Abstract
The identification of beef in animal foods is a major concern not only for the prevention of commercial fraud, but also to avoid safety risks deriving from the presence of prohibited bovine material that might be harmful to both human and animal health. Here we report a novel set of bovine-specific primers, CYTbos1 (forward) and CYTbos2 (reverse), which allow the specific amplification of a 115 base pair fragment of the bovine cytochrome b gene (cytb) between nt 844 (mitochondrial site 15,590) and nt 958 (mitochondrial site 15,704), no cross-reaction being observed with DNA from another 12 frequent commercial meat species. The polymerase chain reaction product obtained is cleaved specifically by endonucleases ScaI and TspE1 to achieve further confirmation evidence. The sensitivity of the proposed method was 0.025%. The CYTbos primers successfully detected bovine DNA in meat samples processed for 20 min at 133 °C/300 kPa or for 2 h at 121 °C. CYTbos primers also detected bovine DNA in heat-processed commercial meat products exhibiting a complex nature, as well as in bovine specific risk materials. The proposed polymerase chain reaction method, aimed at detecting a small and specific fragment of the bovine mitochondrial DNA, may be especially useful for the direct identification of bovine DNA in foodstuffs subjected to severe heating under overpressure conditions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Traditionally, species identification has focused on the prevention of commercial fraud—which involves the substitution of an animal species of higher commercial value, such as beef, by others of less commercial value—also affording a valuable tool for the assessment of safety risks derived from the introduction of any animal material that might be harmful to both human and animal health. In this sense, it is widely accepted that bovine spongiform encephalopathy (BSE) has spread through the consumption of contaminated animal meals by healthy bovines [1]. This critical situation has moved the European Union (EU) to enforce a ban on feeding animals with any material of animal origin [2, 3], with a view to preventing the spread of the syndrome. Moreover, bovine remains, including nervous tissue and viscera, are now considered specific risk materials (SRMs) in Europe, and they must be removed from the food chain and stained with a dye that facilitates the follow-up of such material to avoid its inclusion in any food product destined for human or animal consumption [4]. In addition, and with a view to ensure food safety, the EU authorities require that animal by- products are processed at 133 °C at 300 kPa for 20 min [5].
Recently, heat-stable proteins have been reported to be useful targets for both the detection of animal remains and species identification in animal foods, such as meat [6] and fish products [7]. However, methods based on DNA amplification are preferred, since they are not so affected by industrial processing. Among such DNA-based methods, the analysis of mitochondrial DNA (mtDNA) has been reported to be a powerful tool for identifying beef with respect to that of other land animal species [8, 9] because (1) its presence in multiple copies per cell (as many as 2,500 copies in a postmitotic tissue such as skeletal muscle), increases the probability of achieving a positive result even in the case of samples undergoing intense DNA fragmentation due to severe processing conditions [10], and (2) its large variability compared with nuclear sequences, which undergo a less rapid evolution, facilitates authenticity studies [11].
Among mitochondrial targets, the cytochrome b (cytb) gene has frequently been considered one of the preferential DNA targets for identification purposes. Thus, universal primers based on cytb sequences have been widely used in polymerase chain reaction (PCR)-restriction fragment length polymorphism and DNA sequencing studies in vertebrates [12–15]. However, the reliability of this approach may be hampered by the existence of polymorphic DNA sequences that can complicate restriction analysis, as has been found for bovine DNA in previous studies [16]. To overcome this problem, a PCR-based method aimed at the specific amplification of bovine-specific mtDNA sequences from animal feeds has recently been tested [17–19] and validated [20] as a way to ensure the exclusion of bovine materials from animal feeds. Nevertheless, the mean size of PCR products obtained with most primers previously developed for the specific identification of bovine DNA is often higher than 250 base pairs (bp), and this might limit the success of DNA amplification in samples exhibiting intense DNA degradation caused by heat processing under overpressure conditions [21]. Thus, although the amplification of a 265 bp PCR product in a ruminant feed heated at temperatures up to 141.9 °C has been reported [22], the need to improve existing methods and to develop newer PCR-based methods with potential application to a wide variety of heated products has recently been highlighted [21, 23].
