Abstract
The hydrochemistry and the microbial diversity of a pristine aquifer system near Garzweiler, Germany, were characterized. Hydrogeochemical and isotopic data indicate a recent activity of sulfate-reducing bacteria in the Tertiary marine sands. The community structure in the aquifer was studied by fluorescence in situ hybridization (FISH). Up to 7.3 × 105 cells/mL were detected by DAPI-staining. Bacteria (identified by the probe EUB338) were dominant, representing 51.9% of the total cell number (DAPI). Another 25.7% of total cell were affiliated with the domain Archaea as identified by the probe ARCH915. Within the domain Bacteria, the β-Proteobacteria were most abundant (21.0% of total cell counts). Using genus-specific probes for sulfate-reducing bacteria (SRB), 2.5% of the total cells were identified as members of the genus Desulfotomaculum. This reflects the predominant role these microorganisms have been found to play in sulfate-reducing zones of aquifers at other sites. Previously, all SRB cultured from this site were from the spore-forming genera Desulfotomaculum and Desulfosporosinus.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Cultivation-based techniques for assessing microbial diversity in natural environments are limited in their ability to identify all contributing microorganisms. Only a small proportion of the microbial population is revealed by the isolation of pure cultures, no matter whether they have been obtained from enrichment cultures or dilution series. Since molecular tools became available, a much higher microbial diversity has been detected, reflecting most of the natural microbial communities in various environments including hot springs, soils, sewage sludge, intestines of higher organisms, marine habitats, freshwater systems, and contaminated groundwater systems [11, 20, 25, 34, 39, 49, 51]. Compared to these habitats, knowledge about structure and function of the microbial community in non-contaminated subsurface environments is rather limited (for review, see [26]). Until recently the microbial communities in subsurface environments were mostly described by cultivation-based methods or by cloning of 16S rRNA genes [9, 11, 30, 36, 37, 48]. Nowadays FISH represents a powerful tool for the identification and quantification of different phylogenetic groups in environmental samples [2, 3, 35].
The investigated groundwater system, situated near Cologne, western Germany, is geologically well characterized [8, 42, 43]. Marine sands of Tertiary age with intercalated lignite seams confine a multiple aquifer system. Information obtained before this investigation, on the groundwater chemistry, pointed strongly toward recent metabolic activity of microorganisms. A strong isotopic discrimination signature of dissolved sulfate and the occurrence of methane already indicated complete anoxic conditions and the presence of sulfate-reducing bacteria (SRB) and methanogenic microorganisms [14]. Therefore, we used group- and genus-specific oligonucleotide probes to gain an insight into the microbial diversity of this ecosystem.
Materials and Methods
Study Site and Sampling Procedure
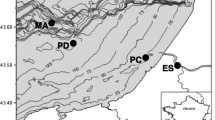
The study site is situated in the Lower Rhine Embayment, Germany next to the open-pit lignite mine Garzweiler I. The general lithostratigraphic column comprises a succession of up to 250 m of marine sands with a Tertiary age covered by up to 100 m of Quaternary fluvial sediments. A clay horizon (Reuver Ton) and up to three lignite seams (Morken, Frimmersdorf, Garzweiler) are intercalated, resulting in up to five different aquifers. The uppermost aquifer is unconfined while all lower aquifers are confined.
Sampling of groundwater monitoring wells was performed using a standard method as described in detail elsewhere [16]. Samples were taken after pumping for ≥40 min and after parameters such as temperature, pH, redox potential, oxygen and conductivity of the groundwater had remained stable for ≥15 min due to recharge of aquifer water. For chemical analysis samples were stored under anoxic conditions at +4°C in the dark.
Chemical and Isotopic Measurements
Basic hydrochemical information about the groundwater in the study area was obtained using standard procedures as outlined elsewhere [8, 42, 43]. The main objective of this study was a characterization of the isotopic compositions of carbon and sulfur components within the aquifer system with special emphasis on evidence for biologically driven processes. Analytical procedures and data were outlined previously [14].
