Abstract
Congenital heart disease (CHD) usually occurs sporadically, with only a minority of cases associated with a known genetic mechanism. Cardiac-specific transcription factors NKX2-5 and GATA4 play key roles in the mammalian heart development, and the affected cardiac tissues of CHD patients are prone to somatic mutations which thus participate in the pathogenesis of CHD. We collected 98 patients with sporadic CHD, extracted genomic DNA from cardiac tissues and blood, and then screened NKX2-5 and GATA4 genes using PCR-direct sequence analysis. A novel heterozygous missense mutation (c.907G > A, p.V303I) of NKX2-5 gene was identified in a patient with tetralogy of Fallots. Functional assay revealed that this mutant was associated with significantly reduced transcriptional activity. In addition, we found two known single-nucleotide polymorphisms (SNPs) (rs2277923, rs3729753) in NKX2-5 and two known SNPs (rs56166237, rs3729856) in GATA4. All variations identified in cardiac tissues were consistent with those of peripheral blood, and no somatic mutations were found in cardiac tissues. Our study shows no evidence of NKX2-5 and GATA4 somatic mutations playing a role in the pathogenesis of sporadic CHD.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Congenital heart disease (CHD) is a common human birth defect, with an incidence of 10 in 1000 live births [1]. It mostly occurs sporadically, and the pathogenesis remains unclear. In the past years, an increasing number of germline mutations of cardiac development-related transcription factor genes have been found in familiar and sporadic patients with CHD. NKX2-5 and GATA4, as crucial transcription factors, are involved in the morphogenesis of heart. Considerable mutations of NKX2-5 and GATA4 have been found in a variety of CHDs [2]. However, most of them were identified in familiar cases but not sporadic ones.
In recent years, many somatic mutations of NKX2-5, GATA4, and other cardiac transcription factor genes have been identified in the affected cardiac tissues of patients with CHD, suggesting that such mutations may play key roles in the pathogenesis of CHD [3,4,5,6,7,8]. Until now, such mutations have been identified in DNA extracted from formalin-fixed tissues for more than 20 years, without using corresponding peripheral blood as control. Besides, somatic mutations have never been found in the fresh frozen tissues of CHD patients hitherto [9,10,11,12,13,14,15,16].
To investigate whether somatic mutations of NKX2-5 and GATA4 genes participate in the pathogenesis of CHD, we collected the cardiac tissues and corresponding peripheral blood from CHD patients and detected the sequences of these two genes.
Materials and Methods
Patients
We recruited 98 unrelated patients with sporadic CHD who underwent surgery in Nanjing Children’s Hospital from June 2010 to December 2012, including tetralogy of Fallots (TOF, n = 26), atrial septal defects (ASD, n = 17), ventricular septal defects (VSD, n = 21), ASD with VSD (n = 15), total anomalous pulmonary venous connection (TAPVC) with ASD (n = 5), complete atrioventricular canals (n = 4), stenosis of right ventricular outflow tract (SRVOT, n = 4), double-chambered right ventricle (DCRV) with VSD (n = 3), Ebstein anomaly (n = 1), transposition of the great arteries (d-TGA, n = 1), and double outlet of right ventricle (DORV, n = 1). All patients were evaluated by echocardiography, and the diagnosis was confirmed by echocardiography, 64-slice spiral CT, and surgical intervention. Patients with chromosome anomalies, unknown syndromes, or 22q11.2 micro-deletion were excluded from this study. Meanwhile, 200 normal healthy individuals were collected as control. All included subjects had written informed consent, and this study was approved by the institutional ethical committee of Nanjing Medical University.
Sample Collection and Storage
Heart tissues were resected during cardiac surgery, cleaned using normal saline, then placed in liquid nitrogen immediately, and stored at − 80 °C at last. Tissues of the right auricle as unaffected tissues were obtained from all patients. Affected tissues including the right ventricular outflow tract (RVOT) were collected from patients with TOF, SRVOT, DCRV/VSD, and DORV; atrial septum tissues were collected from patients with TAPVC/ASD and d-TGA; and atrial tissue was obtained from a patient with Ebstein anomaly. All the tissues resected during surgery were immediately snap frozen in liquid nitrogen and then stored in a freezer at − 80 °C. Corresponding peripheral blood samples were collected prior to surgery, and stored at − 80 °C.
