Abstract
Studies on genetic connectivity are essential for the design of management strategies for coral reef fisheries. In this study we used a mitochondrial DNA marker to investigate population structure of the reef-associated parrotfish, Scarus ghobban, from four countries, Kenya, Mauritius, Seychelles and Tanzania, in the western Indian Ocean. We obtained nucleotide sequences of the mitochondrial control region for 117 individuals. Measures of haplotype diversity were relatively high. Pairwise population differentiation (F ST) was low, but not always non-significant. Analysis of molecular variance (AMOVA) showed genetic differentiation between groups, when the data was partitioned into two groups consisting of samples from Mauritius and Tanzania in one group, and samples from Kenya and Seychelles in another group. Direction of gene flow was estimated using a Bayesian approach. Migration was sometimes asymmetric or directional, coinciding with the flow of major oceanic and coastal currents in the region. Mismatch distributions, based on the observed number of differences among haplotype pairs, produced a unimodal distribution, indicative of recent demographic expansion. Phylogenetic analyses revealed three clades without any geographic structure, suggesting recent migration between historically isolated lineages. We reconstructed the historical demography of S. ghobban and examined it in the context of Pleistocene climate stages and changes in relative sea level. Overall, these results showed that populations of S. ghobban are genetically diverse and have relatively high gene flow, with some genetic structuring in the western Indian Ocean.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Coral Reefs are globally under threat from habitat destruction and degradation from wide-ranging causes, including climate-change-associated coral bleaching (Glynn 1983; Brown 1997; Douglas 2003; Hoegh-Guldberg 1999), over-fishing (Valentine and Heck 2005), pollution (Dubinsky and Stambler 2006) and coastal development (Rogers 1990). The decline of reef-associated fisheries has been linked to the deterioration in the general health and condition of coral reefs (Graham et al. 2007; Munday et al. 2008).
Genetic studies have emerged as an essential tool for the conservation of coral reef ecosystems. The information from such studies provides estimates of genetic diversity, a measure of an organism’s adaptive potential and ability to survive in the face of environmental change (Ryman et al. 1995; Schmitt and Hewitt 2004). Furthermore, marine propagules are notoriously difficult to physically monitor, and genetic markers can be used to track the dispersal of larvae from source to sink reefs, providing an indication of how distant reefs may be ‘connected.’ This is especially important, since knowing the likely location from where a degraded or depleted population recruits allows for management or protection of the source population, with benefits downstream. This type of reasoning has stimulated a number of studies on genetic connectivity and its implications for the design, i.e., number, size and placement, of marine protected areas (MPAs) (Palumbi 2003).
Dispersal of reef fish between populations is primarily achieved during their pelagic larval phase, as most adult reef fish do not move far from their home range. Dispersal may take place over large distances, particularly if larvae drift passively with the ocean currents (Roberts 1997), sometimes resulting in enormous capacity for dispersal and high levels of genetic connectivity across ocean basins (Craig et al. 2007). Despite the expectation that increased pelagic larval duration corresponds to increased levels of connectivity, this prediction has not always been realized (Jones et al. 2005; Taylor and Hellberg 2005), in part due to the recognized swimming ability of the pelagic larvae of many fish species in response to physical and environmental cues. This often results in the settlement of larvae relatively close to their natal reefs (Kingsford et al. 2002; Leis 2002; Simpson et al. 2004).
To date there have been few studies of genetic connectivity of coral reefs or reef-associated fisheries in the western Indian Ocean (WIO), an area that suffered from catastrophic coral bleaching during the 1997–1998 El-Niño Associated global coral bleaching event (Wilkinson 1999; Goreau et al. 2000). A notable exception was the study by Dorenbosch et al. (2006), in which they used amplified fragment length polymorphism (AFLP) to investigate the genetic structure of the Dory snapper, Lutjanus fulviflamma, from sites along the East African coastline (Kenya and Tanzania) and the Comoros Archipelago. Their results revealed low genetic differentiation among populations, and the authors suggested that this species may be genetically connected over large distances in the WIO.
Here, in a study of the blue-barred parrotfish, Scarus ghobban, sampled across a larger geographical scale from the region, we report on its population structure and genetic variability from 10 sites among four countries, Kenya, Mauritius, Seychelles and Tanzania, in the WIO.
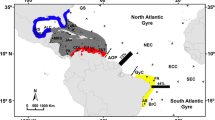
Ocean currents usually play a key role in the distribution and dispersal of marine organisms and the availability of nutrients. The South Equatorial Current (SEC), at about 10° south of the Equator, is the major oceanic current in the Indian Ocean, flowing from the eastern Indian Ocean to northern Madagascar and Africa in southern Tanzania and northern Mozambique (Richmond 2002). On making contact with the continent, the current splits to give rise to two main continental currents, the south-flowing Mozambique current, and the north-flowing East African Coastal Current (EACC). The latter flows to southern Somalia (Richmond 2002). Another important current in the region is the Equatorial Counter Current (ECC), at between 2 and 5°S. This current flows eastwards from northern Kenya towards the eastern Indian Ocean (Benny 2002).
A prominent shallow and mid-ocean feature influencing bathymetry and hydrodynamics in the region is the submerged Mascarene Plateau that extends for approximately 2,300 km between the Seychelles Bank and Mauritius (Fisher et al. 1967; Payet 2005).
Based on the above description of the oceanographic conditions in the WIO, particularly the major oceanic and continental currents, there are at least four hypotheses that could be tested relating to the pattern of dispersal for S. ghobban: (1) Self-recruitment hypothesis: populations from each country could be isolated in a fine geographical scale. (2) Stepping stone hypothesis: populations could disperse to adjacent areas following isolation-by-distance model. (3) Hypothesis of long distance dispersal through ocean currents, further divided into two parts: (3a) Oceanic islands versus continental mainland: oceanic populations (e.g., samples from Mauritius in this study) could be relatively isolated from mainland populations that are dispersed by continental current e.g., EACC. (3b) Equatorial current systems: The SEC and ECC could play a role in the dispersal of S. ghobban in the WIO. In this study, we test these hypotheses for discriminating the population structure and dispersal model of S. ghobban through population genetic analysis using a highly variable marker, the mitochondrial DNA (mtDNA) control region. In addition we carry out phylogenetic analyses to test for the presence of genetic lineages, and also reconstruct the demographic history of S. ghobban in the region and discuss it in the context of historical changes in climate and relative sea level.
