Abstract
To establish a method for accurate quantitation of circulating cell-free mitochondrial DNA (ccf-mtDNA) in plasma by droplet digital PCR (ddPCR), we designed a ddPCR method to determine the copy number of ccf-mtDNA by amplifying mitochondrial ND1 (MT-ND1). To evaluate the sensitivity and specificity of the method, a recombinant pMD18-T plasmid containing MT-ND1 sequences and mtDNA-deleted (ρ0) HeLa cells were used, respectively. Subsequently, different plasma samples were prepared for ddPCR to evaluate the feasibility of detecting plasma ccf-mtDNA. In the results, the ddPCR method showed high sensitivity and specificity. When the DNA was extracted from plasma prior to ddPCR, the ccf-mtDNA copy number was higher than that measured without extraction. This difference was not due to a PCR inhibitor, such as EDTA-Na2, an anti-coagulant in plasma, because standard EDTA-Na2 concentration (5 mM) did not significantly inhibit ddPCR reactions. The difference might be attributable to plasma exosomal mtDNA, which was 4.21 ± 0.38 copies/μL of plasma, accounting for ∼19% of plasma ccf-mtDNA. Therefore, ddPCR can quickly and reliably detect ccf-mtDNA from plasma with a prior DNA extraction step, providing for a more accurate detection of ccf-mtDNA. The direct use of plasma as a template in ddPCR is suitable for the detection of exogenous cell-free nucleic acids within plasma, but not of nucleic acids that have a vesicle-associated form, such as exosomal mtDNA.

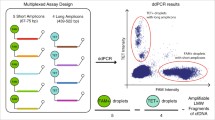
Designs of the present work. *: Module 1, #: Module 2, &: Module 3
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The existence of circulating cell-free DNA (ccfDNA) has been known for over 60 years [1]; however, the clinical significance of ccfDNA was not recognized until 1977 [2]. Thus far, ccfDNA has been investigated as a potential biomarker in noninvasive diagnosis and prognosis, as well as disease monitoring. It was suggested that the release of ccfDNA into the bloodstream occurs from different sources, such as hematopoietic and stromal cells in normal individuals, or tumor cells in cancer patients. In addition, active secretion of DNA carried within exosomes has been demonstrated [3], but it is still unclear whether this is a relevant or negligible source of ccfDNA.
CcfDNA includes nuclear DNA (nDNA) and mitochondrial DNA (mtDNA). Distinct from nDNA, mtDNA exists as a circular, double-stranded nucleic acid with a high copy number. Variations in the copy number of circulating cell-free mtDNA (ccf-mtDNA) have been found in the plasma and serum of patients with various cancers [4–7], trauma [8, 9], diabetes [10], and acute infection [11]. Therefore, assessing the level of ccf-mtDNA could prove significant for monitoring these diseases.
Despite its potential, there are two major hindrances in clinical application of ccf-mtDNA, which are in sample processing method and in the differences of both the DNA extraction methods and quantification technology. Those also are the main hindrances in the determination of ccfDNA [12]. Many extraction methods and commercial kits can be used in ccfDNA studies at present. However, it was demonstrated that the extraction methods for the isolation of ccfDNA from plasma or serum differed in the efficiency. In the quantification technology, to date, the copy number of mtDNA has been detected only by quantitative real-time PCR (qPCR) [13], which depends on the establishment of a standard curve derived from amplification of serially diluted, target gene-containing recombinant plasmid. Plasmid copy number depended on the Cq value and calculated based on the concentration of plasmid DNA; therefore, it can be affected by the accuracy of the concentration measurements or dilution of the plasmid. Moreover, the variability of qPCR is increased, leading to further imprecision and bias at low copy numbers. Therefore, the low copy number of ccf-mtDNA demands a greater quantity of plasma in the quantification by using qPCR.
Droplet digital PCR (ddPCR) is a form of digital PCR that uses a water–oil emulsion droplet system [14], which is capable of detecting rare mutation, and performing absolute quantification of gene copy number and gene expression analysis. With the advantages of high sensitivity, precision, and reproducibility, ddPCR has been applied to noninvasive prenatal testing (NIPT), cancer research, transplantation medicine, and virology [15]. DdPCR relies on an end-point threshold to score each reaction as either positive or negative. The absolute template quantity can then be determined by Poisson statistics. Therefore, for copy number determination, ddPCR is capable of quantitating the sample directly without using a standard curve, requiring less sample input. These features indicate that ddPCR may be suitable for detecting ccf-mtDNA in plasma.