Isolation of Sulfate-Reducing Bacteria
The isolates used as references in this study have been isolated from the same habitat using media and protocols described previously [14]. The sequences of their 16S rRNA encoding genes were obtained using standard protocols [14]. The nearly complete sequences were loaded into the 16S rRNA sequence database of the Technical University of Munich using the program package ARB [47]. The tool ARB_Align was used for sequence alignment. The alignment was visually inspected and corrected manually. Sequences of Desulfotomaculum or other closely related genera of relevance not included in the database were obtained from the NCBI database.
Fluorescence in Situ Hybridization (FISH)
Samples for FISH were fixed with 4% (w/v) paraformaldehyde solution on polycarbonate filters and stored as described in detail by Glöckner et al. [18]. Cy3-labeled oligonucleotides were purchased from Interactiva (Ulm, Germany). Hybridization and microscopy counts of hybridized and 4′,6′-diamidino-2-phenylindole (DAPI)-stained cells were performed as described previously [45]. Means were calculated from 10 to 20 randomly chosen fields on each filter section, corresponding to 800–1000 DAPI stained cells. Counting results were always corrected by subtracting signals observed with the probe NON338. Formamide concentrations and oligonucleotide probes used are given in Table 2. Probes BET42a, GAM42a, PLA886, CREN499R, EURY498R, and BONE23a were used with competitor oligonucleotides as described previously [3, 10, 28, 33].
Results
Groundwater
The hydrochemical and microbiological parameters of the investigated groundwater samples are summarized in Table 1. Water samples were retrieved from four groundwater monitoring wells at a depth of 120–125 m and had a temperature between 11.3 and 13.2°C. Samples of well 2 and well 3 displayed circumneutral pH. Wells 1 and 4 had pH values of 7.6 to 7.7. All water samples except for those from well 2 were anoxic and contained only minor concentrations of nitrate. In contrast to water samples of well 1, all other samples showed comparatively low concentrations of sulfate, paralleled by strongly positive δ34S values, indicative of a recent activity of SRB.
All samples contained dissolved organic carbon (DOC). Negative δ13C-values, indicative of microbial degradation of organic matter, were lowest in water samples of well 3. This groundwater contained the highest cell numbers (7.3 × 105 cells/mL) and additionally showed the highest percentage of FISH-detected cells with domain-specific probes EUB338 and ARCH915 (77.6% vs 28.5% in groundwater well 4). This indicates that well 3 harbored the most active microbial population of all groundwater wells sampled. Groundwater samples from well 3 were therefore chosen for further characterization of the microbial diversity in this aquifer.
Microbial Diversity
FISH results are summarized in Fig. 1. Of the total cell counts indicated by DAPI-staining, 77.6% hybridized with probes specific for the domains Bacteria and Archaea. The Bacteria were more abundant than the Archaea (51.9% of total cells, Table 1). Among the cells detected with the bacterial probe EUB338, Proteobacteria were most abundant (29.0% of total cells). Within the Proteobacteria, members of the β-group were dominant (21.0% of total cells), whereas the α-group played a minor role (1.0% of total cells). A significant fraction of the β-Proteobacteria was affiliated with the β-1-subgroup (8.2% of total cells). Members of the γ- and ε-Proteobacteria, the Cytophaga–Flavobacterium cluster, Planctomycetales, and Gram-positives with high G+C DNA (Table 2) were below the detection limit for FISH. The majority of cells detected with the archaeal probe ARCH915 were affiliated with Euryarchaeota (21.5% of total cells).
Sulfate-Reducing Microorganisms
The general probe SRB385 hybridized with 7.0% of total cells in samples from groundwater well 3. This probe covers most bacteria of the δ-Proteobacteria, but also targets cells from other taxa (e.g., Clostridia and other Gram-positive bacteria) and might therefore cause an overestimation of this group. More specific probes for certain genera of Gram-negative SRB (see Table 2) showed no hybridization with cells in the samples. Members of the genera Desulfobotulus, Desulfobulbus, Desulfomicrobium, Desulfomonile, Desulfovibrio, and Syntrophus, which have been frequently isolated from freshwater habitats [29, 35], were not found. The only positive result for SRB was with probe DTM229, which targets members of the genus Desulfotomaculum. Only Desulfotomaculum acetoxidans, Desulfotomaculum alcaliphilum, and Desulfosporosinus orientis (formerly Desulfotomaculum orientis-cluster) are not targeted by this probe [19].
