Abstract
We and others recently identified an almost-intact human endogenous retrovirus (HERV), termed HERV-K(HML-2.HOM), that is usually organized as a tandem provirus. Studies on HERV proviral loci commonly rely on the analysis of single alleles being taken as representative for a locus. We investigated the frequency of HERV-K(HML-2.HOM) single and tandem alleles in various human populations. Our analysis revealed that another HERV-K(HML-2) locus, the so-called HERV-K(II) provirus, is also present as a tandem provirus allele in the human population. Proviral tandem formations were identified in various nonhuman primate species. We furthermore examined single nucleotide polymorphisms (SNPs) within the HERV-K(HML-2.HOM) proviral gag, prt, and pol genes, which all result in nonsense mutations. We identified four proviral haplotypes displaying different combinations of gag, prt, and pol SNPs. Haplotypes harboring completely intact proviral genes were not found. For the left provirus of the tandem arrangement a haplotype displaying intact gag and prt genes and a mutated pol was found in about two-thirds of individuals from different ethnogeographic origins. The same haplotype was always found in the right provirus. The various haplotypes point toward multiple recombination events between HERV-K(HML-2.HOM) proviruses. Based on these findings we derive a model for the evolution of the proviral locus since germ line integration.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Human endogenous retroviruses (HERVs) were formed in an ancestral genome millions of years ago by integration of exogenous retroviruses in the germ line, subsequent inheritence of proviruses from parent to offspring, and fixation within the population. Approximately 8% of the human genome is of retroviral origin, and a number of distinct HERV families were defined (International Human Genome Sequencing Consortium 2001; Li et al. 2001). Following integration, proviruses retained replicative capabilities leading to copy number increases of up to several thousand for some HERV families. Over time, proviral open reading frames (ORFs) and long terminal repeats (LTRs) accumulated mutations, resulting in coding-deficient proviruses. However, a few HERV families still include coding-competent proviruses, e.g., the HERV-K(HML-2) family displaying ORFs for gag, protease, polymerase, and/or envelope (Lower et al. 1996; Tonjes et al. 1996). Previously, we described a locus on human chromosome 7 containing an almost-intact HERV-K(HML-2) provirus, named HERV-K(HML-2.HOM), that was subsequently also identified by others (Barbulescu et al. 1999; Mayer et al. 1999; Tonjes et al. 1999). Only a mutation within the conserved YXDD motif in the proviral polymerase (pol) renders the reverse transcriptase domain presumably inactive. We subsequently reported that, in most cases, HERV-K(HML-2.HOM) displays a tandem organization, with a central LTR shared by both proviruses. Furthermore, proviruses displaying intact YXDD motifs were identified (Reus et al. 2001). A recent study reported two full-length, one of them apparently completely intact HERV-K(HML-2), proviruses in a fraction of the human population (Turner et al. 2001). Taken together, these data provide evidence for potentially important single nucleotide polymorphisms (SNPs), as well as the presence/absence of HERV proviruses in the human population.
It was recently pointed out by J.P. Stoye that the majority of HERV-related investigations considers a single sequenced HERV locus as representative of that HERV locus (Stoye 2001). Potentially consequential sequence variations for a particular HERV locus were, as of yet, not investigated in detail. However, as evidenced by the above examples, such variations may change an individual’s HERV content significantly regarding, for instance, HERV protein coding capacity. In the present study we examined occurence of distinct HERV-K(HML-2.HOM) SNPs, as well as the frequency of proviral tandem and single alleles in the human population. Results for the latter part of our study may have direct implications for two recently published studies of the HERV-K(HML-2.HOM) tandem provirus (Hughes and Coffin 2004; Macfarlane and Simmonds 2004). We identified a limited number of SNP haplotypes that furthermore varied between different ethnogeographic groups. Based on the available data we derived a model for the evolution of the HERV-K(HML-2.HOM) locus that indicates multiple recombination events since initial provirus formation.
Methods
Collecting HERV-K(HML-2.HOM) Variants
We compiled different HERV-K(HML-2.HOM) provirus isolates from the literature (see Introduction). In addition, we retrieved from GenBank corresponding proviral sequences produced in the course of the human genome project. HERV proviral sequences were multiple aligned to identify sequence variations.
DNA Samples
DNA samples from African, Asian Indian, Southeast Asian, and Papua New Guinean individuals were kindly provided by Norbert Kienzle and Catherine Stolle. Total genomic DNA from additional Caucasian, African, and Southeast Asian donors was prepared according to standard procedures (Sambrook 1989). Genomic DNA from Aborigine individuals was obtained from the European Collection of Cell Cultures (ECACC).
