Abstract
Infectious diseases are a long-standing and severe global public health problem. The rapid diagnosis of infectious diseases is an urgent need to solve this problem. MicroRNA (miRNA) plays an important role in the intervention of some infectious diseases and is expected to become a potential biomarker for the diagnosis and prognosis of infectious diseases. It is of great significance to develop rapid and sensitive methods for detecting miRNA for effective control of infectious diseases. In this study, a simple and highly sensitive homogeneous electrochemical method for microRNAs using enzyme-driven cascaded signal amplification has been developed. In the presence of target miRNA, the reaction system produced plenty of MB-labeled single-nucleotide fragments (MB-MF) containing a few negative charges, which can diffuse to the negative surface of the ITO electrode easily, so an obvious electrochemical signal enhancement was obtained. Without the target, MB-HP contains a relatively large amount of negative charges due to the phosphates on the DNA chain, which cannot be digested by the enzyme and cannot diffuse freely to the negatively charged ITO electrode, so only a small signal was detected. The enhanced electrochemical response has a linear relationship with the logarithm of miRNA concentration in the range of 10 fM to 10 nM and the limit of detection as low as 3.0 fM. Furthermore, the proposed strategy showed the capability of discriminating single-base mismatch and performed eligibly in the analysis of miRNA in cell lysates, exhibiting great potential for disease diagnosis and biomedical research.

Graphical abstract
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
MicroRNAs (miRNAs), a class of noncoding RNA consisting of 18–25 nucleotides, play an essential role in physiological and pathological processes [1,2,3]. MiRNAs have received extensive attention by researchers because of their abnormal expression in a variety of diseases and have been regarded as valuable biomarkers that can be used in the diagnosis and treatment of some cancers [4,5,6]. In recent years, increasing studies showed that miRNAs play an important role in the intervention of some viral infectious diseases (such as HIV/AIDS, West Nile disease, Ebola virus disease, etc.) [7, 8]. The miRNAs of infected cells can inhibit the replication and spread of the virus and achieve the purpose of antiviral therapy. Given that miRNAs are prone to be degraded, have low abundance in samples, and have high sequence similarity among family members, it is necessary to develop simple but sensitive methods for miRNAs detection.
The electrochemical detection technique has received increasing attention owing to its intrinsic characteristics of low cost, fast response, high sensitivity, and simple operation [9]. Many electrochemical biosensors for miRNAs detection have been developed [10,11,12,13,14]. For example, Xu et al. developed an electrochemical biosensor that can realize simultaneous detection of two miRNAs [15]. Liu and his coworkers prepared the ultrathin Aunps-MoS2 nanoflakes on the FTO electrodes by combining atomic layer deposition, greatly increasing the detection sensitivity of the miRNA sensing platform [16]. Most of these miRNA electrochemical biosensors required the synthesis of complex materials or the immobilization of DNA probes on the electrode surface, this step is cumbersome, time-consuming, and may even greatly affect the reproducibility of the proposed biosensor [17, 18]. Furthermore, due to the steric hindrance effect of the electrode surface and the loss of conformational freedom of immobilized DNA, the hybridization efficiency of DNA probe and miRNA at the electrode/solution interface is relatively low [19]. Homogeneous electrochemical detection as an immobilization-free strategy had been put forward based on the difference in electrostatic repulsion between DNA of different strand lengths and the negatively charged electrode surface [20, 21], which can solve the abovementioned problems well. The principle has been applied to the detection of various analytical targets, including various small molecules, metal ions, enzymes and proteins, etc. [22,23,24], which had also been applied to detect miRNAs. For example, Hou and colleagues designed a signal-off homogeneous electrochemical sensor for the detection of miRNA using a hybridization chain reaction [25]. Our group earlier reported a homogeneous electrochemical biosensor using a duplex-specific nuclease (DSN) cycle target to achieve signal amplification for miRNA detection [26]. But the sensitivity of these biosensors is not high enough for real sample detection since the signal amplification technique had been used. There is still a great challenge for developing more sensitive methods for practical miRNA quantification.
Enzyme-driven cascade signal amplification strategy cleverly utilizes the functional properties of DNA exonucleases and DNA polymerases to achieve multiple amplification of the response to the target [27, 28]. The ultra-high signal amplification efficiency can meet the needs for the detection of low-abundance targets, making it particularly suitable for the detection of biomarkers. For example, by combining Bst. DNA polymerase, Nt. BstNBI, ribonuclease H, and other enzymes, Wang et al. developed a fluorescence strategy that can detect uracil-DNA glycosylase at the single-cell level [29]. Xiong and coworkers also designed a label-free electrochemical sensor with exonuclease III–assisted cascade signal amplification, and the detection limit of this method for its target DNA can be as low as femtomolar [30].
