Abstract
Variation in GC content is assumed to correlate with various processes, including mutation biases, recombination, and environmental parameters. To date, most genomic studies exploring the evolution of GC content have focused on nuclear genomes, but relatively few have concentrated on organelle genomes. We explored the mechanisms maintaining the GC content in angiosperm plastomes, with a particular focus on the hypothesis of phylogenetic dependence and the correlation with deletion mutations. We measured three genetic traits, namely, GC content, A/T tracts, and G/C tracts, in the coding region of plastid genomes for 1382 angiosperm species representing 350 families and 64 orders, and tested the phylogenetic signal. Then, we performed correlation analyses and revealed the variation in evolutionary rate of selected traits using RRphylo. The plastid GC content in the coding region varied from 28.10% to 43.20% across angiosperms, with a few non-photosynthetic species showing highly reduced values, highlighting the significance of functional constraints. We found strong phylogenetic signal in A/T tracts, but weak ones in GC content and G/C tracts, indicating adaptive potential. GC content was positively and negatively correlated with G/C and A/T tracts, respectively, suggesting a trade-off between these two deletion events. GC content evolved at various rates across the phylogeny, with significant increases in monocots and Lamiids, and a decrease in Fabids, implying the effects of some other factors. We hypothesize that variation in plastid GC content might be a mixed strategy of species to optimize fitness in fluctuating climates, partly through influencing the trade-off between AT → GC and GC → AT mutations.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
DNA base composition (GC content), defined as the proportion of cytosines and guanines relative to the total number of nucleotides in the genome, is a key feature of genome architecture and believed to play an important role in genome evolution and species biology (Nishio et al. 2003; Vinogradov 2003; Šmarda et al. 2014). Hence, exploring the dynamics of GC content evolution and drivers/causes could provide insight into genome adaptation in response to ecological fluctuation. In recent genomic studies, variation in GC content has often been shown to correlate with various factors, including phylogenetic relatedness (Stackebrandt and Liesack 1993), GC-biased gene conversion (gBGC, a process associated with recombination, Holmquist 1992; Eyre-Walker 1993; Duret and Galtier 2009; Muyle et al. 2011), mutational biases (Filipski 1987; Suoeka 1988), chromosome/genome structure (e.g., isochore, Eyre-Walker and Hurst 2001; Duret and Arndt 2008; Glémin et al. 2014), and ecological selection (Bernardi and Bernardi 1986; Eyre-Walker 1999; Hildebrand et al. 2010). In addition, researchers have also found that DNA methylation (Smith et al. 2009; Mugal et al. 2015), life-history traits (e.g., plant growth form, Trávníček et al. 2019), and genome size (Veselý et al. 2012; Lipnerová et al. 2013) have also influenced the GC content. To date, the dynamics of GC content evolution has been extensively studied in prokaryotes, vertebrates, and plants (Bentley and Parkhill 2004; Mann and Chen 2010; Eyre-Walker and Hurst 2001; Šmarda et al. 2014). However, most of these studies focused on nuclear genomes, and relatively few have concentrated on organelle genomes (i.e., chloroplast and mitochondrial genomes), in which the GC content is believed to be maintained independently of nuclear genomes (Kusumi and Tachida 2005).
The plastid GC content varies greatly among species, with the lowest value of 22.67% found in a parasitic plant Pilostyles hamiltonii (Bellot and Renner 2015), and the highest 56.50% in spikemoss Selaginella remotifolia (Zhang et al. 2019). A few hypotheses to explain this variation have been proposed (Wicke et al. 2013, 2016; Mower et al. 2019; Yu et al. 2020). Wicke et al (2013, 2016) noted that a reduced GC content accompanied by a lifestyle-specific shift to heterotrophy was caused by relaxed functional constraints on codon usage or nutrient economy. In addition, the highest plastid GC content in spikemoss is probably a consequence of a large number of RNA editing sites and reduced AT-mutation pressure (Smith 2009; Mower et al. 2019), although there is still a lack of empirical data supporting the latter hypothesis. In a recent phylogenomic study of liverworts, Yu et al (2020) pointed out that variation in plastid GC content not only reflected phylogenetic relatedness but also correlated with the diversity of poly-(G)/(C) tracts (G/C tracts, deletions/replications caused by DNA polymerase slippage, Viguera et al. 2001). Nevertheless, little is known about whether such observations are lineage specific or a more widespread phenomenon across all plants. Here, we explored the dynamics of GC content evolution and putative mechanisms in plastid genomes of angiosperms using a robust phylogeny and broad taxon sampling, with a particular focus on the hypothesis of phylogenetic dependence and the correlation with deletion mutations (both G/C tracts and A/T tracts).