Chemicals, Reagents and Culture Medium
HEK-293 cells were purchased from the Cell Bank of the Chinese Academy of Sciences. Dulbecco’s modified Eagle’s medium (DMEM), fetal bovine serum, and penicillin–streptomycin solution were purchased from Gibco (Thermo Fisher Scientific, Waltham, MA, USA). Trypsin was purchased from Boguang Biotech Co., Ltd. (Shanghai, China). Trizol was purchased from Takara Bio (Dalian, China).
Genetic Analysis
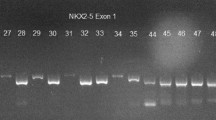
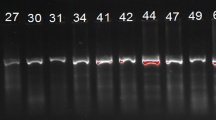
Genomic DNA was extracted from blood samples using TIANamp blood DNA kit (Tiangen, Beijing, China) and from cardiac tissues using QIAmp DNA mini kit (QIAGEN, Hilden, Germany) according to the manufacturers’ instructions. Primers were designed using Primer3 (v. 0.4.0) software (http://frodo.wi.mit.edu/primer3/) based on the cDNA sequences available in GenBank. All coding sequences and flanking introns of NKX2-5 (GenBank NM_004387) and GATA4 (GenBank NM_002052) genes were amplified by PCR. The PCR system contained 1× PCR buffer, 0.2 mmol/L dNTPs, 0.4 µmol/L of each primer, 100 ng genomic DNA, and 1 U Taq DNA polymerase (Takara, Tokyo, Japan) in a final volume of 25 µL. PCR was performed under the following conditions: 5 min at 95 °C, followed by 40 cycles of 30 s at 95 °C, 30 s at 58 °C, and 45 s at 72 °C. The PCR products were purified and directly sequenced using Big Dye Terminator (Applied Biosystems, Foster City, CA, USA) on an ABI Prism 3130 genetic analyzer (Applied Biosystems, Foster City, CA, USA).
Expression Plasmids and Site-directed Mutagenesis
Wild-type full-length human NKX2-5 cDNA (GenBank NM_001166175) was purchased and cloned into pcDNA3.1-3xFlag vector using ClonExpress Entry one-step cloning kit (Vazyme, China). The identified mutation was introduced into the wild-type NKX2-5 gene using the PCR-based Dpn I treatment method (Vazyme, China). The mutant was confirmed by bidirectional sequencing to exclude any other sequence variations.
Reporter Gene Assays
Human ANF promoter was inserted into pGL4.23 vector (Promega, Madison, USA) to produce ANF-Luc. ANF-Luc and an internal control reporter plasmid pGL4.74 (hRluc/TK; Promega) were used in transient transfection assay to evaluate the transcriptional activity of the NKX2-5 mutant. HEK-293 cells were seeded onto a 24-well plate and cultured in DMEM supplemented with 10% fetal bovine serum. The cells were transfected with 0.5 µg of wild-type or mutant NKX2-5, 1.0 µg of ANF-luc, and 0.1 µg of pGL4.74 using PolyJet transfection reagent (SignaGen). Firefly luciferase and Renilla luciferase activities were measured using the Dual-Glo luciferase assay system (Promega) 48 h after transfection. The activity of ANF promoter was presented as the fold activation of firefly luciferase relative to Renilla luciferase. Three independent experiments were performed.
Statistical Analysis
Statistical analysis was performed using SPSS V16.0. Two-sided χ2-test was used to evaluate the frequency of genotype and allele between case and control groups. P < 0.05 was considered statistically significant.
Results
Clinical Features
We collected a total of 98 sporadic, non-syndromic CHD patients consisting of 59 boys and 39 girls, with a mean age of (25.3 ± 32.9) months upon surgery. The youngest was only 2 days, and the oldest was 13.75 years old. This cohort included multiple types of CHD (Table 1).