In this study, we applied ddPCR technology to quantify plasma ccf-mtDNA copy number by amplifying human mitochondrial ND1 (MT-ND1). To evaluate the sensitivity and specificity of the method, a recombinant pMD18-T plasmid with MT-ND1 and DNA from mtDNA-deleted (ρ0) HeLa cells were used as the templates, respectively. Considering the clinical application, we attempted to utilize plasma directly, instead of the samples with DNA extraction on the basis of accurate detection of mtDNA by ddPCR. Therefore, we designed three different experimental modules to evaluate the accuracy and feasibility of utilizing plasma directly in ddPCR reactions, which was significant for clinical detection. Subsequently, we investigated the detection of plasma exosomal mtDNA with very low copy numbers, which also was a valuable attempt for the studies of exosomes and clinical application.
Materials and methods
Construction of recombinant plasmid
The MT-ND1 gene was amplified from human mitochondrial DNA by PCR using the following primers: forward 5′ CAGCCGCTATTAAAGGTTCG 3′, reverse 5′ AGAGTGCGTCATATGTTGTTC 3′, which were synthesized by Genscript Corporation (Nanjing, China). With these PCR primers, the fragment length of the amplified product was 1041 bp [nt3017-4057 in Homo sapiens mtDNA (GenBank No. NC_012920.1)].
The reaction volume of 20 μL contained 10 μL of 2 × PCR Buffer, 1.5 μL of 25 mM MgCl2, 2.0 μL of a dNTP mixture (25 mM each), 0.1 μL of Ex Taq (5 U/μL) (TaKaRa, Japan), 0.2 μL of each 10 μM primer, 4.0 μL of H2O, and 2.0 μL of DNA template. PCR was performed on a Bio-Rad S1000™ Thermal Cycler (Bio-Rad, USA). The PCR reaction conditions were as follows: 1 cycle at 95 °C for 5 min and 35 cycles at 95 °C for 30 s, 56 °C for 45 s, 72 °C for 1 min, and finally 1 cycle at 72 °C for 6 min. Subsequently, MT-ND1 was cloned into the pMD18-T vector by using a pMD™18-T Vector Cloning Kit (TaKaRa, Japan), and the ligation product was transformed into Escherichia coli DH5α.
Determination of plasmid copy number
Plasmids were extracted from E. coli DH5α with the MiniBEST Plasmid Purification Kit (TaKaRa, Japan), and the concentration was measured on a NanoDrop2000 (Thermo Scientific, USA). The plasmid copy number was calculated as follows:
Formula 1: MWDNA(daltons) = (pMD18 − T vector + insert) × 660 (daltons/bp) = (2692 bp + 1041 bp) × 660 (daltons/bp) = 2.46 × 106(daltons)
Formula 2: CNDNA(copies/μL) = CDNA(ng/μL) × 6.02 × 1014/MWDNA = CDNA(ng/μL) × 6.02 × 1014/2.46 × 106 = CDNA(ng/μL) × 2.45 × 108
- (CNDNA :
-
Plasmid copy number
- CDNA :
-
Plasmid DNA concentration
- MWDNA :
-
Plasmid molecular weight)
Then, the plasmids were serially diluted to 104, 103, 102, 10, and 1 copies/μL as the templates.
Quantification of mtDNA copy number by qPCR
The copy number of mtDNA was quantified using qPCR by amplification of the MT-ND1 gene using the following primers: forward 5′ CCCTAAAACCCGCCACATCT 3′, reverse 5′ GAGCGATGGTGAGAGCTAAGGT 3′; fragment length was 69 bp (Homo sapiens mtDNA nt3485-3553). The fluorescent probe used was 5′ (FAM) CCATCACCCTCTACATCACCGCCC (TAMRA) 3′. The probe and primers were synthesized by Genscript Corporation (Nanjing, China).
The qPCR reaction volume of 20 μL contained 10 μL of 2 × HS qPCR Master Mix (TaKaRa, Japan), 0.5 μL of each 10 μM primer, 0.5 μL of the 10 μM probe, 6.5 μL of H2O, and 2.0 μL of DNA template, which was 1 to 104 copies/μL of the recombinant pMD18-T plasmid with the MT-ND1 insert.
PCR was performed on a Bio-Rad CFX96 Touch™ Real-Time PCR Detection System (Bio-Rad, USA). The reaction conditions were as follows: 1 cycle at 95 °C for 2 min, 40 cycles at 95 °C for 10 s, and 60 °C for 1 min.
ddPCR
The primers and probe utilized in the ddPCR analysis were listed previously, though the quencher was changed to BHQ1. The ddPCR reaction mixture volume of 20 μL contained 10 μL of 2 × Supermix (no dUTP), 1.8 μL of each 10 μM primer, 1 μL of the 10 μM probe, 2 μL of DNA template and/or plasma, and 3.4 μL of H2O. 1 to 104 copies/μL of recombinant plasmids were used for testing the sensitivity. Specificity of the mtDNA assay was determined by using DNA from the ρ0 HeLa cells that lacks mtDNA. As a negative control, 2 μL of H2O was added separately to the ddPCR reaction instead of template DNA.