As the most successful land plants and an important component of terrestrial ecosystems, angiosperms have been the focus of a large number of plastid genomic and phylogenomic studies (Qiu et al. 1999; Moore et al. 2007; Davis et al. 2014; Gitzendanner et al. 2018; Li et al. 2019). In these studies, a large number of plastid genomes were documented. As of the end of September 2020, a total of 4116 angiosperm plastid genomes are available in GenBank (https://www.ncbi.nlm.nih.gov/genome/organelle/). These reports provide not only a large number of genetic characters, enabling the investigation of the origin and diversification of angiosperms (Davis et al. 2014; Gitzendanner et al. 2018; Li et al. 2019), but also considerable information on the plastome architecture and assembly (Smith 2009; Wicke et al. 2013; Li et al. 2016; Niu et al. 2017; Mower et al. 2019). To achieve the above aim, we measured three genetic traits, namely, GC content, G/C tracts, and A/T tracts (per kb), in the coding region of plastid genomes for 1382 angiosperm species representing 350 families and 64 orders (APG IV 2016). Using a well-resolved phylogeny of angiosperms published recently (Li et al. 2019), we tested the phylogenetic signal of three selected traits and performed correlation analyses. Then, we reveal variation in the evolutionary rate of the selected traits across the phylogeny using a modified phylogenetic comparative analysis-RRphylo, and discuss possible drivers or causes in biological and ecological contexts.
Materials and Methods
Sampling, Phylogeny, and Data Collection
The chloroplast phylogenomic study of angiosperms performed by Li et al (2019) provided not only the most likely family-level backbone of this group to date, but also a suitable genome-scale dataset-consisting of 80 plastid genes, allowing evolution of GC content to be explored without accounting for the noisy signals arising from frequent reshuffling in noncoding regions (Glémin et al. 2014). To reduce the biases caused by missing data, we pruned the maximum clade credibility (MCC) tree of Li et al. to include 1382 species (Table S1) representing 350 families (84% of family diversity) and 64 orders (100% of order diversity, APG IV 2016) using the function “drop.tip” (package ape; Paradis et al. 2004; Paradis 2012). Only one species with the longest sequence length from each genus was sampled. The sampling represents not only the greatest order and family diversity of angiosperms to date, but also ecological, lifestyle, and life-history diversity. Using the dataset of Li et al., we estimated three genetic traits, namely, GC content, G/C tracts, and A/T tracts [the number of poly-(dN) tracts per kb, only poly-(dN) tracts with a length ≥ 3b were calculated; Table S1], for all samples used in the phylogeny.
Test of Phylogenetic Signal, Correlations, and Evolutionary Rate Variation
We tested the phylogenetic signal (a tendency of related species to resemble each other more than species drawn at random from the phylogeny; the concept follows Münkemüller et al. 2012) of three genetic traits using two statistics: Blomberg’s K (Blomberg et al. 2003) and Pagel’s λ (Pagel 1999). The K-statistic provides a reliable effect size measure and performs well in all conditions (Münkemüller et al. 2012). K is estimated as the ratio of the observed mean-squared errors and the mean-squared errors using the variance–covariance matrix derived from a given phylogeny under the assumption of Brown motion model (BM, Blomberg et al. 2003). K < 1 indicates that a trait has less phylogenetic signal than expected under BM. The λ-statistic is most suitable to capture the effect of changing evolutionary rates in simulation experiments (Münkemüller et al. 2012). λ is defined as the transformation of the phylogeny that fits trait data best to BM (Pagel 1999; Freckleton et al. 2002). λ < 1 indicates that relatives show less similarity than expected, while λ > 1 suggests the opposite. To assess the effects of topological uncertainty on the estimated phylogenetic signal (Revell et al. 2008), we randomly selected 100 trees from the posterior sampling after excluding burn-in trees. Both statistics were performed in the MCC and 100 randomly selected trees using the function “phylosig” (package phytools; Pagels 1999; Blomberg et al. 2003; Ives et al. 2007; Revell 2012). All trees are deposited in Figshare Digital Repository (https://doi.org/10.6084/m9.figshare.12901517).
DNA polymerase slippage could cause both G/C tracts and A/T tracts (Viguera et al. 2001). To determine whether GC content accelerates accumulation of G/C tracts or all deletion events, we tested the correlation between GC content and two deletions events separately and as a whole, using the IPC analysis. Furthermore, we performed the correlation analyses in all angiosperms as well as three major subclades, namely, monocots, superrosids, and superasterids, using the PDAP plugin (Midford et al. 2005) for Mesquite (Maddison and Maddison 2018).
RRphylo is a modified phylogenetic comparative method recently developed by Castiglione et al (2018). This method has the advantage of assigning an evolutionary rate to each branch of a phylogeny, dealing with both extinct and extant phylogenies, and low Type I and Type II error rates (Castiglione et al. 2018). Recently, this method has been successfully applied in macroevolutionary studies of animals and humans (Piras et al. 2018; Raia et al. 2018; Sansalone et al. 2020). In this study, we used RRphylo to reveal the variation in evolutionary rate of three selected traits and to identify potential rate shifts. This analysis was performed using the functions “RRphylo” and “search.shift” (Castiglione et al. 2018; Piras et al. 2018).
