Abstract
The theory and practice of proton microspeciation based on NMR–pH titrations are surveyed. Principles of bi-, tri-, tetra-, and n-protic microequilibrium systems are discussed. Evaluation methods are exemplified by case studies on bi- and tetraprotic biomolecules. Selection criteria and properties of ‘reporter’ NMR nuclei are described. Literature data on complete microspeciations of small ligands and site-specific basicity characterizations of peptides and proteins are critically reviewed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Microscopic acidity/basicity constants and microspecies concentrations are corresponding physicochemical and analytical terms. Information on either one assumes the knowledge of the other. Their determination is a subtle speciation task (also called microspeciation) in analytical chemistry, with special significance in biological systems. In fact, complementarity of the reactants in highly specific biochemical reactions is achieved via interactions of particular microspecies of the participating biomolecules.
Today, the chief methodology of microspeciation is NMR–pH titration, the process in which concentration changes of microspecies are transformed into NMR parameters. In particular, the chemical shift of NMR-active nuclei is governed by several factors such as the inductive effect of constitutionally proximate substituent groups, electric field effects or magnetic anisotropy of sterically neighboring moieties, solvation effects, and last but not least the protonation state of acidic/basic sites [1, 2].
The basics of NMR–pH titrations can be illustrated by the example of a monobasic ligand (L). Since protonation decreases the local electron density, a selected nucleus in the vicinity of the protonating site senses different electronic environments and thus it exhibits different chemical shifts in the neutral (δ L) and protonated (\(\delta _{{{\text{HL}}^{ + } }} \)) states of the ligand. Protonation of “small molecules” in aqueous solution takes place usually with rate constants near to the diffusion limit (interesting exceptions can be found in refs. [3, 4, 5]), therefore these reactions are instantaneous on the NMR time scale. In the fast-exchange regime, a common resonance of two species can be observed at the weighted average of δ L and \(\delta _{{{\text{HL}}^{ + } }} \), the limiting chemical shifts [6, 7, 8]:
The weighting factors, x L and \(x_{{{\text{HL}}^{ + } }} \) are the pH-dependent mole fractions of L and HL+, respectively. They can be expressed in terms of the actual hydrogen ion concentration and the protonation constant (K=[HL+]/[H+][L]) or acid dissociation constant (K a=[H+][L]/[HL+]) of the ligand,
In the present review, ionization equilibria of acids and bases will be uniformly characterized in the direction of proton binding. Equivalent formalisms based on dissociation constants along with the necessary interconversion formulae can be found, for example, in ref. [9].
In the monoprotic case, the δ obsd=f(pH) function described by Eqs. (1) and (2) has the familiar sigmoid shape, with an inflection point at pH=logK. From the beginning of the 1960s, NMR titrations have been extensively used to measure protonation constants.
In the case of polyprotic molecules with well-separated protonation steps (ΔlogK=logK i−logK i+1>3), Eqs. (1) and (2) apply separately for each protonation stage. If, however, protonation of two or more groups takes place in overlapping pH intervals, the logK i values describe the equilibria at the macroscopic level, showing only the stoichiometry, not the site of protonation:
The totally free and fully protonated ligands are species that do not exhibit protonation isomerism (i.e., none and all of the binding sites are occupied by proton(s), respectively). In contrast, the “intermediate” macrospecies are actually mixtures of microspecies that hold identical number of protons but differ in the site of protonation [10]. Microspecies (in other words, protonation isomers) are in continuous interconversion, they cannot be separated analytically and their intensive spectral characteristics can be determined indirectly only. Microscopic protonation constants (microconstants) characterize the group-specific basicity at defined ionization states of all other groups [10, 11, 12, 13]. Microconstants are highly detailed parameters to quantitate equilibria at the submolecular level, which are surpassed only by their rotamer-specific counterparts [13, 14], which are discussed in a separate paper in this issue of Analytical and Bioanalytical Chemistry. Other fundamental details on microscopic protonation equilibria are discussed in several publications [9, 10, 11, 12, 13, 15, 16, 17, 18, 19, 20].
The experimental determination of microconstants relies upon the principle that protonation of each (or “each minus one”, see later) basic center is followed selectively [15]. Before the advent of NMR spectroscopy, mainly UV spectroscopy was applied to monitor the proton binding of chromophore groups such as thiolate and phenolate through absorbance changes. Application of non-NMR methods to microconstant determination has been summarized in recent reviews [13, 20].
Since NMR-active nuclei such as 1H or 13C are ubiquitous in organic ligands, NMR spectroscopy is almost universally applicable. NMR titration exploits the fact that the chemical shift of nuclei adjacent to a basic center changes with its fractional protonation and, in favorable cases, is virtually independent of the degree of protonation of more distant basic sites [21, 22]. Due to the excellent resolution and selectivity of modern, in part multidimensional NMR methods, NMR–pH titrations have become the most powerful tool to probe site-specific acid–base properties [13, 20, 23].
The past few years have provided evidence of considerable progress in the mathematical evaluation of NMR–pH titration curves [17, 19, 20, 24, 25], paving the way to profound microequilibrium analysis for systems up to dendrimers [17, 19, 20, 24, 25, 26, 27, 28]. The present review aims to give an overview of the current state of the NMR titration methodology and microconstant calculation strategies, along with a survey of its successful applications to both “small molecules” and proteins.
Experimental aspects of NMR–pH titrations
The most important medium of acid–base equilibria is water, and the vast majority of NMR–pH titrations have therefore been carried out in aqueous solutions. To avoid an overwhelming H2O peak and the concomitant dynamic range problems in 1H NMR spectroscopy, the use of deuterium oxide as solvent has long been the only reasonable solution with all its known drawbacks (e.g., losing resonances of exchanging amide protons in proteins). Commercial glass electrodes function properly in D2O [29, 30], but when calibrated with H2O-based buffers, a correction of 0.40 has to be added to pH meter readings to get pD values [29]. Protonation constants in D2O are typically 0.3–0.7 logarithmic units larger than the corresponding \(\log K^{{{\text{H}}_{{\text{2}}} {\text{O}}}} \) values due to several reasons. Intrinsically, the zero-point energies of the species DL+ and HL+ are different and the same applies to D3O+ and H3O+ [31]. Also, pD and pH scales fixed by NIST primary standard buffer solutions do not exactly match either [32]. Thus, several empirical correlations have been set up to convert H2O and D2O-based protonation constants [33, 34]. Although the precision of \(\log K^{{{\text{D}}_{{\text{2}}} {\text{O}}}} \) predictions is claimed to be commensurable with the magnitude of experimental errors [34], the most precise way still remains the experimental redetermination of protonation constants for any new solvent (mixture) of interest. To eliminate the ambiguities due to solvent isotope effects, today it is possible to conduct NMR–pH titrations in H2O, which means, however, that field–frequency lock cannot be applied on spectrometers with standard configuration. Combining advantages of light and heavy water, the solvent mixture H2O/D2O (90/10 v/v) represents a widely accepted compromise. A multitude of water suppression methods have been developed to reduce the solvent resonance peak [35, 36, 37, 38]. At any choice of solvent, details regarding solvent composition, electrode calibration, and the pH scale should always be precisely reported; hence comparisons are only meaningful when protonation constants are based on a common pH (or pD) scale.
Another complication of 1H NMR titrations, especially for larger biopolymers, is the significant overlap of resonance multiplets. In these cases, a pH-dependent series of appropriate two-dimensional (2D) spectra should be acquired to obtain the necessary chemical shift versus pH titration profiles. Examples have been reported for pH-dependent HOHAHA [39], NOESY [40], DQF–COSY [41], 13C–1H HSQC [42], 15N–1H HSQC [43], and TOCSY [44, 45] spectral series.