Materials and methods
Fish samples
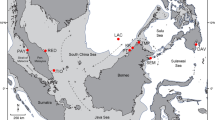
S. ghobban is a mass-spawning reef fish with a pelagic larval phase (Leis and Carson-Ewart 2000). It has a complex social structure, and is a sequential hermaphrodite (Allsop and West 2003). The fish is widespread and abundant in the Indo-Pacific Ocean region with moderate-to-high economic value to the artisanal fishery in countries from the WIO region (Sousa and Dias 1981) in which the study was carried out. Tailfin clips of S. ghobban were obtained from fishermen at local fish landing sites, shown in Fig. 1, and preserved in 70% ethanol. A total of 83 samples were collected between May and July in 2007 from 2 sites in Kenya and 3 sites each in Tanzania and Mauritius, and further collections of 34 samples from 2 sites in Seychelles were made between February and April in 2009 (Fig. 1). The topography of bathymetric data in this region was based on Smith and Sandwell (1997) and was created by using GeoMapApp (http://www.geomapapp.org/).
DNA extraction, PCR amplification and sequencing
Total DNA was extracted by dissolving 25 mg of tail fin in a solution of 100 μL of lysis buffer with 3 μL of ProK (10 mg/mL). The samples were incubated at 56°C for 4 h, and vortexed and spun down in a bench top centrifuge at 60 min intervals during the incubation. DNA was precipitated using 200 μL of freezing 99% EtOH and 20 μL of NaAc (3 M). The pellet was then spun down and washed with 100 μL of 70% EtOH. Excess EtOH was removed and the DNA was dried over night at ambient temperature and dissolved in 50 μL of 1× TE buffer. Extracted DNA was visualised on 1% agarose gels. Polymerase Chain Reaction (PCR) was carried out to amplify 351 bp at the 5′ end of mitochondrial control region, using the primer pair, ScarusDloopF: 5′ AATTCTCACCCCTAGCTCCCAAAG 3′ and ScarusDloopR: 5′ CCTGAAGTAGGAACCAGATG 3′, designed from the complete DNA sequence of the mitochondrial control region of Thallasoma hardwicki (C. A. Chen, unpublished data). A PC-960 thermal sequencer (Corbett Research, Australia) was used to perform PCR with the following thermal cycle: 1 cycle at 94°C (3 min); 33 cycles at 94°C (30 s), 50°C (30 s) and 72°C (1 min), followed by 1 cycle at 72°C (5 min), before cooling to 25°C. Between 50 and 200 ng of template DNA was used for the PCR in a 50 μL volume reaction using the buffer supplied with the enzyme and under conditions recommended by the manufacturer. Each 50 μL reaction mix contained 1.5 mM MgCl2, 0.2 mM dNTPs, 0.2 μM of each primer, 1 unit of Taq polymerase (Invitrogen) and approximately 100 ng of genomic DNA. The PCR products were electrophoresed in a 1% agarose (Invitrogen) gel to check the yield. The nucleotide sequences were determined for both ends of the PCR products using an ABI 377 Genetic Analyzer. Following an initial alignment with CLUSTAL W 1.7 (Thompson et al. 1994), all S. ghobban sequences were manually edited using SeqApp 1.9 (Gilbert 1994).
Population structure
Population comparison (F ST) based on genetic distances between pairs of populations (Reynolds et al. 1983; Slatkin and Hudson 1991) was calculated using ARLEQUIN v3.1 (Schneider et al. 2000). The significance threshold of pairwise comparisons (P < 0.05) was always adjusted by sequential Bonferroni correction (Rice 1989).
The matrix of distances was calculated using the Kimura 2-parameter model. The standard variance and haplotypic correlation measures (Φ statistics) could be used to reveal the extent of population subdivision (Schneider et al. 2000). The significance of the resultant statistics and variance components was tested with 10,000 permutations. Unique haplotypes were recorded and haplotypic diversity, h, nucleotide diversity, π (Nei 1987), and their standard deviations were calculated with DNASP 4.5 (Rozas et al. 2003).
The extent of population structuring was then examined using four sets of hierarchical AMOVA (Excoffier et al. 1992). Based on the prevailing oceanographic conditions in the WIO, notably the major oceanic and continental currents, we developed four hypotheses of how the different samples may group together into different populations. First, we divided the samples according to the country from where they were obtained. Second, samples were divided into a theoretical population from the mainland (Kenya and Tanzania combined) and two isolated populations from the oceanic islands (Mauritius and Seychelles). This was based on the assumption that the EACC would serve to homogenise S. ghobban from Kenya and Tanzania. We also tested a hypothesis that divides the samples into two groups, with Mauritius and Tanzania comprising one group and samples from northern Kenya and Seychelles into another. This was in order to test the possible influence of the SEC and the ECC in dispersal of recruits of S. ghobban between Mauritius and Tanzania, and between Kenya and Seychelles respectively. Finally, we examined the possibility that S. ghobban from Tanzania were connected to Kenya, possibly via the EACC, and that Mauritius and Seychelles were connected, possibly via the Mascarene Plateau.
To test the stepping stone hypothesis, the slope and intercept of isolation by distance (IBD) relationship based on reduced major axis (RMA) regression was calculated by using Isolation By Distance Web Service (IBDWS) version 3.15 (Jensen et al. 2005). A Mantel test was used (30,000 permutations) to assess whether the association between logarithm of genetic distance and logarithm of geographic distance was statistically significant. Genetic distance of zero or negative value was set to 0.0001 because zero or negative values could not be log-transformed and any negative values of F ST were considered as infinite gene flow between populations. As geographic distances between population pairs were greater than habitat width, F ST values were adjusted with the following transformation: Rousset’s genetic distance F ST/(1 − F ST) (Rousset 1997). Geographic distances between samples were measured according to the minimum coastline distance by using Google Earth™ (http://earth.google.com/).