Results
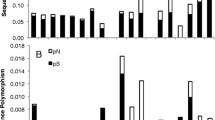
Three genetic traits have a wide variation range, including 28.10%–43.20% in GC content, 8.27–23.37 per kb in G/C tracts, and 34.93–65.32 per kb in A/T tracts (Fig. 1, Table S1). Interestingly, the amount of A/T tracts was almost three times that of G/C tracts (the mean ratio of A/T tracts to G/C tracts was 2.7, Table S1). Using the K-statistic, only A/T tracts displayed a strong phylogenetic signal, with a K = 1.30 [1.25, 1.34] (p < 0.001), while the other two did not, as indicated by K < 0 (K = 0.64 [0.64, 0.68], p < 0.001 in GC content, and 0.61 [0.59, 0.63], p < 0.001 in G/C tracts, Table S2). In contrast, using the λ-statistic, all three genetic traits showed a nearly BM pattern, with λ ≈ 1.0 (λ = 0.962 [0.960, 0.966], p < 0.001 in GC content, 0.988 [0.987, 0.990], p < 0.001 in G/C tracts, and 0.954 [0.951, 0.957], p < 0.001 in A/T tracts, Table S2).
Variation of GC content, and A/T and G/C tracts per kb in the coding region of plastid genomes across 1382 angiosperm species using the dataset of Li et al (2019). Box bars show the minimum-to-maximum range (whiskers), interquartile range (blue boxes), median (black bars), and outliers (empty circles)
The GC content was positively correlated with G/C tracts with a coefficient > 0.6 (p < 0.001) but negatively correlated with A/C tracts with a coefficient < − 0.75 (p < 0.001) and the total deletion mutations with a coefficient < − 0.50 (p < 0.001, Table 1). These correlations were consistently supported in all angiosperms and three major subclades (Table 1). In addition, the negative correlation between G/C tracts and A/T tracts was only found in monocots.
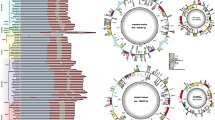
Across the angiosperm phylogeny, we found significant increases in three traits on the internal branches of monocots especially Poales but decreases in superrosids (e.g., Fabids). The additional increases in GC content and decreases in A/T tracts occurred in superasteridae (e.g., Lamiids, Fig. 2).
Reconstructing evolution of GC content, and the diversity of A/T and G/C tracts across the phylogeny of angiosperms generated recently by Li et al. (2019). Colored circles indicate significant increase and decrease (p < 0.01) in each trait identified using RRphylo. Branches in red subtend to non-photosynthetic genera
Discussion
Variation in Plastid GC Content, G/C Tracts, and A/T Tracts Across the Phylogeny
The chloroplast genome contains a subset of genes encoding proteins that are crucial for photosynthesis and some other metabolic processes, such as the cytochrome b6f complex and ATP synthase (Martin et al. 2002). Hence, understanding its evolution is fundamental to comprehend the adaptation, diversification, and ecomorphospace evolution of modern plants. A comparative analysis of 1382 plastomes with focus on the coding region from species spanning the breadth of extant angiosperms revealed some important characteristics of the dynamic evolution of GC content and deletion mutations. The relatively low GC content occurring in a few non-photosynthetic species coincides with massive gene arrangements (e.g., partial or complete loss or transfer to other genomes, Wicke et al. 2013, 2016; Schneider et al. 2018a, b; Wicke and Naumann 2018). These findings not only reconfirmed the hypothesis of functional constraints on photosynthesis as previously proposed (Wicke and Naumann 2018) but also implied that some other processes that determined which genes or proteins were retained in plastids, such as inefficient protein import and regulatory coupling of genes, may be responsible for variation in plastid GC content across non-green plants (Daley and Whelan 2005; Barbrook et al. 2006; Wicke et al. 2016).
The accumulation of both G/C tracts and A/T tracts is regulated by the DNA polymerase/mismatch repair system (Akashi and Yoshikawa 2013), but these two events showed great variation in diversity: the amount of A/T tracts was nearly three times that of G/C tracts (Table 1). This observation suggested a bias toward accumulation of A/T tracts over accumulation of G/C tracts in the coding region of angiosperm plastomes. A similar conclusion was also reached in previous studies (Eyre-Walker 1999; Smith and Eyre-Walker 2001; Massouh et al. 2016). This pattern was considered to be a consequence of the lower biochemical stability and higher energy cost of a G/C pair compared with an A/T pair, the limitation of available resources (Rocha and Danchin 2002; Akashi and Yoshikawa 2013), and selections for protection against inactivation and high mutability, considering the relative mutation rates of mononucleotide repeats (Boyer et al. 2002; Gragg et al. 2002).