Ligand concentration is also an important issue. As compared to electronic and vibrational spectroscopies, NMR spectroscopy is an insensitive technique, despite the continuous efforts to improve probeheads and acquisition electronics to make studies on samples of low concentrations feasible. While monitoring protonation through 1H, 31P, or 19F sensor nuclei is relatively cost-effective, 13C or 15N titrations require either more acquisition time, which may be improved by 2D techniques with inverse detection [46] or by isotopic enrichment [47, 48]. In the case of highly hydrophobic (poly)carboxylic acids like bilirubin derivatives, 13C labeling of the carboxylate group and use of DMSO-d6 as cosolvent enabled macroscopic protonation constants to be determined via 13C NMR–pH titrations in the 10–4–10–6 M concentration range [47, 48]. When increasing ligand concentration to achieve better signal-to-noise ratio, complications due to ionic strength fluctuations or ligand self-association may arise that bias the protonation constants. Nearly constant ionic strength can be maintained by adding a large amount of inert, strong electrolyte (e.g., KCl or NaClO4), which in turn reduces the amount of bulk solvent. Ionic strength in the range 0.05–0.3 M is therefore often used. It should also be noted that the counter ion of the inert salt might influence the resonance frequencies and linewidths of polyfunctional ligands by weak complexation or non-specific interaction. The self-aggregation affinity of the compound should be checked prudently prior to the NMR–pH titration by simpler methods like pH potentiometry at varying ligand concentrations.
It is essential that the compound used as chemical shift reference should exhibit no titration shift in the pH range studied. In aqueous 1H or 13C NMR spectroscopy, tert-butanol and dioxane have been extensively used. The most recommended reference substance is the sodium salt of 3-trimethylsilyl-1-propanesulfonate (DSS), which protonates at “pH −6” only [49]. The use of the analogous propionate salt (TSP) should be avoided because it exhibits a protonation shift of 0.019 ppm around pH 5.0 [50]. In the case of other heteronuclei it is customary to use external referencing (e.g., 85% H3PO4 in 31P NMR).
In the usual way of sample preparation, the components are mixed in separate vessels, pH is measured under well-stirred conditions with a typical error of approximately 0.02 pH units, and then the solution is transferred into separate NMR tubes. Although the sample composition can be controlled precisely, this procedure is time- and ligand-consuming. It is better to conduct the whole titration in a single NMR tube, adding the titrant from a fine syringe and homogenizing the solution [51, 52]. The pH measurement in the NMR tube with a long, thin glass electrode is not too precise due to difficulties in stirring. It is more straightforward to use indicator molecule(s) for in situ monitoring of pH [53]. A recent breakthrough in this field was the introduction of multicomponent titrations [59, 60, 61, 62]. In this method, the ligand under study and an additional monobasic compound of known basicity are co-titrated in the same NMR tube with an appropriate titrant. Through eliminating pH values from the calculations, this technique enables basicity differences (relative logK values) to be determined with an improved precision and accuracy, surpassing such classical methods as potentiometric titrations. Perrin et al. also developed a novel device to deliver small aliquots of titrant directly into the NMR tube in the magnet [63]. Another advanced high-accuracy experimental setup is the on-line coupling of a potentiometric titrator apparatus to the NMR spectrometer. This computer-controlled hyphenated technique was introduced in 1988 and is under continuous development in the Hägele research group [54, 55, 56, 57]. The titration-controlled NMR spectroscopy also means sophisticated data evaluation methods [56, 57, 58].
Selecting NMR nuclei to monitor site-specific protonation
A fundamental question of every NMR study on molecules of two or more adjacent basic sites is how the protonation fractions of individual sites are derived from the experimental chemical shift versus pH profiles. Our current knowledge on this issue is summarized below.
The chemical shift of the jth nucleus of a given molecule is defined as the relative difference of its resonance frequency with regard to the reference compound (e.g., DSS):
where σ ref and σ j are the respective shielding constants. They can be further decomposed into three parts [2]:
The diamagnetic contribution σ diamagn.>0 describes the shielding effects of electrons and is thus proportional to the local electron density [64]. The paramagnetic term σ paramagn. <0 is connected to electron excitations to low-energy unoccupied orbitals [65]. The last term σ other summarizes all other effects [66] such as magnetic anisotropy of sterically proximate groups, electric field effects of anisotropic C–X bonds, van der Waals interactions, solvation or conformational influences [67]. The relative proportions of the three terms in Eq. (5) are different for various nuclei, enabling them to differently monitor site-specific protonation equilibria.
1H NMR titrations
In 1H NMR spectroscopy, the diamagnetic contribution dominates over the paramagnetic one [68]. This makes non-exchanging, carbon-bound protons a good probe of local electron density and thus group-specific ionization. The range of influence of a protonating moiety can be studied on homologous series of monoprotic systems. For instance, upon protonation of a carboxylate group, the methylene hydrogens at the α, β, and γ positions are deshielded by approximately 0.2, 0.03, and 0.02 ppm, respectively [69]. Ionization seems to have practically no impact on the chemical shift of more remote aliphatic protons. It has been stated that when two basic sites are isolated by more than four covalent bonds, the adjacent, nonlabile protons can be used to follow site-specific protonation to a good approximation. This precondition usually holds for the side-chains of polypeptides and proteins [23]. In cases of two or more nearby basic sites, the influence of every protonating group must be taken into account (Sudmeier–Reilly approach, see below).
In some peculiar examples of large molecules, the effect of protonation can apparently reach protons located even 25 bonds away, as documented for coenzyme A [71], but such interactions usually occur through space or via conformational changes. Contrary to aliphatic systems, the π electrons of (hetero)aromatic molecules may transmit charge density changes to remote protons through bonds [72, 73, 74].
We must also recall that most protons are situated at the “outer border” of the molecule and are thus exposed to outer influences such as solvation, conformational changes, or aggregation phenomena to a greater extent than the “buried” heavy atoms in the molecular backbone, like 13C. Protonation usually causes a downfield shift of neighboring C–H protons. Upfield, “wrong way” 1H protonation shifts indicate special effects, like pH-dependent conformational changes [74] or the redistribution of protons among the binding sites [25].
13C NMR titrations
The chemical shift of the 1/2-spin 13C isotope (1.108% natural abundance, 0.0159 sensitivity relative to 1H) is also extensively used to monitor protonation equilibria. The advantages of 13C NMR titrations arise from the broader ppm scale: spectral overlap is rare, influences of outer circumstances (ionic strength, temperature) are smaller [75], and protonation shifts are usually larger than in 1H NMR. Nevertheless, the dominance of the paramagnetic term in Eq. (5) makes the 13C chemical shift often sensitive to long-range effects such as protonation on distant groups.
The protonation of a carboxylate group is monitored most sensitively by the carboxylic 13C atom itself: a shielding of approximately 4–5 ppm can usually be observed [69, 70]. Upfield protonation shifts of aliphatic carboxylic acids for the α, β, γ, and δ carbons are typically 3–4, 1.5, 0.6, and 0.2 ppm, respectively [69]. An anomalous trend has been recorded for α-amino acids: upon protonation of the amino group, larger 13C shifts are detected for the carboxylic and β-carbons than for the amino-bearing α-carbon [76, 77, 78, 79]. This β-effect could be in part rationalized in terms of the linear electric field shift (LEFS) theory [76, 77]. Moreover, it has also been stated that for amino acids and derivatives protonation of (up to 5 bonds) remote sites can influence the 13C chemical shift [80, 81], although not monotonously. In polyprotic molecules, protonation shifts of opposite sign can also be observed [82].