The PCR reaction conditions were as follows: 1 cycle at 95 °C for 10 min, 40 cycles at 95 °C for 30 s, 60 °C for 1 min, and 98 °C for 10 min. Droplet reading was performed on a Bio-Rad QX200 droplet digital PCR system (Bio-Rad, USA), and analysis was performed with Bio-Rad QuantaSoft software version 1.3.2.
Subjects
To apply the ddPCR method on plasma samples, 38 healthy volunteers, 20 males and 18 females, aged from 20–30 years, were recruited from the Medical Center in the First Affiliated Hospital of Wenzhou Medical University of Zhejiang Province in China. The study was approved by the hospital’s Ethics Committee, and methodologies conformed to the standards set by the Declaration of Helsinki.
Plasma cell-free DNA extraction
Blood samples were collected from the healthy volunteers into EDTA-Na2-containing tubes, and the plasma was separated. Plasma DNA was extracted from 250 μL of plasma using a Plasma Cell-Free DNA Isolation Kit (Maibo Biotech, China), which uses a magnetic bead method.
Briefly, 250 μL of plasma was incubated with 20 μL of 20 mg/mL Proteinase K at 37 °C for 15 min; then, 300 μL of lysis buffer, 5 μL of acryl carrier, and 20 μL of magnetic beads were added, followed by mixing for 30 min. The tube was then loaded on the magnetic separator (Sigma, USA) for 1 min to capture the beads, and the supernatant was discarded. Next, 550 μL of washing buffer was used to wash the beads three times. Finally, the DNA was eluted from the beads into 50 μL of elution buffer.
Evaluation of plasma ccf-mtDNA detection by ddPCR
To evaluate the potential of utilizing plasma directly as the source of ddPCR templates, plasma preconditioning was performed first. Briefly, the plasma samples from 10 subjects were centrifuged at 1600g for 10 min; then, the supernatant from each sample was divided into two aliquots. One of the aliquots was centrifuged at 16,000g for 10 min (method 1), and the other was ultrafiltered through a 0.22-μm filter (Millipore, USA, method 2).
Further, to confirm the accuracy of the results obtained when plasma was utilized as the template source in ddPCR reactions, we set three experimental modules.
-
Module 1:
EDTA-Na2 is a commonly used anticoagulant, and it is also an inhibitor of PCR. In the clinical lab, the standard concentration of EDTA-Na2 as an anticoagulant in plasma is 5 mM. Therefore, to evaluate the effects of EDTA-Na2 on ddPCR, 50, 100, 150, 200, and 250 mM EDTA-Na2 was added separately into recombinant plasmid samples (100 copies/μL) at a ratio of 1:9 and diluted to a final EDTA-Na2 concentration of 5, 10, 15, 20, or 25 mM.
-
Module 2:
To evaluate the effects of plasma samples on ddPCR, known copy numbers of plasmid, 12.41, 83.45, 318.15, or 1050.20 copies/μL (Log10: 1.09, 1.92, 2.50, 3.02 copies/μL), as determined previously by ddPCR, were added separately into plasma samples from 10 subjects at a ratio of 1:1, as group 1, 2, 3, and 4, respectively. Then, these four groups of plasma samples and the original plasma samples were detected simultaneously by ddPCR. The different copy number values were calculated according to Formula 3 and compared with that of the added plasmid.
Formula 3: Log [Calculated CNmtDNA] = Log [CNmtDNAin (plasma + added plasmid) − CNmtDNAin (plasma + H2O)]
- CNmtDNA :
-
mtDNA copy number
-
Module 3:
To evaluate the feasibility of plasma as the template in ddPCR, plasma samples from the same subjects, with or without DNA extraction, were analyzed by ddPCR and compared.