Discussion
Microbial Activity
In this communication we describe the community composition of the groundwater-associated microbial population. Hydrochemical data of the characterized groundwater monitoring wells indicated anaerobic processes in three of the four sampled wells (wells 2, 3, and 4). Positive δ34S-values paralleled by low concentrations of dissolved sulfate in wells 3 and 4 are pointing strongly toward recent bacterial sulfate reduction [42, 43]. Nevertheless, sulfate might not be used exclusively as terminal electron acceptor in this system. The presence of high microbial activity is supported by the finding of the highest cell numbers and highest percentage of FISH-detected cells in these samples. Some cells may be undetectable due to spore-formation, starvation, dormancy, or cell death and are therefore considered to be an indicator for low in-situ activity [6]. Only groundwater samples from well 3 contained a microbial population with >75% hybridizable cells. In samples from other wells, <30% of DAPI-stained cells hybridized with EUB 338 and ARCH915 probes even though the other samples were retrieved from a similar depth (Table 1). Obviously, aquifer systems display a significant heterogeneity with respect to their microbial activity.
Microbial Diversity
FISH can be a suitable method for the quantification of phylogenetic groups in environmental samples. Nevertheless, this method has limitations when cell walls are impermeable, when target sites are poorly accessible, or when the cellular rRNA content is too low. This makes it unsuitable for samples where cellular activity is low. However, it would seem reasonable to use FISH for well 3 as metabolic activity is obviously high. By using FISH, 77.6% of the total cell number in samples from well 3 could be affiliated with specific domains, giving an overview of structural and functional aspects of the microbial population in this aquifer.
As in other freshwater systems, a large proportion of the abundant microorganisms belonged to the β-Proteobacteria [13, 17, 37]. The Comamonas–Variovorax group (β-1 subgroup) seems to be of particular importance. Microorganisms belonging to these taxa were frequently isolated from subsurface environments [7, 11, 22, 36, 37, 52]. Obviously β-Proteobacteria are generally well adapted to freshwater conditions and might be important for the oxidation of dissolved organic carbon.
The abundance of Euryarchaeota can most probably be attributed to methanogens. This finding correlates well with the detection of biogenic methane as identified by typical δ13C signatures [42, 44]. The probe EURY498R also hybridizes with members of the genus Archaeoglobus, but the presence of these thermophilic sulfate-reducing archaea (temperature range for growth between 60 and 95°C) at this low temperature is very unlikely. Since sulfate concentrations are very low, carbon dioxide is important as an electron acceptor in this aquifer system. Fe3+ as electron acceptor was important only at lower depths, whereas Mn4+ was not relevant in this system [8]. In contrast to hydrogen-dependent methanogenesis, which would be outcompeted by sulfate reduction, the acetate-dependent methanogenesis would probably not be affected, because of the low ability of sulfate-reducing bacteria to compete for acetate as electron donor in nonmarine habitats [1]. The coexistence of acetoclastic methanogenic archaea, homoacetogenic bacteria, and sulfate-reducing bacteria has been demonstrated in deep granitic rock groundwater [23, 31].