Various empirical structure–chemical shift correlations have been established through substituent additivity equations. The most relevant ones are those of Sarneski et al. [83], Rabenstein et al. [80], and Hague et al. [84] which enable the prediction of 13C chemical shifts at various protonation states, thus furnishing C coefficients for the Sudmeier–Reilly approach (see below).
31P NMR titrations
The 1/2-spin nucleus 31P (100% natural abundance, 0.0663 relative sensitivity to protons) is widely applied in protonation equilibrium studies. However, the theoretical understanding of 31P chemical shifts of P(V) oxyacids is incomplete. Several possible electronic effects, potentially of opposite signs, hydrogen bonds, O–P–O bond angles can affect δ P in a complex way [85, 86, 87]. For instance, the protonation of phosphates leads to an upfield shift of the 31P resonance signal, whereas downfield shifts of thiophosphates were observed [88]. In α-aminoalkyl phosphonates and phosphinates, the nitrogen protonation gives rise to the 31P peak downfield shift greater than the upfield shift upon the phosphonate oxygen protonation [54, 89, 90]. Thus, 31P chemical shifts are amenable to respond to protonation effects of remote groups. Nevertheless, their applicability to follow site-specific protonation has to be judged individually for each class of compounds.
The Spiess research group has gained considerable evidence that the C–O–31PO3 2– moiety of most inositol (poly)phosphates selectively monitors its own protonation state; the 31P chemical shift usually undergoes a 3.8- to 4.0-ppm upfield shift.
On the other hand, there is experimental evidence that the 31P NMR titrations alone cannot be used with certainty to identify the proton-binding phosphates in nucleoside di- and triphosphates. 17O NMR proved to be a more direct approach (see below).
Hägele and other authors have studied a large number of phospha-analogues of α- and β-amino acids, peptides, and polycarboxylic acids. In the absence of other ionizable sites, the 31P resonance peak exhibits a downfield protonation shift in phosphonates, phospinates, and 1-hydroxyphosphonates. Fluorine bound to the carbon skeleton can change the sign of the 31P protonation shift. In polyfunctional molecules, the 31P chemical shift can be affected by protonation events up to four bonds away [56].
15N NMR titrations
The chemical shift of the less abundant (0.37%), 1/2-spin 15N nucleus (1.04×10–3 relative sensitivity to protons) is also sensitive to acid–base equilibria [91, 92]. Natural abundance 15N NMR spectroscopy represents a straightforward approach to study the protonation of amine groups, since the basic atom is directly observed. Amino sugars and antibiotics are most commonly titrated, reporting downfield shifts ranging from 7 to 14 ppm upon NH2 protonation [93, 94]. In the case of an aromatic, sp2-hybridized 15N nucleus, such as in pyridine derivatives, an upfield shift is sometimes observed due to the dominance of the paramagnetic term. This upfield shift also carries over to some of the carbons. 15N NMR titrations can be performed more conveniently on isotopically enriched enzyme samples [95].
Uncommon NMR nuclei in microconstant determinations
To follow the proton coordination to oxyacid functional groups (carboxylate, phosphonate, phosphate ester group, etc.), it is advantageous to monitor the chemical shift of the proton-binding oxygen directly. Unfortunately, the 5/2-spin, quadrupolar 17O nucleus has a natural abundance of 0.037% only, though its sensitivity relative to protons is 0.0291. In very systematic and instructive studies, Gerlt and co-workers demonstrated that for phosphorus oxyacids the 17O chemical shift varies linearly with the partial charge of the oxygen atoms [96, 97, 98]. The protonation shift is approximately 50 ppm per charge neutralization for phosphates, phosphonates, di- and triphosphates, and their thioderivatives.
The 1-spin nucleus 14N (99.63% natural abundance, 1.01×10–3 relative sensitivity to protons) provides in principle the most direct means to study the protonation of individual amino or amine groups as well as heterocyclic nitrogen atoms. Due to the low sensitivity, there are only a few examples of its application. Gajda and co-workers observed an upfield shift of 10–60 ppm upon 14N-protonation of imidazole derivatives [99].
The 1/2-spin 19F nucleus (100% natural abundance, 0.83 relative sensitivity to protons) can also be used to monitor protonation equilibria [57, 100, 101]. Fluorine atoms, however, are scarcity in drugs and especially in biomolecules. To the best of our knowledge, no microconstants have been determined by 19F NMR–pH titration.
Principles of microequilibria
Biprotic systems
The fundamentals to evaluate microconstants from NMR–pH titration curves are exemplified by case studies on bi- and tetraprotic systems. Scheme and principles of tri- and n-protic microspeciation are discussed here at the theoretical level. As one of the numerous biprotic microequilibrium systems, the dipeptide cysteinylglycine (CysGly) is shown. The tetraprotic example is the reduced form of the neurohypophyseal peptide hormone arginine vasopressin, which contains analogous moieties with CysGly. The NMR–pH titrations of both compounds have been carried out in D2O using the pD scale [102]. Since we focus our attention on evaluation principles, all equilibria will be treated as protonation ones, instead of deuteration. Also, the symbols pH and [H+] are used for simplicity.
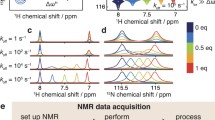
Cysteinylglycine has three basic sites, the amino (N), thiolate (S), and the carboxylate (C) groups (see Fig. 1). The apparent contradiction that a molecule with three protonating sites is the prototype in this section entitled “Biprotic systems” is resolved by the NMR–pH profile of the CysGly α-proton in Fig. 2. The curve indicates two merged, major downfield shifts between pH 11 and 6, and a separate, minor one near pH 4. This allows division of the complete titration curve into a diprotic and a monoprotic one. Chemical evidence leaves no doubt that former and latter belong to the aminothiolate and carboxylate protonations, respectively.
NMR titration curve of the Hα nucleus of cysteinylglycine, indicating the chemical shifts of individual macrospecies H i L. B Individual protonation fraction curves (f) of the amino (N), thiolate (S) and the separately protonating carboxylate (C) groups [102]
Thus, biprotic evaluation contains data in the 14–5 pH range and the corresponding 3.33–4.24 ppm chemical shift range.
Evaluation usually starts at the macroscopic level, confining considerations to the stoichiometry of protonation, ignoring the sites of proton binding.
K 1 and K 2 are the stepwise (successive) macroscopic protonation constants of CysGly, designated here as the ligand (L), and charges on the species are omitted:
The β i cumulative macroscopic constants are especially useful in multiprotic systems and they are products of the stepwise ones:
The observed, pH-dependent chemical shift of any protonation-sensitive nucleus can be formulated by appropriately extending Eq. (1):
where concentration of the mono- and diprotonated species can be expressed in terms of [L], [H+], and cumulative macroconstants, which yields after rearrangements:
where δ L and \( \delta _{{{\text{H}}_{2} {\text{L}}}} \) directly reads from the NMR–pH titration curve. Contrary to that, δ HL can be obtained from nonlinear parameter estimation. In addition, the species HL is a mixture of amino- and thiolate-protonated ones, the ratio of which changes heavily with solvent, ionic strength, and temperature, calling forth significant sensitivity of δ HL to the solution circumstances.