Long-PCR Primers and Conditions
The left provirus of the HERV-K(HML-2.HOM) tandem repeat was amplified from total genomic DNA by long-PCR using primers 5′flank2FOR (5′TACCCACAGCACCCAAGAGATG3′) and envendREV (5′GTCATCATGGCCCGTTCTCG3′). PCR cycling conditions were as follows: 93°C for 5 min; 9 cycles of 93°C for 15 s, 54°C for 30 s, and 68°C for 9 min; followed by 19 cycles of 93°C for 15 s, 54°C for 30 s, and 68°C for 11 min; and final incubation for 10 min at 68°C. The right provirus was amplified by long-PCR using primers 5′gagFOR (5′CGCTCGGAAGAAGCT AGG3′) and 3′flankREV (5′GCTTTCGGGACTTGAACATTG G3′). PCR cycling conditions were as above. Single proviruses were amplified from total genomic DNA by long-PCR using primers 5′flank2FOR and 3′flankREV with cycling conditions as above. Tandem proviruses were amplified by an extended long-PCR using primers 5′flank2FOR and 3′flankREV. PCR cycling conditions were as follows: 93°C for 5 min; 9 cycles of 93°C for 15 s, 54°C for 30 s, and 68°C for 12 min; followed by 19 cycles of 93°C for 15 s, 54°C for 30 s, and 68°C for 12 min; and final incubation for 12 min at 68°C. HERV-K(HML-2.HOM) and HERV-K(II) tandem provirus portions, spanning a 292-bp region within pol–env and the central LTR, were amplified with primers HK290FOR (5′GGGGAGAGGTTTTGCTTGT3′) and LTRgagREV2. Cycling conditions were as follows: 94°C for 5 min; 10 cycles of 94°C for 15 s, 58°C for 30 s, and 68°C for 4 min; 20 cycles of 94°C for 15 s, 58°C for 30 s, and 72°C for 4 min (+5 s); and 72°C for 7 min. The TripleMaster PCR-System (Eppendorf), was used for long-PCR reactions as recommended by the manufacturer. Long-PCR products were TA-cloned employing the pGEM T Easy Vector System (Promega), and inserts were sequenced using vector-specific primers.
PCR Primers and Conditions
For verification of target site duplications a sequence portion flanking the left provirus was amplified from total genomic DNA by PCR using primers 5′flank2FOR and LTRgagREV2 (5′TTTGCCCCATTATCACCCTA3′). PCR cycling conditions were as follows: 94°C for 5 min; 30 cycles 94°C for 1 min, 54°C for 1 min, and 72°C for 2 min; and final incubation for 1 min at 72°C. A sequence portion flanking the right provirus was PCR-amplified from total genomic DNA using primers 3′flankREV and ENV8406 (5′GTCTGCAGGTGTACCCAACAG3′). PCR cycling conditions were as follows: 94°C for 5 min; 30 cycles of 94°C for 1 min, 57°C for 1 min, and 72°C for 2 min; and final incubation for 1 min at 72°C. The central LTR in the tandem provirus was amplified from total genomic DNA using primers ENV8406F and LTRgagREV2. PCR cycling conditions were as follows: 94°C for 5 min; 30 cycles of 94°C for 1 min, 54°C for 1 min, and 72°C for 2 min; and final incubation for 1 min at 72°C. Taq DNA polymerase was used for all those PCRs as recommended by the manufacturer (Invitrogen).
ARMS-PCR
ARMS-PCR (Amplification Refractory Mutation System) (Newton et al. 1989) was applied to genotype SNPs in the HERV-K(HML-2.HOM) gag, prt, and pol genes. ARMS-PCR is based on the fact that oligonucleotides with a mismatched 3′ residue will not function as primers in the PCR under stringent conditions. Each SNP is genotyped by two separate PCR reactions amplifying the two different SNP alleles. Therefore, each sample was tested with an appropriate “normal” or “mutant” forward primer for the respective proviral gene, paired with a universal reverse primer. Specificity of ARMS-PCR was optimized by varying Mg2+ concentrations and annealing temperature, until the “normal” forward primer was refractory to PCR on “mutant” template DNA, and vice versa. Purified and diluted PCR product from the long-PCRs was used as template for ARMS-PCR. Sequence analysis of a number of long-PCR products confirmed the ARMS-PCR results. As controls, we included in each experiment two ARMS-PCRs containing template with known SNP sequences. Primers gagARMS1for (C-allele) (5′GTCTCTCTCAC CCTCTCAATTTTTAC3′), gagARMS2for (T-allele) (5′GTCTCT CTCACCCTCTCAATTTTTAT3′), and gagARMSrev (5′GCTTT ATGCATAGCTCCTCCG3′) were used to characterize the SNP in the proviral gag gene. PCR parameters were as follows: 94°C for 5 min; 25 cycles of 94°C for 30 s, 64°C for 30 s, and 72°C for 25 s; and final incubation for 10 min at 72°C. Reaction mixtures contained 0.75 mM MgCl2. Primers polARMS1for (G allele) (5′AAG TTTTCAGACTGTTATATTATTCATTG3′), polARMS2for (A allele) (5′AAGTTTTCAGACTGTTATATTATTCATTA3′), and polARMSrev (5′ACTGTGAGGAAGGAATGACCA3′) were selected to characterize the pol gene SNP. PCR parameters were as for gag. Reactions included 1.2 mM MgCl2. Since the mutation in the prt gene is a deletion, but no substitution, we restricted the PCR to detection of the nonmutated allele. Presence of the mutated allele in those samples failing to amplify a PCR product under stringent conditions was confirmed by PCR under less stringent conditions. We designed forward primer prtARMS for (5′CTTCCAGGGGA GCCCCC3′), corresponding to the nonmutated allele, and reverse primer prtARMSrev (5′GCTATATGATG GAGCGGGC3′). PCR parameters were as follows: 94°C for 5 min; 25 cycles of 94°C for 30 s, 67°C for 30 s, and 72°C for 25 s; and a final incubation of 10 min at 72°C. Reactions included 0.5 mM MgCl2. All ARMS-PCRs furthermore contained a 100 μM concentration of each dNTP, and 2.5 U of Taq polymerase in standard PCR buffer (Gibco, BRL, Life Technologies).