Phylogenetic Signal, Correlations, and Variation in the Evolutionary Rate
The taxonomic value of GC content has been widely recognized in taxonomic studies of micro-organisms (Stackebrandt and Liesack 1993; Johnson and Whitman 2007; Tindall et al 2010) and phylogenomic studies of plants (Šmarda et al. 2014; Yu et al. 2020). However, it was also argued recently that distinct species living in the same environmental conditions tend to show similar GC content (Foerstner et al. 2005; Mann and Chen 2010). In this study, we failed to detect strong phylogenetic signals in GC content and G/C tracts in plastid genomes of angiosperms using the K-statistic (Table 1), indicating that close relatives are less similar than expected under a Brownian motion model of trait evolution. This pattern could be resulted either from adaptive radiations in which close relatives rapidly differentiate to fill new niches or from convergent evolution (Kamilar and Cooper 2013). In contrast, evolution of A/T tracts (per kb) showed a strong phylogenetic signal, making it an informative feature that could be used in the taxonomy of flowering plants. Nevertheless, we cannot completely rule out the possibility of “measurement errors” using the present dataset (Blomberg et al. 2003), in relation to disproportional sampling, topological uncertainty, and errors in branch length.
The mutation biases are considered as being among the major causes for variation in GC content (Filipski 1987; Suoeka 1988), and high GC content in turn was assumed to accelerate the rate of all mutations, including single base substitutions and deletions/replications (Kiktev et al. 2018). These assumptions were partly supported in this study, as we found evidence supporting that GC content was positively correlated with G/C tracts. However, the negative correlation with A/T tracts simultaneously identified in this study raised the possibility of a trade-off between accumulation of the two deletion events, G/C and A/T tracts, a process that was probably associated with competition for limited energy/resources (Rocha and Danchin 2002; Hellweger et al. 2018). Under this assumption, the accumulation of G/C tracts is directly affected by GC content, while the accumulation of A/T tracts depend heavily on the availability of energy/resources. In addition, the heterogeneity of energy costs for different base pairs (i.e., higher energy cost for a G/C pair than for an A/T pair) may be responsible for the reduced number of all deletion events as GC content increases, as long as the above trade-off is taken into account. In this respect, variation in plastid GC content might be a mixed strategy for species to optimize fitness in fluctuating environments, partly through influencing the trade-off between GC → AT and AT → GC mutations (both single base substitution and deletions). Nevertheless, little is currently known about how GC content responds to ecological fluctuation.
The heterogeneity of the evolutionary rate of plastid GC content across angiosperms suggested that some other factors, rather than functional constraints on photosynthesis, have shaped the evolution of this trait. One possibility is selection for a broader tolerance range. In monocots, Šmarda et al. (2014) proposed that increased GC content in grasses (Poaceae) may facilitate complex gene regulation, and consequently, favor these groups to grow in seasonally cold and dry climates. If so, an increase in plastid GC content of grass family may also be a response to such stressful environments, given the significant functions of plastid genes in energy and material metabolism, as well as the connections between plastid and nuclear genomes in gene regulation and assembly (Martin et al. 2002). Another possibility is plant size. This trait is often associated with mutation rate across angiosperms (Lanfer et al. 2013) and ferns (Barrera-Redondo et al. 2018): taller vascular plants tend to have slower substitution rates than smaller ones. One line of evidence consistent with this hypothesis is that taller palms showed both slower mutation rate (Barret et al. 2016) and lower GC content than their herb relatives, e.g., Poales (Table S1). Coincidently, in this study, we found a significant increase of GC content in small-mean-sized angiosperm groups, for example, Poales and Lamiids, in which most members are herbs, and a decrease in a few large-mean-sized clades, for example, Rosids, in which most members are taller trees. Apart from these two factors, some others that have been proposed to explain the variation in plastid GC content in several lineages, should also be taken into account, such as the frequency of RNA editing (Smith 2009) and gBGC (Niu et al. 2017). In general, the above hypotheses still need to be verified across a broad range of species diversity.
This study explored the dynamic evolution of GC content in the coding region of plastid genomes across angiosperms using a comprehensive phylogeny and a large taxon-character dataset. Our results not only provide evidence to support the hypothesis of adaptive evolution of GC content and G/C tracts but also revealed the complex correlations between GC content and diversity of mononucleotide repeats. This work also implies that variation in plastid GC content of angiosperms may be attributed to a combination of various factors, such as functional constraints on photosynthesis, selection for a broad tolerance range, competition for available energy/resources, and plant size. Nevertheless, some crucial issues about the biological and ecological significance of plastid GC content remain unknown, such as whether variation in plastid GC content could reflect the ecological distribution range, or mutation rate; whether plastid GC content has evolved independently from floral/lifestyle traits, and to what extent the variation in GC content is heritable.
