Abstract
Purpose
To determine the frequency of N-acetyltransferase 2 (NAT2) polymorphisms, the NAT2 acetylation profile and its relation to the incidence of gastrointestinal adverse drug reactions (ADRs), anti-tuberculosis (TB) drug-induced hepatotoxicity, and the clinical risk factors for hepatotoxicity in a population from Brazil.
Methods
Two hundred and fifty-four Brazilian TB patients using isoniazid (INH), rifampicin (RMP), and pirazinamide (PZA) were tested in a prospective cohort study. NAT2 genotyping was performed by direct PCR sequencing. The association between gastrointestinal ADRs/hepatotoxicity and the NAT2 profile genotype was evaluated by univariate analysis and multiple logistic regression.
Results
Of the 254 patients analyzed, 69 (27.2%) were slow acetylators and 185 (72.8%) were fast acetylators. Sixty-five (25.6%) patients were human immunodeficiency virus (HIV)-positive. Thirty-three (13%) and 14 (5.5%) patients developed gastrointestinal ADR and hepatotoxicity, respectively. Of the 14 hepatotoxicity patients, nine (64.3%) were slow acetylators and five (35.7%) were fast acetylators. Sex, age, presence of hepatitis C virus, alcohol abuse, and baseline aminotransferases were not found to be risk factors for hepatotoxicity. However, logistic regression analysis revealed that slow acetylator status and the presence of HIV (p < 0.05) were independent risk factors for hepatotoxicity.
Conclusions
Our findings show that HIV-positive patients that have the slow acetylation profile are significantly associated with a higher risk of developing hepatotoxicity due to anti-TB drugs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Throughout history, tuberculosis (TB) has assumed a prominent role as a disease that has affected society. The current most effective control is to cure the infection by treating the patient with anti-TB drugs. The three main drugs used to treat TB are isoniazid (INH), rifampin (RMP) and pyrazinamide (PZA), used in combination for 6 or more months [1]. These anti-TB drugs are also the principal responsible agents for a significant number of cases of hepatotoxicity, or potential cases, with INH being the main drug to induce these adverse drug reactions (ADRs) [1]. Hepatotoxicity caused by anti-TB drugs is associated with high morbidity and mortality as well as with increased costs during treatment [2, 3]. While the occurrence of drug-induced hepatotoxicity is difficult to predict, it has been observed that certain patients are at higher risk of developing drug-induced hepatotoxicity during the course of anti-TB chemotherapy than others. These include patients with pre-existing liver diseases, particularly those associated with chronic viral infection due to hepatitis B and C, immunodeficiency virus (HIV), sex (female), and advanced age [4–6].
Genetic factors have also been described as risk factors for hepatotoxicity [7–10]. The acetylation polymorphism was discovered over 50 years ago following differences being observed in TB patients in terms of INH toxicity [11]. These differences were subsequently attributed to genetic variability in N-acetyltransferase 2 (NAT2), a cytosolic phase II conjugation enzyme primarily responsible for the deactivation of INH [12, 13]. Genetic differences in N-acetylation capacity confer corresponding differences in the biotransformation of INH. The classical N-acetylation polymorphism results from variant NAT2 alleles yielding fast and slow acetylator phenotypes [14]. The frequency of NAT2 alleles and acetylator phenotype varies remarkably with ethnic origin [15].
Earlier studies demonstrated an association between the NAT2 acetylation polymorphism and higher incidences and/or severity of ADRs to INH [7, 8, 10, 16, 17]. Although the acetylator status (fast or slow) of the individuals has been suspected as a potent risk factor for INH-induced hepatotoxicity, considerable controversy and uncertainty still exists given the wide variability of the results of these earlier studies [9, 14, 18–20].
The aim of the study reported here was to determine the frequency of the NAT2 polymorphisms, the NAT2 acetylation profile and its relation to the incidence of gastrointestinal ADRs and/or anti-TB drug-induced hepatotoxicity, and the clinical risk factors for hepatotoxicity through a prospective study of a population from Porto Alegre, Rio Grande do Sul State, Brazil.
Patients, materials and methods
The protocol used in the present study was approved by the Research Ethics Committee of the School of Public Health, Rio Grande do Sul State (protocol number 156/05) and by the Fundação Estadual de Produção e Pesquisa em Saúde-FEPPS (protocol number 18/2006). All patients recruited in the present study provided an informed written consent.