In this study, coupled with the convenience of the homogeneous strategy of DNA-based electrochemical detection and high amplification efficiency of enzyme-driven cascaded signal amplification, a homogeneous electrochemical biosensor for miRNA detection (miRNA-155 has been chosen as a model target, which is a common disease marker) had been proposed. The biosensor shows excellent anti-interference in the detection of complex samples and has been successfully applied to the detection of miRNA in different cell lysates with satisfactory results.
Experimental section
Reagents
Klenow fragment polymerase (3′ → 5′ exo, KF polymerase), T7 exonuclease (T7 exo), 10× NEBuffer 4, and dNTPs were bought from New England Biolabs (Ipswich, MA). RNase inhibitor was obtained from Bio Basic Inc. (Markham Ontario, Canada). DSN and 10× DSN master buffer were bought from Evrogen (Moscow, Russia). HEK-293 cells, HeLa cells, MCF-7 cells, miRNA and HPLC-purified DNA oligonucleotides used in this work were provided by Sangon Biotech Co., Ltd. (Shanghai, China), and the relevant sequences (5′ → 3′) are shown below:
cDNA probe: AGATTCGAACACTGCCATGACCCCTATCACGATTAGCATTAA
MB-HP: MB-ATCCTCACTGCAGACTCCATGGCAGTGTTCGCTGA
None-labeled HP: ATCCTCACTGCAGACTCCATGGCAGTGTTCGCTGA
miRNA-155: UUAAUGCUAAUCGUGAUAGGGGU
SM: UUAAGGCUAAUCGUGAUAGGGGU
SM-3′: UUAAGGCUAAUCGUGAUAGGGGA
SM-m: UUAAGGCUAAUGGUGAUAGGGGU
TM: UUAAGGCUAAUAGUGAUAUGGGU
miRNA-21: UAGCUUAUCAGACUGAUGUUGA
miRNA-141: UAACACUGUCUGGUAAAGAUGG
let-7a: UGAGGUAGUAGGUUGUAUAGUU
NC: UUGUACCUACACAAAAGUACUG
Electrochemical detection system
A CHI660A electrochemical workstation (Chenhua Instruments, Shanghai, China) was used for differential pulse voltammetry (DPV) measurements. The three-electrode electrochemical system used in the experiment consisted of an ITO electrode (the working area was 5 mm × 3 mm) as the working electrode, an Ag/AgCl electrode as the reference electrode, and a platinum wire as the counter electrode. The purchased ITO electrodes need to be ultrasonic sequentially in 10 g/L alconox solution, isopropanol, and ultrapure water for 15 min respectively to obtain a clean surface negatively charged ITO electrode. The DPV signals in the range of 0 ~ − 0.6 V were recorded under the electrochemical measurement parameters with an amplitude of 50 mV, quiet time of 2 s, pulse period of 20 ms, and pulse width of 50 ms.
Procedures for miRNA detection
First, 0.5 μM cDNA probe, different concentrations of miRNA, 0.1 U DSN, and 10 U RNase inhibitor were added into 1× DSN master buffer and incubated at 55 °C for 1 h. Second, the DSN enzyme was inactivated by adding 5 mM EDTA into the above mixture and incubating at 55 °C for 5 min. After that, 1.5 μM methylene blue-labeled hairpin DNA (MB-HP), 5 U KF polymerase, 4 mM dNTPs, 10 U T7 exo and 2 μL 10× NEBuffer 4 were added. The final total volume of the reaction system was 50 μL and incubated at 37 °C for 1.5 h. Finally, the electrochemical detection of the system was carried out and the DPV signal was recorded after the above reaction. Each sample was tested three times, and the average value of the three measurements was used for quantitative analysis.
Gel electrophoresis
The hybridization of the cDNA probe and the miRNA and DSN digestion process were explored by 12% non-denaturing polyacrylamide gel electrophoresis (PAGE). The working voltage was set to 80 V, and the electrophoresis was performed in 0.5× Tris-borate-EDTA running buffer (pH 8.4) for 2 h. The obtained gel was imaged by a gel image system (JS-2012, Peiqing Science & Technology Co., Ltd., Shanghai, China).