Characterization of Homo- and Heterozygotes
Homozygotes are characterized by a PCR product in only one of two reactions required to type each SNP (Fig. 4). Presence of PCR products in both reactions potentially results from heterozygosity. We then cloned the long-PCR product and performed ARMS-PCR onto plasmid DNA from several independent clones as template, confirming the presence of both alleles.
DNA Sequencing and Analysis
Cloned PCR products were sequenced using the SequiTherm Excel II DNA Sequencing Kit-LC (Biozym) on an automated DNA sequencer (Licor 4000-L, MWG). Sequence data were analyzed using the Sequencher software (Gene Codes Corp. Inc.).
Results
Frequency of Tandem and Single Proviral Alleles
As opposed to the common HERV-K(HML-2.HOM) tandem arrangement where a central LTR is shared by both proviruses, we recently identified two individuals from Papua New Guinea (PNG) that harbor single HERV-K(HML-2.HOM) proviruses in a homozygous state (Reus et al. 2001). Here we examined 67 human DNA samples, representing 134 chromosomes, from various ethnogeographic origins for the presence of tandem and single proviruses. Tandem proviruses were identified by PCR using a forward primer located in the retroviral env gene and a reverse primer in the gag gene, encompassing the central LTR. A PCR product of 1216 bp indicated the presence of the tandem allele. Single proviruses were amplified by long PCR using primers located in the 5´′ and 3´′ flanking regions, resulting in a 9891-bp PCR product. Among the 69 DNA samples included in this study (67+2 PNG) we identified five more samples to harbor single proviruses in a homozygous state, totaling 10% of individuals. Sixteen samples (23%) harbored tandem proviruses in a homozygous state, and 46 individuals (67%) were heterozygous for single and tandem alleles. Single proviruses were found in all ethnogeographic groups. Homozygous states for single proviruses were found in two individuals from Papua New Guinea (Reus et al. 2001), one African, and four Asian individuals.
Another HERV-K(HML-2) Tandem Provirus in the Human Population
The observed frequencies of single and tandem alleles of the HERV-K(HML-2.HOM) locus appeared different from those predicted by the Hardy–Weinberg equilibrium. Heterozygous individuals were significantly more frequent, and individuals carrying single proviruses in a homozygous state were observed less often than expected. We evaluated statistical significance of observed versus expected frequencies, employing a G-test. A G-value of 15.694 indicated that observed genotype frequencies differ significantly (p < 0.001) from the Hardy–Weinberg equilibrium. The observed disequilibrium is not due to PCR artifacts. It is imaginable that false-positive PCR products indicative of single proviruses may be due to annealing and extension of partly overlapping DNA strands that actually stem from a tandem provirus. Such PCR products could feign a single provirus. We verified the presence of single and tandem proviruses by an extended long-PCR approach able to amplify both tandem and single alleles. Long-PCR results confirmed previous findings for 10 representative samples supposedly homozygous or heterozygous for single and/or tandem proviruses. PCR products with lengths characteristic for single and tandem alleles were amplified. Importantly, homozygous tandem alleles never resulted in artifactual PCR products characteristic of single proviruses (Fig. 1).
Representative results for amplification of single and tandem HERV-K(HML-2.HOM) proviruses, employing an extended long-PCR approach. Single proviruses in homozygous and heterozygous states are shown in lanes 1 and 2, while lanes 3 and 4 show results for tandem proviruses in a homozyous state. Note that the latter PCRs did not yield products characteristic of single proviruses, excluding possible PCR artifacts (see text). PCR products were separated by electrophoresis in a 0.8% TAE agarose gel.
