thank Dr David Badía and Dr. Francisco Pugnaire for the localization of Ononis populations. We thank Monika Oggerin for assistance with phylogenetic analysis. This work was supported by grants GR/AMB/0735/2004 and S-0505/AMB/0321 from the Comunidad de Madrid (Spain). A. Rincón holds a I3P postdoctoral fellowship awarded by the Consejo Superior de Investigaciones Científicas (CSIC, Spain), and F. Arenal was awarded a postdoctoral grant by the Comunidad de Madrid.
Author information
Authors and Affiliations
Corresponding author
Additional information
A. Rincón and F. Arenal contributed equally to this work.
Rights and permissions
About this article
Cite this article
Rincón, A., Arenal, F., González, I. et al. Diversity of Rhizobial Bacteria Isolated from Nodules of the Gypsophyte Ononis tridentata L. Growing in Spanish Soils. Microb Ecol 56, 223–233 (2008). https://doi.org/10.1007/s00248-007-9339-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-007-9339-6