Study subjects
This was a prospective cohort study carried out between August 2005 and June 2007. A total of 254 unrelated patients with newly diagnosed TB from the outpatient section of Hospital Sanatório Partenon, a public TB reference hospital located in Porto Alegre, RS, were consecutively entered into the study. Inclusion criteria consisted of: adult patients (>18 years) who were newly diagnosed with active TB and who had been treated daily with INH, RMP, and PZA for the first 2 months followed by INH and RMP daily for 4 additional months, as recommended by the Brazilian National TB Program [1]. Drug dosages used were calculated according to patient’s weight [21] (weight <45 kg: RMP 300 mg, INH 200 mg, PZA 1000 mg; 45–55 kg: RMP 450 mg, INH 300 mg, PZA 1500 mg; more than 55 kg: RMP 600 mg, INH 400 mg, PZA 2000 mg). Exclusion criteria were: patients presenting clinically and laboratory-confirmed liver chronic disease, patients using anti-TB drugs prior to enrollment in the study, patients presenting results of liver function tests prior to the beginning of treatment higher that were twofold the upper normal limit and refusal to participate of the study.
The levels of aspartate aminotransferase (AST), alanine aminotransferase (ALT), and direct and total bilirubin were measured prior to anti-TB therapy, 30 and 60 days after the beginning of therapy or when the physician suspected hepatotoxicity. Serological testing for hepatitis B virus (HBV), hepatitis C virus (HCV) and HIV were also carried out prior to the initiation of anti-TB therapy. Clinical and epidemiological data, such as age, sex, skin color (self-reported), alcohol abuse (according to CAGE criteria [22]), and use of highly active antiretroviral therapy (HAART) or another co-medication, were collected using a standardized questionnaire at an interview and the review of medical records of each patient. Treatment adherence was evaluated by a pill count, regularity of attending medical appointments, and information obtained from medical records related to the regularity of pill taking.
Liver function tests and serology
Serum liver biochemical tests were measured using an automatic analyzer Dimension AR (DADE Behring, Germany) of the kinetic colorimetrics, and the upper limit normal (ULN) results were confirmed by the same method in Cobas Integra 400 Plus equipment (Roche, Germany). The comparison of liver functions tests were carried out by comparing the baseline levels (prior treatment) with the peak value (the highest level after commencing treatment). Hepatitis markers [hepatitis B surface antigen (HbsAg) and anti-HCV antibodies] and anti-HIV antibodies were analyzed using enzyme-linked immunosorbent analysis (ELISA) kits. All tests were carried out according the manufacturer’s protocols.
Gastrointestinal ADRs and definition of hepatotoxicity
Anorexy, nausea, vomiting, and/or abdominal pain were considered to be gastrointestinal ADRs. Criteria for the diagnosis of hepatotoxicity was an elevation in liver function tests, AST and/or ALT of more than threefold the ULN (reference: 40 and 65 U/L, respectively) and/or in total bilirubin up to >2.0 mg/dL in the presence of such gastrointestinal symptoms as anorexy, nausea, vomiting and/or jaundice, with a normalization of serum ALT level after discontinuation of the anti-TB drugs. These criteria are routinely used by pneumologists and gastroenterologists of HSP and are consistent with the recommendations of the Brazilian Tuberculosis Consensus [23]. Analysis of hepatotoxicity based on the criteria of the International Consensus meeting [24] (ALT of more than twofold the ULN) for drug-induced hepatotoxicity was also carried out.
Records of patients who developed hepatotoxicity were reviewed in detail in a search for risk factors as well as the subsequent consequences. In the group of patients with the diagnosis of hepatotoxicity, all drugs were withdrawn, and aminotransferases and bilirubin were measured weekly until they returned to normal levels. After the hepatotoxicity-related symptoms had disappeared and the liver function tests had returned to normal levels, anti-TB drugs were reintroduced in all patients studied. All patients who presented hepatotoxicity had their regimens modified as described in the Technical Recommendations of Tuberculosis Control Policy from Rio Grande do Sul State [21]; they received streptomycin (SM), INH, and ethambutol (EMB) for the first 3 months, followed by INH and EMB for an additional 9 months, after the liver function tests had returned to normal levels. None of the patients had a recurrence of hepatotoxicity after reintroduction of the anti-TB drug therapy.