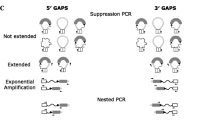
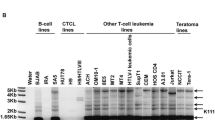
The observed disequilibrium—more tandem provirus alleles than expected—could also be explained by the presence of a tandem provirus other than the HERV-K(HML-2.HOM) locus in the human population. Another tandem provirus could equally produce a 1.2-kb PCR product when testing for the presence of a central LTR (see above). We therefore cloned the 1.2-kb PCR product obtained from various individuals, indicative of a tandem provirus allele, and sequenced a number of clones each. Indeed, several cloned PCR products were clearly different from the HERV-K(HML-2.HOM) provirus sequence. Further sequence comparison showed that those sequences matched a HERV-K(HML-2) provirus located on chromosome 3q12.3, which was previously reported by others as HERV-K(II) (Sugimoto et al. 2001). Further sequence comparisons show that the central LTR of the HERV-K(II) tandem provirus arrangement comprises the proviral 5′LTR that displays a number of differences compared to the 3′LTR (Fig. 2). We further confirmed the presence of a HERV-K(II) tandem provirus in human individuals by PCR. We made use of the fact that HERV-K(II) is a type 1 provirus, lacking a 292-bp sequence within the pol–env boundary that is present within the type 2 HERV-K(HML-2.HOM) provirus (Lower et al. 1995; Ono et al. 1986). Thus, a PCR product spanning that region and the central LTR distinguishes HERV-K(II) from HERV-K(HML-2.HOM) tandem provirus alleles by a 292-bp size difference. Indeed, PCR primers HK290FOR and LTRgagREV2 yielded in some individuals PCR products of sizes characteristic for either HERV-K(II) or HERV-K(HML-2.HOM) tandem alleles (Fig. 2). We therefore conclude that the human population harbors not only a tandem allele of the HERV-K(HML-2.HOM) provirus, but also a tandem allele of the HERV-K(II) provirus. The observed higher number of tandem proviruses can be fully explained by this finding (Fig. 2 and Discussion).
Detection of an additional HERV-K(HML-2) tandem provirus allele in the human population. Top A multiple alignment of a diagnostic LTR sequence portion from various PCR products, indicative of the presence of a tandem provirus (see text), with the corresponding portion of the HERV-K(HML-2.HOM) and HERV-K(II) proviruses. The latter sequence was obtained from the most recent human genome sequence. HERV-K(II) 5′ and 3′ LTRs are also included in the alignment to demonstrate the higher sequence similarity of PCR products with the HERV-K(II) 5′ LTR rather than the 3′ LTR. Numbers above the alignment indicate the location of the regarded sequence region with respect to the HERV-K(HML-2.HOM) sequence (GenBank accession no. AF074086). Middle Representative results of a PCR from Caucasian and from African-American individuals that specifically distinguishes HERV-K(HML-2.HOM) from HERV-K(II) tandem proviruses, owing to the lack of a 292-bp sequence within the amplicon of HERV-K(II) (see text). PCR products from the HERV-K(HML-2.HOM) and the HERV-K(II) tandem provirus are expected to be 3257 and 2965 bp, respectively, in size. DNA marker bands (bp) are indicated on the left. Bottom Summary of expected genotype frequencies when a second tandem provirus allele is considered. Pale gray fields indicate that corresponding genotypes (human DNA samples) will appear in corresponding PCR tests as heterozygous for the HERV-K(HML-2.HOM) tandem allele, although they may be heterozygous for HERV-K(HML-2.HOM) single proviruses, because PCR products actually stem from the HERV-K(II) tandem provirus. Dark gray fields indicate homozygosity for tandem proviruses. Genotypes and expected and observed frequencies are summarized at the bottom.
We furthermore note that formation of HERV-K(HML-2) tandem proviruses is obviously not restricted to humans. We also examined various primate species using the above-mentioned env forward and gag reverse primers. Besides a PCR product characteristic of tandem alleles in Pan troglodytes (Reus et al. 2001), we obtained PCR products from genomic DNA from Gorilla gorilla, Macaca mulatta, and Macaca fascicularis. Sequences of PCR products obtained from Gorilla gorilla and Macaca fascicularis (GenBank accession nos. AY425962 and AY425963) confirmed env–LTR and LTR–gag boundaries. No PCR products were obtained from Pan paniscus, Pongo pygmaeus, Hylobates lar, Mandrillus sphinx, and Colobus guereza (data not shown).
HERV-K(HML-2.HOM) SNPs
The HERV-K(HML-2.HOM) proviral locus has been repeatedly isolated by us and others in specific studies and in the course of the human genome sequencing project (Barbulescu et al. 1999; International Human Genome Sequencing Consortium 2001; Mayer et al. 1999; Tonjes et al. 1999). We identified sequence variants among those different proviral isolates. We found that the different reported proviral sequences displayed single nucleotide polymorphisms (SNPs) within the gag, prt, and pol genes, resulting in a stop codon, a frameshift, and a mutation within the polymerase YXDD motif, respectively. With respect to HERV-K(HML-2.HOM), a C/T exchange at nt 2204 resulted in a stop codon in gag. An extra C at nt 3158 resulted in a frameshift and a premature stop codon in prt. An A/G exchange at nt 4462 altered a highly conserved reverse transcriptase motif from YIDD to CIDD. Differences between various HERV-K(HML-2.HOM) isolates are summarized in Fig. 3. SNPs in gag, prt, and pol occurred in different combinations. The recently reported HERV-K(HML-2.HOM) proviral sequence, isolated from an Asian sample, displayed mutated gag and prt genes and an intact YXDD motif (GenBank accession no. AF490464) (Reus et al. 2001). Barbulescu et al. (1999) reported a similarly mutated proviral sequence (AF261945). The provirus isolated by Tonjes et al. (1999) (Y17832) harbored an intact gag but mutated prt and pol genes. The corresponding chromosome 7 proviral entry from the human genome sequencing project (AC072054) displays intact gag and prt genes but a mutated pol. Additional nonsense mutations were not detected in the respective genes.