In the investigated aquifer system SRB are present and active as indicated by the isotopic signature of the residual sulfate, depletion of sulfate and formation of sulfide in the groundwater [14]. Although many of the oligonucleotide probes which have been used seem to be restricted to marine SRB, the absence of Desulfovibrio spp. and Desulfomicrobium spp. in the groundwater was surprising. The positive results with SRB385 but negative results using more specific probes for SRB could be explained by the presence of members of the Geobacteraceae. These microorganisms use sulfur or ferric iron as electron acceptor or are growing by fermentation. The dominant population of SRB in our aquifer system are spore-forming members of the genus Desulfotomaculum (Fig. 1). These results perfectly reflect the isolates we have obtained from this aquifer system which were spore-forming SRB affiliated with Desulfotomaculum spp. and Desulfosporosinus spp. as indicated by comparative analysis of the 16S rRNA sequences (Fig. 2; [14]). All of these isolates are able to use a large variety of organic compounds as electron donors and show similar physiological properties as has been demonstrated for other members of this taxa (e.g., [24]). In contrast to Desulfovibrio and Desulfomicrobium, they are not restricted to a few organic compounds and hydrogen as electron donors. Desulfotomaculum spp. seem to be widespread in subsurface environments [7, 9, 12, 21]. Many Desulfotomaculum spp. can cope with low levels or complete absence of sulfate, because they can grow by fermentation of organic compounds or by homoacetogenesis using hydrogen and carbon dioxide [24]. It is possible that hydrogen, the favored electron donor for Desulfovibrio spp. and Desulfomicrobium spp., is not present together with sulfate at sufficient levels in the aquifer, or hydrogen is effectively removed by homoacetogenesis as is the case in similar environments [1, 23, 35]. Additionally, the formation of spores by Desulfotomaculum spp. allows survival under varying redox conditions. Generally Gram-positive bacteria play an important role in the subsurface [52, 21]. Members of the genera Desulfosarcina and Desulfococcus are widespread in marine environments [38, 41]. However, because of their universalistic physiological traits Desulfotomaculum spp. seem to be the counterpart in this niche in the subsurface.
Phylogenetic position of sulfate-reducing isolates (in bold type) from a pristine aquifer. Desulfotomaculum spp., which are not detected with DFM229, are indicated with an asterisk. The tree was constructed using 1255 unambiguously aligned nucleotides using the maximum-likelihood method. All sequences belong to the Gram-positive bacteria except the Desulfovibrios, which are members of the δ-Proteobacteria and function as an outgroup.
References
C Achtnich A Schuhmann T Wind R Conrad (1995) ArticleTitleRole of interspecies H2 transfer to sulfate and ferric iron-reducing bacteria in acetate consumption in anoxic paddy soil. FEMS Microbiol Ecol 16 61–70 Occurrence Handle10.1016/0168-6496(94)00070-D Occurrence Handle1:CAS:528:DyaK2MXjt1Smu70%3D
R Amann F-O Glöckner A Neef (1997) ArticleTitleModern methods in subsurface microbiology: in situ identification of microorganisms with nucleic acid probes. FEMS Microbiol Rev 20 191–200 Occurrence Handle10.1016/S0168-6445(97)00007-7 Occurrence Handle1:CAS:528:DyaK2sXltl2msL0%3D