Exosomal isolation, identification, and DNA extraction
Plasma samples from 18 healthy volunteers were collected and preconditioned using method 1 (centrifugation at 1600g for 10 min and 16,000g for an additional 10 min) described above and then ultrafiltered with a 0.22-μm filter (Millipore, USA). Subsequently, 250 μL of pretreated plasma was incubated with 2.5 μL of 500 U/mL thrombin (SBI, USA) at 37 °C for 5 min and then centrifuged at 10,000g for 5 min. Next, ExoQuick exosome precipitation solution (SBI, USA) was added at a ratio of 4:1 and incubated overnight at 4 °C, after which, the solution was centrifuged at 1500g for 30 min, and the supernatant was discarded. The precipitate was centrifuged again for 5 min, and any remaining supernatant was removed. To obtain pure exosomes, the precipitate was resuspended in 200 μL of PBS. Subsequently, 50 μL of ExoQuick exosome precipitation solution was added to isolate exosomes a second time, with final resuspension in 250 μL PBS.
To identify the exosomal fraction, exosomes were observed with a transmission electron microscope (TEM) by negative staining. Exosome-specific markers Tsg101 and CD9 were analyzed by Western blotting.
Before the extraction of exosomal DNA, one unit of DNase I was added to the isolated exosomal precipitates resuspended in PBS, and the reaction mixtures were incubated at 37 °C for 1 h to digest extra-exosomal DNA. Then, DNA was extracted with the Plasma Cell-Free DNA Extraction kit by the magnetic bead method as described before. Finally, DNA was eluted into 50 μL elution buffer.
Statistical analysis
Statistical analysis was performed by SPSS 17.0. All copy numbers were converted to Log10 (copy numbers) per microliter of samples and expressed as the mean with standard deviation. Linear regression was applied to analyze the relationship between the qPCR and ddPCR results and between the results for plasma, with or without prior DNA extraction. The Mann–Whitney rank test was used for data with an abnormal distribution; a P value of < 0.05 was considered statistically significant.
Results
Copy number determination of recombinant plasmids by qPCR and ddPCR
The MT-ND1 gene (1041 bp) was amplified by PCR (Fig. 1a), cloned into the pMD18-T vector, and verified by sequencing (Fig. 1b).
Verification of the recombinant plasmid with the MT-ND1 gene insert. A Agarose gel electrophoresis of MT-ND1 PCR products, lines 1–3 are the PCR products from three different E. coli colonies separately. B Partial sequence of the MT-ND1 insert DNA, np3485-3553 (69 bp), which corresponds to the primers and probe sequence for detecting mtDNA copy number by qPCR and ddPCR
By utilizing specific primers and a probe (as shown in Fig. 1b), qPCR and ddPCR were performed to detect the MT-ND1 sequences of the recombinant plasmid from 1 to 104 copies/μL, as shown in Fig. 2a–c, and a linear correlation was seen between the two methods (Fig. 2d). However, when the plasmid copy number was 10 and 1 copies/μL, the Cq value was as much as 31.4 and 34.4, respectively (Fig. 2b). This discrepancy indicated that, at low copy number, qPCR did not detect the plasmid accurately, but ddPCR did detect it quantitatively (Fig. 2c). Additionally, DNA from ρ0 HeLa cells was not amplified, confirming the specificity of the ddPCR detection method (Fig. 2e).
Quantification of the mtDNA copy number by qPCR and ddPCR. A, B Amplification and standard curves of the recombinant plasmids by qPCR. C 1-D plots of the ddPCR assays with recombinant plasmids as templates. NTC: negative control. Blue dots represent positive droplets containing a DNA template, black dots represent negative droplets without a DNA template, and the pink line is the threshold line. D Linear correlation between the mtDNA copy numbers by qPCR and ddPCR. E 1-D plots of ddPCR assays for the negative control, DNA from ρ0 cell line, and extracted plasma DNA
Preconditioned plasma mtDNA copy number detection by ddPCR
To determine the effect of different preconditioning methods on the analysis, we compared mtDNA copy number detection from plasma after two preconditioning methods. There was no significant difference between the two methods in detecting mtDNA copy number from 50 μL plasma DNA. With method 1, there were 157.50 ± 24.16 (Log10: 2.17 ± 0.06) copies/μL, and with method 2, we detected 168.83 ± 31.95 (Log10: 2.19 ± 0.09) mtDNA copies/μL (Fig. 3a).
mtDNA copy number of plasma samples and extracted plasma DNA detected by ddPCR. A mtDNA copy number of plasma with two preconditioning methods. B mtDNA copy number of plasma samples with or without DNA extraction from the same subjects (n = 10). C 1-D plot of plasmids with different concentration of EDTA-Na2. D mtDNA copy number of added plasmid, experimental and calculated plasmid values for the plasma sample
Detection of plasma ccf-mtDNA copy number by ddPCR
In the module 1 experiments, plasmid samples with different final concentrations (5, 10, 15, 20, and 25 mM) of EDTA-Na2 were used to evaluate the potential inhibitory effect of EDTA-Na2 on ddPCR. The results indicated that EDTA-Na2 did not inhibit amplification by ddPCR until the concentration reached 20 mM or higher, well over the standard of 5 mM (Fig. 3c). In the experiments of module 2, the initial and calculated plasmid copy number difference was similar in all groups except group 1, which had a low plasmid copy number (Log10: 1.090 copies/μL) (Fig. 3d). The results from module 3 experiments indicated that the amount of mtDNA was higher in plasma samples with extraction than those without extraction (Fig. 3b).