References
Akashi M, Yoshikawa H (2013) Relevance of GC content to the conservation of DNA polymerase iii/mismatch repair system in gram-positive bacteria. Front Microbiol 4:266. https://doi.org/10.3389/fmicb.2013.00266
Barbrook AC, Howe CJ, Purton S (2006) Why are plastid genomes retained in non-photosynthetic organisms? Trends Plant Sci 11:101–108. https://doi.org/10.1016/j.tplants.2005.12.004
Barrera-Redondo J, Ramírez-Barahona S, Eguiarte LE (2018) Rates of molecular evolution in tree ferns are associated with body size, environmental temperature, and biological productivity. Evolution 72:1050–1062. https://doi.org/10.1111/evo.13475
Barrett CF, Baker WJ, Comer JR, Conran JG, Lahmeyer SC, Leebens-Mack JH, Li J, Lim GS, Mayfield-Jones DR, Perez L, Medina J, Pires JC, Santos C, Wm Stevenson D, Zomlefer WB, Davis JI (2016) Plastid genomes reveal support for deep phylogenetic relationships and extensive rate variation among palms and other commelinid monocots. New Phytol 209:855–870. https://doi.org/10.1111/nph.13617
Bellot S, Renner SS (2015) The plastomes of two species in the endoparasite genus Pilostyles (apodanthaceae) each retain just five or six possibly functional genes. Genome Biol Evol 8:189–201. https://doi.org/10.1093/gbe/evv251
Bentley SD, Parkhill J (2004) Comparative genomic structure of prokaryotes. Ann Rev Genet 38:771–792. https://doi.org/10.1146/annurev.genet.38.072902.094318
Bernardi G, Bernardi G (1986) Compositional constraints and genome evolution. J Mol Evol 24:1–11. https://doi.org/10.1007/bf02099946
Blomberg SP, Garland T Jr, Ives AR (2003) Testing for phylogenetic signal in comparative data: Behavioral traits are more labile. Evolution 57:717–745. https://doi.org/10.1111/j.0014-3820.2003.tb00285.x
Boyer JC, Yamada NA, Roques CN, Hatch SB, Riess K, Farber RA (2002) Sequence dependent instability of mononucleotide microsatellites in cultured mismatch repair proficient and deficient mammalian cells. Hum Mol Genet 11:707–713. https://doi.org/10.1093/hmg/11.6.707
Castiglione S, Tesone G, Piccolo M, Melchionna M, Mondanaro A, Serio C, Di Febbraro M, Raia P (2018) A new method for testing evolutionary rate variation and shifts in phenotypic evolution. Methods in Ecol Evol 9:974–983. https://doi.org/10.1111/2041-210x.12954
Daley DO, Whelan J (2005) Why genes persist in organelle genomes. Genome Biol 6:110. https://doi.org/10.1186/gb-2005-6-5-110
Davis CC, Xi ZX, Mathews S (2014) Plastid phylogenomics and green plant phylogeny: Almost full circle but not quite there. BMC Biol 12:11. https://doi.org/10.1186/1741-7007-12-11
Duret L, Arndt PF (2008) The impact of recombination on nucleotide substitutions in the human genome. PLoS Genet 4:e1000071. https://doi.org/10.1371/journal.pgen.1000071
Duret L, Galtier N (2009) Biased gene conversion and the evolution of mammalian genomic landscapes. Annu Rev Genom Hum G 10:285–311. https://doi.org/10.1146/annurev-genom-082908-150001
Eyre-Walker A (1993) Recombination and mammalian genome evolution. P Roy Soc B-Biol Sci 252:237–243. https://doi.org/10.1098/rspb.1993.0071
Eyre-Walker A (1999) Evidence of selection on silent site base composition in mammals: Potential implications for the evolution of isochores and junk DNA. Genetics 152:675–683
Eyre-Walker A, Hurst L (2001) The evolution of isochores. Nat Rev Genet 2:549–555. https://doi.org/10.1038/35080577
Filipski J (1987) Correlation between molecular clock ticking, codon usage, fidelity of DNA repair, chromosome banding and chromatin compactness in germline cells. FEBS Lett 217:184–186. https://doi.org/10.1016/0014-5793(87)80660-9
Foerstner KU, von Mering C, Hooper SD, Bork P (2005) Environments shape the nucleotide composition of genomes. EMBO Rep 6:1208–1213. https://doi.org/10.1038/sj.embor.7400538
Freckleton RP, Harvey PH, Pagel M (2002) Phylogenetic analysis and comparative data: A test and review of evidence. Am Nat 160:712–726. https://doi.org/10.1086/343873
Gitzendanner MA, Soltis PS, Wong GKS, Ruhfel BR, Soltis DE (2018) Plastid phylogenomic analysis of green plants: A billion years of evolutionary history. Am J Bot 105:291–301. https://doi.org/10.1002/ajb2.1048
Glémin S, Clément Y, David J, Ressayre A (2014) GC content evolution in coding regions of angiosperm genomes: A unifying hypothesis. Trends Genet 30:263–270. https://doi.org/10.1016/j.tig.2014.05.002