The program MIGRATE 3.0.3 (Beerli and Felsestein 2001) was used for estimations of the Theta parameter (Θ = N efμ, N ef is effective population size, μ is mutation rate per site per generation) for each population and gene flow (M = m/μ, is a measure of how much immigration is over mutation to bring new variants into the population; m is migration rate) between multiple populations. A Bayesian search strategy (Beerli 2006) was selected with three replicates of four chains each running for 1 million generations. Universal substitution rate of mitochondrial control region of teleost as 3.6% per million years was used (Donaldson and Wilson 1999). Number of migrants per generation was calculated as the equation, N ef m ji = Θ i × M ji and P value of comparison of pair of N efm was estimated by using ANOVA.
Historical demography
The historical demography of S. ghobban was examined with mismatch distributions based on the observed and simulated differences against the number of differences among all pairs of haplotypes by using Arelequin 3.1 (Slatkin and Hudson 1991; Roger and Harpending 1992). The sum of square deviation (SSD) and Harpending’s raggedness index were used to test for fit to a unimodal mismatch distribution indicating a population was under sudden expansion.
We also reconstructed the past population demography for S. ghobban using the Bayesian skyline plot approach of Drummond et al. (2005) implemented in the program BEAST 1.48 (Drummond and Rambaut 2007). Markov Chain Monte Carlo (MCMC) sampling procedures were used to estimate a posterior distribution of effective population size through time from a sample of gene sequences, given a previously specified nucleotide substitution model (Drummond et al. 2002). We performed three MCMC runs for 10 million iterations, sampling genealogy and population size parameters every 2,000 iterations and discarding the first 10% as burn-in. We used the GTR +Γ as most appropriate substitution model selected by Akaike Information Criterion (AIC) in Modeltest 3.07 (Posada and Crandall 1998). Although a fixed substitution rate was applied (3.6% per million years) (Donaldson and Wilson 1999), uncorrelated lognormal model was used to account for rate variation among lineages (Drummond et al. 2006). Demographic history, i.e., effective population size, through time was reconstructed and plotted using the software Tracer version 1.4.1 (Rambaut and Drummond 2007). This data was interpreted by comparison with two sets of geological information; The relative sea level (RSL) data over the past 432 Ka was estimated from global δ18O data (Waelbroeck et al. 2002), and Pleistocene climate stages in the tropical Indian Ocean were identified based on weight percentage (wt) % of CaCO3 coarse fraction over past 0.8 Ma from Ocean Drilling Program (ODP) Site 758 (5°23′N, 90°21′E: 2,925 m water depth) (Farrell and Janecek 1991). To check if there were different effects on demography of coastal populations of S. ghobban derived from oceanic islands or the continental mainland, we also constructed individual Bayesian skyline plots for mainland populations from Kenya and Tanzania and oceanic populations from Mauritius and Seychelles.
Phylogenetic analyses
Phylogenetic analyses of S. ghobban including outgroups, S. rivulatus (Genbank Accession AY324603.1), S. dimidiatus (AY324591.1) and S. rubroviolaceus (AY324604) were conducted with two independent algorithms; PhyML (Guindon and Gascuel 2003) and Mr. Bayes 3.12 (Ronquist and Huelsenbeck 2003). For the maximum likelihood (ML) analysis, we used the GTR +Γ substitution model and gamma shape parameter (Γ = 0.323). The ML analysis with 500 bootstrap replicates was performed using PhyML ver. 3.0 (Guindon and Gascuel 2003), via a web server (http://www.atgc-montpellier.fr/phyml/). The best solution among two approaches, NNIs and Subtree Pruning and Regrafting (SPR) moves was used in tree topology search. An approximate likelihood ratio test (Anisimova and Gascuel 2006), the non-parametric, Shimodaira-Hasegawa-like branch test was used to estimate the reliability of each node.
MrBayes 3.12 was performed for four independent runs of a MCMC analysis with four chains running for 3 million generations. We saved trees every 300 generations for each run and set the substitution type as GTR (nst = 6), alpha parameter of the gamma distribution for rate heterogeneity (uniform 0.00–200) using 4 categories and tree topology parameters (uniform over all possible topologies). To ensure that stationarity was reached, the first 15% (0.45 million) generations from each run as burn-in were discarded. A majority-rule consensus tree calculated from the post burn-in trees was constructed and used to determine the posterior probabilities of clades.
Results
DNA variation
After trimming unreadable signals flanking the primers at either end, a nucleotide sequence of 351 bp at the 5′ end of mitochondrial control region was obtained for 117 Scarus ghobban individuals collected from 10 sampling sites in 4 countries (refer to map in Fig. 1). Measures of genetic diversity, i.e., haplotype and nucleotide diversity are displayed in Table 1. A total of 102 haplotypes were identified, of which 76 were unique or endemic to the sampling sites. In general, total haplotypic diversity was high (0.988 ± 0.005). Of the 10 sites, 4 had a haplotype diversity of 1; S. ghobban did not share an identical haplotype with another individual sampled from the same site.
Population structure
The population structure of S. ghobban was estimated with samples collected from 2 sites each in the north Kenya coastline and in Seychelles and 3 sites each in Mauritius and Tanzania. All pairwise F ST values were low and non-significant with the exception of a site in Seychelles, Mahe, being significantly genetically differentiated from 2 sites in Tanzania, Kilwa and Tanga (Table 2).
Results of AMOVA, shown in Table 3 in which four hypotheses of sample groupings were tested, indicate significant genetic structure between the two groups based on the Equatorial Currents, (Kenya, Seychelles) and (Mauritius, Tanzania) (P = 0.032). Within-population variations were all significant in the 4 comparisons, with the greatest variation in genetic structure in all cases arising from the ‘within populations’ component of the AMOVA.