Equation (10) can be generalized to systems of arbitrary number of basic groups:
where n is the total number of protonation sites of the ligand in question and β 0=1 by definition.
The nonlinear fit based on the NMR–pH titration in Fig. 2A resulted in macroconstants logK 1=9.85, logK 2=7.58, and logK 3=3.64, where the last of these refers to the carboxylate protonation, whereas the first two quantitate compositely the thiolate-amino proton binding, which needs to be decomposed into microconstants.
Microscopic protonation, microspeciation
In order to assign protonation to binding sites, equilibria have to be considered at the microscopic (or submolecular) level. The microscopic protonation scheme of CysGly is depicted in Fig. 3.
Macroscopic and microscopic protonation scheme of cysteinylglycine. N and S represent the amino- and thiolate groups, K and k denote macroscopic and microscopic protonation constants, respectively [102]
The amino (N) and thiolate (S) groups are represented as parts of a two-armed symbol, to which protons attach in all possible sequences. The microspecies S, N, and SN are labeled by their protonated sites. L is the nonprotonated ligand. The superscript on microconstant k indicates the group protonating in the equilibrium in question, whereas the subscript (if any) refers to already protonated group(s). For instance, the microconstants k S and \(k^{{\text{S}}}_{{\text{N}}} \) characterize the following equilibrium reactions:
Other nomenclatures for microconstants and microspecies have also been introduced [20, 24, 25, 74, 103].
Since microspecies L, N, NS, and S form a Hessian, thermodynamic cycle, the four unknown microconstants k N, k S, \(k^{{\text{N}}}_{{\text{S}}} \) and \(k^{{\text{S}}}_{{\text{N}}} \) in Fig. 3 are not independent, they are interrelated via the following constraint:
Microspecies S and N are of the same stoichiometry, holding the proton at different sites. They are therefore protonation isomers, the concentration ratio of which is independent of both the pH and total concentration:
Solvent, ionic strength, temperature, and conditions that modify k N or k S, however, can cause dramatic changes in the ratio of protonation isomers. The scheme in Fig. 3 shows that both k N and \(k^{{\text{N}}}_{{\text{S}}} \) refer to the amino protonation. Similarly, both k S and \(k^{{\text{S}}}_{{\text{N}}} \) characterize the thiolate basicity. Differences between microconstants of the same site arise from the protonation state of the neighboring site(s). Protonation of a neighboring site normally exerts an anticooperative effect. For example, protonation of the thiolate group decreases the electron density everywhere in the molecule, including the amino site, thereby reducing its basicity. This reciprocal effect can be quantified in E NS, the pair-interactivity parameter, as follows:
Thus, E<1 in the vast majority of cases, indicating anticooperativity. The strength of the interaction is usually inversely proportional to the number of chemical bonds between the protonation sites. The closer the sites, the stronger the inductive effects and the anticooperative interaction, the smaller the value of E. On the other hand, remote protonation sites can exist without any real interaction, due to the consecutive isolating effects of the intervening bonds, resulting in E≅1. Significance of this is shown in the tetraprotic case.
Relationships between the macro- and microconstants [9, 13, 15] can be deduced from the facts that [HL]=[N]+[S] and [H2L]=[NS], as follows:
Evaluation of microconstants
Microconstants can be calculated from f, site-specific protonation mole fractions. For example, f N, the protonation fraction of the amino group is given by the sum of relative concentration of those microspecies in which site N is protonated:
where T L denotes the total (analytical) molar concentration of the ligand, T L=[L]+[N]+[S]+[NS].
Similar equation holds for the thiolate protonation fraction f S:
At every pH, the sum of group-specific protonation degrees gives the cumulative protonation degree of the ligand, in other words, the Bjerrum \(\bar{n}_{{\text{H}}} \) function, which is a function of β macroconstants only:
Thus, for a biprotic molecule, it is sufficient to follow the protonation of only one group selectively (e.g., by UV or NMR spectroscopy); the other one can be expressed from Eq. (21). This fact will be exploited below.
Site-specific f N and f S-type functions, however, cannot always be extracted directly from NMR–pH profiles. In fact, every NMR nucleus of CysGly reflects to some extent the protonation state of both the amino and thiolate sites. In general, the extent (contribution) of an individual site to the total protonation shift of a given NMR nucleus is known a priori in limiting cases only, when the nucleus in question is influenced by the electron density of one single protonation site and it is independent of all others. In cases of small molecules of bond-mediated interactions, composite NMR–pH profiles are observed, when site-specific f functions can be obtained using the Sudmeier–Reilly approach.
Composite NMR titration curves from interacting sites: the Sudmeier–Reilly model
In CysGly the pH-dependent chemical shift of Hα between pH 6 and 12 (Fig. 2A) is influenced by the protonation degree f of both the amino (N) and thiolate (S) groups. In the Sudmeier–Reilly model, the two contributions are assumed to be additive:
where the protonation shift coefficient \(C_{{{\text{H}}^{\alpha } ,{\text{S}}}} \) describes the change in the Hα chemical shift caused by the complete protonation of the thiolate group.
Equation (22) can be further simplified by eliminating f N using Eq. (21). This leads to the following expression:
It is now widely recognized that simultaneous calculation [105] of the C and f values from Eqs. (22) or (23) is an ill-fated idea [107, 108, 109, 110], since these variables are highly correlated, linearly dependent ones. Thus, either protonation fractions can be determined if the C coefficients are known (e.g., imported from protonation shifts of structurally similar compounds of reduced number of sites) or accurate C coefficients can be obtained provided that f S is measured by an independent technique. For CysGly, the second type of evaluation is used.
The sum of C coefficients equals the total protonation shift of Hα (see also Fig. 2A):
Combination of Eqs. (19), (20), (21), (22), and (23) leads to the following master equation to fit the NMR–pH titration curve of Hα:
The two unknown parameters \(C_{{{\text{H}}^{\alpha } ,{\text{S}}}} \) and k S are highly correlated and cannot be simultaneously obtained by direct fitting of Eq. (25) to the experimental 1H NMR titration curve of Hα.
The thiolate microconstant k S can be determined by using UV pH titration, an independent technique. The thiolate group exhibits a UV absorbance at 234 nm which diminishes upon protonation [111]. Since there are no additional pH-dependent absorbance changes at this wavelength, the protonation fraction of the thiolate group can be assessed from the measured absorbances by using the following equation:
where A 234 nm, L and \(A_{{234{\text{nm}},{\text{ H}}_{2} {\text{L}}}} \) are the limiting absorbance values corresponding to the fully deprotonated (pH=12) and protonated (pH=6) species, respectively. The resulting f S versus pH curve is shown in Fig. 2B. k S is then calculated by fitting Eq. (20) to this dataset to yield logk S=9.72. Equation (14) enables the calculation of the remaining three microconstants. The macroconstants and microconstants of CysGly are collected in Table 1.
With knowledge of k S, the single unknown parameter \(C_{{{\text{H}}^{\alpha } ,{\text{S}}}} \) can be obtained reliably by fitting Eq. (25) to the experimental NMR–pH profile of Hα, leading to the following protonation shift coefficients: \(C_{{{\text{H}}^{\alpha } ,{\text{S}}}} = 0.228\;{\text{ppm}}\) and \(C_{{{\text{H}}^{\alpha } ,{\text{N}}}} = 0.682\;{\text{ppm}}{\text{.}}\) As expected from the molecular structure in Fig. 1, the more closely spaced amino group has about three times larger influence on the chemical shift of Hα, but the impact of the thiolate protonation is far from negligible.