In previous work we reported on the optimisation of an extraction method for the recovery, amplification and species-specific analysis of DNA from hard animal tissues, such as bone and derived bone meals [15]. In the present work, the main goal was to develop a novel PCR-based method aimed at detecting a small—115-bp—specific fragment of the mitochondrial cytb gene of cattle DNA and to evaluate its usefulness for the specific detection of bovine DNA in foodstuffs and SRMs subjected to severe heat-processing under overpressure conditions.
Materials and methods
Design of bovine-specific primers based on mitochondrial cytb sequences
The nucleotide sequences of the cytb gene from different animal species of interest in the food sector were retrieved from NCBI databases, the accession numbers being listed in the following. Sequences from cattle (Bos taurus V00654 and J01394), pig (Sus scrofa domesticus X56295 and 4220565), wild boar (Sus scrofa scrofa AB015082), chicken (Gallus gallus AF028795 and X52392), turkey (Meleagris gallopavo L08381), duck (Cairina moschata L08385), quail (Coturnix coturnix L08377), deer (Dama dama AJ000022 and Cervus elaphus AJ000021), roe deer (Capreolus capreolus Y14951), ostrich (Struthio camelus U76055 and NC002785), rabbit (Oryctolagus cuniculus NC001913), sheep (Ovis aries AF034730 and X56284), goat (Capra hircus AF217254 and AB044308), buffalo (Bubalus bubalis D88637 and D82892) and horse (Equus caballus D82932 and D32190) were considered.
Novel bovine-specific primers were designed by means of GENEFISHER software [24], according to the following premises: (1) special attention was focused on a less studied variable cytb region comprising nt 437 (mitochondrial site 15,183) to nt 1,140 (mitochondrial site 15,886) and located in the mitochondrial genome downstream from the most frequent cytb universal targets from previous PCR studies; (2) novel primers should permit the amplification of a cytb fragment only in the case of bovine DNA; (3) such amplification products should be smaller than 150 bp, with a view to increasing the possibilities of obtaining positive results with materials subjected to intense heat-processing conditions.
The following bovine-specific primers were designed: CYTbos1 (forward 5′-CGATCAATCCCCAACAAACTA-3′), and CYTbos2 (reverse 5′-GAAGCATAATATTCCGACCAC-3′), which theoretically amplify a 115-bp region of the cytb gene comprised between nt 844 (mitochondrial site 15,590) and nt 958 (mitochondrial site 15,704) only in the case of bovine DNA. Table 1 displays the specificity of these primers for bovine DNA, showing the mismatches with respect to other animal species of interest in the food sector. Prediction of restriction sites inside the 115-bp PCR product was carried out using DNASIS software (Hitachi Software Engineering Co., Japan).
DNA extraction and purification from raw and heated skeletal muscle of cattle and other animal species
Meat samples were obtained from the skeletal muscle of cattle, sheep, goat, pig, wild boar, chicken, turkey, duck, quail, roe deer, ostrich, rabbit and horse. DNA was extracted from portions of 250 mg of each sample, either raw or heated, by means of a commercial kit (DNeasy Tissue Isolation kit, QIAGEN, Darmstadt, Germany). Heat-processing of skeletal muscle was performed at 121 °C (15 min, 30 min, or 2 h) or at 133 °C (20 min) in a laboratory autoclave (Raypa, model AE 75 TIC, Sterilmatic, Barcelona, Spain). DNA concentrations in the purified extracts were determined by fluorimetry using an LS 50 fluorimeter (Applied Biosystems, Foster City, CA) after mixing with Hoechst 33258 reagent (Sigma Chemical Co., St. Louis, MO). When required, DNA extracts were concentrated using a Microcon YM-100 centrifugal filter system (Millipore, Bedford, MA), following the manufacturer’s instructions. The sensitivity of the PCR method was explored in mixtures containing 0.01, 0.025, 0.05, 0.1, 0.25, 0.5, 1, 2, 5, 10 and 25% of beef in commercial wheat flour. A sample of wheat flour without bovine material was included as a negative control.