Somatic Mutations
By direct sequencing, we identified a heterozygous missense mutation in exon 2 of NKX2-5 gene (c.907G > A, p.V303I) in both RVOT and auricula dextra tissues of a patient with TOF, leading to the substitution of valine by isoleucine, which was absent from the 200 controls. This mutation is localized in a conserved sequence and has not been reported before (Fig. 1). Then we performed computational predictive analysis using two algorithms, i.e., SIFT and PolyPhen. The SIFT analysis predicted mutation c.907G > A was TOLERATED with a score of 0.85, while the PolyPhen analysis gave a score of 0.869 (POSSIBLY DAMAGING).
A novel heterozygous mutation detected in NKX2-5 gene. a A heterozygous c.907G > A mutation of NKX2-5 detected in both cardiac tissue and peripheral blood of a patient with TOF. b Normal sequence of NKX2-5 as control. c Amino acid sequence comparison shows that Val 303 of NKX2-5 has high conservation between different species
Since the sequences of NKX2-5 and GATA4 fragments from the blood samples and fresh frozen tissues of the 98 CHD cases were identical, there was no somatic mutation.
Sequence Variations
Four reported single-nucleotide polymorphisms (SNPs) were found, i.e., rs2277923 c.63 A > G (p.E21E) and rs3729753 c.606 G > C (p.L202L) in NKX2-5 gene, and rs56166237 c.99G > T (p.A33A) and rs3729856 c.1129A > G (p.S377G) in GATA4 gene. All the variations were detected in the genomic DNA from both cardiac tissues and blood (Table 2). No significant differences were found in the allele frequencies of these SNPs between CHD patients and normal controls, accompanied by similar frequencies in the SNP database (http://www.ncbi.nlm.nih.gov/projects/SNP/) (Table 3).
Diminished Transcriptional Activity of NKX2-5 Mutant
To determine whether p.V303I affected the transcriptional activity of NKX2-5, ANF promoter luciferase reporter together with either wide-type or mutant NKX2-5 plasmid were co-transfected to HEK-293 cells. The transfection efficiency was quantified by Western blot. As shown in Fig. 2, p.V303I plasmid significantly attenuates the transactivation of ANF promoter (P < 0.05), demonstrating that target gene-induced activation of the mutant protein was suppressed.
Discussion
Cardiac transcription factors NKX2-5 and GATA4 play crucial roles in cardiac morphogenesis. Up to now, many germline mutations of NKX2-5 and GATA4 genes have been identified in CHD patients, especially in familial ones, but rare in sporadic CHD cases [17,18,19,20].
Somatic mosaicism means genetic variation in somatic cell occurs after fertilization, which is well-established to dominate in cancer, aging, and other diseases [21]. In recent years, somatic mutation has been involved in variety of cardiovascular diseases, including long-QT syndrome [22], vascular malformative/overgrowth disorders [23], and idiopathic atrial fibrillation [24, 25]. Interestingly, Reamon-Buettner et al. reported frequent somatic mutations of cardiac transcription factors NKX2.5, TBX5, GATA4, HEY2, and HAND1 in malformed regions but not unaffected ones of the heart, as a novel mechanism for CHD [3,4,5,6,7,8]. However, all the somatic mutations have been identified in cardiac tissues fixed by formalin for over 20 years, and no corresponding blood samples have been employed as control.
In this study, we used fresh frozen tissues instead of formalin-fixed tissues. RVOT, atrial septum, and atrial tissues were considered to be malformed, whereas the right atrial appendage was considered unaffected. We failed to found any somatic mutations of NKX2-5 or GATA4 gene. In addition, we identified a novel missense mutation of NKX2-5 gene in RVOT tissue of a patient with TOF, which was confirmed in the right atrial appendage and peripheral blood of the same case. Additionally, two known SNPs were identified in cardiac tissues and the peripheral blood DNA of corresponding patients. All the variations identified in cardiac tissues were confirmed in blood samples. Overall, there was no somatic variant.
Similarly, several recent studies were unable to find any somatic mutation in a series of transcription factors NKX2-5, GATA4, TBX20, and HAND1 using fresh frozen heart tissues of patients with CHD [9,10,11,12,13,14,15,16]. The most important difference between these studies and those of Reamon-Buettner et al. is the storage method of heart tissues, i.e., fresh frozen or formalin fixation. Long-term fixation in formalin causes DNA degradation and jeopardizes its quality. Furthermore, formalin may remain in the DNA solution, affecting PCR results [26]. Quach et al. reported that formalin damaged bases and affected PCR, which may induce mutation [27]. Therefore, a large number of mutations detected in the studies of Reamon-Buettner et al. may be ascribed to low-quality DNA [26, 27].