Calculation of microspecies distribution
Macroconstants and microconstants enable the distribution of macrospecies and microspecies to be calculated as a function of pH. The distribution curve of HL is decomposed into those of protonation isomers N and S. For instance, the mole fraction of microspecies S is given at an arbitrary pH by the following equation:
Figure 4 clearly shows that microspecies S dominates over N at each pH, so the major pathway of protonation includes the microspecies L→S→NS. The pH-independent concentration ratio of microspecies S and N is given by the following equation: [S]/[N]=k S/k N=2.9.
Distribution curves of the macrospecies (H i L) and microspecies (N, S) of cysteinylglycine [102]
As mentioned above, the thiolate and amino groups of CysGly modulate their basicity mutually. Indeed, the amino protonation decreases the thiolate basicity in CysGly significantly, by a factor of 35 (E NS=0.0286, or pE NS=1.55 on the logarithmic scale) and vice versa.
Triprotic systems
Figure 5 shows the general scheme of protonation of a trivalent base. Examples of bio and drug molecules that bind three protons in an overlapping and interacting fashion are DOPA (dihydroxyphenylalanine), dopamine, and γ-carboxyglutamic acid [104].
Designation of microconstants in Fig. 5 are analogous with that of the biprotic system. Bi- and triprotic microequilibrium systems are the only ones that allow determination of all the microconstants from site-selective NMR–pH titrations without further assumptions [17, 25]. The remarkable difference between the bi- and triprotic microequilibrium systems lies in the number of microspecies and microconstants. Both numbers further increase in tetra- and n-protic systems. Fortunately, the overwhelming complexity of tetra- and n-protic systems can often be simplified into sets of mono-, bi-, and triprotic subsystems, as shown below.
Tetraprotic microequilibria: a case study of reduced arginine vasopressin (rAVP)
The structure of the reduced arginine vasopressin (rAVP) is shown in Fig. 6. In the studied pH range 2–13, the ligand is capable of binding four protons to sites as follows: amino (N) and thiolate (S) of the terminal cysteine, phenolate (O) of tyrosine2, and thiolate (S′) of cysteine6.
The complete assignment of the 1H NMR resonances of rAVP has been achieved by Larive and Rabenstein by using COSY, TOCSY, and ROESY spectra [37]. The 1H NMR titration resulted in chemical shift versus pH profiles for each observed carbon-bound protons [102]. Figure 7 shows the 1H NMR titration curves for those hydrogens that are used in the evaluation. Note the unusual behavior of one of the δ-methylene protons of Pro7, which undergoes an upfield shift upon protonation of the neighboring thiolate S′, suggesting a concomitant conformational change.
A 1H NMR titration curves of the aromatic 3,5-protons of Tyr2, the Cys1 α-CH proton and one of the Pro7 δ-CH2 protons of reduced arginine vasopressin. B individual protonation fraction curves (f) of the Cys1 amino (N) and thiolate (S), the Tyr2 phenolate (O) and the Cys6 thiolate (S′) groups [102]. Note that f O and f S′ are regular sigmoids, representing sites of no intermoiety interactions. Contrary to that, f S and f N are non-sigmoidal curves due to thiolate-amino interaction in Cys1
The 1H NMR–pH profiles were fitted to the tetraprotic analogue of Eq. (10) to yield the chemical shifts of each macrospecies H i L and the following logK macroconstants: 10.70, 9.30, 8.65, and 7.02.
In principle, the complete microequilibrium scheme of a four-basic ligand contains 24=16 microspecies and 4×23=32 unknown microconstants (Fig. 8) [17]. The theory and practice used to analyze a genuine tetraprotic system has recently been published [18]. However, the complexity of the full analysis of rAVP can be reduced significantly by “decoupling” the independently protonating phenolate (O) and Cys6 thiolate (S′) sites from protonation equilibria of the strongly interacting Cys1 amino (N) and thiolate (S) groups in the evaluation procedure.
Macroscopic and microscopic protonation scheme of reduced arginine vasopressin, a tetrabasic molecule. The groups are labeled as follows: Cys6 thiolate (S′), Tyr2 phenolate (O), Cys1 amino (N) and thiolate (S). In the microequilibrium scheme (bottom), the arrows represent the 32 protonation microconstants
Evaluation of “selective” NMR titration curves
The Tyr2 Hφ and the Pro7 Hδ hydrogens are separated by more than nine isolating covalent bonds from each other and from the amino and thiolate groups of Cys1. Therefore, these nuclei can well be assumed to be selective sensors (“unique resonances” [105]) to monitor the protonation of the phenolate (O) and Cys6 thiolate (S′) sites, respectively. Of course, the best reporters for S′ would be the Cys6 CH2 and CH protons. Unfortunately, their resonances are obscured by several other peaks, but the Hδ nucleus of the neighboring Pro7 residue that goes the “wrong way” offers a convenient means to follow the ionization of Cys6 selectively.
The 1H NMR–pH profile of the Tyr2 Hφ protons in Fig. 7A fits well to the monoprotic model,
from which the group constant logk O=10.70 results. Group constants represent a limiting case of microconstants [11, 12, 13, 17]. The single subscript O indicates that the intrinsic basicity of the phenolate group is independent of the protonation state of the remaining S, S′, and N sites. Note that group constants do not exclude through-space interactions of the sites by coulombic forces or eventual hydrogen bonds. Comparison of group constants to basicities of model compounds can indicate such interactions (see the discussion of protein residues below).
Similarly, the group constant of the Cys6 thiolate site is obtained by fitting Eq. (28) to the NMR–pH titration curve of the “indicator” Hδ proton of Pro7, leading to logk S′=8.65.
Evaluation of “non-selective” NMR titration curves: the Sudmeier–Reilly approach with imported protonation shifts
The N-terminal thiolate (S) and amino (N) groups are in close vicinity and they interact through covalent bonds. Preconditions of the group constant treatment are not at all valid. Rather, a biprotic microequilibrium system has to be considered which can be represented with the same symbols as for CysGly in Fig. 3, because both molecules contain the same molecular fragment.
Similarly to CysGly, the pH-dependent chemical shift of the Cys1 α-CH proton of rAVP is also assumed to obey the Sudmeier–Reilly relationship stated in Eqs. (22) and (25). Again, the unknown parameters, \(C_{{{\text{H}}^{\alpha } ,{\text{S}}}} \) and k S are highly correlated and cannot be obtained simultaneously by nonlinear regression. Instead, the value of \(C_{{{\text{H}}^{\alpha } ,{\text{S}}}} \) is “imported” from the model compound CysGly. That means that \(C_{{{\text{H}}^{\alpha } ,{\text{S}}}} \) is assumed to be equal to \(C^{{{\text{CysGly}}}}_{{{\text{H}}^{\alpha } ,{\text{S}}}} = 0.228\;{\text{ppm}}{\text{.}}\) The validity of this assumption can be tested [81] by comparing the total protonation shift of CysGly, \(\Delta \delta ^{{{\text{CysGly}},\max }}_{{{\text{H}}^{\alpha } }} = 0.910\;{\text{ppm}}\) to that of rAVP, \(\Delta \delta ^{{\max }}_{{{\text{H}}^{\alpha } }} = 0.857\;{\text{ppm}}{\text{.}}\) The small discrepancy of these values can be taken into correction by a proportional adjustment of \(C^{{{\text{CysGly}}}}_{{{\text{H}}^{\alpha } ,{\text{S}}}} \) as follows:
With this value, Eq. (25) is fitted to the experimental NMR titration curve of the rAVP Hα nucleus (Fig. 7A), resulting in logk S=9.18. The remaining three microconstants are calculated from Eq. (14) to be logk N=8.69, \(\log k^{{\text{S}}}_{{\text{N}}} = 7.63, \) and \( \log k^{{\text{N}}}_{{\text{S}}} = 7.14. \) The pair-interactivity parameter pE NS=1.55 characterizes the mutual basicity-decreasing effect of the amino and thiolate sites through bonds. Virtually the same interactivity parameter has been found for CysGly, Cys methyl ester, and reduced oxytocin, which contain both groups in the same distance and intramolecular environment [102]. Indeed, the interactivity parameter of a pair of functional groups proved to be less perturbed by actual molecular environment and thus it is a more transferable parameter between molecules than microconstants [18, 112].