Detection of plasma exosomal mtDNA by ddPCR
After purification of exosomes, we observed 30–100 nm diameter microvesicles by TEM (Fig. 4a). Additionally, the exosome-specific markers Tsg101 and CD9 were detected by Western blotting, as shown in Fig. 4b.
Plasma ccf-mtDNA and exosomal mtDNA detected by ddPCR. A Plasma exosomes observed by TEM. B exosome-specific marker, Tsg101, CD63, and CD9 analyzed by Western blotting; lanes 1–4 represent plasma exosomes from four healthy volunteers. C 1-D plot of extracted plasma ccf-mtDNA and exosomal mtDNA. P: plasma, F: free mtDNA, E: exosomal mtDNA
In these validated exosome samples from 18 subjects, the mean copy number of exosomal mtDNA was 4.21 ± 0.38 (Log10: 0.59 ± 0.04) copies/μL plasma. For determining the proportion of exosomal mtDNA in plasma ccf-mtDNA, the plasma ccf-mtDNA copy number in the samples was detected simultaneously (Fig. 4c). The results demonstrated that the mean copy number of plasma ccf-mtDNA was 23.52 ± 2.14 (Log10: 1.34 ± 0.04) copies/μL, and the proportion of exosomal mtDNA in plasma ccf-mtDNA was 0.193 ± 0.018.
Discussion
To date, the use of ddPCR in mtDNA analysis has predominantly focused on the detection of deletions and rare heteroplasmic mutations [16, 17], with few studies focused on ccf-mtDNA levels [18]. Compared with the detection of mutations, ccf-mtDNA copy number quantification showed easier in technology. Moreover, ccf-mtDNA has higher copy numbers than nuclear genes in the cells, which means the mild variations of the mtDNA copy number may denote the changes in the body efficiently.
However, the quantitative determination process of ccf-mtDNA still lacks more convenient and precise methods in DNA extraction and detection. QPCR-based absolute quantification is the most commonly used in ccf-mtDNA quantitative method, which needs the establishment of a standard curve as well as the accuracy and stability of qPCR decreased in the detection of low copy numbers. Compared with traditional qPCR, ddPCR is more convenient, allowing direct quantitation without a standard curve, therefore alleviating the need for reference material required. ddPCR as well demonstrated high sensitivity and precision even in analyzing the trace nuclear acid which is very significant for clinical analysis.
Considering these advantages of ddPCR, in this study, we established a ddPCR method to detect ccf-mtDNA copy number directly from plasma and showed that ddPCR could even detect exosomal mtDNA in very low copy numbers, relevant to disease monitoring and prognosis.
To evaluate the validity of the ddPCR method, the sensitivity and specificity were tested first in our present study. A total of 1∼104 copies/μL of MT-ND1-pMD18 recombinant plasmids were detected by qPCR and ddPCR separately for testing the sensitivity. The results showed that ddPCR was capable of detecting 1 copy/μL of template, beyond the threshold of accurate detection by qPCR (Fig. 2a–d) [19], demonstrating high sensitivity and accuracy for the ddPCR method of detecting mtDNA. Specificity was proved by the data that DNA from the ρ0 cells was undetectable by ddPCR (Fig. 2e). Therefore, in further experiments, we attempted to detect plasma ccf-mtDNA from healthy volunteers by using the ddPCR method.
Plasma is a complex biological fluid containing a variety of proteins, ions, nutrients, hormones, and so on, at least some of which can inhibit amplification by PCR. Thus, plasma DNA is extracted routinely prior to analysis by PCR. However, considering the high inhibitor tolerance of ddPCR [20], we performed ddPCR utilizing plasma without DNA extraction, directly as a source of template. To remove residual cells or large vesicles that might affect the formation of 250-μm-diameter droplets in ddPCR, plasma was preconditioned first through centrifugation or by ultrafiltration (method 1 or method 2). The results indicated that there were no significant differences between the data of the two methods (Fig. 3a). Though DNA extraction process might lead the loss of DNA, our data showed that the mtDNA copy numbers obtained by using plasma directly in the ddPCR reaction were lower than the copy numbers obtained by using extracted plasma DNA (Fig. 3b), suggesting factors in the plasma samples might affect the results, and the common PCR inhibitors or other underlying factors were considered first.