Pairwise logarithm of geographic distances plotted against pairwise logarithm of genetic distances among 10 populations shows a positive correlation suggesting isolation by distance (r = 0.305) (Fig. 2). The Mantel test confirmed that the association between logarithm of genetic distance [log F ST/(1 − F ST)] and logarithm of geographic distance (log Km) was statistically significant (P = 0.0018).
We used the program MIGRATE to estimate effective population sizes, and mean number of migrants per generation of female S. ghobban between countries (Table 4), and between the clades identified by phylogenetic analysis (Fig. 5). The effective female population size in Mauritius (1.71 × 106) was estimated to be lower than the effective population sizes in each of the other three countries under study, which ranged between 2.27 and 2.40 × 106. Asymmetric gene flow, occurred in 3 instances; (a) There were more than twice as many migration events per generation from Mauritius to Kenya (24.4) as in the reverse direction (9.4; P < 0.05). (b) Gene flow from Mauritius to Tanzania was strongly directional, with the greatest estimated value from our study for number of migrants per generation between countries of 55.9, contrasting to only 3.7 migrants in the opposite direction (P < 0.001). (c) Gene flow from Tanzania to Kenya (45.2 migrants per generation) was more than twice that from Kenya to Tanzania (19.9; P < 0.001). There were no significant differences between the number of migrants per generation from each of the other 3 countries to and from Seychelles. The largest contributor of migrants to each country was as follows; Mauritius (26.6 migrants from Seychelles), Seychelles (31 migrants from Mauritius), Kenya (45.2 migrants from Tanzania), Tanzania (55.9 migrants from Mauritius). The mean number of migrants per generation between each of the 3 clades from phylogenetic analysis was generally low, ranging between 0.6 and 4.2, and not directional.
Historical demography
The historical demography of S. ghobban was analysed by mismatch distributions based on the observed number of differences among all pairs of haplotypes. The frequency of mismatches (Fig. 3) more closely resembles a unimodal distribution than multimodal, indicative of populations that have undergone recent demographic expansions within the time horizon of the marker with a non-significant P value (P = 0.36) of the sum of squared deviations (SSD) and low Harpending’s raggedness index (r = 0.0036) (Slatkin and Hudson 1991; Roger and Harpending 1992).
Bayesian skyline plot using BEAST shows that before 250 Ka, the overall population size of S. ghobban was relatively stable, or even gradually expanding, despite glacial periods or sea level fluctuations to 120 m below present level or more (Fig. 4). All S. ghobban populations underwent two major demographic bottleneck events during the long-term glacial periods at climate stage 6 (CS6; 130–190 Ka) and the last glacial maximum (LGM, CS2; 19–23 Ka) to the present date. There were two stages of population expansion of S. ghobban: an earlier and gradually occurring one at 190–400 Ka and a recent one characterised by rapid growth at 70–130 Ka. The S. ghobban derived from each country had similar demographic curves exhibiting dramatic population expansion during 70–130 Ka.
Climate change and the demographic history of Scarus ghobban over the past 1 Ma. (a) Composite relative sea level (RSL) curves over the past 400 Ka (Waelbroeck et al. 2002). The major glacial intervals are characterized as even gray strips of climate stages from Farrell and Janecek (1991) in the tropical Indian Ocean for the last 0.8 Ma. (b) The Bayesian skyline plots (Drummond et al. 2005) are derived from mtDNA control region sequences for Scarus ghobban in the western Indian Ocean, and the y-axis is equal to the product of female effective population size and generation time in years. (c) Bayesian skyline plot for each group. M Mauritius, K Kenya, T Tanzania, S Seychelles. Pairs of thin lines indicate the boundaries of 95% highest posterior density (HPD) interval
Phylogenetic analyses
The maximum likelihood (ML; Online Resource 1 which can be found in the supplementary materials) and Bayesian consensus (Fig. 5) phylogenetic trees constructed with S. ghobban mtDNA D-loop sequences exhibited three consistent clades. Clade1 and Clade 2 were identified with confident Bayesian posterior probabilities but without strong bootstrap support in the ML tree. Subclades of Clade 1 and 2 were also consistent within both ML and Bayesian trees, and the clustering is therefore reliable. However, clade 3 could comprise different subclades owing to the inconsistent topologies between ML and Bayesian trees. Additionally, there was no geographic relationship among taxa, i.e., each clade comprised individuals from each of the countries from where samples were derived, with the exception of clade 3 which did not have any individuals from Mauritius. This suggests that there has been recent migration between genetic lineages that were previously isolated.
Bayesian inference tree based on Scarus ghobban mtDNA D-loop sequences. Individuals are labelled according to site abbreviations, as listed in Table 1. Posterior probability of Bayesian inference tree is shown on the branch near nodes. Only above 0.70 of posterior probability is shown. Clades are identified by the majority consensus cluster of taxa. Outgroups used for the tree are Scarus rivulatus (Sri; Genbank Accession AY324603.1), S. dimidiatus (Sdi; AY324591.1) and S. rubroviolaceus (Sru; AY324604)
Discussion
High gene flow and ocean currents
The relatively high level of gene flow that we report for S. ghobban in the WIO is consistent with the general pattern of high gene flow for reef fish with a pelagic larval phase (Chen et al. 2004; Craig et al. 2007). Indeed there are numerous examples of marine organisms that exhibit no significant population structure over distances spanning thousands of kilometres (Palumbi 1992, 1994; Benzie 1999), including many species of fish (Waters et al. 2000; Riginos and Victor 2001; Planes and Fauvelot 2002, Craig et al. 2007; Klanten et al. 2007; Horne et al. 2008). Our results are not unexpected, especially in light of a study by Lessios and Robertson (2006) in which a wide range of reef associated fish species, including S. ghobban and two other species of parrotfish, were found to sporadically traverse the eastern Pacific barrier (Ekman 1953), a unique and vast expanse of uninterrupted deep water that stretches for ca 5,000 km, and separates the eastern Pacific from the central Pacific Ocean.