Results and discussion
Principle of the homogeneous electrochemical biosensor for miRNA detection
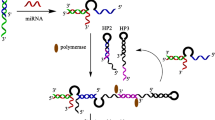
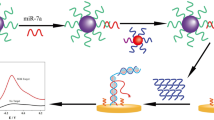
The principle of the proposed enzyme-assisted cascaded signal amplification homogeneous electrochemical miRNA biosensor is shown in Scheme 1. When the target miRNA is present, the miRNA hybridizes with a portion of the cDNA probe to form a DNA-RNA duplex. The duplex can be recognized and digested by DSN, thereby releasing miRNA and the remaining DNA fragments (released DNA). Meanwhile, the released target miRNA can combine another cDNA probe and repeat the cyclic cleavage reaction, leading to produce a great deal of released DNA (cycle I). Subsequently, the released DNA can be used to open up methylene blue–labeled hairpin DNA (MB-HP). The polymerization could be initiated by polymerase, resulting in an extended double-strand DNA with blunt 5′ terminus, which was then recognized and digested by T7 exo, generating short MB-labeled mononucleotide fragments (MB-MFs). In addition, a new single-strand DNA extended by KF polymerase (extended DNA) was released and can be used to open another MB-HP probe and generated a double-strand DNA with a 5′ blunt end, which was also recognized and digested by T7 exo (cycle II). The process produced a large amount of MB-MFs, which can easily diffuse to the negatively charged ITO electrode surface due to their low negative charge, so that an obvious DPV signal can be obtained. But in the absence of the target, nearly no released DNA had been obtained, so the MB-HP had not been opened and which cannot be digested by T7 exo. The hairpin-structured MB-HP and the duplex formed by hybridization with the cDNA probe both have strong electrostatic repulsion against the negatively charged ITO electrode (since the backbones of DNA contains negative charges) and are not easy to diffuse to the electrode surface, so only a weak signal can be detected. The electrochemical signal strength enhanced by adding target miRNA is related to the concentration of miRNA in the system. By this means, a simple, ultrasensitive, and specific homogeneous electrochemical miRNA biosensor can be developed.
Feasibility study
A gel electrophoresis experiment was employed firstly to verify the process of hybridization and DSN digestion. As shown in Fig. 1a, when only a cDNA probe was present, a short and bright band (lane a) was observed. After the addition of the target miRNA, a new band with a larger molecular weight was detected (lane b), indicating that the cDNA probe hybridized with miRNA to obtain a DNA-RNA duplex. Further addition of DSN results in the disappearance of the DNA-RNA duplex band. Since the 19-base released DNA and the 23-base miRNA were greatly overlapped on the band, only a short and shallow new band can be observed (lane c), indicating the DSN digestion reaction occurred.
A PAGE analysis of DSN enzyme–driven target cycle. Lane M: DNA marker; lane a: 1.0 μM cDNA probe; lane b: 1.0 μM cDNA probe + 0.5 μM miRNA; Lane c: 1.0 μM cDNA probe + 0.5 μM miRNA + 0.1 U DSN. B DPV responses under different conditions. (a) with cDNA probe, DSN, MB-HP, KF polymerase, dNTPs, and T7 exo; (b) with 1.0 pM miRNA, cDNA probe, DSN, MB-HP, KF polymerase, dNTPs, and T7 exo
Next, electrochemical detection had been applied to verify the feasibility of the proposed strategy. As displayed in Fig. 1b, when the target miRNA was not present, the DPV signal detected was quite weak (curve a). But in the presence of miRNA, DPV signal was significantly enhanced (curve b). That because the hybridization of miRNA with cDNA probe to form a RNA-DNA duplex, which can be cleaved by DSN, and the product released DNA can be used to open MB-HP, thereby eliciting KF polymerase extension reaction and the subsequent process of T7 exo cycling digestion, achieving signal amplification. All these results verify the feasibility of the assumption.