Gragg H, Harfe BD, Jinks-Robertson S (2002) Base composition of mononucleotide runs affects DNA polymerase slippage and removal of frameshift intermediates by mismatch repair in saccharomyces cerevisiae. Mol Cell Biol 22:8756–8762. https://doi.org/10.1128/MCB.22.24.8756-8762.2002
Hellweger FL, Huang YJ, Luo HW (2018) Carbon limitation drives GC content evolution of a marine bacterium in an individual-based genome-scale model. Isme J 12:1180–1187. https://doi.org/10.1038/s41396-017-0023-7
Hildebrand F, Meyer A, Eyre-Walker A (2010) Evidence of selection upon genomic GC-content in bacteria. PLoS Genet 6:e1001107. https://doi.org/10.1371/journal.pgen.1001107
Holmquist GP (1992) Chromosome bands, their chromatin flavors, and their functional features. Am J Hum Genet 51:17–37
Ives AR, Midford PE, Garland T Jr (2007) Within-species variation and measurement error in phylogenetic comparative methods. Syst Biol 56:252–270. https://doi.org/10.1080/10635150701313830
Johnson JL, Whitman WB (2007) Similarity analysis of DNAs. In: Reddy CA, Beveridge TJ, Breznak JA, Marzluf GA, Schmidt TM, Snyder LR (eds) Methods for General and molecular Microbiology. American Society for Microbiology, Washington, DC, pp 624–652
Kamilar JM, Cooper N (2013) Phylogenetic signal in primate behaviour, ecology and life history. Phil Trans R Soc B 368:20120341. https://doi.org/10.1098/rstb.2012.0341
Kiktev DA, Sheng ZW, Lobachev KS, Petes TD (2018) GC content elevates mutation and recombination rates in the yeast saccharomyces cerevisiae. Proc Natl Acad Sci U S A 115:E7109–E7118. https://doi.org/10.1073/pnas.1807334115
Kusumi J, Tachida H (2005) Compositional properties of green-plant plastid genomes. J Mol Evol 60:417–425. https://doi.org/10.1007/s00239-004-0086-8
Lanfear R, Ho SYW, Jonathan Davies T, Moles AT, Aarssen L, Swenson NG, Warman L, Zanne AE, Allen AP (2013) Taller plants have lower rates of molecular evolution. Nat Commun 4:1879. https://doi.org/10.1038/ncomms2836
Li FW, Kuo LY, Pryer KM, Rothfels CJ (2016) Genes translocated into the plastid inverted repeat show decelerated substitution rates and elevated GC content. Genome Biol Evol 8:2452–2458. https://doi.org/10.1093/gbe/evw167
Li HT, Yi TS, Gao LM, Ma PF, Zhang T, Yang JB, Gitzendanner MA, Fritsch PW, Cai J, Luo Y, Wang H, van der Bank M, Zhang SD, Wang QF, Wang J, Zhang ZR, Fu CN, Yang J, Hollingsworth PM, Chase MW, Soltis DE, Soltis PS, Li DZ (2019) Origin of angiosperms and the puzzle of the Jurassic gap. Nat Plants 5:461–470. https://doi.org/10.1038/s41477-019-0421-0
Lipnerová I, Bureš P, Horová L, Šmarda P (2013) Evolution of genome size in Carex (Cyperaceae) in relation to chromosome number and genomic base composition. Ann Bot 111:79–94. https://doi.org/10.1093/aob/mcs239
Maddison, W. P. and D.R. Maddison (2018) Mesquite: a modular system for evolutionary analysis. Version 3.51. http://www.mesquiteproject.org.
Mann S, Chen YP (2010) Bacterial genomic G+C composition-eliciting environmental adaptation. Genomics 95:7–15. https://doi.org/10.1016/j.ygeno.2009.09.002
Martin W, Rujan T, Richly E, Hansen A, Cornelsen S, Lins T, Leister D, Stoebe B, Hasegawa M, Penny D (2002) Evolutionary analysis of Arabidopsis, cyanobacterial, and chloroplast genomes reveals plastid phylogeny and thousands of cyanobacterial genes in the nucleus. Proc Natl Acad Sci U S A 99:12246–12251. https://doi.org/10.1073/pnas.182432999
Massouh A, Schubert J, Yaneva-Roder L, Ulbricht-Jones ES, Zupok A, Johnson MT, Wright SI, Pellizzer T, Sobanski J, Bock R, Greiner S (2016) Spontaneous chloroplast mutants mostly occur by replication slippage and show a biased pattern in the plastome of Oenothera. Plant Cell 28:911–929. https://doi.org/10.1105/tpc.15.00879
Midford PE, Garland Jr T, Maddison WP (2005) PDAP Package of Mesquite. Version 1.07. http://mesquiteproject.org/pdap_mesquite/index.html
Moore MJ, Bell CD, Soltis PS, Soltis DE (2007) Using plastid genome-scale data to resolve enigmatic relationships among basal angiosperms. Proc Natl Acad Sci U S A 104:19363–19368. https://doi.org/10.1073/pnas.0708072104
Mower JP, Ma PF, Grewe F, Taylor A, Michael TP, VanBuren R, Qiu YL (2019) Lycophyte plastid genomics: Extreme variation in GC, gene and intron content and multiple inversions between a direct and inverted orientation of the rRNA repeat. New Phytol 222:1061–1075. https://doi.org/10.1111/nph.15650