Sequence comparison of different HERV-K(HML-2.HOM) provirus isolates, as previously reported by us and others (for references, see text). Relevant gag, prt, and pol regions and corresponding reading frame changes are shown. Polymorphisms in gag and pol result in a stop codon and an amino acid exchange, respectively. An extra C in prt causes a premature stop. SNP positions are numbered with respect to the HERV-K(HML-2.HOM) sequence. Correct and mutated reading frames are indicated. Asterisk indicates a stop codon.
We next examined the occurrence of the above-mentioned gag, prt, and pol gene SNPs in the human population. Using ARMS-PCR as an allele-specific PCR approach (Newton et al. 1989), we analyzed 67 human DNA samples, representing 134 chromosomes, from various ethnogeographic origins. Since the HERV-K(HML-2.HOM) provirus is commonly organized in tandem (Reus et al. 2001), we separately amplified and analyzed the left and the right portion of the tandem provirus.
We amplified the left provirus from the HERV-K(HML-2.HOM) tandem by long-PCR using a forward primer in the 5′´ flanking chromosomal sequence and a reverse primer in env.
Gel-purified long-PCR products served as templates for ARMS-PCR that consisted of two reactions, each containing an allele-specific forward primer and a conserved reverse primer. A PCR product in only one reaction indicated homozygosity of the analyzed SNP while PCR products in both reactions indicated heterozygosity (Fig. 4). The latter was further confirmed by ARMS-PCR analysis of several independent plasmids cloned from long-PCR products. Furthermore, sequencing of the SNP-containing gene regions of three retroviral inserts confirmed the ARMS-PCR results, and excluded additional mutations altering gag, prt, and pol for those clones.
Representative ARMS-PCR results for detection of HERV-K(HML-2.HOM) gag SNPs. The left portion of the top and bottom panels shows control reactions employing sequences with either thymine (T) (lanes 1 and 2) or cytosine (C) (lanes 3 and 4) in the relevant position. The right portions of both panels each show analyzed genomic DNA samples. Samples 1, 2, and 4 display a homozygous state, while sample 3 is heterozygous for the regarded SNP.
As for the left provirus, ARMS-PCR identified 27 chromosomes (20%) displaying the stop mutation within gag, 28 (21%) having the frameshift within prt, and 92 (69%) showing a defective YXDD motif. The identified single nucleotide changes could be summarized in four different haplotypes, in the following referred to as haplotypes A, B, C, and D. Haplotype A shows mutant gag and prt genes and a pol with an intact YXDD motif. Haplotype B shows an intact gag but mutated prt and pol. In haplotpye C only pol is mutated. In haplotype D all three genes carry mutations. Haplotype C was most frequent (92 chromosomes; 69%), followed by haplotype A (24; 18%), haplotype D (5; 3.7%) and haplotype B (1; 0.7%) (Table 1). Notably, we did not identify any of the other four possible haplotypes. Of further importance, a haplotype displaying ORFs for all retroviral genes was not detected either.
The right provirus was amplified from the tandem arrangement by long PCR using a forward primer located at the start of gag and a reverse primer located in the 3′ flanking region. SNP analysis was as described above. We chose eight DNA samples that displayed different haplotypes in the left provirus and that were of different ethnogeographic origin. The 16 analyzed chromosomes uniformly displayed haplotype C, that is, intact gag and prt, but a mutated YXDD-motif in pol. ARMS-PCR results are summarized in Table 1. All HERV-K(HML-2.HOM) variants identified in this study are summarized in Fig. 5.
Summary of HERV-K(HML-2.HOM) haplotypes identified in different individuals. Tandem and single proviruses are illustrated. Haplotypes are indicated above each provirus. Rectangles represent LTRs. Boundaries between proviral gag, prt, pol, and env genes are depicted by short vertical lines. Haplotypes 6–8 are heterozygous or homozygous for tandem and single proviruses, respectively. See text and Table 1 for haplotype details.
We furthermore analyzed haplotype distribution depending on ethnogeographic groups included in this study (Table 2). Haplotype C was most frequent in all groups, followed by haplotype A. Compared to Caucasians, Southeast Asian and African individuals displayed greater numbers of haplotype A. Notably, Aborigine and Papua New Guinea individuals exclusively displayed haplotype C. Haplotypes B and D were found in African individuals in one case each, and haplotype D was also present in Southeast Asians. As revealed by Fisher’s exact test for 3×2 tables, haplotype distributions were significantly different (p=0.0051) between Caucasians and Southeast Asians and between Aborigine and Southeast Asian individuals (p=0.005). Sample numbers for other groups were too small to determine statistical significance.