Microconstants and group constants allow the construction of distribution curves for all 16 microspecies of rAVP (Fig. 9).
A Distribution curves of the macrospecies H i L of reduced arginine vasopressin [102]. B Logarithmic distribution curves of all 16 microspecies of reduced arginine vasopressin
n-Protic microspeciation systems: problems and solutions
Effective parametrization schemes to describe large systems
The number of basic sites, n, is in exponential relationship with the number of microspecies (2n) and k microconstants (n2n–1) [12, 13]. An increase in the number of basic sites gives rise to an especially overwhelming increase in the number of k microconstants, causing serious problems in the evaluation and even in the formal description of the system. In order to alleviate difficulties in the formalism of description, we introduced cumulative microconstants [17, 18], designated by κ, the number of which is 2n−1. This Hessian-type microconstant is assigned to every microspecies containing at least one proton, and it unifies the various, alternative microscopic routes of microspecies formation in one single parameter [17, 18]. Taking the example in Eq. (14), it reads:
Advantages of cumulative microconstants become obvious at tri- and higher-protic systems.
Even using this compact parametrization, a fundamental difference was revealed between three-group and larger microequilibrium systems in 1999 [17]. For bi- and tridentate ligands of any arbitrary symmetry, every microconstant can in principle be unambiguously calculated from the f protonation fraction curves of the individual groups. However, in cases of systems of four or more nonidentical basic sites, the f curves do not contain sufficient information to obtain a unique set of microconstants. By systematically examining the influence of symmetry up to the hexaprotic case, equivalence of protonation sites has been shown to reduce the number of unknown parameters significantly, allowing unique solutions for the total symmetrical cases [17]. Four years later, Ullmann came to the same conclusion [25] on the basis of the decoupled site representation (DSR) [24].
In polyprotic molecules of lower symmetry, it is more straightforward to parametrize the system in terms of n ”core” microconstants, describing site-specific basicity of noninteracting sites and of n(n−1)/2 pair-interaction parameters [17]. With this choice of unknown parameters, all 32 microconstants describing the four carboxylates of oxidized glutathione (GSSG) could be calculated from the 1H NMR–pH titration curves [18]. In this case, the pairwise interaction of two carboxylates is assumed to be independent of the protonation state of the remaining ones. Even this assumption can be released by introducing interaction parameters for group triplets, quartets, etc. in the framework of the more general cluster expansion method introduced by Borkovec and Koper [19, 20, 27, 28]. Today, this formalism represents one general tool to assess microequilibria of very large systems (e.g., dendrimers) [26, 27].
The other alternative is the decoupled site representation (DSR), developed by Onufriev and Ullmann [24, 25]. This is based on quasisite constants (pK′) describing non-interacting sites. It can be easily shown that quasisite constants are in fact identical to the earlier introduced group constants [11, 12], used also in our case study above. From the quasisite constants, a unique set of cumulative microconstants of the real sites is obtained through a linear transformation [24, 25]. The power of this methodology is illustrated by decomposition of irregular NMR–pH profiles into site-specific f protonation fraction curves for rubredoxin [24] and DTPA [25] (see below).
Uniqueness of microconstant sets
It is absolutely fundamental that the microequilibrium analysis results in microconstants and microspecies concentrations that are the one and only solutions of the system.
For this purpose, both chemical and mathematical criteria must be met. The chemical criterion is that the f function reflects the protonation state of one single basic site. This can be achieved by selecting specific “reporter” nuclei, or by sorting out the interference of other sites, by means of the Sudmeier–Reilly relationship. In order to check the site-specificity of f functions, three methods are mentioned below.
In the first method [114], the sum of the site-specific protonation fractions f is compared at each pH to the overall degree of protonation (the \(\bar{n}_{{\text{H}}} \) Bjerrum function), determined by independent potentiometric titration. Good agreement between these datasets should be obtained. If microconstants have been calculated without the implicit use of macroconstant values, relations between macroconstants and microconstants [10, 13] can be tested. This test is essentially identical with the previous one. The second checking method is useful for multinuclear studies on protein residues. If several intraresidue nuclei (1H, 13C, 15N) in the vicinity of a basic site exhibit the same sigmoid titration curve treatable with the single-logk model Eq. (28), they can be treated as specific “reporters” for group constant-type evaluation [42, 43, 115]. A third method to check pairs of selective nuclei [53] will be published in the near future.
Site-specificity of f functions is a necessary but not sufficient requirement to obtain a unique set of microconstants and microspecies concentrations.
Concerning mathematical considerations, both DSR and cluster expansion methods attempt to obtain microequilibrium parameters and NMR protonation shifts (equivalent to C shielding constants) simultaneously from the measured NMR–pH titration curves. While such an approach can yield unique solutions in special, favorable cases [27], the failure to calculate C Sudmeier–Reilly coefficients together with microequilibrium constants has been demonstrated several times [107, 108, 109]. Thus, model calculations and a subsequent, rigorous statistical analysis [18] of the evaluated microscopic and NMR parameters are inevitable measures to unravel possible linear dependence (correlation) of the NMR protonation shift and equilibrium parameters, which leads to nonuniqueness of the solution.
Finally, a few remarks from the viewpoint of classical thermodynamics, which says that no isothermal, pH-dependent spectral series contains sufficient information to unambiguously determine all microconstants of a polyprotic molecule, without making further assumptions of structural nature [21]. The assumptions can be generally recognized, trivial ones that justify microspeciation. For example, such an assumption is that a spin-active nucleus is the specific probe of protonation for one particular group [22]. The Sudmeier–Reilly model takes the effect of distant groups into account, assuming the perfect additivity of protonation influences [106]. Sophisticated assumptions are being developed even today (see, e.g., ref. [113] for UV pH titrations), leading hopefully to more profound understanding of microequilibrium systems and improved microconstant determination strategies in the future.
NMR studies on protonation microequilibria of bioligands
The following literature survey will focus on those studies where microscopic protonation constants or at least pH-dependent fractional protonation of individual basic sites have been derived from NMR–pH titration curves of “small molecules”. The structures of some of the discussed ligands are given in Table 2. NMR studies that identify the site of protonation but supply no quantitative information on its relative basicity to other proton-binding groups in the molecule will not be covered here.
Natural amino acids, oligopeptides, and simple derivatives
Amino acids, di-, and tripeptides were among the first compounds characterized in terms of protonation microconstants. In favorable cases, the overlapping protonating groups are separated by more than four covalent bonds, allowing the use of the adjacent carbon-bound protons as selective, “reporter” nuclei. Based on this principle, microconstants have been determined by 1H NMR titration for lysine [105], l-3,4-dihydroxyphenylalanine (DOPA, structure 1 in Table 2) [116], histidine or histamine-containing dipeptides [99, 117], glycylglycylhistamine [118], reduced glutathione (γ-GluCysGly) [21], oxidized glutathione [18], and the γ-methylphosphino analogue of glutamic acid (phosphinothricin, 2) [56].