Extraction of DNA from commercial meat products and SRMs
DNA was also extracted from portions of 250 mg of both commercial meat products and SRM, as described earlier. Thus, four precooked foods—tortellini with veal meat, ravioli with veal meat, dehydrated soup with veal meat and pasteurised/smoked beef sausage—and four commercial sterilised product samples—two types of commercial meat balls and two types of canned beef—were purchased from local supermarkets. All these eight commercial food samples investigated reputedly contained beef as the only meat ingredient and included a variety of different products subjected to either pasteurisation or sterilisation temperatures, respectively. In addition, two different bovine SRMs were obtained from a local slaughter house and were subjected to heat processing for 20 min at 133 °C/300 kPa prior to DNA extraction.
DNA amplification
All amplification assays comprised 100 ng of template DNA, 25 μl of a master mix (BioMix, Bioline, London, UK)—this including reaction buffer, deoxyribonucleosidetriphosphates, magnesium chloride and Taq DNA polymerase–PCR water (Genaxis, Montigny le Bretonneaux, France), and 25 pmol of each oligonucleotide primer to achieve a final volume of 50 μl. All DNA extracts were tested in parallel for the absence of amplification inhibitors by the performance of an amplification assay with the universal primers CYTb1 and CYTb2, designed by Kocher et al. [12], this yielding in all cases a 359-bp PCR product of the cytb mitochondrial gene. The conditions of this amplification assay have been previously described [15]. Briefly, a previous denaturing step at 94 °C for 1.5 min was coupled to 25 cycles of denaturation (94 °C for 10 s), annealing (55 °C for 30 s), and extension (72 °C for 40 s), and to a final extension step at 72 °C for 15 min.
Once the absence of amplification inhibitors had been checked, amplification conditions with the novel primers CYTbos1 and CYTbos2 were as follows: a previous denaturing step at 94 °C for 1.5 min was coupled to 25 cycles of denaturation (94 °C for 10 s), annealing (61 °C for 30 s), and extension (72 °C for 10 s), and to a final extension step at 72 °C for 15 min. All PCR assays were carried out using a GeneAmp 2700 thermal cycler (Applied Biosystems).
DNA electrophoretic separation and image analyses
PCR products were processed in homemade 2.5% horizontal agarose (MS-8, Pronadisa, Madrid, Spain) gels in 1X tris(hydroxymethyl)aminomethane–acetate–ethylenediaminetetraacetate buffer. The agarose gels included 0.5 μg/ml ethidium bromide (Merck, Darmstadt, Germany) for visualisation purposes at 254 nm, running the gels at 100 V. Sizing of the PCR products was accomplished by comparison with a ladder consisting of a MspI-digest of plasmid pUC18 run in parallel (Sigma). Gels were photographed with a DC290 Zoom Digital Camera (Kodak, Edinburgh, UK) and image analysis was carried out by means of the 1-D Manager software (TDI, Madrid, Spain).
DNA sequencing and computer analyses
Both light and heavy strands of the 115-bp PCR product were sequenced to improve the reliability of the sequencing data by means of an automated DNA sequencing system (model 3100, Applied Biosystems). The DNA sequences were carefully reviewed by eye using Chromas software (Griffith University, Queensland, Australia). Prediction of restriction sites was carried out with DNASIS software (Hitachi Software Engineering Co., Japan).