NAT2 Genotyping method
A 5-mL volume of venous blood from each participant was collected in a tube containing EDTA. The leukocyte layer was separated by centrifugation and stored at −20°C. Genomic DNA was isolated from 500 μL of the leukocyte layer using the salting out method [25]. After extraction, DNA samples were stored at −20°C until genotyping. The complete NAT2 gene (870 bp) was amplified with primers NAT2-EF (5′-TTAGTCACACGAGGAAATCAAA-3′) and NAT2-IR (5′-TGGTCCAGGTACCAGATTCC-3′) and NAT2-IF (5′ACCATTGACGGCAGGAATTA-3′) and NAT2-ER (5′-AAATGCTGACATTTTTATGGATGT-3′), resulting in two fragments. The first fragment is 560 bp long and contains part of the promoter region, starting 72 bp upstream of the start codon and ending at nucleotide 488 of the gene. The second (685 bp) starts at nucleotide 336 of the gene and ends 148 bp downstream of stop codon. The two primer sets used in the PCR analysis and/or for sequencing have been described by Teixeira et al. [26]. Amplification was performed in a thermocycler PTC 200 DNA Engine, (MJ Research, Waltham, MA) as follows: 20 pmol of each oligonucleotide, 2.5 mM MgCl2, 0.25 mM each dNTP, 5 U Taq polymerase (Cenbiot, UFRGS, Porto Alegre, RS, Brazil), 10 mM Tris-Cl (pH 8.3), 50 mM KCl, 200 ng of genomic DNA in a 50 μL reaction volume. Samples were incubated at 94°C for 5 min, followed by 35 cycles of 94°C for 1 min, 54°C for 1.5 min and 72°C for 1 min, with a final extension at 72°C for 5 min. The amplification products were analyzed by electrophoresis in 1.5% agarose gels stained with ethidium bromide (0.5 μg/mL). The PCR products were purified with 7.5 M ammonium acetate and used for direct sequencing. Sequencing was carried out on a MegaBACE 1000 DNA Analysis System (Molecular Dynamics, Sunnyvale, CA) as recommended by the manufacturer. The obtained sequences were analyzed for the identification of single nucleotide polymorphism (SNP) by alignment with the reference sequence (GenBank accession X14672) using programs PREGAP and GAP4 from STADEN software package ver. 10.0. Nucleotide sequences with Phred values >20 were considered for analysis. The nomenclature of the NAT2 genotype is given in accordance to that described in http://louisville.edu/medschool/pharmacology/NAT.html. The acetylator profile of NAT2 was predicted according to the number of mutant alleles observed: presence of any two mutant alleles for NAT2 variations 191A, 341C, 590A, and/or 857A was defined as a slow-acetylator profile, whereas a rapid acetylator presents one or two wild-type NAT2*4 alleles. The physicians were not informed about the NAT2 genotypes of their patients and applied treatment as usual.
Statistical analysis
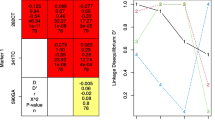
Allele frequencies at individual SNPs were estimated by counting. The maximum likelihood estimate of haplotype frequencies and genotype with unknown phase was calculated with multiside marker data using the Multiple Locus Haplotype Analysis software program, ver. 2.0 [27–29]. Linkage disequilibrium (D) and D′ (the relative magnitude of D as compared to its theoretical maximum) were calculated also using the software described above.
All statistical analyses were performed using the SPSS ver. 12.0 statistical program (SPSS, Chicago, IL) and EPiInfo ver. 6.04d (Centers for Disease and Control, Atlanta, GA). Values are expressed as means ± standard deviation (SD) or as numbers and percentages. Group comparisons for categorical variables were carried out using the χ2 test, while Student's t test was used for the analysis of continuous variables. Odds ratios (OR) and confidence intervals (CI = 95%) were also calculated. Multiple logistic regression analyses were carried out using the backward model. All statistical tests were based on a two-tailed probability, and a p value ≤ 0.05 was considered significant. The multivariate model was generated using variables with a p value > 0.20.