Discussion
There is limited information on HERV polymorphisms. A recent study reported full-length retroviruses, termed HERV-K113 and HERV-K115, in 15% and 29% of the human population, respectively (Turner et al. 2001). A relatively recent formation of these proviruses in evolution may explain why HERV-K113 and HERV-K115 are not fixed in humans. Hughes and Coffin (2004) revealed several polymorphic HERV-K(HML-2) loci in the human population. Polymorphic solitary LTRs in the HLA-DQ locus were previously reported (Kambhu et al. 1990). As for single nucleotide polymorphisms (SNPs) within particular proviral sequences, it was shown that a SNP within the ERV-3 env gene results in a stop codon. About 1% of Caucasian individuals are homozygous for the defective allele, thus coding deficient for the Env protein for which a role in placental development had been postulated (de Parseval and Heidmann 1998). Furthermore, SNPs in the so-called HERV-K18 provirus were analyzed in diabetes mellitus type 1 patients but were found not to be correlated with the disease (Kinjo et al. 2001). We recently reported HERV-K(HML-2.HOM) variants with and without intact YXDD motifs within the reverse transcriptase domain, a tandem organization of HERV-K(HML-2.HOM) in a majority of human genomic DNA samples, and a single provirus in two samples from Papua New Guinea (Reus et al. 2001). From several independent investigations it became apparent that the HERV-K(HML-2.HOM) locus displays defined sequence variations, SNPs, within the gag, prt, and pol genes that effect the coding capacity of those genes by introducing stop mutations or altering a conserved catalytic motif. We present here a systematic investigation of those HERV-K(HML-2.HOM) polymorphisms in various ethnogeographic groups.
When we investigated the presence of HERV-K(HML-2.HOM) single versus tandem provirus alleles, observed frequencies significantly differed from Hardy–Weinberg equilibrium (p < 0.001). Further analysis revealed that the so-called HERV-K(II) provirus on chromosome 3q12.3 is likewise present in the human population as a tandem provirus allelic variant. The HERV-K(II) tandem allele produced similar-sized PCR products when human DNA samples were tested for the env–central LTR–gag arrangement, and thus feigned HERV-K(HML-2.HOM) tandem provirus alleles. While the overall structure of the HERV-K(II) tandem provirus is currently not known, the presence of a second tandem provirus allele in the human population explains very well the genotype frequencies observed in our analysis. In our PCR-based analysis, we tested for single HERV-K(HML-2.HOM) proviruses and for tandem alleles (see Methods). Thus, the presence of a second tandem provirus will increase the overall number of genotypes heterozygous for single and tandem proviruses and will reduce the overall number of genotypes homozygous for single proviruses. Indeed, our observed genotype frequencies match very well the expected frequencies (Fig. 2). In detail, 25% of genotypes are expected to represent single HERV-K(HML-2.HOM) proviruses in a homozygous state. However, of those genotypes with two HERV-K(HML-2.HOM) single proviruses, three will contain at least one HERV-K(II) tandem provirus (Fig. 2). In PCR analysis, those genotypes will thus appear as heterozygous for a HERV-K(HML-2.HOM) single and tandem allele. Hence, the overall number of homozygous single HERV-K(HML-2.HOM) provirus alleles is reduced from 4 of 16 (=25%) to 1 of 16 (=6.25%) since the other 3 of 16 (=18.75%) will appear as heterozygous for a single and a tandem allele of HERV-K(HML-2.HOM). Indeed, we observe 10% of individuals to be homozygous for HERV-K(HML-2.HOM) single proviruses. Accordingly, the 3 of 16 genotypes with homozygous HERV-K(HML-2.HOM) single proviruses, but at least one HERV-K(II) tandem provirus, will add to the number of expected heterozygous single/tandem alleles, increasing their frequency from 50% to 68.75%. Indeed, we observed 67% of individuals to contain single and tandem proviruses in a heterozygous state.
We note in this context that two previous studies on the HERV-K(HML-2.HOM) provirus allele did not reveal a second HERV-K(HML-2) tandem provirus allele in the human population (Hughes and Coffin 2004; Macfarlane and Simmonds 2004). Since the detection of a tandem provirus by means of PCR amplification of an env–central LTR–gag portion was very similar, or identical, to our methodology, it is possible that some of the observed HERV-K(HML-2.HOM) tandem allele frequencies, that likewise deviate from Hardy–Weinberg equilibrium, actually stem from HERV-K(II) tandem proviruses. It is currently not known whether the human population harbors HERV-K(HML-2) tandem proviruses in addition to the two examined/revealed in our study. As the recent study by Hughes and Coffin (2004) shows considerable allelic variation for several HERV-K(HML-2) proviruses in different chromosomal locations, it seems possible that other HERV-K(HML-2) tandem provirus alleles indeed exist.