Mugal CF, Arndt PF, Holm L, Ellegren H (2015) Evolutionary consequences of DNA methylation on the GC content in vertebrate genomes G3(5):441–447. https://doi.org/10.1534/g3.114.015545
Münkemuller T, Lavergne S, Bzeznik B, Dray S, Jombart T, Schiffers K, Thuiller W (2012) How to measure and test phylogenetic signal. Methods Ecol Evol 3:743–756. https://doi.org/10.1111/j.2041-210X.2012.00196.x
Muyle A, Serres-Giardi L, Ressayre A, Escobar J, Glemin S (2011) GC-biased gene conversion and selection affect GC content in the Oryza genus (Rice). Mol Biol Evol 28:2695–2706. https://doi.org/10.1093/molbev/msr104
Nishio Y, Nakamura Y, Kawarabayasi Y, Usuda Y, Kimura E, Sugimoto S, Matsui K, Yamagishi A, Kikuchi H, Ikeo K, Gojobori T (2003) Comparative complete genome sequence analysis of the amino acid replacements responsible for the thermostability of Corynebacterium efficiens. Genome Res 13:1572–1579. https://doi.org/10.1101/gr.1285603
Niu ZT, Xue QY, Wang H, Xie XZ, Zhu SY, Liu W, Ding XY (2017) Mutational biases and GC-biased gene conversion affect GC content in the plastomes of dendrobium genus. Int J Mol Sci 18:2307. https://doi.org/10.3390/ijms18112307
Pagel M (1999) Inferring the historical patterns of biological evolution. Nature 401:877–884. https://doi.org/10.1038/44766
Paradis E (2012) Analysis of Phylogenetics and Evolution with R, 2nd edn. Springer, New York
Paradis E, Claude J, Strimmer K (2004) APE: analyses of phylogenetics and evolution in R language. Bioinformatics 20:289–290. https://doi.org/10.1093/bioinformatics/btg412
Piras P, Silvestro D, Carotenuto F, Castiglione S, Kotsakis A, Maiorino L, Melchionna M, Mondanaro A, Sansalone G, Serio C, Vero VA, Raia P (2018) Evolution of the sabertooth mandible: A deadly ecomorphological specialization. Palaeogeogr Palaeoclimatol Palaeoecol 496:166–174. https://doi.org/10.1016/j.palaeo.2018.01.034
Qiu YL, Lee J, Bernasconi-Quadroni F, Soltis DE, Soltis PS, Zanis M, Zimmer EA, Chen Z, Savolainen V, Chase MW (1999) The earliest angiosperms: Evidence from mitochondrial, plastid and nuclear genomes. Nature 402:404–407. https://doi.org/10.1038/46536
Raia P, Boggioni M, Carotenuto F, Castiglione S, Di Febbraro M, Di Vincenzo F, Melchionna M, Mondanaro A, Papini A, Profico A, Serio C, Veneziano A, Vero VA, Rook L, Meloro C, Manzi G (2018) Unexpectedly rapid evolution of mandibular shape in hominins. Sci Rep 8:7340. https://doi.org/10.1038/s41598-018-25309-8
Revell LJ (2012) Phytools: An R package for phylogenetic comparative biology (and other things). Methods Ecol Evol 3:217–223. https://doi.org/10.1111/j.2041-210X.2011.00169.x
Revell LJ, Harmon LJ, Collar DC (2008) Phylogenetic signal, evolutionary process, and rate. Syst Biol 57:591–601. https://doi.org/10.1080/10635150802302427
Rocha EP, Danchin A (2002) Base composition bias might result from competition for metabolic resources. Trends Genet 18:291–294. https://doi.org/10.1016/S0168-9525(02)02690-2
Sansalone G, Castiglione S, Raia P, Archer M, Dickson B, Hand S, Piras P, Profico A, Wroe S (2020) Decoupling functional and morphological convergence, the study case of fossorial mammalia. Front in Earth Sci 8:112. https://doi.org/10.3389/feart.2020.00112
Schneider AC, Braukmann T, Banerjee A, Stefanovic S (2018a) Convergent plastome evolution and gene loss in holoparasitic Lennoaceae. Genome Biol Evol 10:2663–2670. https://doi.org/10.1093/gbe/evy190
Schneider AC, Chun H, Stefanovic S, Baldwin BG (2018b) Punctuated plastome reduction and host-parasite horizontal gene transfer in the holoparasitic plant genus Aphyllon. P Roy Soc B-Biol Sci 285:20181535. https://doi.org/10.1098/rspb.2018.1535
Šmarda P, Bureš P, Horová L, Leitch IJ, Mucina L, Pacini E, Tichý L, Grulich V, Rotreklová O (2014) Ecological and evolutionary significance of genomic GC content diversity in monocots. Proc Natl Acad Sci U S A 111:E4096-4102. https://doi.org/10.1073/pnas.1321152111
Smith DR (2009) Unparalleled GC content in the plastid DNA of Selaginella. Plant Mol Biol 71:627–639. https://doi.org/10.1007/s11103-009-9545-3
Smith NG, Eyre-Walker A (2001) Synonymous codon bias is not caused by mutation bias in G+C-rich genes in humans. Mol Biol Evol 18:982–986. https://doi.org/10.1093/oxfordjournals.molbev.a003899