Optimization of the experimental conditions
The reaction conditions, such as the concentration of MB-HP, the dosage of KF polymerase and T7 exo, and the reaction time, play a critical role in the performance of the biosensor. These conditions had been optimized to obtain the best sensing performance. The experimental signal originates from methylene blue in the system. Excessive MB-HP will cause a high background signal, while insufficient MB-HP will affect the sensitivity of the system. So, the concentration of MB-HP was optimized firstly. The effect of the concentration of MB-HP (0.5 to 2.0 μM) on the DPV signal of the system was investigated when the concentration of miRNA was 1.0 pM. Figure 2a demonstrates that the DPV signals of the system increased as the MB-HP concentration increased from 0.5 to 1.5 μM, and tend to be stable when the MB-HP concentration exceeds 1.5 μM. When the MB-HP concentration is lower than 1.5uM, the background signal hardly increased as MB-HP increased. However, when the MB-HP concentration exceeds 1.5 μM, the background signal increased greatly. Thus, 1.5 μM MB-HP was consequently used for the subsequent studies.
The employed dosages of KF polymerase and T7 exo were also optimized. Figure 2b shows that the ΔI (the difference in the DPV peak current in the presence and absence of target) gradually increased with the dosage of KF polymerase elevated from 0 to 5.0 U firstly and ΔI almost unchanged if the dosage of KF polymerase was higher than 5 U. Therefore, 5 U was chosen as the optimal dosage of KF polymerase. As shown in Fig. 2c, the ΔI increased with the enhancing of the dosage of T7 exo from 0 to 10 U and then reached a plateau, so T7 exo of 10 U was chosen for the following experiments. Moreover, the reaction time of T7 exo had also been optimized, as shown in Fig. 2d; ΔI increased with the prolonged reaction time firstly and reached the platform at 90 min, indicating that 90 min was sufficient for the cleavage of T7 exo.
Performance of the proposed biosensor
Figure 3a shows that as the miRNA concentration increased, the DPV signal increased. The value of ΔI and the logarithm of miRNA concentration in the range of 10 fM to 10 nM exhibit a good linear relationship (shown in Fig. 3b). The linear regression equation is ΔI (μA) = − 0.2050 − 0.0835 lgCmiRNA, (R2 = 0.9952). The limit of detection (LOD) was calculated to be 3.0 fM (S/N = 3), which is comparable to or better than the reported methods for miRNA detection (Table 1).
MiRNA-141, miRNA-21, let-7a, single-base mismatched (SM), single-base mismatched in middle (SM-m), single-base mismatched in 3′-end (SM-3′) three-base mismatched (TM), and non-complementary (NC) were selected as interferences to explore the selectivity of the proposed biosensor for miRNA detection. Especially deserving to be mentioned, the concentrations of interferences are 100-fold higher than the target miRNA (1.0 pM). As illustrated in Fig. 4, a remarkable electrochemical signal had been observed when the target was added, while when the reaction system was incubated with these interfering substances, no obvious electrochemical signal enhancement was obtained compared with the background signal. Considering the above results, it can be concluded that this biosensor possesses high selectivity toward miRNA detection. In addition, five replicate experiments were conducted in the presence of 10 pM target miRNA, and the relative standard deviation (RSD) was 3.5%. The good repeatability also proves the reliability of the proposed sensing strategy for miRNA detection.
Real sample analysis
In view of the high sensitivity and good selectivity, the proposed homogeneous electrochemical biosensor has been applied to detect miRNA-155 in biological samples. The target in 10-fold diluted cell lysates of HEK-293 cell, HeLa cell, and MCF-7 cell were detected and the standard addition recoveries were tested to verify the validity of the proposed method. The results were shown in Table 2, the recovery rates of 94.0–108.2% were obtained when spiked with miRNA in the linear range, which indicated that the interference of cell lysates could be overcome, and the RSDs were in the range of 2.1–4.3%. Therefore, the proposed strategy is effective and reliable for miRNA detection and it is expected to realize the detection of miRNA in real biological samples.
Conclusion
A novel homogeneous electrochemical biosensor for miRNA determination based on the cascade signal amplification strategy had been developed, owing to the excellent ability of enzyme-powered cascaded signal amplification strategy and simple homogeneous electrochemical detection. This method successfully realized the simple and ultrasensitive homogeneous electrochemical quantitative detection of miRNA. The proposed method avoids the tedious procedure of probe immobilization. Furthermore, high sensitivity had been reached because of the cascaded signal amplification strategy. The proposed biosensor shows an excellent ability to measure miRNA in cell lysates and is expected to be applied in the field of disease diagnosis and biomedical research.
References
Lu J, Getz G, Miska EA, Alvarez-Saavedra E, Lamb J, Peck D, et al. MicroRNA expression profiles classify human cancers. Nature. 2005;435(7043):834–8.