R Amann W Ludwig K Schleifer (1995) ArticleTitlePhylogenetic identification and in situ detection of individual microbial cells without cultivation. Microbiol Rev 59 143–169 Occurrence Handle1:CAS:528:DyaK2MXkvVGmurk%3D Occurrence Handle7535888
R Amann J Snaidr M Wagner W Ludwig K-H Schleifer (1996) ArticleTitleIn situ visualization of high genetic diversity in a natural bacterial community. J Bacteriol 178 3496–3500 Occurrence Handle1:CAS:528:DyaK28XjsFGls7Y%3D Occurrence Handle8655546
RI Amann BJ Binder RJ Olson SW Chisholm R Devereux DA Stahl (1990) ArticleTitleCombination of 16S rRNA-targeted oligonucleotide probes with flow cytometry for analyzing mixed microbial populations. Appl Environ Microbiol 56 1919–1925 Occurrence Handle1:CAS:528:DyaK3cXkvFCgsrs%3D Occurrence Handle2200342
P Amy (1997) Microbial dormancy and survival in the subsurface. P Amy D Haldeman (Eds) The Microbiology of the Terrestrial Deep Subsurface Lewis Boca Raton, FL 185–203
D Balkwill R Reeves G Drake J Reeves F Crocker M King D Boone (1997) ArticleTitlePhylogenetic characterization of bacteria in the subsurface microbial culture collection. FEMS Microbiol Rev 20 201–216 Occurrence Handle10.1016/S0168-6445(97)00008-9 Occurrence Handle1:CAS:528:DyaK2sXltl2msbo%3D Occurrence Handle9299704
Bergmann, A (1999) Hydrogeochemische Untersuchungen anoxischer Redoxprozesse in tiefen Porengrundwasserleitern der Niederrheinischen Bucht im Umfeld des Tagebaus Ph.D. thesis. Garzweiler I, Ruhr-Universität, Bochum
V Boivin-Jahns R Ruimy A Bianchi S Daumas R Christen (1996) ArticleTitleBacterial diversity in a deep-subsurface clay environment. Appl Environ Microbiol 62 3405–3412 Occurrence Handle1:CAS:528:DyaK28XlsFeqtr8%3D Occurrence Handle8795233
S Burggraf T Mayer A Amann S Schadhauser C Woese K Stetter (1994) ArticleTitleIdentifying members of the domain Archaea with rRNA-targeted oligonucleotide probes. Appl Environ Microbiol 60 3112–3119 Occurrence Handle1:CAS:528:DyaK2cXlslWmu7c%3D Occurrence Handle7524440
D Chandler S Li C Spadoni G Drake D Balkwill J Fredrickson F Brockman (1997) ArticleTitleA molecular comparison of culturable aerobic heterotrophic bacteria and 16S rDNA clones derived from a deep subsurface sediment. FEMS Microbiol Ecol 23 131–144 Occurrence Handle10.1016/S0168-6496(97)00021-4 Occurrence Handle1:CAS:528:DyaK2sXkvFaksLc%3D
F Colwell T Onstott M Delwiche D Chandler J Fredrickson Q Yao J McKinley D Boone R Griffiths T Phelps D Ringelberg D White L LaFreniere D Balkwill R Lehman J Konisky P Long (1997) ArticleTitleMicroorganisms from deep, high temperature sandstones: constraints on microbial colonization. FEMS Microbiol. Rev. 20 425–435 Occurrence Handle10.1016/S0168-6445(97)00024-7 Occurrence Handle1:CAS:528:DyaK2sXltl2msLs%3D
B Crump V Virginia Armbrust J Baross (1999) ArticleTitlePhylogenetic analysis of particle-attached and free-living bacterial communities in the Columbia River, its estuary, and the adjacent coastal ocean. Appl Environ Microbiol 65 3192–3204 Occurrence Handle1:CAS:528:DyaK1MXktlemu78%3D Occurrence Handle10388721
J Detmers U Schulte H Strauss J Kuever (2001) ArticleTitleSulfate reduction at a lignite seam: microbial abundance and activity. Microb Ecol 42 238–247 Occurrence Handle10.1007/s00248-001-1014-8 Occurrence Handle1:CAS:528:DC%2BD38XksFc%3D Occurrence Handle12024249
R Devereux MD Kane J Winfrey DA Stahl (1992) ArticleTitleGenus- and group-specific hybridization probes for determinative and environmental studies of sulfate-reducing bacteria. System Appl Microbiol 15 601–609 Occurrence Handle1:CAS:528:DyaK3sXksVGisL4%3D