Results
The mean age of participating patients was 36.5 ± 12.2 years (range 18–83 years). Of the 254 patients, 66.9% were male, 57% were white, and 25.6% were HIV-positive. Co-medication during the TB treatment was used by 102 patients (40.2%). Table 1 of the Electronic Supplementary Material summarizes the co-medication used by the patients of this study. Of the 65 HIV-positive patients, 34 (52.3%) used co-medication, and five of these (7.7%) used sulfamethoxazole–trimetropin concomitant with anti-TB drugs. None of these patients developed hepatotoxicity. Of the 254 patients studied, 207 (81.5%) presented no ADR during anti-TB therapy. During the first 2 months of TB treatment, 33 (13%) patients presented a gastrointestinal ADR and 14 (5.5%) developed hepatotoxicity.
SNP frequency of the NAT2 gene
Sequencing analysis of the 254 patients enrolled in the study showed that ten different SNPs were present in this population, seven of which have been described previously in other studies as being the most frequent in different ethnic groups. Distribution of the allele frequencies of the seven more frequent SNPs, their presumed influence on the protein sequence, and their possible association with the hepatotoxicity outcome are summarized in Table 1. Among the seven most frequent SNPs identified in the studied population, variant 803G was present at the highest frequency (64.5%), while variant 191A (cluster NAT2*14) was the least frequent (1.4%). Three additional variants were found at frequencies lower than 0.4%: 345T, 578T, and 609T. The results of a linkage disequilibrium test carried out with the most frequently identified SNPs are shown in Table 2. The SNPs in the position 282T (cluster NAT2*13), 341C (cluster NAT2*5), 481T and 590A (cluster NAT2*6) were found to be significantly associated with anti-TB drug hepatotoxicity (p < 0.05). No significant association was observed between the SNPs evaluated and the occurrence of gastrointestinal ADR patients.
Hepatotoxicity and NAT2 genotype/ predicted phenotype
According to our classification, 69 (27.2%) and 185 (72.8%) patients were slow and fast acetylators, respectively. Genotype NAT2*12/5 (31.4%), which contains polymorphisms 341C and 803G that are characteristic of the fast acetylator profile, was the most prevalent among the 19 different genotypes observed (Table 3).
Among the 14 patients with hepatotoxicity, nine (64.3%) were slow acetylators and five (35.7%) were fast acetylators, with only one patient (20%) presenting two alleles for fast acetylation. Figure 1 shows the relationship between the NAT2 acetylator status and hepatotoxicity/gastrointestinal ADR patients. There was a noticeably significant association between the frequency of the slow acetylation profile and hepatotoxicity (p = 0.003; OR 5.5; CI 95% 1.6–19.8). Based on the criteria of the International Consensus, there was no observed significant association between hepatotoxicity and the acetylation profile in our patient cohort (p = 0.23).
The NAT2*6/6 genotype was significantly more frequent among patients that developed hepatotoxicity than among those without hepatotoxicity (OR 5.7; 95% CI 1.3–23) (Table 3). However, in gastrointestinal ADR patients, there was neither a significant association with the slow acetylation profile (p = 0.69) nor with a specific genotype.
Evaluation of clinical and genetic factors in relation to hepatotoxicity
Clinical, laboratory, and genetic information were analyzed. The mean age of patients who developed hepatotoxicity was 38.9 ± 12.8 years (range 19–66 years); 50% were female, 57.1% were HIV positive, 15.4% were HCV positive, and six (42.9%) were taking co-medication. Age, sex, alcohol abuse, presence of HCV and/or HBV, use of co-medication (antiretroviral drugs and others), aminotransferases, and bilirubin baseline were not significantly different among patients with and without hepatotoxicity, neither were they significant among patients with the fast or slow acetylator profile. Human immunodeficiency virus-positive and -negative patients also did not present any significant difference for the characteristics analyzed (data not shown).
In terms of hepatotoxicity, the HIV and slow acetylator profile were found to be risk factors (p < 0.01) in this population. After logistic multiple regression, HIV and slow acetylation profile remained as independent risk factors of hepatotoxicity (Table 4).
In an attempt to eliminate the confounding effect of HIV on hepatotoxicity, we subsequently restricted the association analysis between hepatotoxicity and acetylation profile to the 181 HIV-negative patients in our sample. Fifty (27.6%) HIV-negative patients were slow acetylators and 131 (72.4%) were fast acetylators. Six (3.3%) of the 181 HIV-negative patients developed hepatotoxicity, of whom four (66.7%) were slow acetylators and two (33.3%) were fast acetylators (presenting one allele for slow acetylation) (p = 0.05; OR 5.6; 95% CI 0.8–45.8).