EDTA-Na2 is commonly used as an anticoagulant in clinical labs. However, it can also act as a PCR inhibitor by virtue of its ability to chelate Mg2+ ions, and it is possible that residual EDTA-Na2 may be present in plasma samples. Dingle et al. [20] showed that ddPCR was more tolerant to inhibition caused by SDS and heparin than qPCR. To evaluate the effect of EDTA-Na2 on ddPCR, the MT-ND1-pMD18 plasmids were quantitated in the presence of various concentrations of EDTA-Na2 in our study. The data showed that DNA amplification by ddPCR was not inhibited below 20 mM of EDTA-Na2 in the detection of 100 copies/μL of plasmid (Fig. 3c). This is higher than the concentration that is typically used clinically, suggesting that ddPCR can tolerate the conventional dose of EDTA-Na2 without affecting DNA copy number quantification. The data also revealed that this PCR inhibitor was not the main factor leading to the difference in detection of mtDNA from plasma with and without DNA extraction.
Further, in the experiments of Module 2, the MT-ND1-pMD18 plasmids at a known copy number were added to plasma samples. From the collected data, the calculated copy number difference (according to Formula 3) was consistent with the copy number of the added plasmid (Fig. 3d), which indicated that the composition of plasma has little effect on the linearity of the ddPCR reactions. On the other point of view, it suggested that ddPCR might be allowed to detect exogenous nucleic acids in plasma, such as microbial DNA. It is significant to monitor infectious diseases in a timely and cost-effective manner because DNA extraction is not required.
Considering that plasma ccfDNA might be found in free form and a vesicle-associated form [21], the vesicle-associated mtDNA in plasma was evaluated next in our study. Exosomes [22–24] are one type of extracellular vesicles, 30–100 nm in diameter, assembled in the cytoplasm, and released into the extracellular environment upon fusion with the plasma membrane [25]. Exosomes contain proteins, RNA, and DNA, including mtDNA [26, 27], which could contribute to the overall level of ccf-mtDNA in plasma. The copy number of exosomal mtDNA was considered previously to be very low [28]. In our study, it was possible to detect low copy number mtDNA by ddPCR, indicating that ddPCR has the advantage over conventional qPCR in detecting trace amounts of DNA because of its high sensitivity. Our results demonstrated that the proportion of exosomal mtDNA in plasma ccf-mtDNA was 0.193 ± 0.018, and thus, it was an important component of ccf-mtDNA, more prevalent than described in a previous study [28]. Reasons for this difference might be the application of different protocols for isolating exosomes [29] and/or the use of different methods for detecting mtDNA.
Consequently, when using plasma samples directly without a DNA extraction step, mtDNA may not be released from exosomes completely at the denaturing temperature used in PCR reaction (such as 95 °C) and hence will be undetectable in ddPCR. Likewise, this could partly explain the higher level of mtDNA in plasma with prior DNA extraction compared to plasma without prior extraction.
In our study, we considered several common potential impacts and evaluate the effectiveness of the ddPCR method comprehensively, which may be valuable in future clinical application. Due to its high sensitivity, ddPCR method may be especially applicable on analyzing rapid increase of ccf-mtDNA and monitoring the disease, such as the early stage of transplantation [30], or some chronic disease, like diabetes, cancers, and aging, which demonstrated higher level of ccf-mtDNA, relevant to the chronic inflammation and development of the diseases [10, 31]. Ccf-mtDNA results from cellular apoptosis and/or necrosis [32]. Given that severe mitochondrial damage can lead to cell death by necrosis or activation of the apoptotic signaling pathway, increased ccf-mtDNA might be associated with mitochondrial dysfunction or oxidative stress. Therefore, the quantitative assessment of ccf-mtDNA by ddPCR may also allow sensitive evaluation of mitochondrial damage. Additionally, the clinical value of detecting exosomal mtDNA should be taken into account. Increased plasma exosomal mtDNA may be correlated with the release of exosomes, which has been observed under some conditions, such as oxidative stress and myocardial infarction [33, 34].
Conclusion
In summary, ddPCR can give the initial copy number directly, independent on the Cq value, and without establishment of standard curve. Therefore, ddPCR was proven to be simpler and more sensitive than qPCR and suitable for quantitatively detecting plasma ccf-mtDNA, even plasma exosomal mtDNA, which is present in low quantity. In addition, ddPCR was not affected by inhibitors, such as EDTA-Na2. Plasma was not suitable for use directly in ddPCR for the detection of nucleic acids with a vesicle-associated form, such as exosomal mtDNA. As such, it was more feasible to perform a DNA extraction on plasma and then use the extracted DNA in the ddPCR reaction. However, plasma without extraction could be used directly in ddPCR for detecting exogenous nucleic acids.