This large capacity for dispersal is principally brought about by ocean currents (Roberts 1997). In the WIO, these currents are dominated by the SEC, an oceanic current that may serve to potentially transport larvae from the islands in the WIO to the East African Mainland, and the EACC, that may then ferry larvae northwards along the continent. The ECC, which flows between northern Kenya towards the eastern Indian Ocean and which is more pronounced during the northeast monsoon period (December to April; Benny 2002), one of the two peaks of spawning of East African reef fish (Nzioka 1979), may also influence dispersal patterns. The AMOVA results from this study validate the Equatorial Currents hypothesis; samples grouped on this basis have reduced genetic exchange, because one Seychelles site (Mahe) and two Tanzania sites (Kilwa and Tanga) are genetically differentiated. However, it does not hold for the other samples (Kenya and Mauritius) or for the other sites within Seychelles (Praslin) and Tanzania (Bagamoyo).
Convincing support for the role of regional currents in the dispersal of S. ghobban also comes from our estimates on the direction and number of migrants per generation between locations. Asymmetric or directional gene flow, favouring migrants from Mauritius to Tanzania, and Tanzania to Kenya, proceeds downstream with the SEC and the EACC, respectively. These analyses on migration rates also highlight three further points; (a) Secondary or indirect currents may also strongly influence dispersal. For example, migrants from Mauritius to northern Kenya would initially be under the primary influence of the SEC, followed by the secondary influence of the EACC. Our results indicate that gene flow between Mauritius and Kenya is asymmetric, with twice as many migrants from Mauritius to Kenya as in the other direction. (b) The ECC, a seasonally pronounced current, does not seem to have a large effect on dispersal since gene flow between northern Kenya and Seychelles occurs at about the same rate in either direction. (c) In the absence of a major current system between Seychelles and Mauritius, the Mascarene Plateau serves as an important stepping stone, connecting the two countries. This is evident from the observation that the greatest contributor of migrants in Seychelles is Mauritius, and vice versa.
Some studies have shown that, despite the absence of any apparent barriers to dispersal, some reef fish with a high dispersal capacity demonstrate high genetic divergence among populations (Bell et al. 1982; Planes et al. 1994; McMilliam and Palumbi 1995; Shulman and Bermingham 1995; Waters et al. 2000; Riginos and Victor 2001; Fauvelot and Planes 2002; Planes and Fauvelot 2002; Taylor and Hellberg 2003). This implies that whereas there may be relatively high gene flow among the study sites as suggested by the relatively low and primarily non-significant F ST values from our data, it is vital to include more study sites from the region before arriving at a general conclusion on the level of genetic connectivity of S. ghobban in the entire WIO. In addition, studies carried out on species ranging from the tiger prawn, Penaeus monodon (Duda and Palumbi 1999; Benzie et al. 2002) and the striped mullet, Mugil cephalus (Rossi et al. 1998) to bigeye tuna, Thunnus obesus (Appleyard et al. 2002), have reported genetic partitioning of stocks between the western and eastern Indian Ocean. Bay et al. (2004) have also described a similar pattern of a western-eastern Indian Ocean genetic division in the bullethead parrotfish Chlorurus sordidus using mtDNA. An extension of our work on S. ghobban to include samples from the eastern Indian Ocean is therefore an important and opportune direction for future research.
Climate change and historical demography
Population demography of S. ghobban underwent perturbations during long-term glacial periods during 130–190 Ka and the last glacial maximum (LGM; 19–23 Ka). There are two possible factors that affected the demography of reef fish during these glacial periods. A 5°C-cooling would exceed the survival limits of most corals (Crowley 2000) and consequently reduce the amount of available habitat for reef fish. However, during the LGM the sea surface temperature (SST) only dropped by ~1°C in the tropical WIO (Barrows and Juggins 2005). This is not only within the temperature tolerance range of scleractinian corals, but also tropical reef fish. Many reef-associated species appear able to survive long-term exposure as long as the rate of temperature change does not exceed their capacity for acclimation. For example, Mangal-associated bathygobiids and squaretail mullet tolerate temperatures between about 16 and 38°C (Eme and Bennett 2009). Although little is known on the thermal tolerance of S. ghobban, a wide range of thermal tolerance can be inferred from their widespread latitudinal distribution of between 30° north and south of the equator. (FishBase, http://www.fishbase.org/). Therefore, the effect of SST changes during the most recent glacial period would not directly have exerted a large impact on S. ghobban’s habitat and demography in the tropical WIO.
Alternatively, a strong decline of coral reefs caused by sea level changes during glacial maxima would inevitably be accompanied by drastic reductions in populations of reef fish and invertebrates. This is quite evident from a meta-analysis of fish monitoring studies, which indicated that 62% of fish species declined within 3 years of a reduction of at least 10% coral coverage (Wilson et al. 2006). Our data indicates that S. ghobban has a high dispersal ability of at least 2,500 km, and previous studies show that S. ghobban occupies a depth range of between 3 and 36 m (Humann and Deloach 1993). Adjacent areas on an extensive scale could have served as potential refugia when surface of the sea dropped 60 m below present level during glacial periods. However, during the glacial maxima of CS2 and CS6, sea levels dropped to 120 m below present level. During these periods, S. ghobban would have undergone dramatic decline owing to shortage of habitats, as the area of ocean floor with a bathymetric depth range of 120–180 m, equivalent to a depth range of 0–60 m during CS2 and CS6, is exceedingly less than the current area occupied by the upper 60 m in the WIO (Figs. 1, 4). It is plausible therefore, that populations would have been disturbed by a drop in sea levels of 120 m, but not by changes in sea level less severe than that. We recognize that when sea level previously dropped by more than 120 m below present level ~370 Ka during CS10, S. ghobban population remained stable, and therefore additional factors influencing demography are implicated. A possible explanation lies in the fact that S. ghobban is much less dependent on coral reefs than other coral reef fish, and these herbivores can survive by scraping algae from rock or coral (Humann and Deloach 1993). Therefore, populations may have persisted through sea level drops as great as 120 m below present sea level, during which they would have survived on the algae that blossomed on rocky reefs at such times.