Results and discussion
Specificity of CYTbos1 and CYTbos2 primers for the detection of bovine DNA
Although other authors have recently reported bovine-specific PCR methods aimed at detecting other target sequences from the 1,709 satellite DNA [25, 26], the lactoferrin gene [27], the 18S ribosomal RNA (rRNA) gene [27], the 12S rRNA gene [28], the glial fibrillary acidic protein-encoding gene [29], or short interspersed elements [30], this work was aimed, for the reasons described earlier, at developing a new PCR method based on cytb sequences.
With this purpose in mind, the oligonucleotides CYTbos1 (21 nt) and CYTbos 2 (21 nt), proposed in this work, were designed for the specific detection of bovine DNA in foodstuffs. The 115-bp PCR product displayed a G+C content of 46%. Although the predicted Tm was 58 °C, the annealing temperature was adjusted to 61 °C to avoid possible cross-amplification of related animal species. Extension was carried out at 72 °C for only 10 s, this being considered sufficient time to complete copies of the 73 nt flanking region comprised between both primers.
Figure 1 shows the suitability of CYTbos1 and CYTbos2 for the direct identification of bovine DNA, no cross-reaction being observed with skeletal muscle from other common commercial meat species. Thus, the novel primers designed in this work annealed specifically to bovine DNA sequences (Fig. 1a, lane 3) but not to any of the other 11 animal species checked; i.e., goat, sheep, pig, chicken, turkey, quail, rabbit, horse, wild boar, roe deer and ostrich (Fig. 1a, lanes 4–14). Interestingly, no cross-amplification was observed even in the case of animal species showing only one or two mismatches in the primer sequences (Table 1), such as the ruminant species goat (lane 4) or sheep (lane 5) (Fig. 1a). As stated previously, all these DNA extracts were tested for the absence of amplification inhibitors, by performing an amplification assay with CYTb1 and CYTb2 universal primers that in all cases yielded the expected 359-bp DNA fragment from the cytb gene (Fig. 1b). Accordingly, the lack of amplification of DNA extracts obtained from species other than cattle with CYTbos primers was clearly a consequence of the specificity of these primers for bovine DNA.
a Specificity of CYTbos primers for the detection of bovine DNA. Lane 1 molecular weight marker; lane 2 negative control; lane 3 cattle; lane 4 goat; lane 5 sheep; lane 6 pig; lane 7 chicken; lane 8 turkey; lane 9 quail; lane 10 rabbit; lane 11 horse; lane 12 wild boar; lane 13 roe deer; lane 14 ostrich. b Absence of amplification inhibitors in the DNA extracts as checked by amplification with CYTb1 and CYTb2 universal primers: lanes are as in a.
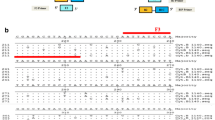
Although the size of the PCR product obtained for bovine DNA is only 115 bp, further evidence concerning the specificity of the primers for bovine DNA can be achieved by means of specific cleavage with endonucleases ScaI and TspE1. Thus, the nucleotide sequence of the 115-bp PCR product amplified from the cattle mtDNA was obtained (Fig. 2). Computer analyses predicted that ScaI recognises a unique AGT↓ACT sequence, specific to the 115-bp PCR product amplified from bovine DNA, this causing cleavage at nt 876 (mitochondrial site 15,619) and leading to two restriction fragments of 86 and 29 bp, only in the case of cattle (Fig. 2). Likewise, TspE1 recognises an ↓AATT sequence at nt 897 (mitochondrial site 15,643), this affording two restriction fragments of 62 and 53 bp only in the case of bovine DNA (Fig. 2).