DVWK (1992) “Entnahme und Untersuchungsumfang von Grundwasserproben. DVWK-Regeln zur Wasserwirtschaft.” DVWK, Hamburg, Berlin
F Glöckner B Fuchs R Amann (1999) ArticleTitleBacterioplankton compositions of lakes and oceans: a first comparison based on fluorescence in situ hybridization. Appl Environ Microbiol 65 3721–3726 Occurrence Handle10427073
FO Glöckner R Amann A Alfreider J Pernthaler R Psenner K Trebesius K-H Schleifer (1996) ArticleTitleAn in situ hybridization protocol for detection and identification of planktonic bacteria. Syst Appl Microbiol 19 403–406
K Hristova M Mau D Zheng R Aminov R Mackie H Gaskins L Raskin (2000) ArticleTitleDesulfotomaculum genus and sub-genus-specific 16S rRNA hybridization probes for environmental studies. Environ Microbiol 2 143–160 Occurrence Handle10.1046/j.1462-2920.2000.00085.x Occurrence Handle1:CAS:528:DC%2BD3cXjslSnsbw%3D Occurrence Handle11220301
P Hugenholtz C Pitulle KL Hershberger NR Pace (1998) ArticleTitleNovel division level bacterial diversity in a Yellowstone hot spring. J Bacteriol 180 366–376 Occurrence Handle1:CAS:528:DyaK1cXlsVaqsA%3D%3D Occurrence Handle9440526
K Ishi S Takii S Fukunaga K Aoki (2000) ArticleTitleCharacterization by denaturating gardient gel electrophoresis of bacterial communities in deep groundwater at the Kamaishi Mine, Japan. J Gen Appl Microbiol 46 85–93 Occurrence Handle12483595
L Jimenez (1990) ArticleTitleMolecular analysis of deep-subsurface bacteria. Appl Environ Microbiol 56 2108–2113 Occurrence Handle1:CAS:528:DyaK3cXltFahtbo%3D Occurrence Handle2389933
S Kotelnikova K Pedersen (1997) ArticleTitleEvidence for methanogenic archaea and homoacetogenic bacteria in deep granitic rock aquifers. FEMS Microbiol Rev 20 339–349 Occurrence Handle10.1016/S0168-6445(97)00016-8 Occurrence Handle1:CAS:528:DyaK2sXltl2ms7g%3D
J Kuever F Rainey H Hippe (1999) ArticleTitleDescription of Desulfotomaculum sp. Groll as Desulfotomaculum gibsoniae, sp. nov. Int J Syst Bacteriol 49 1801–1808 Occurrence Handle1:CAS:528:DyaK1MXnsFGiur0%3D Occurrence Handle10555363
W Liesack E Stackebrandt (1992) ArticleTitleOccurrence of novel groups of the domain Bacteria as revealed by analysis of genetic material isolated from an Australian terrestrial environment. J Bacteriol 174 5072–5078 Occurrence Handle1:CAS:528:DyaK38XlsVKqu7k%3D Occurrence Handle1629164
E Madsen (2000) ArticleTitleNucleic-acid characterization of the identity and activity of subsurface microorganisms. Hydrogeol J 8 112–125 Occurrence Handle10.1007/s100400050012 Occurrence Handle1:CAS:528:DC%2BD38Xkt1GrtL8%3D
W Manz R Amann W Ludwig M Vancanneyt K-H Schleifer (1996) ArticleTitleApplication of a suite of 16S rRNA-specific oligonucleotide probes designed to investigate bacteria of the phylum Cytophaga–Flavobacter–Bacteroides in the natural environment. Microbiology 142 1097–1106 Occurrence Handle1:CAS:528:DyaK28XjtFyju7c%3D Occurrence Handle8704951
W Manz R Amann W Ludwig M Wagner K-H Schleifer (1992) ArticleTitlePhylogentic oligodeoxynucelotide probes for the major subclasses of proteobacteria: problems and solutions. Syst Appl Microbiol 15 593–600
W Manz M Eisenbrecher TR Neu U Szewzyk (1998) ArticleTitleAbundance and spatial organization of Gram-negative sulfate-reducing bacteria in activated sludge investigated by in situ probing with specific 16S rRNA targeted oligonucleotides. FEMS Microbiol Ecol 25 43–61 Occurrence Handle10.1016/S0168-6496(97)00082-2 Occurrence Handle1:CAS:528:DyaK1cXislymtA%3D%3D