When acetylation profile was compared to skin color, among 145 white patients, 68.3% were fast acetylators and 31.7% were slow acetylators (p = 0.04). The skin color was not significantly different between patients with and without hepatotoxicity (p = 0.40).
The effects of TB treatment on the levels of serum aminotransferases are shown in Table 5. During the peak, the serum levels of aminotransferases increased more than threefold ULN in 13% of the patients that presented two alleles for slow acetylation; this elevation was observed in only 0.5% of patients without slow alleles (p < 0.05). The elevated level of aminotransferases in patients that presented only one allele for slow acetylation was almost twofold higher than that in patients without alleles for slow acetylation, although the difference is not statistically significant (AST p = 0.2; ALT p = 0.1).
The mean dosage of RMP was 9.2 ± 1.2 mg/kg per day, INH, 6.1 ± 0.8 mg/kg per day, and PZA, 23.7 ± 3.2 mg/kg per day. These means were neither significantly different among fast and slow acetylators nor among patients with and without hepatotoxicity.
Discussion
We have determined the frequency of the NAT2 polymorphisms, the NAT2 acetylation profile and its relation to the occurrence of gastrointestinal ADRs and anti-TB drug-induced hepatotoxicity, and the clinical risk factors for hepatotoxicity. Our major finding is the association of the slow acetylation profile with anti-TB drug-induced hepatotoxicity in a population of TB patients from Southern Brazil (p < 0.005; OR 5.5; 95% CI 1.6 – 19.8), which is in accordance with results described by other researchers [7, 8, 10, 17]. The incidence of elevated levels of serum aminotransferase was significantly higher in slow acetylators than in fast acetylators (p < 0.05). This finding is in accordance with that observed by Ohno et al. [7]. Some studies have found that fast acetylators were more susceptible to developing anti-TB drug hepatotoxicity [14], while other studies have reported an increased risk of drug-induced hepatotoxicity among slow acetylators [7, 8, 10]. Huang et al. [8] studied 224 patients, with an incidence of 6.3% of hepatotoxicity, and observed that, in comparison to fast acetylators, slow acetylators had a higher incidence of hepatotoxicity as well as being more prone to developing more serious liver injury [8]. In contrast, a number of other studies did not find any relationship between acetylator status and drug-induced hepatotoxicity [9, 19, 30]. Vuilleumier et al. [9] studied a population of 89 Caucasian patients using only INH and did not observe a significant association between hepatotoxicity and the slow acetylation profile. This clear discrepancy among the results of previous studies on acetylation status and anti-TB hepatotoxicity may be due to the different designs of the studies, especially in terms of the methodology for NAT2 typing [8, 14], the anti-TB drugs used [7–9], and the criteria for defining anti-TB drug hepatotoxicity. The NAT2*6/6 genotype has also been found to be significantly associated with hepatotoxicity [8]. Higuchi et al. [10] studied 100 patients from Japan and reported that the NAT2*6A allele as well as the NAT2*6A/7B genotype were related to a higher incidence of hepatotoxicity. All of these findings support the hypothesis that the acetylation profile plays an important role in the pathogenesis of anti-TB drug-induced hepatotoxicity. Although many studies have described clinical aspects as risk factors for hepatotoxicity development [5, 31], INH and its metabolic intermediates have been indicated as a cause of hepatotoxicity [31]. Isoniazid is inactivated by NAT2, resulting in acetylisoniazid, which is hydrolyzed to acetylhidrazine. It has been proposed that acetylhidrazine is oxidized into hepatototoxic intermediates by cytochrome P4502E1 (CYP2E1). The other metabolic pathway to generate toxic intermediates is the direct hydrolysis of INH to hydrazine, a potent hepatotoxin [32]. Rifampin also induces hepatic amidase, which catabolizes acetylisoniazid into acetylhidrazine [33].