Given that mosaicism may reduce the detection rate of somatic mutations in affected tissue, the technique of sequence analysis and the clone area of mutant cells are determinative [28]. Therefore, whether somatic mutation exists in cardiac tissue upon CHD is still controversial, requiring further in-depth studies. In our CHD cohort, NKX2-5 or GATA4 gene had no somatic mutations. More sensitive mutation detection approaches may be necessary to find low-frequency somatic variants.
Although there were no somatic mutations of transcription factor NKX2-5 or GATA4, we identified a novel heterozygous missense mutation of NKX2-5 gene, with weakened transcriptional activity.
Conclusion
In summary, we sequenced the variations of NKX2-5 and GATA4 genes in the cardiac tissues of CHD patients using PCR-direct sequencing technique, finding no somatic mutations though. Given the detection methods and the sizes of tissues, further studies are needed to confirm whether somatic mutations are the pathogenesis of CHD. Moreover, a novel heterozygous missense mutation (c.907G > A, p.V303I) was identified in NKX2-5 in a TOF patient. Our findings expand the mutation spectrum of NKX2-5 and provide a new clue for clarifying the mechanism of CHD.
Abbreviations
- CHD:
-
Congenital heart disease
- TOF:
-
Tetralogy of Fallots
- ASD:
-
Atrial septal defects
- VSD:
-
Ventricular septal defects
- TAPVC:
-
Total anomalous pulmonary venous connection
- SRVOT:
-
Stenosis of right ventricular outflow tract
- DCRV:
-
Double-chambered right ventricle
- d-TGA:
-
Transposition of the great arteries
- DORV:
-
Double outlet of right ventricle
- RVOT:
-
Right ventricular outflow tract
- SNP:
-
Single-nucleotide polymorphism.
References
van der Linde D, Konings EE, Slager MA et al (2011) Birth prevalence of congenital heart disease worldwide: a systematic review and meta-analysis. J Am Coll Cardiol 58:2241–2247
Wessels MW, Willems PJ (2010) Genetic factors in non-syndromic congenital heart malformations. Clin Genet 78:103–123
Reamon-Buettner SM, Borlak J (2004) Somatic NKX2-5 mutations as a novel mechanism of disease in complex congenital heart disease. J Med Genet 41:684–690
Reamon-Buettner SM, Hecker H, Spanel-Borowski K, Craatz S, Kuenzel E, Borlak J (2004) Novel NKX2-5 mutations in diseased heart tissues of patients with cardiac malformations. Am J Pathol 164:2117–2125
Reamon-Buettner SM, Borlak J (2005) GATA4 zinc finger mutations as a molecular rationale for septation defects of the human heart. J Med Genet 42:e32
Reamon-Buettner SM, Borlak J (2006) HEY2 mutations in malformed hearts. Hum Mutat 27:118
Reamon-Buettner SM, Borlak J (2004) TBX5 mutations in non-Holt-Oram syndrome (HOS) malformed hearts. Hum Mutat 24:104
Reamon-Buettner SM, Ciribilli Y, Traverso I, Kuhls B, Inga A, Borlak J (2009) A functional genetic study identifies HAND1 mutations in septation defects of the human heart. Hum Mol Genet 18:3567–3578
Orjuela Quintero DC, Nunez F, Caicedo V, Pachon S, Salazar Salazar M (2014) Mutations in the GATA4 gen in patients with non-syndromic congenital heart disease. Invest Clin 55:207–216
Zheng J, Li F, Liu J et al (2015) Investigation of somatic NKX2-5 mutations in Chinese children with congenital heart disease. Int J Med Sci 12:538–543
Sabina S, Pulignani S, Rizzo M et al (2013) Germline hereditary, somatic mutations and microRNAs targeting-SNPs in congenital heart defects. J Mol Cell Cardiol 60:84–89
Esposito G, Butler TL, Blue GM et al (2011) Somatic mutations in NKX2-5, GATA4, and HAND1 are not a common cause of tetralogy of Fallot or hypoplastic left heart. Am J Med Genet A 155A:2416–2421
Salazar M, Consoli F, Villegas V et al (2011) Search of somatic GATA4 and NKX2.5 gene mutations in sporadic septal heart defects. Eur J Med Genet 54:306–309