D Martino E Grossman G Ulrich K Burger J Schlichenmeyer J Suflita J Ammerman (1998) ArticleTitleMicrobial abundance and activity in a low-conductivity aquifer system in East-Central Texas. Microb Ecol 35 224–234 Occurrence Handle10.1007/s002489900078 Occurrence Handle1:CAS:528:DyaK1cXivFCntbs%3D Occurrence Handle9569280
H Meier R Amann W Ludwig K-H Schleifer (1999) ArticleTitleSpecific oligonucleotide probes for in situ detection of a major group of Gram-positive bacteria with low DNA G+C content. Syst Appl Microbiol 22 186–196 Occurrence Handle1:CAS:528:DyaK1MXks1Smtbs%3D Occurrence Handle10390869
Neef, A (1997) Anwendung der in situ-Einzelzell-Identifizierung von Bakterien zur Populations-Analyse in komplexen mikrobiellen Biozönosen. Ph.D. thesis, Technische Universität München
A Neef R Amann H Schlesner K-H Schleifer (1998) ArticleTitleMonitoring a widespread bacterial group: In situ detection of planctomycetes with 16S rRNA-targeted probes. Microbiology 144 3257–3266 Occurrence Handle1:CAS:528:DyaK1MXisVOrtg%3D%3D Occurrence Handle9884217
V Orphan L Taylor D Hafenbradl E Delong (2000) ArticleTitleCulture-dependent and culture-independent characterization of microbial assemblages associated with high-temperature petroleum reservoirs. Appl Environ Microbiol 66 700–711 Occurrence Handle10.1128/AEM.66.2.700-711.2000 Occurrence Handle1:CAS:528:DC%2BD3cXhtFeqs7w%3D Occurrence Handle10653739
K Pedersen (1997) ArticleTitleMicrobial life in deep granitic rock. FEMS Microbiol Rev 20 399–414 Occurrence Handle10.1016/S0168-6445(97)00022-3 Occurrence Handle1:CAS:528:DyaK2sXltl2msLw%3D
K Pedersen J Arlinger S Ekendahl L Hallbeck (1996a) ArticleTitle16S rRNA gene diversity of attached and unattached bacteria in boreholes along the access tunnel to the Äspö hard rock laboratory, Sweden. FEMS Microbiol Ecol 19 249–262 Occurrence Handle10.1016/0168-6496(96)00017-7 Occurrence Handle1:CAS:528:DyaK28XjsVOktb8%3D
K Pedersen J Arlinger L Hallbeck C Petterson (1996b) ArticleTitleDiversity and distribution of subterranean bacteria in groundwater at Oklo in Gabon, Africa, as determined by 16S rRNA gene sequencing. Mol Ecol 5 427–436 Occurrence Handle10.1046/j.1365-294X.1996.d01-320.x Occurrence Handle1:CAS:528:DyaK28Xktlynt78%3D
K Ravenschlag K Sahm C Knoblauch BB Jørgensen R Amann (2000) ArticleTitleCommunity structure, cellular rRNA content and activity of sulfate-reducing bacteria in marine arctic sediments. Appl Environ Microbiol 66 3592–3602 Occurrence Handle10.1128/AEM.66.8.3592-3602.2000 Occurrence Handle1:CAS:528:DC%2BD3cXlsFarsbc%3D Occurrence Handle10919825
K Ravenschlag K Sahm J Pernthaler R Amann (1999) ArticleTitleHigh bacterial diversity in permanently cold marine sediments. Appl Environ Microbiol 65 3982–3989 Occurrence Handle1:CAS:528:DyaK1MXlvFeqtbs%3D Occurrence Handle10473405
C Roller M Wagner R Amann W Ludwig K-H Schleifer (1994) ArticleTitleIn situ probing of gram-positive bacteria with high DNA G+C content using 23S rRNA-targeted oligonucleotides. Microbiology 140 2849–2858 Occurrence Handle1:CAS:528:DyaK2MXhslGlu7Y%3D Occurrence Handle8000548
K Sahm B MacGregor BB Jørgensen D Stahl (1999) ArticleTitleSulphate reduction and vertical distribution of sulphate-reducing bacteria quantified by rRNA slot-blot hybridization in a coastal marine sediment. Environ Microbiol 1 65–74 Occurrence Handle10.1046/j.1462-2920.1999.00007.x Occurrence Handle1:CAS:528:DyaK1MXit1Wks7Y%3D Occurrence Handle11207719
