Abstract
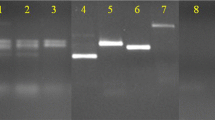
PCR fingerprinting offers a practical molecular means to quickly and reliably differentiate bacteria for microbial products screening. A combination of low resolution and high resolution PCR fingerprinting provides a hirarchical system which allows the discrimination of bacteria at species and subspecies level within 7 h. DNA was extracted from cells by incubating them in water at 95°C for 30 min. A sample of 1 μl of the cell-free aqueous extract then was used as a source of template DNA in the PCR. The PCR products were separated by electrophoresis on an acrylamide gel and visualized by ethidium bromide staining. The band patterns generated for each different culture were unique, reproducible, and independent of cultivation conditions. Band patterns may be compared visually or by using imaging and pattern matching software. In our laboratory, bacteria such as actinomycetes, Gram-negative and Gram-positive soil eubacteria, and photosynthetic non-sulfur bacteria have been differentiated using PCR fingerprinting.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Anzai Y, T Okuda and J Watanabe. 1994. Application of the random amplified polymorphic DNA using the polymerase chain reaction for efficient elimination of duplicate strains in microbial screening. II. Actionomycetes. J Antibiot 47: 183–193.
Brousseau R, A Saint-Onge, G Prefontaine, L Masson and J Cabana. 1993. Arbitrary primer polymerase chain reaction, a powerful method to identifyBacillus thuringiensis serovars and strains. Appl Environ Microbiol 59: 114–119.
Cote R (ed). 1984. ATCC Media Handbook. American Type Culture Collection, Rockville, MD.
Devos KM and MD Gale. 1992. The use of random amplified polymorphic DNA markers in wheat. Theor Appl Genet 84: 567–572.
Fujimori F and T Okuda, 1994. Application of the random amplified polymorphic DNA using the polymerase chain reaction for efficient elimination of duplicate strains in microbial screening. I. Fungi. J Antibiot 47: 173–182.
Goodfellow M and AG O'Donnell.1993. Roots of bacterial systematics. In: Handbook of New Bacterial Systematics (Goodfellow M and AG O'Donnell, eds), pp 3–54, Academic Press, New York, NY.
Jensen MA, JA Webster and N Straus. 1993. Rapid identification of bacteria on the basis of polymerase cahin reaction-amplified ribosomal DNA spacer polymorphisms. Appl Environ Microbiol 59: 945–952.
Johnson JL. 1984. Nucleic acids in bacterial classification. In: Bergey's Manual of Systematic Bacteriology (Krieg NR and JG Holt, eds), 1, pp 8–11, Williams and Wilkins, Baltimore, MD.
Johnson JL. 1994. Similarity analysis of DNAs. In: Methods for General and Molecular Bacteriology (Gerhardt P, ed), pp 655–682, American Society for Microbiology, Washington, DC.
Kenerly ME, EA Morgan, L Post, L Lindahl and M Nomura. 1977. Characterization of hybrid plasmids carrying individual ribosomal ribonucleic acid transcription units ofEscherichia coli. J Bacteriol 132: 931–949.
Kiss A, B Sain and P Venetainer. 1977. The number of rRNA genes inEscherichia coli. FEBS Lett 79: 77–79.
Kostman JR, TD Edlind, JJ LiPuma and TL Stull. 1992. Molecular epidemiology ofPseudomonas cepacia determined by polymerase chain ribotyping. J Clin Microbiol 30:2084–2084.
Logan NA and RCW Berkeley. 1984. Identification ofBacillus using the API system. J Gen Microbiol 130: 1871–1882.
MacPherson JM, PE Eckstein, GJ Scoles and AA Gajadhar. 1993. Variability of the random amplified polymorphic DNA assay among thermal cyclers, and effects of primer and DNA concentration. Molec Cell Probes 7: 293–299.
Meunier J-R and PAD Grimont. 1993. Factors affecting reproducibility of random amplified polymorphic DNA fingerprinting. Res Microbiol 144: 373–379.
Sneath PHA. 1986. Endospore-forming gram-positive rods and cocci. In: Bergey's Manual of Systematic Bacteriology (Sneath PHA, ed), 2, p 1131, Williams and Wilkins, Baltimore, MD.
Tanaka H, S Sawairi and T Okuda. 1994. Application of the random amplified polymorphic DNA using the polymerase chain reaction for efficient elimination of duplicate strains in microbial screening. III. Bacteria. J Antibiot 47: 194–200.
Vauterin L, J Swings and K Kersters. 1993. Protein electrophoresis and classification: In: Handbook of New Bacterial Systematics (Goodfellow M and AG O'Donnell, eds), pp 251–280, Academic Press, New York, NY.
Visuvanathan S, MT Moss, JL Stanford, J Hermon-Taylor and J McFadden. 1989. Simple enzymic method for isolation of DNA from diverse bacteria. J Microbiol Methods 10: 59–64.
Welsh J and M McClelland. 1991. Genomic fingerprints produced by PCR with consensus tRNA gene primers. Nucleic Acids Res 19: 861–866.
Welsh J and M McClelland. 1992. PCR-amplified length polymorphisms in tRNA intergenic spacers for categorizing staphylococci. Mol Microbiol 6: 1673–1680.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Hirsch, C.F., Sigmund, J.M. Use of polymerase chain reaction (PCR) fingerprinting to differentiate bacteria for microbial products screening. Journal of Industrial Microbiology 15, 85–93 (1995). https://doi.org/10.1007/BF01569805
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF01569805