Stackebrandt E, Liesack W, Nucleic acid and classification, (1993). In: Goodfellow M, O’Donnell AG (eds) Handbook of new bacterial systematics. Academic Press, London, pp 151–194
Sueoka N (1988) Directional mutation pressure and neutral molecular evolution. Proc Natl Acad Sci U S A 85:2653–2657. https://doi.org/10.1073/pnas.85.8.2653
The Angiosperm Phylogeny Group IV (2016) An update of the angiosperm phylogeny group classification for the orders and families of flowering plants: APG IV. Bot J Linn Soc 181:1–20. https://doi.org/10.1111/boj.12385
Tindall BJ, Rosselló-Móra R, Busse HJ, Ludwig W, Kämpfer P (2010) Notes on the characterization of prokaryote strains for taxonomic purposes. Int J Syst Evol Microbiol 60:249–266. https://doi.org/10.1099/ijs.0.016949-0
Trávníček P, Čertner M, Ponert J, Chumová Z, Jersáková J, Suda J (2019) Diversity in genome size and GC content shows adaptive potential in orchids and is closely linked to partial endoreplication, plant life-history traits and climatic conditions. New Phytol 224:1642–1656. https://doi.org/10.1111/nph.15996
Veselý P, Bureš P, Šmarda P, Pavlíček T (2012) Genome size and DNA base composition of geophytes: The mirror of phenology and ecology? Ann Bot 109:65–75. https://doi.org/10.1093/aob/mcr267
Viguera E, Canceill D, Ehrlich SD (2001) Replication slippage involves DNA polymerase pausing and dissociation. EMBO J 20:2587–2595. https://doi.org/10.1093/emboj/20.10.2587
Vinogradov AE (2003) DNA helix: The importance of being GC-rich. Nucleic Acids Res 31:1838–1844. https://doi.org/10.1093/nar/gkg296
Wicke S, Muller KF, de Pamphilis CW, Quandt D, Wickett NJ, Zhang Y, Renner SS, Schneeweiss GM (2013) Mechanisms of functional and physical genome reduction in photosynthetic and nonphotosynthetic parasitic plants of the broomrape family. Plant Cell 25:3711–3725. https://doi.org/10.1105/tpc.113.113373
Wicke S, Muller KF, de Pamphilis CW, Quandt D, Bellot S, Schneeweiss GM (2016) Mechanistic model of evolutionary rate variation en route to a nonphotosynthetic lifestyle in plants. Proc Natl Acad Sci U S A 113:9045–9050. https://doi.org/10.1073/pnas.1607576113
Wicke S, Naumann J (2018) Molecular evolution of plastid genomes in parasitic flowering plants. In: Chaw SM and Jansen R (eds) Plastid genome evolution. Adv Bot Res 85: 315–347. https://doi.org/https://doi.org/10.1016/bs.abr.2017.11.014
Yu Y, Yang JB, Ma WZ, Pressel S, Liu HM, Wu YH, Schneider H (2020) Chloroplast phylogenomics of liverworts: A reappraisal of the backbone phylogeny of liverworts with emphasis on Ptilidiales. Cladistics 36:184–193. https://doi.org/10.1111/cla.12396
Zhang HR, Xiang QP, Zhang XC (2019) The unique evolutionary trajectory and dynamic conformations of DR and IR/DR-coexisting plastomes of the early vascular plant selaginellaceae (lycophyte). Genome Biol Evol 11:1258–1274. https://doi.org/10.1093/gbe/evz073
Acknowledgements
This study was financially supported by the National Natural Science Foundation of China (31970227 and 41571049) and the Science and Technology Basic Resources Investigation Program of China (2019FY100900).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
All authors declare that they have no conflict of interest.
Additional information
Handling Editor: Alexander Platt.
Supplementary Information
Below is the link to the electronic supplementary material.
239_2020_9987_MOESM1_ESM.xlsx
Supplementary file1 Table S1 Three genetic traits in protein-coding regions of plastomes were measured for 1382 angiosperm species representing 350 families and 64 orders using the data set of Li et al (2019). "*" indicates non-photosynthetic plants, including holoparasitic and mycroheterotropic species (Merckx and Freudenstein 2010; Nickrent 2020) (XLSX 194 KB)
239_2020_9987_MOESM2_ESM.xlsx
Supplementary file2 Table S2 Test of phylogenetic signal of three genetic traits using 101 trees, including Li et al.’s MCC and 100 randomly selected trees, using two statistics: Blomberg's K and Pagel's λ. p < 0.001 is considered significant (XLSX 22 KB)
Rights and permissions
About this article
Cite this article
Yu, Y., Li, HT., Wu, YH. et al. Correlation Analysis Reveals an Important Role of GC Content in Accumulation of Deletion Mutations in the Coding Region of Angiosperm Plastomes. J Mol Evol 89, 73–80 (2021). https://doi.org/10.1007/s00239-020-09987-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-020-09987-5