In the Sudmeier–Reilly approach, the protonation effects of remote groups are also considered (see Eq. (22) above). In lysine and ornithine (3) [107], GlyHis, GlyHisGly, and GlyHisLys [51], the number of separating bonds is greater than four; therefore, the C “cross-terms” are found to be small (<0.03 ppm). In DOPA, adrenaline [108], and cysteine [119], the protonating groups are closer to each other and the influence of both functional groups on 1H protonation shifts become comparable. The Sudmeier–Reilly approach has been applied to obtain microconstants from 13C NMR–pH titration of lysine and δ-hydroxylysine (with ε-aminocaproic acid and norleucine as model compounds to obtain C j,N shielding constants, [81]), aspartic acid (using asparagine as model, [14]), and α- and β-alaninehydroxamic acid (4) (using propionohydroxamic acid as model [110]).
Special problems arise in the investigation of microequilibria of histidine (5), histamine, and some derivatives like carnosine (β-AlaHis). First, the low occurrence of the minor microspecies holding proton at the imidazole nitrogen precludes the determination of reliable microconstants by conventional NMR–pH titrations [99, 120]. Instead, microconstants have been derived from the kinetics of deuteron exchange of the imidazole C2–H proton in D2O [120]. Investigating the effect of charged groups on the imidazole basicity leads to empirical logk prediction relationships [121]. Other complications arise from the N1–H and N3–H tautomerism of the imidazole ring (see 5 in Table 2). By using N-alkylated model compounds, tautomeric ratios have been determined for imidazole, histidine, and histamine derivatives including peptides and proteins by 1H NMR titrations [122, 123, 124], 15N NMR titrations [125, 126, 127], and 13C NMR titrations in aqueous [122, 128, 129, 130, 131, 132] and solid phases [132, 133]. Microconstants of individual N1–H and N3–H tautomers of histidine derivatives have been estimated using model compounds [122] and directly from the 14N NMR–pH titration of glycylhistamine and sarcosylhistamine [99].
Nucleic bases, nucleosides
The correlative electron structure and additional tautomeric equilibria render microconstants of nucleic bases difficult to determine [13], despite the fact that it would be important to understand their base-pairing and metal-ion-complexing ability [134, 135]. The predominant site(s) of protonation can be studied by 13C and 15N NMR titrations [136].
Group constants of nucleosides are usually 0.2–0.5 logk units smaller than those of the corresponding nucleic bases, due to the electron-withdrawing effect of the hydroxyl groups on the carbohydrate subunit [13, 134].
Nucleotides, inositol phosphates, and other phosphate esters
Going from nucleosides to nucleotides, nitrogen sites with logk<7 do not alter their basicity significantly. For more basic nitrogens, an increase of approximately 0.1–0.4logk units is observed owing to the basicity-increasing effect of the dianionic phosphate group [13, 134]. Another effect of the phosphate group protonation near pH 6.3 is the “wrong way” upfield shift of the H8 and H6 nuclei in purines and pyrimidines, respectively, which are adjacent to the sugar-substituted nitrogen atoms [134].
Gerlt and co-workers have shown that 17O offers a better alternative to 31P to follow the protonation of individual phosphate groups [96, 97, 98], though no microconstant values have been derived.
Crisponi et al. analyzed the pH-dependent chemical shifts of the adenine 13C, 1H and the phosphate 31P nuclei of adenosine-5′-trisphosphate (ATP) to derive microconstants for the overlapping protonating adenyl and gamma-phosphate groups [137]. Further complications arise when sodium ions are present in the solution: as proved by 23Na NMR, Na+ ions compete with protons for binding the triphosphate site, which leads to exchange-broadening of 31P signals in certain pH regions [138].
15N NMR titration has been applied to characterize the basicity of N1 site of adenine in A–G and A–C mispairs of oligonucleotides [139].
Microconstants of the Schiff base composed of 2-amino-3-phosphonopropionic acid and pyridoxal 5′-phosphate have been determined by combined use of potentiometric, UV, 31P and 1H NMR titrations [140].
The microscopic acid–base properties of the thiol (CoASH), homodisulfide (CoASSCoA) and heterodisulfide (CoASSG) forms of coenzyme A 6 have been investigated by 1H and 31P NMR titrations [71]. The group constant of the adenine N1, cysteamine thiolate, and 3′-phosphate sites has been obtained from the NMR–pH profiles of the aromatic CH2, cysteamine methylene protons, and \({}^{{31}}{\text{PO}}^{{2 - }}_{3} ,\) respectively. The through-space titration shifts of the panteteine protons provided valuable information about the solution structures of these molecules [71].
1d-myo-Inositol 1,4,5-trisphosphate (Ins(1,4,5)P3, 7) plays a major role as secondary messenger in transmembrane signaling [141]. Since pH influences the binding of inositol phosphates (IPs) to Ins(1,4,5)P3 receptors [142], Spiess and co-workers have used 31P NMR–pH titrations to determine microconstant for a large number of IPs and their analogues. In the pH range 2–12, each \({\text{C}} - {\text{O}} - {\text{PO}}^{{2 - }}_{3} \) group coordinates one proton, giving rise to biprotic [143] and triprotic [74, 114, 144, 145, 146, 147, 148] microequilibrium systems. The microconstants and interactivity parameters indicate that hydrogen bonding and hydration are the main factors to determine the basicity of individual phosphate groups. The concomitant “wrong way” protonation shifts in 1H and 31P NMR lead to the proposal of a C–H...O–C hydrogen bond for Ins(1,4,5)P3 in aqueous solution [147].
The hexaprotic inositol phosphate, phytic acid has been the subject of several 31P NMR titrations [149, 150] and represents a challenging microequilibrium system. The complete resolution of microconstants could not be attained in these studies, and only a tentative sequence of protonation has been proposed, where intramolecular hydrogen bonds between vicinal phosphate groups seem to play an important role [149].
Open-chain polyamines
Bencini et al. published a comprehensive review discussing the typical protonation patterns of linear, macrocyclic, and macropolycyclic polyamines [151]. The present survey covers only those articles in which fractional protonation of individual nitrogen atoms or microconstants have been determined by NMR–pH titrations.
In linear polyamines, 1H and 13C resonances of the methylene groups are affected by protonation of both neighboring amine centers [83]. To obtain site-specific protonation information from these composite NMR–pH titration curves, a general approach was introduced by Sudmeier and Reilly in 1964 [106]. By investigating the 1H NMR protonation shifts of mono- and symmetric bifunctional amines and carboxylates with unambiguous states of protonation, substituent constants have been calculated for (CH2) n NH2, (CH2) n COO− groups and their protonated counterparts (n=0–2). With the aid of these increments, C protonation shift coefficients could be assembled for methylene groups of complex polyamines (and polyaminopolycarboxylates, see the next section). With knowledge of the C coefficients, the measured protonation shifts \( \Delta \delta _{j} \) can be converted into protonation fraction curves f i of each amine group by the Sudmeier–Reilly equation [106]:
The pH-dependent f functions characterize the distribution of protons among the basic sites quantitatively and most NMR–pH studies of these compounds terminate at this stage. The final step is, however, the calculation of microconstants and/or interactivity parameters from the f i versus pH functions by nonlinear regression [17, 22]. In the most recent methodologies like the cluster expansion technique [27], the f functions are not calculated explicitly, and microscopic equilibrium constants are obtained by fitting analogues of Eq. (25) directly to the experimental data.