This aspect about the ecology of this species has likely permitted it to persist and maintain high genetic diversity despite massive historical changes in coral reef habitat in the WIO (also see Hughes and Stachowicz 2004). Intra-specific genetic diversity is critical for the adaptive potential of species (Ryman et al. 1995; Schmitt and Hewitt 2004), and the observation from this study of high diversity of mtDNA in S. ghobban bodes well for the long-term survival and adaptability of this species in the WIO.
The demographic profiles of skyline plot curves for each individual country (Fig. 4c), regardless of whether it is an oceanic or coastal reef or the total area of reef structure in each country, are strikingly similar. This likely indicates that S. ghobban in the WIO shares a common demographic history due to their high capacity for dispersal. Recently, climate change has become a serious cause of concern for coral reef organisms, in part due to rising sea level and elevated temperatures. With respect to this, our demographic studies serve to illustrate that reef fish with a greater capacity for dispersal have higher resilience.
Phylogenetic partition and evolutionary history
The phylogeny of S. ghobban d-loop exhibits three clades without geographic relationship (Fig. 5). “Clade 3” forms a polytomy cluster at the base of the tree, which are all sister to clades 1 and 2. This suggests that clades 1 and 2 are more recent than the other clades, which presumably acted as the source populations giving rise to the clade 1 and 2 populations, all of which have since dispersed extensively in the region leading to the observed lack of geographic structure. The composition of “clade 3” includes samples from Kenya, Tanzania and Seychelles, but none from Mauritius. One explanation is that Mauritian S. ghobban might originate from the other three areas. Alternative explanations are that the observed pattern may have resulted either from insufficient sampling, or lineage extinction in Mauritius.
Absence of spatial partitioning could reflect a recent demographic expansion of S. ghobban in the WIO, as suggested by the data from mismatch distribution and BEAST analysis. It is possible that geographic partitioning may have existed historically, and that spatial expansion would have led to the loss of geographic relationships, but the retention of genetically partitioned lineages or clades, as evident in the phylogenetic trees that we present for S. ghobban.
Conclusion and implications for management
Pertinent to the issue of management of fish stocks is the distinction between historical and contemporary gene flow. When interpreting patterns of gene flow, it is important to consider the geological and oceanographic history of the region (Barber et al. 2000; Fauvelot and Planes 2002). If there has been a relatively recent divergence between populations of S. ghobban in the region then the rate of evolution of the mitochondrial marker used for this study may not be sufficiently high to detect that divergence. Any future studies should capitalise on recent advances in methods (Hellberg 2007) that allow for estimates of gene flow over ecologically significant timescales of a few generations.
References
Allsop DJ, West SA (2003) Constant relative age and size at sex change for sequentially hermaphroditic fish. J Evol Biol 16:921–929
Anisimova M, Gascuel O (2006) Approximate likelihood-ratio test for branches: a fast, accurate, and powerful alternative. Syst Biol 55:539–552
Appleyard SA, Ward RD, Grewe PM (2002) Genetic stock structure of bigeye tuna in the Indian Ocean using mitochondrial DNA and microsatellites. J Fish Biol 60:767–770
Barber PH, Palumbi SR, Erdmann MV, Moosa MK (2000) A marine Wallace’s line? Nature 406:692–693
Barrows TT, Juggins S (2005) Sea-surface temperatures around the Australian margin and Indian Ocean during the last glacial maximum. Quat Sci Rev 24:1017–1047
Bay LK, Choat JH, van Herwerden EL, Robertson DR (2004) High genetic diversities and complex genetic structure in an Indo-Pacific tropical reef fish (Chlororus sordidus): evidence of an unstable evolutionary past? Mar Biol 144:757–767
Beerli P (2006) Comparison of Bayesian and maximum-likelihood inference of population genetic parameters. Bioinformatics 22:341–345
Beerli P, Felsestein J (2001) Maximum likelihood estimation of a migration matrix and effective population sizes in n subpopulations by using a coalescent approach. Proc Natl Acad Sci USA 98:4563–4568
Bell JL, Moyer JT, Numachi K (1982) Morphological and genetic variation in Japanese populations of anemonefish Amphriprion clarkii. Mar Biol 72:99–108
Benny PN (2002) Variability of western Indian Ocean currents. West Indian Ocean J Mar Sci 1:81–90
Benzie JAH (1999) Genetic structure of coral reef organisms: ghosts of dispersal past. Am Zool 39:131–145
Benzie JAH, Ballment E, Forbes AT, Demetriades NT, Sugama NT, Haryanti S, Moria S (2002) Mitochondrial DNA variation in Indo-Pacific populations of the giant tiger prawn, Penaeus monodon. Mol Ecol 11:2553–2569
Brown BE (1997) Coral bleaching: causes and consequences. Coral Reefs 16:S129–S138
Chen CA, Ablan MCA, McManus JW, Bell JD, Tuan VS, Cabanban AS, Shao K-T (2004) Population structure and genetic variability of six-bar wrasse (Thallasoma hardwicki) in northern South China Sea revealed by mitochondrial control region sequences. Mar Biotechnol 6:312–326
Craig MT, Eble JA, Bowen BW, Robertson DR (2007) High genetic connectivity across the Indian and Pacific Oceans in the reef fish Myripristis berndti (Holocentridae). Mar Ecol Prog Ser 334:245–254
Crowley TJ (2000) Causes of climate change over the past 1000 years. Science 289:270–277