The frequency of the slow acetylator profile observed in our study was 27.2%, which is lower than that observed in many previous studies including well-characterized Caucasian populations where the slow acetylation profile observed was around 50% [34]. Teixeira et al. [26] studied 404 patients from two different populations from Brazil, one from Rio de Janeiro and another from Goiás. They predicted the NAT2 phenotype in 50% of the population studied, i.e. only in those homozygous for all SNPs found or heterozygous for only one SNP. Within the confines of this description, Teixeira et al. reported that 49% of patients were slow acetylators. The low frequency of the slow acetylation profile presented in our report could be related with the low frequency of SNP 590A (10.2%) (cluster NAT2*6). The frequency of SNP 590A (27%) reported by Teixeira et al. [26] was significantly higher than that found in our study (10.4%) (p < 0.01). The frequencies of other SNPs that characterize slow acetylation profile (341C and 857A) were not statistically different between our study and that conducted by Teixeira et al. [26]. The study carried out by Bailliet et al. [35] in which 90 individuals of different populations from Argentina and Paraguay were evaluated, identified as Amerindians on the basis of their geographic location, found that SNP 590A was present at a frequency of 9% only in Argentinean populations. The frequency observed in our study (10.4%) is similar to this value. Studies including European descendants reported the frequency of SNP 590A to be around 26% [34].
The Brazilian population is one of the most heterogeneous in the world due to the ethnic admixture of people from three continents: the European colonizers, the African slaves, and the native Amerindians [36]. This admixture of ethnicities is spread throughout the country, with predominance of certain groups depending on the region [26]. Caucasian from the Azores Islands founded Porto Alegre in 1752, and at present this population still has many Portuguese descendants, but Italians, Spanish and Germans have also contributed to its gene pool [37] as have those of African descent. For this reason, the pharmacogenetics data obtained for the population described in our study cannot be compared to other Brazilian populations or with well-characterized Caucasian populations.
The incidence of anti-TB drug-induced hepatotoxicity in our study was 5.5%, and 64.3% of these patients were slow acetylators. Following logistic multiple regression analysis, HIV and the and slow acetylation profile remained as independent risk factors for hepatotoxicity. In contrast to other studies [7–9, 38], which were able to exclude patients with chronic viral infection (HCV and HIV) as well as alcohol abusers to avoid confusion, we were unable to exclude these patients due to the high incidence of these co-infections and of alcohol abuse in the study population (HCV 18.6%, HIV 26.4%, alcohol abuse 16.5%). In addition, the treatment of patients with these characteristics is a daily routine in the outpatient clinic of Sanatorio Partenon Hospital and in all TB outpatient clinics from Porto Alegre.
In summary, the data presented here describe the frequency of SNPs for NAT2 and the NAT2 acetylation profile in a Southern Brazilian population and, for the first time, provide evidence of an association between the NAT2 slow acetylation profile and hepatotoxicity due to anti-TB drugs in Brazilian patients. However, further prospective studies must be carried out to determine the NAT2 acetylation profile and its association with risk to anti-TB drug-induced hepatotoxicity in different Brazilian regions. Given the limitation of the low number cases of hepatotoxicity, additional multi-center studies with a higher number of hepatotoxicity cases will be needed to confirm these findings. Clinical trials could be carried out to evaluate the performance of genotyping test for NAT2 as routine tools in clinical practice to pre-screen patients at high risk for hepatotoxicity in this population.