Although previously reported bovine-specific primers have been based on other mitochondrial cytb sequences, the PCR products obtained in such studies exhibited sizes of 274 [9] or 285 bp [31]. Other authors also reported a bovine-specific PCR method aimed at the amplification of a 113-bp cytb region, although this method was designed and optimised for the identification of bovine milk [32]. The primers presented in this work, whose application for meat products and SRMs will be discussed later, allowed us to obtain in all cases a bovine-specific 115-bp fragment of the cytb mitochondrial gene. Such a small size is desirable to maximise the possibility of getting positive results from samples subjected to severe heat-processing, as has been suggested by other authors [21]. Other bovine-specific methods based on other mitochondrial targets, such as the transfer RNALys/ATPase subunit 8/ATPase subunit 6 genes or the cytochrome oxidase II gene, have also described PCR products of higher size: 271 [17, 18] and 651 bp [33]. Accordingly, the PCR method proposed in this work may be especially useful for the detection of bovine DNA in food samples subjected to intense heat-procesing under overpressure conditions, thus containing highly degraded DNA.
Sensitivity of CYTbos primers for the detection of bovine DNA
Mixtures of beef and commercial wheat flour were prepared at different ratios, as described earlier. A sample of wheat flour without bovine material was included as a negative control. Figure 3 shows the results of the sensitivity assays. The sensitivity of the method was at least 0.025%, this corresponding to 2.5 pg of bovine DNA (Fig. 3). Other studies have reported detection limits of 0.005% [30], more than 0.02% [27], 0.04% [17, 18], 0.0625% [22], 0.13% [34], 0.25% [35], 0.5% [29], or 1% [28]. In comparison with the results of those studies, the results of the method proposed in this work, based on specific cytb mitochondrial sequences, revealed a significantly high sensitivity for the detection of bovine DNA.
Sensitivity of CYTbos primers for the detection of bovine meat. Lane 1 molecular weight marker; lane 2 commercial wheat flour (negative control); lane 3 100% (positive control); lane 4 25%; lane 5 10%; lane 6 5%; lane 7 2%; lane 8 1%; lane 9 0.5%; lane 10 0.25%; lane 11 0.1%; lane 12 0.05%; lane 13 0.025%; lane 14 0.01%
Usefulness of CYTbos1 and CYTbos2 primers for the detection of bovine DNA in heated products
The identification and/or differentiation of animal species has proved to be difficult, particularly in samples of complex composition and subjected to intense processing [36]. Thus, although DNA exhibits fairly high thermal stability it is well known that intense heat coupled with overpressure conditions may cause severe DNA degradation, this affecting the quality of the DNA recovered [21].
In our work, portions of beef were aseptically fragmented to very small particles and subjected to heat-processing under overpressure conditions in an autoclave, as previously described. The meat was ground to facilitate the penetration of heat in order to achieve homogeneous and severe heating of the samples. Figure 4 shows the results obtained at different temperatures and heating times. Thus, bovine-specific PCR-based detection with the CYTbos primers was successful even after heating for 2 h at 121 °C, or for 20 min at 133 °C (Fig. 4). In light of this, the size of the bovine-specific DNA target proposed here—115 bp—may be sufficiently small to avoid amplification problems derived from severe DNA fragmentation in foodstuffs subjected to intense heating conditions.
Usefulness of CYTbos primers for the detection of bovine meat heated either at 121 or at 133 °C. a Lane 1 molecular weight marker; lane 2 negative control; lane 3 unheated bovine meat (positive control); lane 4 121 °C/15 min; lane 5 121 °C/30 min; lane 6 121 °C/2 h. b Lane 1 molecular weight marker; lane 2 negative control; lane 3 133 °C/20 min
The next goal of our work was to investigate the usefulness of the CYTbos primers to specifically detect bovine DNA in commercially heated meat products. The detection of beef in processed commercial foods is an important issue since it may be fraudulently substituted by other types of meat of less commercial value [37]. However, the processing conditions to which commercial foods are subjected may involve either the presence of additives—which may inhibit DNA polymerase—or intense heating conditions—which may degrade DNA to such an extent that amplification may not be possible [21]. Accordingly, the CYTbos primers were evaluated with different pasteurised and sterilised meat products including different additives and manufacturing processes. The results are shown in Fig. 5. The presence of bovine DNA was successfully identified in the four pasteurised meat products tested, amplification not being affected by the composition or processing conditions (Fig. 5a). Moreover, when the CYTbos primers were evaluated for the identification of beef in commercial meat products subjected to sterilisation, the results were also successful in all four cases (Fig. 5b). Accordingly, the primers proposed in this work proved to be useful for the direct and specific identification of beef in meat products, even in the case of complex foods subjected to intense heating under overpressure conditions.