Ruan K, Fang X, Ouyang G. MicroRNAs: novel regulators in the hallmarks of human cancer. Cancer Lett. 2009;285(2):116–26.
Pritchard CC, Cheng HH, Tewari M. MicroRNA profiling: approaches and considerations. Nat Rev Genet. 2012;13(5):358–69.
Tricoli JV, Jacobson JW. MicroRNA: potential for cancer detection, diagnosis, and prognosis. Cancer Res. 2007;67(10):4553.
Ryan BM, Robles AI, Harris CC. Genetic variation in microRNA networks: the implications for cancer research. Nat Rev Cancer. 2010;10(6):389–402.
Saito M, Schetter AJ, Mollerup S, Kohno T, Skaug V, Bowman ED, et al. The association of microRNA expression with prognosis and progression in early-stage, non–small cell lung adenocarcinoma: a retrospective analysis of three cohorts. Clin Cancer Res. 2011;17(7):1875.
Swaminathan G, Rossi F, Sierra L-J, Gupta A, Navas-Martin S, Martin-Garcia J. A role for microRNA-155 modulation in the anti-HIV-1 effects of toll-like receptor 3 stimulation in macrophages. PLoS Pathog. 2012;8(9):e1002937.
Natekar JP, Rothan HA, Arora K, Strate PG, Kumar M. Cellular microRNA-155 regulates virus-induced inflammatory response and protects against lethal West Nile virus infection. Viruses. 2020;12(1).
Liu S, Lin Y, Wang L, Liu T, Cheng C, Wei W, et al. Exonuclease III-aided autocatalytic DNA biosensing platform for immobilization-free and ultrasensitive electrochemical detection of nucleic acid and protein. Anal Chem. 2014;86(8):4008–15.
Hu T, Zhang L, Wen W, Zhang X, Wang S. Enzyme catalytic amplification of miRNA-155 detection with graphene quantum dot-based electrochemical biosensor. Biosens Bioelectron. 2016;77:451–6.
Zeng D, Wang Z, Meng Z, Wang P, San L, Wang W, et al. DNA tetrahedral nanostructure-based electrochemical miRNA biosensor for simultaneous detection of multiple miRNAs in pancreatic carcinoma. ACS Appl Mater Interfaces. 2017;9(28):24118–25.
Tran HV, Piro B, Reisberg S, Huy Nguyen L, Dung Nguyen T, Duc HT, et al. An electrochemical ELISA-like immunosensor for miRNAs detection based on screen-printed gold electrodes modified with reduced graphene oxide and carbon nanotubes. Biosens Bioelectron. 2014;62:25–30.
Zhang J, Hun X. Electrochemical determination of miRNA-155 using molybdenum carbide nanosheets and colloidal gold modified electrode coupled with mismatched catalytic hairpin assembly strategy. Microchem J. 2019;150:104095.
Duan F, Guo C, Hu M, Song Y, Wang M, He L, et al. Construction of the 0D/2D heterojunction of Ti3C2Tx MXene nanosheets and iron phthalocyanine quantum dots for the impedimetric aptasensing of microRNA-155. Sens. Actuators, B. 2020;310:127844.
Xu S, Chang Y, Wu Z, Li Y, Yuan R, Chai Y. One DNA circle capture probe with multiple target recognition domains for simultaneous electrochemical detection of miRNA-21 and miRNA-155. Biosens Bioelectron. 2020;149:111848.
Liu L, Zhu S, Wei Y, Liu X, Jiao S, Yang J. Ultrasensitive detection of miRNA-155 based on controlled fabrication of AuNPs@MoS2 nanostructures by atomic layer deposition. Biosens Bioelectron. 2019;144:111660.
Wang X, Jiang A, Hou T, Li F. A sensitive and versatile “signal-on” electrochemical aptasensor based on a triple-helix molecular switch. Analyst. 2014;139(23):6272–8.
Zhou L, Ou L-J, Chu X, Shen G-L, Yu R-Q. Aptamer-based rolling circle amplification: a platform for electrochemical detection of protein. Anal Chem. 2007;79(19):7492–500.
Liu A, Wang K, Weng S, Lei Y, Lin L, Chen W, et al. Development of electrochemical DNA biosensors. Trac-trend Anal Chem. 2012;37:101–11.
Xuan F, Luo X, Hsing IM. Ultrasensitive solution-phase electrochemical molecular beacon-based DNA detection with signal amplification by exonuclease III-assisted target recycling. Anal Chem. 2012;84(12):5216–20.