Abbreviations
- ccfDNA:
-
Circulating cell-free DNA
- ccf-mtDNA:
-
Circulating cell-free mitochondrial DNA
- ddPCR:
-
Droplet digital PCR
- mtDNA:
-
Mitochondrial DNA
- MT-ND1 :
-
Mitochondrial ND1 gene
- NIPT:
-
Noninvasive prenatal testing
References
Mandel P, Metais P. C R Seances Soc Biol Fil. 1948;142(3–4):241–3. Les acides nucleiques du plasma sanguin chez l’homme.
Leon SA, Shapiro B, Sklaroff DM, Yaros MJ. Free DNA in the serum of cancer patients and the effect of therapy. Cancer Res. 1977;37(3):646–50.
Thakur BK, Zhang H, Becker A, Matei I, Huang Y, Costa-Silva B, et al. Double-stranded DNA in exosomes: a novel biomarker in cancer detection. Cell Res. 2014;24(6):766–9.
Yu M. Circulating cell-free mitochondrial DNA as a novel cancer biomarker: opportunities and challenges. Mitochondrial DNA. 2012;23(5):329–32.
Huang CY, Chen YM, Wu CH, Tsai CM, Lee YC, Perng RP, et al. Circulating free mitochondrial DNA concentration and its association with erlotinib treatment in patients with adenocarcinoma of the lung. Oncol Lett. 2014;7(6):2180–4.
Ellinger J, Muller DC, Muller SC, Hauser S, Heukamp LC, von Ruecker A, et al. Circulating mitochondrial DNA in serum: a universal diagnostic biomarker for patients with urological malignancies. Urol Oncol. 2012;30(4):509–15.
Fernandes J, Michel V, Camorlinga-Ponce M, Gomez A, Maldonado C, De Reuse H, et al. Circulating mitochondrial DNA level, a noninvasive biomarker for the early detection of gastric cancer. Cancer Epidemiol Biomarkers Prev. 2014;23(11):2430–8.
McIlroy DJ, Bigland M, White AE, Hardy BM, Lott N, Smith DW, et al. Cell necrosis-independent sustained mitochondrial and nuclear DNA release following trauma surgery. J Trauma Acute Care Surg. 2015;78(2):282.
Zhang Q, Raoof M, Chen Y, Sumi Y, Sursal T, Junger W, et al. Circulating mitochondrial DAMPs cause inflammatory responses to injury. Nature. 2010;464(7285):104–7.
Lee HK, Song JH, Shin CS, Park DJ, Park KS, Lee KU, et al. Decreased mitochondrial DNA content in peripheral blood precedes the development of non-insulin-dependent diabetes mellitus. Diabetes Res Clin Pract. 1998;42(3):161–7.
Nakahira K, Kyung SY, Rogers AJ, Gazourian L, Youn S, Massaro AF, et al. Circulating mitochondrial DNA in patients in the ICU as a marker of mortality: derivation and validation. PLoS Med. 2013;10(12):e1001577. discussion e.
Devonshire AS, Whale AS, Gutteridge A, Jones G, Cowen S, Foy CA, et al. Towards standardisation of cell-free DNA measurement in plasma: controls for extraction efficiency, fragment size bias and quantification. Anal Bioanal Chem. 2014;406(26):6499–512.
Ajaz S, Czajka A, Malik A. Accurate measurement of circulating mitochondrial DNA content from human blood samples using real-time quantitative PCR. Methods Mol Biol. 2015;1264:117–31.
Manoj P. Droplet digital PCR technology promises new applications and research areas. Mitochondrial DNA. 2014.
Hudecova I. Digital PCR, analysis of circulating nucleic acids. Clin Biochem. 2015.
Rebolledo-Jaramillo B, Su MS, Stoler N, McElhoe JA, Dickins B, Blankenberg D, et al. Maternal age effect and severe germ-line bottleneck in the inheritance of human mitochondrial DNA. Proc Natl Acad Sci U S A. 2014;111(43):15474–9.
Taylor SD, Ericson NG, Burton JN, Prolla TA, Silber JR, Shendure J, et al. Targeted enrichment and high-resolution digital profiling of mitochondrial DNA deletions in human brain. Aging Cell. 2014;13(1):29–38.