Schulte, U (1998) Isotopengeochemische Untersuchungen zur Charakterisierung biologisch gesteuerter Redoxprozesse in Aquiferen der Niederrheinischen Bucht. Ph.D. thesis, Ruhr-Universität, Bochum
U Schulte H Strauss A Bergmann P Obermann (1997) ArticleTitleIsotopenverhältnisse der Schwefel-und Kohlenstoffspezies aus Sedimenten und tiefen Grundwässern der Niederrheinischen Bucht. Grundwasser 3 103–110 Occurrence Handle10.1007/s767-1997-8531-4
U Schulte H Strauss J Detmers J Kuever (1999) ArticleTitleCharacterization of an aquifer system—stable isotopes and microbiology. J Conf Abstr 4 578
J Snaidr R Amann I Huber W Ludwig K-H Schleifer (1997) ArticleTitlePhylogenetic analysis and in situ identification of bacteria in activated sludge. Appl Environ Microbiol 63 2884–2896 Occurrence Handle1:CAS:528:DyaK2sXktlekt7Y%3D Occurrence Handle9212435
D Stahl R Amann (1991) Development and application of nucleic acid probes in bacterial systematics. ESaM Goodfellow (Eds) Nucleic Acid Techniques in Bacterial Systematics John Wiley & Sons Chichester, UK 205–248
O Strunk O Gross B Reichel M May S Hermann N Stuckmann M Nonhoff M Lenke A Ginhart A Vilbig T Ludwig A Bode K-H Schleifer W Ludwig (1999) ARB: a software environment for sequence data Department of Microbiology, Technische Universität München Munich, Germany
G Ulrich D Martino K Burger J Routh E Grossman J Ammerman J Suflita (1998) ArticleTitleSulfur cycling in the terrestrial subsurface: commensal interactions, spatial scales, and microbial heterogeneity. Microb Ecol 36 141–151 Occurrence Handle10.1007/s002489900101 Occurrence Handle1:CAS:528:DyaK1cXlsVOnurY%3D Occurrence Handle9688776
H Urukawa K Kita-Tsukamoto K Ohwada (1999) ArticleTitleMicrobial diversity in marine sediments from Sagami Bay and Tokyo Bay, Japan, as determined by 16S rRNA gene analysis. Microbiology 145 3305–3315 Occurrence Handle10589740
G Wallner R Amann W Beisker (1993) ArticleTitleOptimizing fluorescent in situ hybridization with rRNA-targeted oligonucleotide probes for flow cytometric identification of microorganisms. Cytometry 14 136–143 Occurrence Handle1:CAS:528:DyaK3sXisVylsrs%3D Occurrence Handle7679962
B Zarda G Mattison A Hess D Hahn P Höhener J Zeyer (1998) ArticleTitleAnalysis of bacterial and protozoan communities in an aquifer contaminated with monoaromatic hydrocarbons. FEMS Microbiol Ecol 27 141–152 Occurrence Handle10.1016/S0168-6496(98)00061-0 Occurrence Handle1:CAS:528:DyaK1cXmtl2ntr8%3D
I Zlatkin M Schneider F de Bruijn L Forney (1996) ArticleTitleDiversity among bacteria isolated from the deep subsurface. J Industr Microbiol 17 219–227 Occurrence Handle1:CAS:528:DyaK2sXltF2hsQ%3D%3D
Acknowledgements
This paper represents a publication of the Priority Program 546, “Geochemical processes with long-term effects in anthropogenically-affected seepage- and groundwater.” Financial support provided by the Deutsche Forschungsgemeinschaft is gratefully acknowledged. Jan Detmers, Katrin Knittel, and Jan Kuever were supported by the Max-Planck-Society. R. Amann is acknowledged for critical reading of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Detmers, J., Strauss, H., Schulte, U. et al. FISH Shows That Desulfotomaculum spp. Are the Dominating Sulfate-Reducing Bacteria in a Pristine Aquifer . Microb Ecol 47, 236–242 (2004). https://doi.org/10.1007/s00248-004-9952-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-004-9952-6