Specific detection of bovine DNA in commercial heat-processed foods with CYTbos primers. a Pasteurised food products. Lane 1 molecular weight marker; lane 2 negative control; lane 3 tortellini with veal meat; lane 4 ravioli with veal meat; lane 5 soup with veal meat; lane 6 pasteurised/smoked beef sausage. b Sterilised canned products. Lane 1 molecular weight marker; lane 2 negative control; lane 3 unheated bovine meat (positive control); lanes 4–7 sterilised meat products containing only bovine meat
Identification of bovine SRM with CYTbos primers
CYTbos primers were also evaluated for their ability to identify bovine SRM. As stated earlier, this forbidden material exhibits a significant risk for the spread of BSE [4]. Thus, and although the EU has prohibited all kinds of bovine tissues as ingredients of animal feeds [2, 3], SRMs must be specifically separated from other bovine components at slaughter houses, dyed, and destroyed at high temperatures [4].
In our work, two different SRMs were processed at 133 °C/20 min and then subjected to DNA extraction and amplification under the previously described conditions. As may be observed in Fig. 6, the presence of bovine DNA was successfully detected in SRMs. Thus, the dye used for marking the SRMs did not act as an inhibitor in this amplification assay, and bovine DNA could be successfully amplified with CYTbos primers even from such stained material. In addition, amplification of bovine DNA in the SRMs was achieved even after severe heat-processing at 133 °C/20 min. A real-time method based on CYTbos primers is currently being developed in our laboratory.
Conclusions
The bovine-specific primers CYTbos1 and CYTbos2, based on cytb mitochondrial sequences, have allowed the direct and specific detection of bovine DNA in heated food samples, even in those processed for 2 h at 121 °C or for 20 min at 133 °C/300 kPa. The specificity and sensitivity—0.025%—of the proposed method, and the small size of the specific PCR product, make the proposed PCR method especially advisable for the direct identification of bovine DNA in samples that have suffered intense heat-processing under overpressure conditions, such being the cases of sterilised meat products and SRMs.
References
Wilesmith JW, Wells GAH, Granwell MP, Ryan JBM (1998) Vet Rec 123:638–644
Commission of the European Communities (1994) J Eur Comm L 172:0023–0024
Commission of the European Communities (2001) J Eur Comm L 006:16–17
Commission of the European Communities (2000) J Eur Comm L 158:0076–0082
Council of the European Communities (1999) Off J Eur Comm L 204:0037–0042
Chen FC, Hsieh YHP (2001) J AOAC Int 83:79–85
Piñeiro C, Vázquez A, Figueras A, Barros-Velázquez J, Gallardo JM (2003) J Proteome Res 2:127–135
Kikkawa Y, Amano T, Suzuki H (1995) Biochem Genet 33:51–60
Matsunaga T, Chikuni K, Tanabe R, Muroya S, Shibata K, Yamada J, Shinmura Y (1999) Meat Sci 51:143–148
Verkaar ELC, Nijman IJ, Boutaga K, Lenstra JA (2002) Meat Sci 60:365–369
Partis L, Croan D, Guo Z, Clark R, Coldham T, Murby J (2000) Meat Sci 54:369–376