References
Brasil (2002) Guia de Vigilância Epidemiológica, 1st edn. Fundação Nacional de Saúde/Funasa, Brasília
Ingelman-Sundberg M (2001) Pharmacogenetics: an opportunity for a safer and more efficient pharmacotherapy. J Intern Med 250:186–200
Lundkvist J, Jonsson B (2004) Pharmacoeconomics of adverse drug reactions. Fundam Clin Pharmacol 18:275–280
Pande JN, Singh SP, Khilnani GC, Khilnani S, Tandon RK (1996) Risk factors for hepatotoxicity from antituberculosis drugs: a case-control study. Thorax 51:132–136
Ungo JR, Jones D, Ashkin D, Hollender ES, Bernstein D, Albanese AP, Pitchenik AE (1998) Antituberculosis drug-induced hepatotoxicity. The role of hepatitis C virus and the human immunodeficiency virus. Am J Respir Crit Care Med 157:1871–1876
Yee D, Valiquette C, Pelletier M, Parisien I, Rocher I, Menzies D (2003) Incidence of serious side effects from first-line antituberculosis drugs among patients treated for active tuberculosis. Am J Respir Crit Care Med 167:1472–1477
Ohno M, Yamaguchi I, Yamamoto I, Fukuda T, Yokota S, Maekura R, Ito M, Yamamoto Y, Ogura T, Maeda K et al. (2000) Slow N-acetyltransferase 2 genotype affects the incidence of isoniazid and rifampicin-induced hepatotoxicity. Int J Tuberc Lung Dis 4:256–261
Huang YS, Chern HD, Su WJ, Wu JC, Lai SL, Yang SY, Chang FY, Lee SD (2002) Polymorphism of the N-acetyltransferase 2 gene as a susceptibility risk factor for antituberculosis drug-induced hepatitis. Hepatology 35:883–889
Vuilleumier N, Rossier MF, Chiappe A, Degoumois F, Dayer P, Mermillod B, Nicod L, Desmeules J, Hochstrasser D (2006) CYP2E1 genotype and isoniazid-induced hepatotoxicity in patients treated for latent tuberculosis. Eur J Clin Pharmacol 62:423–429
Higuchi N, Tahara N, Yanagihara K, Fukushima K, Suyama N, Inoue Y, Miyazaki Y et al. (2007) NAT2*6A, a haplotype of the N-acetyltransferase 2 gene, is an important biomarker for risk of anti-tuberculosis drug-induced hepatotoxicity in Japanese patients with tuberculosis. World J Gastroenterol 13:6003–6008
Hughes HB, Biehl JP, Jones AP, Schimdt LH (1954) Metabolism of isoniazid in man as related to the occurrence of peripheral neuritis. Am Rev Tuberc 70:266–273
Evans DAP, White TA (1964) Human acetylation polymorphism. J Lab Clin Med 63:394–401
Weber WW, Hein DW (1979) Clinical Pharmacokinetics of isoniazid. Clin Pharmacokinet 4:401–422
Mitchell JR, Thorgeisson UP, Black M et al. (1975) Increased incidence of isoniazid hepatitis in rapid acetylators: possible relation to hydralazine metabolites. Clin Pharmacol Ther 18:70–79
Lin HJ, Han CH, Lin BK, Hardy S (1993) Slow acetylator mutations in the human polymorphic N-acetyltransferase gene in 786 Asians, Blacks, Hispanics and Whites: application to metabolic epidemiology. Am J Hum Genet 52:827–834
Doll MA, Fretland AJ, Deitz AC, Hein DW (1995) Determination of human NAT2 acetylator genotype by restriction fragment length polymorphism and allele specific amplification. Anal Biochem 231:412–420
Cho HJ, Koh WJ, Ryu YJ, Ki CS, Nam MH, Kim JW, Lee SY (2007) Genetic polymorphisms of NAT2 and CYP2E1 associated with antituberculosis drug-induced hepatotoxicity in Korean patients with pulmonary tuberculosis. Tuberculosis 87:551–556
Sarma GR, Immanuel C, Kailasan S, Narayana ASL, Venkatesan P (1986) Rifanpin-induced release of hydrazine from isoniazid. A possible cause of hepatitis during treatment of tuberculosis with regimens containing isoniazid and rifanpin. Am Rev Respir Dis 133:1072–1075
Singh J, Garg PK, Thakur VS, Tandon RK (1995) Antitubercular treatment induced hepatotoxicity: does acetylator status matter? Indian J Physiol Pharmacol 39:43–46
Parthasarathy R, Sarma GR, Janardhanam B, Ramachandran P, Santha T, Sivasubramanian S, Somasundaram PR, Tripathy SP (1986) Hepatic toxicity in South Indian patients during treatment of tuberculosis with short-course regimens containing isoniazid, rifampicin and pyrazinamide. Tubercle 67:99–108
Secretaria Estadual da Saúde (2001) Norma Técnica da política de Controle da Tuberculose. Rio Grande do Sul, Porto Alegre