Majumdar R, Yagubyan M, Sarkar G, Bolander ME, Sundt TM 3rd (2006) Bicuspid aortic valve and ascending aortic aneurysm are not associated with germline or somatic homeobox NKX2-5 gene polymorphism in 19 patients. J Thorac Cardiovasc Surg 131:1301–1305
Draus JM Jr, Hauck MA, Goetsch M, Austin EH 3rd (2009) Tomita-Mitchell A and Mitchell ME: investigation of somatic NKX2-5 mutations in congenital heart disease. J Med Genet 46:115–122
Wang J, Lu Y, Chen H, Yin M, Yu T, Fu Q (2011) Investigation of somatic NKX2-5, GATA4 and HAND1 mutations in patients with tetralogy of Fallot. Pathology 43:322–326
Elliott DA, Kirk EP, Yeoh T et al (2003) Cardiac homeobox gene NKX2-5 mutations and congenital heart disease: associations with atrial septal defect and hypoplastic left heart syndrome. J Am Coll Cardiol 41:2072–2076
McElhinney DB, Geiger E, Blinder J, Benson DW, Goldmuntz E (2003) NKX2.5 mutations in patients with congenital heart disease. J Am Coll Cardiol 42:1650–1655
Nemer G, Fadlalah F, Usta J et al (2006) A novel mutation in the GATA4 gene in patients with Tetralogy of Fallot. Hum Mutat 27:293–294
Schluterman MK, Krysiak AE, Kathiriya IS et al (2007) Screening and biochemical analysis of GATA4 sequence variations identified in patients with congenital heart disease. Am J Med Genet A 143A:817–823
Erickson RP (2014) Recent advances in the study of somatic mosaicism and diseases other than cancer. Curr Opin Genet Dev 26:73–78
Priest JR, Gawad C, Kahlig KM et al (2016) Early somatic mosaicism is a rare cause of long-QT syndrome. Proc Natl Acad Sci USA 113:11555–11560
Luks VL, Kamitaki N, Vivero MP et al (2015) Lymphatic and other vascular malformative/overgrowth disorders are caused by somatic mutations in PIK3CA. J Pediatr 166:1048–1054 (e1041–1045)
Gollob MH, Jones DL, Krahn AD et al (2006) Somatic mutations in the connexin 40 gene (GJA5) in atrial fibrillation. N Engl J Med 354:2677–2688
Thibodeau IL, Xu J, Li Q et al (2010) Paradigm of genetic mosaicism and lone atrial fibrillation: physiological characterization of a connexin 43-deletion mutant identified from atrial tissue. Circulation 122:236–244
Coura R, Prolla JC, Meurer L, Ashton-Prolla P (2005) An alternative protocol for DNA extraction from formalin fixed and paraffin wax embedded tissue. J Clin Pathol 58:894–895
Quach N, Goodman MF, Shibata D (2004) In vitro mutation artifacts after formalin fixation and error prone translesion synthesis during PCR. BMC Clin Pathol 4:1
Erickson RP (2010) Somatic gene mutation and human disease other than cancer: an update. Mutat Res 705:96–106
Acknowledgements
We thank all the patients and families who participated in this study. We would like to thank all the cardiothoracic surgeons from Nanjing Children’s Hospital for their support regarding CHD heart tissues, and all the doctors of clinical laboratory.
Funding
This study was funded by the National Natural Science Foundation of China (Grant Nos. 81670284, 81000076), Nanjing Science and Technology Project (Grant No. 201715057), and Nanjing Medical Science and Technique Development Foundation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical Approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Informed Consent
Informed consent was obtained from all individual participants included in the study.
Rights and permissions
About this article
Cite this article
Yin, J., Qian, J., Dai, G. et al. Search of Somatic Mutations of NKX2-5 and GATA4 Genes in Chinese Patients with Sporadic Congenital Heart Disease. Pediatr Cardiol 40, 17–22 (2019). https://doi.org/10.1007/s00246-018-1955-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00246-018-1955-z