Luo X, Lee TM-H, Hsing IM. Immobilization-free sequence-specific electrochemical detection of DNA using ferrocene-labeled peptide nucleic acid. Anal Chem. 2008;80(19):7341–6.
Zhang Y, Xia J, Zhang F, Wang Z, Liu Q. A dual-channel homogeneous aptasensor combining colorimetric with electrochemical strategy for thrombin. Biosens Bioelectron. 2018;120:15–21.
Liu F, Yang L, Yin X, Liu X, Ge L, Li F. A facile homogeneous electrochemical biosensing strategy based on displacement reaction for intracellular and extracellular hydrogen peroxide detection. Biosens Bioelectron. 2019;141:111446.
Fu C, Liu C, Li Y, Guo Y, Luo F, Wang P, et al. Homogeneous electrochemical biosensor for melamine based on DNA triplex structure and exonuclease III-assisted recycling amplification. Anal Chem. 2016;88:10176–82.
Hou T, Li W, Liu X, Li F. Label-free and enzyme-free homogeneous electrochemical biosensing strategy based on hybridization chain reaction: a facile, sensitive, and highly specific MicroRNA assay. Anal Chem. 2015;87(22):11368–74.
Fu C, Liu C, Wang S, Luo F, Lin Z, Chen G. A signal-on homogeneous electrochemical biosensor for sequence-specific microRNA based on duplex-specific nuclease-assisted target recycling amplification. Anal Methods. 2016;8(39):7034–9.
Zhou W, Gong X, Xiang Y, Yuan R, Chai Y. Quadratic recycling amplification for label-free and sensitive visual detection of HIV DNA. Biosens Bioelectron. 2014;55:220–4.
Liu SF, Wang CF, Zhang CX, Wang Y, Tang B. Label-free and ultrasensitive electrochemical detection of nucleic acids based on autocatalytic and exonuclease III-assisted target recycling strategy. Anal Chem. 2013;85(4):2282–8.
Wang L-j, Ren M, Zhang Q, Tang B, C-y Z. Excision repair-initiated enzyme-assisted bicyclic cascade signal amplification for ultrasensitive detection of uracil-DNA glycosylase. Anal Chem. 2017;89(8):4488–94.
Xiong E, Yan X, Zhang X, Liu Y, Zhou J, Chen J. Exonuclease III-assisted cascade signal amplification strategy for label-free and ultrasensitive electrochemical detection of nucleic acids. Biosens Bioelectron. 2017;87:732–630.
Song X, Hou T, Lu F, Wang Y, Liu J, Li F. Homogeneous photoelectrochemical biosensing via synergy of G-quadruplex/hemin catalysed reactions and the inner filter effect. Chem Commun. 2020;56(12):1811–4.
Miao X, Cheng Z, Ma H, Li Z, Xue N, Wang P. Label-free platform for microRNA detection based on the fluorescence quenching of positively charged gold nanoparticles to silver nanoclusters. Anal Chem. 2018;90(2):1098–103.
Borghei YS, Hosseini M, Ganjali MR. Oxidase-like catalytic activity of Cys-AuNCs upon visible light irradiation and its application for visual miRNA detection. Sens. Actuators, B 2018;273:1618–1626.
Funding
This project was financially supported by the National Sciences Foundation of China (21775026, 21974020); the cooperative project of production and study in the University of Fujian Province (2018Y4007); and the Sciences Foundation of Fujian Province (2018J01685, 2018J01682).
Author information
Authors and Affiliations
Contributions
The manuscript was written through contributions of all authors. C.Lin and Z.Lin conceived the projects, and Y. Huang, X. Huang, and H.Z. designed and performed the experiments and collected the data. Y. Huang, X. Huang, H. Zheng, and Z. Lin analyzed and discussed the data. All authors discussed the results and contributed to the writing of this manuscript. All authors have given approval to the final version of the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Published in the topical collection Analytical Chemistry for Infectious Disease Detection and Prevention with guest editors Chaoyong Yang and XiuJun (James) Li.
Rights and permissions
About this article
Cite this article
Huang, Y., Huang, X., Zheng, H. et al. Homogeneous electrochemical biosensor for microRNA based on enzyme-driven cascaded signal amplification strategy. Anal Bioanal Chem 413, 4681–4688 (2021). https://doi.org/10.1007/s00216-020-03027-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-020-03027-3