Besides the frequencies of tandem and single alleles, we also characterized particular SNPs within the gag, prt, and pol genes of the HERV-K(HML-2.HOM) provirus. We specifically amplified the leftmost or rightmost proviruses from human genomic DNA samples and determined their specific SNP status. For the left provirus of the HERV-K(HML-2.HOM) tandem arrangement we identified four haplotypes, of eight possible haplotypes, regarding the presence of intact versus defective gag, prt, and pol genes. Obviously, not all possible haplotypes were created during evolution or fixed in the human population. The majority of proviruses/individuals displayed intact gag and prt genes and a defective YXDD motif, that is, haplotype C. However, a significant number of individuals harbored different haplotypes in that, for instance, gag and prt genes were defective but the YXDD motif was intact. We note in this context that we did not identify HERV-K(HML-2.HOM) proviruses with intact gag, prt, and pol genes, although there was opportunity for creating such a completely intact provirus by recombination events (see below). According to our results such an allele was not fixed in the human population, at least not at higher frequencies. One may hypothesize a selection pressure against a completely intact provirus. However, the effect of such a completely intact provirus seems vague. As a recent report shows, the so-called HERV-K113 provirus—having an allele frequency of 0.19 and displaying ORFs for all retroviral genes and an intact YXDD motif—probably is not deleterious (Turner et al. 2001). On the other hand, more elaborate studies investigating more samples will be required to exclude with higher certainty completely intact HERV-K(HML-2.HOM) alleles in the human population.
Our study further demonstrates that particular HERV loci are not necessarily identical in sequence and coding capacity. Generally, the HERV-K(HML-2) family is characterized by relatively intact retroviral genes. Besides several loci with intact genes (Barbulescu et al. 1999; Mayer et al. 1997a, b, 1999; Tonjes et al. 1999), defective genes are due to one or a few nonsense mutations. As exemplified by the observed polymorphisms within the HERV-K(HML-2.HOM) locus, other HERV-K(HML-2) loci may as well display sequence polymorphisms, and genes with or without ORFs could be rendered defective or intact, respectively, in various individuals. Therefore, each individual could display a particular set of intact and defective HERV-K(HML-2) loci. Hence, some individuals may harbor a greater number of intact HERV-K(HML-2) genes than others. Whether there could be a biological consequence from smaller or greater numbers of intact retroviral genes is currently not clear, though.
The combined results of our haplotype analysis allows us to envisage the evolutionary history of this specific proviral locus (Fig. 6). First, a HERV-K(HML-2) provirus was formed in chromosome 7p22. It is not clear whether that provirus was completely intact or already harbored the transition in pol. However, the pol mutation probably occurred prior to other mutations. A tandem provirus, sharing the central LTR, was probably generated by recombination between 5′ and 3′ LTR of the same locus. This event could also explain the frequent occurrence of haplotype C (mutation in pol) in both the left and the right provirus. Tandem provirus formation implies simultaneous formation of a solitary LTR on the other chromosome. We did not find evidence for loci harboring only a solitary LTR. It is possible that such an allele was not fixed in the human population, at least not at high frequencies. The tandem arrangement seems to be the prevalent of HERV-K(HML-2.HOM). The nucleotide changes in the gag and prt genes were found only in the left provirus, making it unlikely that these mutations occurred before tandem formation. It seems very unlikely that observed haplotypes are due to recurrent mutations. We propose that observed haplotypes other than C were generated by recombination between a HERV-K(HML-2) provirus carrying mutations in gag and prt in the left part of the tandem, with the recombination site excluding the mutation in pol. The resulting tandem provirus then carried haplotype A (mutated gag, prt, and pol) in the left provirus and haplotype C (intact gag and prt, mutated pol) in the right provirus. Likewise, haplotypes B and D can be explained by recombination events. For example, haplotye D in the left provirus could be the result from recombination between haplotype C and haplotype A (Fig. 6). As others recently reported frequent recombination events between HERV-K(HML-2) proviruses resulting in rearrangements of flanking cellular sequences (Hughes and Coffin 2001), we sequenced target site duplication (TSD) regions for several tandem and single allele proviruses. All proviruses displayed identical TSDs (not shown). Thus, our results indicate recombination events within the HERV-K(HML-2.HOM) locus rather than between loci in different genomic regions.
Proposed evolutionary history of the HERV-K(HML-2.HOM) locus indicating multiple recombination events between provirus loci. A Initial provirus formation in chromosome 7p22 followed by a mutation within the pol gene. B Tandem formation by recombination between two haplotype C proviruses. C Generation of haplotype A by interelement recombination with a provirus having mutated gag and prt. The latter may have been present as single or tandem provirus. D Generation of haplotypes B and D from recombination between haplotypes C and A. See Discussion for details.
The identification of several HERV-K(HML-2.HOM) haplotypes provides insight into the evolution of a single HERV locus. Recently, Hughes and Coffin (2001) showed that recombination between proviral HERV-K(HML-2) loci in different genomic regions generated rearrangements in flanking cellular sequences. Our analysis provides further evidence that recombination between HERV-K(HML-2.HOM) proviruses occurred several times since provirus formation. Therefore, exchange of HERV sequence portions by recombination seems to be a frequent event during evolution.