Podlesniy P, Figueiro-Silva J, Llado A, Antonell A, Sanchez-Valle R, Alcolea D, et al. Low cerebrospinal fluid concentration of mitochondrial DNA in preclinical Alzheimer disease. Ann Neurol. 2013;74(5):655–68.
Sedlak RH, Cook L, Cheng A, Magaret A, Jerome KR. Clinical utility of droplet digital PCR for human cytomegalovirus. J Clin Microbiol. 2014;52(8):2844–8.
Dingle TC, Sedlak RH, Cook L, Jerome KR. Tolerance of droplet-digital PCR vs real-time quantitative PCR to inhibitory substances. Clin Chem. 2013;59(11):1670–2.
Chiu RW, Chan LY, Lam NY, Tsui NB, Ng EK, Rainer TH, et al. Quantitative analysis of circulating mitochondrial DNA in plasma. Clin Chem. 2003;49(5):719–26.
Pan BT, Teng K, Wu C, Adam M, Johnstone RM. Electron microscopic evidence for externalization of the transferrin receptor in vesicular form in sheep reticulocytes. J Cell Biol. 1985;101(3):942–8.
Thery C, Zitvogel L, Amigorena S. Exosomes: composition, biogenesis and function. Nat Rev Immunol. 2002;2(8):569–79.
Keller S, Sanderson MP, Stoeck A, Altevogt P. Exosomes: from biogenesis and secretion to biological function. Immunol Lett. 2006;107(2):102–8.
Loyer X, Vion AC, Tedgui A, Boulanger CM. Microvesicles as cell-cell messengers in cardiovascular diseases. Circ Res. 2014;114(2):345–53.
Guescini M, Genedani S, Stocchi V, Agnati LF. Astrocytes and Glioblastoma cells release exosomes carrying mtDNA. J Neural Transm. 2010;117(1):1–4.
Guescini M, Guidolin D, Vallorani L, Casadei L, Gioacchini AM, Tibollo P, et al. C2C12 myoblasts release micro-vesicles containing mtDNA and proteins involved in signal transduction. Exp Cell Res. 2010;316(12):1977–84.
Helmig S, Fruhbeis C, Kramer-Albers EM, Simon P, Tug S. Release of bulk cell free DNA during physical exercise occurs independent of extracellular vesicles. Eur J Appl Physiol. 2015;115(11):2271–80.
Lobb RJ, Becker M, Wen SW, Wong CS, Wiegmans AP, Leimgruber A, et al. Optimized exosome isolation protocol for cell culture supernatant and human plasma. J Extracell Vesicles. 2015;4:27031.
Beck J, Bierau S, Balzer S, Andag R, Kanzow P, Schmitz J, et al. Digital droplet PCR for rapid quantification of donor DNA in the circulation of transplant recipients as a potential universal biomarker of graft injury. Clin Chem. 2013;59(12):1732–41.
Pinti M, Cevenini E, Nasi M, De Biasi S, Salvioli S, Monti D, et al. Circulating mitochondrial DNA increases with age and is a familiar trait: Implications for “inflamm-aging”. Eur J Immunol. 2014;44(5):1552–62.
Jahr S, Hentze H, Englisch S, Hardt D, Fackelmayer FO, Hesch RD, et al. DNA fragments in the blood plasma of cancer patients: quantitations and evidence for their origin from apoptotic and necrotic cells. Cancer Res. 2001;61(4):1659–65.
Zhou Y, Xu H, Xu W, Wang B, Wu H, Tao Y, et al. Exosomes released by human umbilical cord mesenchymal stem cells protect against cisplatin-induced renal oxidative stress and apoptosis in vivo and in vitro. Stem Cell Res Ther. 2013;4(2):34.
Sahoo S, Losordo DW. Exosomes and cardiac repair after myocardial infarction. Circ Res. 2014;114(2):333–44.
Acknowledgments
This work is supported by the National Key Basic Research Program of China (2013CB531200), the National Nature Science Fund of China (81271918), and Specialized Research Fund for the Doctoral Program of Higher Education (20133321110001).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
We state that all blood samples were collected from the individual participants, who were informed and voluntarily participated in this research work.
Conflict of interest
All authors declare that they have no conflicts of interest to this work.
Additional information
Wei Ye and Xiaojun Tang contributed equally to this work.
Rights and permissions
About this article
Cite this article
Ye, W., Tang, X., Liu, C. et al. Accurate quantitation of circulating cell-free mitochondrial DNA in plasma by droplet digital PCR. Anal Bioanal Chem 409, 2727–2735 (2017). https://doi.org/10.1007/s00216-017-0217-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-017-0217-x