Kocher TD, Thomas WK, Meyer A, Edwards SV, Pääbo S, Villablanca FX, Wilson AC (1989) Proc Natl Acad Sci USA 86:6196–6200
Meyer R, Höfelein C, Lüthy J, Candrian U (1995) J AOAC Int 78:1542–1551
Wolf C, Rentsch J, Hübner P (1999) J Agric Food Chem 47:1350–1355
Prado M, Franco CM, Fente CA, Cepeda A, Vázquez BI, Barros-Velázquez J (2002) Electrophoresis 23:1005–1013
Wu J, Smith RK, Freeman AE, Beitz DC, McDaniel BT, Lindberg GL (2000) Biochem Genet 38:323–335
Tartaglia M, Saulle E, Pestalozza S, Morelli L, Antonucci G, Battaglia PA, (1998) J Food Prot 61:513–518
Wang RF, Myers MJ, Campbell W, Cao WW, Paine D, Cerniglia CE (2000) Mol Cell Probes 14:1–5
Krcmár P, Rencová E (2001) J Food Prot 64:117–119
Myers MJ, Friedman SL, Farrell DE, Dove-Pettit DA, Bucker MF, Kelly S, Madzo S, Campbell W, Wang RF, Paine D, Cerniglia CE (2001) J Food Prot 64:564–566
Momcilovic D, Rasooly A (2000) J Food Prot 63:1602–1609
Bottero MT, Dalmasso A, Nucera D, Turi RM, Rosati S, Squadrone S, Goria M, Civera T (2003) J Food Prot 66:2307–2312
Prado M, Casqueiro J, Iglesias Y, Cepeda A, Barros-Velázquez J (2004) J Sci Food Agric 84:505–512
Giegerich R, Meyer F, Schleiermacher C (1994) Proceedings of the fourth international conference on intelligent systems for molecular biology, AAAI Press (ISSN 57735-002-2)
Guoli Z, Mingguang Z, Zhijian Z, Hongsheng O, Qiang L (1999) Meat Sci 51:233–236
Calvo JH, Roderllar C, Zaragoza P, Osta R (2002) J Agric Food Chem 50:5262–5264
Gao HW, Zhang D-B, Pan A-H, Liang W-Q, Liang C-Z (2003) J AOAC Int 86:764–767
Sun Y-L, Lin C-S (2003) J Agric Food Chem 51:1771–1776
Seyboldt AC, v Mueffling JT, Nowak B, Wenzel S (2003) J Food Prot 66:644–651
Walker JA, Hughes DA, Anders BA, Shewale J, Sinha SK, Batzer MA (2003) Anal Biochem 316:259–269
Herman BL (2001) J Dairy Res 68:429–436
Rea S, Chikuni K, Branciari R, Sangamayya RS, Ranucci D, Avellini P (2001) J Dairy Res 68:689–698
Janecek LL, Honeycutt RL, Adkins RM, Davis SK (1996) Mol Phylogen Evol 6:107–119
Andrews CD, Berger RG, Mageau RP, Schwab B, Johnston RW (1992) J AOAC Int 75:572–576
Bellagamba F, Valfrè F, Panseri S, Moretti VM (2003) J Food Prot 66:682–685
Laube I, Spiegelberg A, Butschke A, Zagon J, Schauzu M, Kroh L, Broll H (2003) Int J Food Sci Technol 38:111–118
Pascoal A, Prado M, Castro J, Cepeda A, Barros-Velázquez J (2004) Eur Food Res Technol 218:306–312
Acknowledgements
The authors wish to thank the Fundación Caixa Galicia for the financial support that made this work possible. The authors also thank the PGIDT Biotechnology Program (project PGIDIT02BTF 26102PR) from the Xunta de Galicia for financial support.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Pascoal, A., Prado, M., Calo, P. et al. Detection of bovine DNA in raw and heat-processed foodstuffs, commercial foods and specific risk materials by a novel specific polymerase chain reaction method. Eur Food Res Technol 220, 444–450 (2005). https://doi.org/10.1007/s00217-004-1088-x
Received:
Revised:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00217-004-1088-x