References
Barbulescu M, Turner G, Seaman MI, Deinard AS, Kidd KK, Lenz J (1999) Many human endogenous retrovirus K (HERV-K) proviruses are unique to humans. Curr Biol 9:861–868
de Parseval N, Heidmann T (1998) Physiological knockout of the envelope gene of the single-copy ERV-3 human endogenous retrovirus in a fraction of the Caucasian population. J Virol 72:3442–445
Hughes JF, Coffin JM (2001) Evidence for genomic rearrangements mediated by human endogenous retroviruses during primate evolution. Nat Genet 29:487–489
Hughes JF, Coffin JM (2004) Human endogenous retrovirus K solo–LTR formation and insertional polymorphisms: Implications for human and viral evolution. Proc Natl Acad Sci USA 101:1668–1672
International Human Genome Sequencing Consortium (2001) Initial sequencing and analysis of the human genome. Nature 409:860–921
Kambhu S, Falldorf P, Lee JS (1990) Endogenous retroviral long terminal repeats within the HLA-DQ locus. Proc Natl Acad Sci USA 87:4927–4931
Kinjo Y, Matsuura N, Yokota Y, Ohtsu S, Nomoto K, Komiya I, Sugimoto J, Jinno Y, Takasu N (2001) Identification of nonsynonymous polymorphisms in the superantigen-coding region of IDDMK1,2 22 and a pilot study on the association between IDDMK1,2 22 and type 1 diabetes. J Hum Genet 46:712–716
Li WH, Gu Z, Wang H, Nekrutenko A (2001) Evolutionary analyses of the human genome. Nature 409:847–849
Lower R, Tonjes RR, Korbmacher C, Kurth R, Lower J (1995) Identification of a Rev-related protein by analysis of spliced transcripts of the human endogenous retroviruses HTDV/HERV-K. J Virol 69:141–149
Lower R, Lower J, Kurth R (1996) The viruses in all of us: characteristics and biological significance of human endogenous retrovirus sequences. Proc Natl Acad Sci USA 93:5177–5184
Macfarlane C, Simmonds P (2004) Allelic variation of HERV-K(HML-2) endogenous retroviral elements in human populations. J Mol Evol 59:642–656
Mayer J, Meese E, Mueller-Lantzsch N (1997a) Chromosomal assignment of human endogenous retrovirus K (HERV-K) env open reading frames. Cytogenet Cell Genet 79:157–161
Mayer J, Meese E, Mueller-Lantzsch N (1997b) Multiple human endogenous retrovirus (HERV-K) loci with gag open reading frames in the human genome. Cytogenet Cell Genet 78:1–5
Mayer J, Sauter M, Racz A, Scherer D, Mueller-Lantzsch N, Meese E (1999) An almost-intact human endogenous retrovirus K on human chromosome 7. Nat Genet 21:257–258
Newton CR, Graham A, Heptinstall LE, Powell SJ, Summers C, Kalsheker N, Smith JC, Markham AF (1989) Analysis of any point mutation in DNA. The amplification refractory mutation system (ARMS). Nucleic Acids Res 17:2503–2516
Ono M, Yasunaga T, Miyata T, Ushikubo H (1986) Nucleotide sequence of human endogenous retrovirus genome related to the mouse mammary tumor virus genome. J Virol 60:589–598
Reus K, Mayer J, Sauter M, Scherer D, Muller-Lantzsch N, Meese E (2001) Genomic organization of the human endogenous retrovirus HERV-K(HML-2HOM) (ERVK6) on chromosome 7. Genomics 72:314–20
Stoye JP (2001) Endogenous retroviruses: still active after all these years? Curr Biol 11:R914–R916
Sugimoto J, Matsuura N, Kinjo Y, Takasu N, Oda T, Jinno Y (2001) Transcriptionally active HERV-K genes: identification, isolation, and chromosomal mapping. Genomics 72:137–144
Tonjes RR, Czauderna F, Kurth R (1999) Genome-wide screening, cloning, chromosomal assignment, and expression of full-length human endogenous retrovirus type K. J Virol 73:9187–95
Tonjes RR, Lower R, Boller K, Denner J, Hasenmaier B, Kirsch H, Konig H, Korbmacher C, Limbach C, Lugert R, Phelps RC, Scherer J, Thelen K, Lower J, Kurth R (1996) HERV-K: the biologically most active human endogenous retrovirus family. J Acquir Immune Defic Syndr Hum Retrovirol 13:S261–S267
Turner G, Barbulescu M, Su M, Jensen–Seaman MI, Kidd KK, Lenz J (2001) Insertional polymorphisms of full–length endogenous retroviruses in humans. Curr Biol 11:1531–1535
Acknowledgments
This work was supported by DFG Research Grants Ma2298/2-1 and Me917/16-1 to J.M. and E.M., respectively. Genomic DNA from Southeast Asian individuals was kindly provided by John Nicholls (University of Hongkong). Hans Zischler (University of Mainz, Germany) and Werner Schempp (University of Freiburg, Germany) generously provided primate genomic DNAs. We are grateful to Jochem König (University of Saarland, Germany) for help with statistical analysis.
Author information
Authors and Affiliations
Corresponding author
Additional information
[Reviewing Editor : Dr. Martin Kreitman]
Rights and permissions
About this article
Cite this article
Mayer, J., Stuhr, T., Reus, K. et al. Haplotype Analysis of the Human Endogenous Retrovirus Locus HERV-K(HML-2.HOM) and Its Evolutionary Implications. J Mol Evol 61, 706–715 (2005). https://doi.org/10.1007/s00239-005-0066-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-005-0066-7