Mayfield D, McLeod G, Hall P (1974) The CAGE questionnaire: validation of a new alcoholism screening instrument. Am J Psychiatry 131:1121
Castelo Filho A, Kritski AL, Barreto AW, Lemos ACM, Neto AR, Guimarães CA, Silva CL (2004) II Consenso Brasileiro de Tuberculose: Diretrizes Brasileiras para Tuberculose. J Bras Pneumol 30 Suppl 1
Bénichou C (1990) Criteria of drug-induced liver disorders:report of an international concensus meeting. J Hepatol 11:272–276
Muller AS, Dykes DD, Polesky HF (1988) A simple salting out procedure for extracting DNA from human nucleated cells. Nucleic Acids Res 16:1215
Teixeira RL, Miranda AB, Pacheco AG, Lopes MQ, Fonseca-Costa J, Rabahi MF, Melo HM et al. (2007) Genetic profile of the arylamine N-acetyltransferase 2 coding gene among individuals from two different regions of Brazil. Mutat Res 624:31–40
Long JC, Willians RC, Urbanek M (1995) An E-M algorithm and testing strategy for multiple-locus haplotypes. Am J Hum Genet 56:799–810
Long JC (1999) Multiple locus haplotype analysis, version 2.0. Software and documentation distributed by the autor. Section on population genetics and linkage. Laboratory of Neurogenetics (NIAAA), National Institute of Health, Bethesda
Feterson RJ, Goldman G, Long JC (1999) Effects of worldwide population subdivision on ALDH2 linkage disequilibrium. Genome Res 9:844–852
Gurumurthy P, Krishnamurthy MS, Nasareth O, Parthasarthy R, Samara GR, Somasundaran PR (1984) Lack of relationship between hepatotoxicity and acetylator phenotype in three thousand south Indian patients during treatment with isoniazid for tuberculosis. Am Rev Respir Dis 129:58–61
Fernandez-Villar A, Sopena B, Fernandez-Villar J, Vazquez-Gallardo R, Ulloa F, Leiro V, Mosteiro M, Pineiro L (2004) The influence of risk factors on the severity of anti-tuberculosis drug-induced hepatotoxicity. Int J Tuberc Lung Dis 8:1499–1505
Huang YS (2007) Genetic polymorphisms of drug-metabolizing enzymes and the susceptibility to antituberculosis drug-induced liver injury. Expert Opin Drug Metab Toxicol 3:1–8
Thomas BH, Wong LT, Zeitz W, Solomonarj G (1981) Isoniazid metabolism in the rabbit, and the effect of rifampin pretreatment. Res Commun Chem Pathol Pharmacol 33:235–247
Agundez JAG, Martinez C, Oliveira M, Ledesma MC, Ladero JM, Benitez J (1994) Molecular analysis of the arylamine N-acetyltransferase polymorphism in a Spanish population. Clin Pharmacol Ther 56:202–209
Baillet G, Santos MR, Alfaro EL, Dipierri JE, Demarchi DA, Carnese FR, Bianchi NO (2007) Allele and genotype frequencies of metabolic genes in Native Americans from Argentina and Paraguay. Mutat Res 627:171–177
Parra FC, Amado RC, Lambertucci JR, Rocha J, Antunes CM, Pena SDJ (2003) Color and genomic ancestry in Brazilians. Proc Natl Acad Sci USA 63:623–628
Salzano FM, Freire-Maia N (1970) Problems in human biology. A study of Brazilian populations. Wayne State University Press, Detroit
Hiratsuka M (2002) Development of simplified and rapid detection assay for genetic polymorphisms influencing drug response and its clinical applications. Yakugaku Zasshi 122:451–463
Acknowledgments
The authors wish to thank trial participants for their consent and participation in the study, and we also thank the staff of the outpatient Department and Research and Knowledge Department from Sanatorio Partenon Hospital for their assistance. This work was supported by National Council for Research (CNPq), Laboratório Exame, Novo Hamburgo, RS, and PADCT/ FEPPS. Adalberto R. Santos is supported by CNPq Grant number 308786/2005-0. We declare that the experiments comply with the Brazilian current laws.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Table 1
Analysis of medication type and drug groups in use by patients studied (DOC 62 KB)
Rights and permissions
About this article
Cite this article
Possuelo, L.G., Castelan, J.A., de Brito, T.C. et al. Association of slow N-acetyltransferase 2 profile and anti-TB drug-induced hepatotoxicity in patients from Southern Brazil. Eur J Clin Pharmacol 64, 673–681 (2008). https://doi.org/10.1007/s00228-008-0484-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00228-008-0484-8