Donaldson KA, Wilson RR Jr (1999) Amphi-panamic geminates of snook (Percoidei: Centropomidae) provide a calibration of the divergence rate in the mitochondrial DNA control region of fishes. Mol Phylogenet Evol 13:208–213
Dorenbosch M, Pollux BJA, Pustjens AZ, Rajagopal S, Nagelkerken I, van der Velde G, Moon-van der Staay SY (2006) Population structure of the Dory snapper, Lutjanus fulviflamma, in the western Indian Ocean revealed by means of AFLP fingerprinting. Hydrobiologia 568:43–53
Douglas AE (2003) Coral bleaching—how and why? Mar Pollut Bull 46:385–392
Drummond AJ, Rambaut A (2007) BEAST: Bayesian evolutionary analysis by sampling trees. BMC Evol Biol 7:214
Drummond AJ, Nicholls GK, Rodrigo AG, Solomon W (2002) Estimating mutation parameters, population history and genealogy simultaneously from temporally spaced sequence data. Genetics 161:1307–1320
Drummond AJ, Rambaut A, Shapiro B, Pybus OG (2005) Bayesian coalescent inference of past population dynamics from molecular sequences. Mol Biol Evol 22:1185–1192
Drummond AJ, Ho SYW, Phillips MJ, Rambaut A (2006) Relaxed phylogenetics and dating with confidence. PLoS Biol 4:e88
Dubinsky Z, Stambler N (2006) Marine pollution and coral reefs. Global Chang Biol 2:511–526
Duda TF Jr, Palumbi SR (1999) Population structure of the black tiger prawn, Penaeus monodon, among western Indian Ocean and western Pacific populations. Mar Biol 134:705–710
Ekman S (1953) Zoogeography of the sea. Sidgwick and Jackson Ltd, London
Eme J, Bennett WA (2009) Critical thermal tolerance polygons of tropical marine fishes from Sulawesi, Indonesia. J Therm Biol 34:220–225
Excoffier L, Smouse PE, Quattro JM (1992) Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human DNA mitochondrial data. Genetics 131:479–491
Farrell JW, Janecek TR (1991) Late neogene paleoceanography and paleoclimatology of the northeast Indian Ocean (ODP Site 758). In: Proceedings, Ocean Drilling Program, Scientific results, volume 121, College Station, Ocean Drilling Program, pp 297–355
Fauvelot C, Planes S (2002) Understanding origins of present-day genetic structure in marine fish: biologically or historically driven patterns? Mar Biol 141:773–788
Fisher RL, Johnson GL, Heezen BC (1967) Mascarene plateau, western Indian Ocean. Geol Soc Am Bull 78:1247–1266
Gilbert DC (1994) SeqApp 1.9. A biological sequence editor and analysis program for Macintosh computers. Available via http://ftp.indiana.edu
Glynn PW (1983) Extensive bleaching and death of reef corals on the Pacific coast of Panama. Environ Conserv 10:149–154
Goreau T, McClanahan T, Hayes R, Strong A (2000) Conservation of coral reefs after the 1998 global bleaching event. Conserv Biol 14:5–15
Graham NAJ, Wilson SK, Jennings S, Polunin NVC, Robinson J, Bijoux JP, Daw TM (2007) Lag effects in the impacts of mass coral bleaching on coral reef fish, fisheries and ecosystems. Conserv Biol 5:1291–1300
Guindon S, Gascuel O (2003) A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst Biol 52:696–704
Hellberg ME (2007) Footprints on water: the genetic wake of dispersal among reefs. Coral Reefs 26:463–473
Hoegh-Guldberg O (1999) Climate change, coral bleaching and the future of the world’s coral reefs. Mar Freshw Res 50:839–866
Horne JB, van Herwerden L, Choat JH, Robertson DR (2008) High population connectivity across the Indo-Pacific: congruent lack of phylogeographic structure in three reef fish congeners. Mol Phylogenet Evol 49:629–638
Hughes AR, Stachowicz JJ (2004) Genetic diversity enhances the resistance of a seagrass ecosystem to disturbance. Proc Natl Acad Sci USA 101:8998–9002
Humann P, Deloach N (1993) Reef fish identification. Galapagos. New World Publications Inc, Florida, p 267 p
Jensen JL, Bohonak AJ, Kelley ST (2005) Isolation by distance, web service. BMC Genet 6: 13. v.3.15. Available via http://ibdws.sdsu.edu/
Jones GP, Planes S, Thorrold SR (2005) Coral reef fish larvae settle close to home. Curr Biol 15:1314–1318
Kingsford MJ, Leis JM, Shanks A, Lindeman KC, Morgan SG, Pineda J (2002) Sensory environments, larval abilities and local self-recruitment. Bull Mar Sci 70:309–340
Klanten SO, Choat JH, van Herwerden L (2007) Extreme genetic diversity and temporal rather than spatial partitioning in a widely distributed coral reef fish. Mar Biol 150:659–670
Leis JM (2002) Pacific coral-reef fishes: the implications of behaviour and ecology of larvae for biodiversity and conservation, and a reassessment of the open population paradigm. Environ Biol Fishes 65:99–208
Leis JM, Carson-Ewart BM (eds) (2000) The larvae of Indo-Pacific coastal fishes. An identification guide to marine fish larvae. Brill Academic Publishers, Leiden, Boston, Köln, 850 pp. ISBN 9004115773
Lessios HA, Robertson DR (2006) Crossing the impassable: genetic connections in 20 reef fishes across the eastern Pacific barrier. Proc R Soc B 273:2201–2208
McMilliam WO, Palumbi SR (1995) Concordant evolutionary patterns among Indo-West Pacific butterfly fishes. Proc R Soc Lond B Biol Sci 260:229–236
Munday PL, Jones GP, Pratchett MS, Williams AJ (2008) Climate change and the future for coral reef fishes. Fish Fish 9:261–285
Nei M (1987) Molecular evolutionary genetics. Columbia University Press, New York
Nzioka RM (1979) Observations on the spawning seasons of East African reef fishes. J Fish Biol 14:329–342
Palumbi SR (1992) Marine speciation on a small planet. Trends Ecol Evol 112:319–326
Palumbi SR (1994) Reproductive isolation, genetic divergence, and speciation in the Sea. Annu Rev Ecol Syst 25:547–572
Palumbi SR (2003) Population genetics, demographic connectivity and the design of marine protected areas. Ecol App 13:S146–S158
Payet R (2005) Research, assessment and management on the Mascarene Plateau: a large marine ecosystem perspective. Phil Trans R Soc A 363:295–307
Planes S, Fauvelot C (2002) Isolation by distance and vicariance drive genetic structure of a coral reef fish in the Pacific Ocean. Evolution 56:378–399
Planes S, Borsa P, Galzin R, Bonhomme F (1994) Geographic structure and gene flow in the manini (convict surgeonfish, Acanthurus triostegus) in the South Central Pacific. In: Beaumont AR (ed) Genetics and evolution of aquatic organisms. Chapman & Hall, London, pp 113–122
Posada D, Crandall KA (1998) MODELTEST: testing the model of DNA substitution. Bioinformatics 14:817–818
Rambaut A, Drummond AJ (2007) Tracer v1.4. Available from http://beast.bio.ed.ac.uk/Tracer
Reynolds J, Weir BS, Cockerham CC (1983) Estimation of the coancestry coefficient: basis for a short-term genetic distance. Genetics 105:767–779
Rice WR (1989) Analyzing tables of statistical tests. Evolution 43:223–225
Richmond MD (ed) (2002) A field guide to the seashores of Eastern Africa and the western Indian Ocean islands. Sida/SAREC-USDM, 461 pp. ISBN 91-586-8783-1
Riginos C, Victor BC (2001) Larval spatial distributions and other early life-history characteristics predict genetic differentiation in eastern Pacific blennioid fishes. Proc R Soc Lond B Biol Sci 268:1931–1936
Roberts CM (1997) Connectivity and management of Caribbean coral reefs. Science 278:1454–1457
Roger AR, Harpending H (1992) Population growth makes waves in distribution of pairwise genetic differences. Mol Biol Evol 9:552–569
Rogers CS (1990) Responses of coral reefs and reef organisms to sedimentation. Mar Ecol Prog Ser 62:185–202
Ronquist F, Huelsenbeck JP (2003) MRBAYES 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574
Rossi AR, Capula M, Crosetti D, Sola L, Campton DE (1998) Allozyme variation in global populations of striped mullet, Mugil cepahlus (Pisces: Mugilidae). Mar Biol 131:203–212
Rousset F (1997) Genetic differentiation and estimation of gene flow from F-statistics under isolation by distance. Genetics 145:1219–1228
Rozas J, Sánchez D, Xavier M, Rozas R (2003) DnaSP, DNA Polymorphism analyses by the coalescent and other methods. Bioinformatics 19:2496–2497
Ryman N, Utter F, Laikre L (1995) Protection of intraspecific biodiversity of exploited fishes. Rev Fish Biol Fish 5:417–446
Schmitt T, Hewitt GM (2004) The genetic pattern of population threat and loss: a case study of butterflies. Mol Ecol 13:21–31
Schneider S, Roessli D, Excoffier L (2000) Arlequin version 2.000: a software for population genetics data analysis. Genetics and Biometry Laboratory, University of Geneva, Switzerland
Shulman MJ, Bermingham E (1995) Early life histories, ocean currents, and population genetics of Caribbean reef fishes. Evolution 49:897–910
Simpson SD, Meekan MG, McCauley RD, Jeffs A (2004) Attraction of settlement-stage coral reef fishes to reef noise. Mar Ecol Prog Ser 276:263–268
Slatkin M, Hudson RR (1991) Pairwise comparison of mitochondrial DNA sequences in stable exponentially growing populations. Genetics 129:555–562
Smith WH, Sandwell DT (1997) Global seaflood topography from satellite altimetry and ship depth soundings. Science 277:1957–1962
Sousa MI, Dias M (1981) Catálogo de peixes de Moçambique—Zona Sul. Instituto de Desenvolvimento Pesqueiro Maputo 121 pp
Taylor MS, Hellberg ME (2003) Genetic evidence for local retention of pelagic larvae in a Caribbean reef fish. Science 299:107–109
Taylor MS, Hellberg ME (2005) Marine radiations at small geographic scales: speciation in neotropical reef gobies (Elacatinus). Evolution 59:374–385
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Valentine JF, Heck KL Jr (2005) Perspective review of the impacts of overfishing on coral reef food web linkages. Coral Reefs 24:209–213
Waelbroeck C, Labeyrie L, Michel E, Duplessy JC, McManus JF, Lambeck K, Balbon E, Labracherie M (2002) Sea-level and deep water temperature changes derived from benthic foraminifera isotopic records. Quat Sci Rev 21:295–305
Waters JM, Dijkstra LH, Wallis GP (2000) Biogeography of a southern hemisphere freshwater fish: how important is marine dispersal. Mol Ecol 9:1815–1821
Wilkinson CR (1999) Global and local threats to coral reef functioning and existence: review and predictions. Mar Freshw Res 50:867–878
Wilson SK, Graham NAJ, Pratchett MS, Jones GP, Polunin NVC (2006) Multiple disturbances and the global degradation of coral reefs: are reef fishes at risk or resilient? Global Chang Biol 12:2220–2234
Acknowledgments
This paper greatly benefited from the comments of its anonymous reviewers. We are grateful to the staff of WWF Kiunga Station for assistance with sampling of fish in northern Kenya. Jan Robinson and Rodney Govinden of the Seychelles Fishing Authority provided us with samples from Seychelles. This work was supported by the Western Indian Ocean Marine Science Association (WIOMSA) through their Marine Science for Management (MASMA) Grant program and Academia Sinica Thematic Grants (2006–2007) to CAC. SV is the recipient of a postdoctoral research fellowship from Academia Sinica (2006–2010) and MCY is the recipient of a PhD scholarship from the Biodiversity Research Centre, Academia Sinica (BRCAS; Coral Reef Evolutionary Ecology and Genetics contribution No. 56).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by T. Reusch.
Shakil Visram, Ming-Che Yang are Co-first authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Visram, S., Yang, MC., Pillay, R.M. et al. Genetic connectivity and historical demography of the blue barred parrotfish (Scarus ghobban) in the western Indian Ocean. Mar Biol 157, 1475–1487 (2010). https://doi.org/10.1007/s00227-010-1422-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00227-010-1422-8