Protonation fractions as a function of pH have been determined for ethylenediamine, trien 8, and tetren 9 by 1H [106] and 13C NMR [152], for 3,2,3-tet and 3,3,3-tet (for nomenclature, see ref. [151]) by 13C NMR [155], for spermine (10) and spermidine (11) by 1H [156] and 13C NMR [153, 154], for thermospermine by 15N NMR [157] and for the neuroactive wasp toxin philanthotoxin-343 by 1H and 13C NMR titration [158]. Microconstants have been determined by NMR titration for N-(2-mercaptoethyl)-1,3-diaminopropane [159], tetren [27], spermine, spermidine, and homologues [154, 160].
To summarize these results, electrostatic repulsion of nearest-neighbor NH+ sites determines the protonation sequence of linear polyamines with all-ethylenic chains [151]. Nitrogens isolated by longer alkyl chains protonate more extensively and in a more random fashion [161]. In fact, the biogenic polyamine spermidine is fully protonated at physiological pH to play its role as charge neutralizer of DNA.
Linear polyamines with basic side chains
Microconstants were derived from 1H NMR titration of ethylenediaminemonoacetate, assuming adjacent methylene protons as specific probes of nitrogen protonation [105].
Ethylenediaminetetraacetate (EDTA, 12) is the simplest polyaminopolycarboxylate to exhibit a non-monotonous 1H NMR–pH profile [106]. A possible explanation was proposed by Letkeman and Martell [162]: the first two protons bind to nitrogens creating two–two hydrogen bonds (or, at least, strong electrostatic interactions) with the carboxylates [163], thus fixing their position. Upon addition of a further equivalent acid, the N-attached carboxylates start to protonate and gain rotational freedom near the backbone CH2 protons, thus causing their 1H “wrong way” shift. This sequence of protonation and the proposed hydrogen bonds are in accord with infrared studies in aqueous solution [164, 165].
Another interpretation of the “wrong way” protonation shift is that while the first two protons are bound to the nitrogens, the predominant triprotonated form bears protons at one nitrogen and two remote carboxylates, due to repulsive forces between protonated sites, and the concomitant relative increase of the electron density upon proton migration.
Diethylenetriaminepentaacetic acid (DTPA, 13) also exhibits a “wrong way” 1H shift between pH 7 and 10 [106, 162, 166]. This phenomenon has been attributed to a peculiar protonation sequence of nitrogens [24, 25, 167]: the first proton binds preferentially to the central nitrogen atom Nc. At the second protonation step, Nc loses its proton to a large extent and the two bound hydrogen ions migrate to the two terminal amine groups Nt [167]. This proton migration, which occurs to a lesser extent in the DTPA-bis(amide) derivatives, is favored by the greater separation of positively charged NH+ groups and by formation of two hydrogen bond rings involving each protonated terminal nitrogen and their two attached acetate groups [162, 163]. The non-monotonous fractional protonation of the central nitrogen is reflected in the irregular 1H–pH profile of the Nc–CH2 methylene protons [24, 25, 168, 169]. The same pattern of protonation has been inferred from the 1H NMR titration of DTPA bis(amide) derivatives [168, 169] and 13C NMR titration of BOPTA (14) [170, 171].
Though microconstants have been published for the nitrogens of DTPA and its bis-amides [24, 25, 172], the complete resolution of the carboxylate microequilibria reaching into the strongly acidic interval is still lacking.
Fractional protonation of the “polyamine backbone” nitrogens has been determined by 1H NMR titration for higher complexone homologues as triethylenetetraminehexaacetate (TTHA) [173] and its bis-butylamides as potential radiopharmaceuticals [168, 169], tetraethylenepentamineheptaacetate (TPHA), and pentaethylenehexamineoctaacetate (PHOA) [173]. The protonation sequence is determined by the same factors as for DTPA discussed above.
In general, amine protonation of linear complexones precedes carboxylate protonation [106, 162]. The proton population of nitrogens is determined mainly by minimizing coulombic repulsion between neighboring NH+ groups and by maximizing the hydrogen-bonded rings involving terminal carboxylates. Site-specific protonation can usually be assessed with a pH-independent set of C shielding constants, although small modifications to C coefficients proposed originally by Sudmeier and Reilly become necessary for higher homologues [173].
1H NMR–pH titration curves and protonation sequences of EDTA and ethylenediaminetetrakis(methylenephosphonate) (15) are very similar, albeit the nitrogens in the latter are approximately 3 log units more basic due to the double negative charge of phosphonate groups [174]. Contrary to that, the protonation sequence of diethylenetriaminepentamethylenephosphonate (16) differs significantly from that of the carboxylate analogue DTPA, as revealed by its 1H NMR–pH titration [89]. In the pH interval 14–9, complete protonation of the terminal nitrogens occur, followed by monoprotonation of two terminal phosphonates. Significant protonation of the central nitrogen, the most basic site in DTPA, begins at pH<4 only.
13C NMR–pH titrations are of limited use to study protonation microequilibria of polyaminocarboxylates and polyaminomethylenephosphonates, since 13C protonation shifts are usually small (see ref. [89] and refs. therein).
Cyclic polyamines
Protonation sequences of polyazacycloalkanes are discussed in detail in the reviews of Bencini et al. [151] and Sroczynski et al. [175]. Plotting of overall basicities (ΣlogK) against the number of nitrogen atoms results in parallel lines for linear and cyclic polyamines with all-ethylenic chains, respectively. This fact suggests a common principle governing protonation, namely, the electrostatic repulsion of neighboring ammonium groups [151]. Elongation of the separating alkyl chains reduces this constraint [161]. While the first incoming protons usually build a hydrogen-bonded network in the interior of the macrocycle [151, 176, 177], extensive protonation results in a conformational transition to a more open structure [151]. The additivity of protonation shifts stated in Eq. (31) holds only if the protonating groups maintain a constant average orientation throughout the pH range [106]. In polyazacycloalkanes, the preferred pH-dependent conformations mean that C coefficients also vary with pH [52, 178, 179]. If protonation states of particular nitrogens can uniquely be identified at certain points of the NMR–pH titration curves, this information can be used to obtain a new, compound-specific set of C coefficients [52].
Protonation sequences of N-methylated cyclic polyamines have been derived from 1H NMR titration data [179, 180, 181, 182, 183]. Linewidth variations of methylenic protons as a function of pH have also been observed for methylated cyclic triamines, caused by slow interconversion of various conformations of the partially protonated ring [179]. For a trimethylated oxatriaza macrocycle, irregular 1H NMR–pH profiles with maxima have been observed, suggesting a redistribution of protons as described for DTPA nitrogens above [52].
NMR titrations and protonation sequences of mixed donor macrocycles containing O/S atoms, polyazacyclophanes, cryptands, and aza-cages are also covered in Bencini’s review [151]. Nazarski recently demonstrated that for the “scorpiand” ligand 1-(2′-aminoethyl)-1,4,8,11-tetraazacyclotetradecane (17), besides modern 2D NMR experiments, 1H and 13C protonation shifts could even be of value today to achieve a complete assignment of resonances [184].
Cyclic polyamines with basic pendant arms
Considerable attention has been paid to protonation sequences of cyclic polyamines bearing acetato, propionato, phosphonato, phosphinato, or hydroxamato groups on ring nitrogens.
Several studies have concluded that the parent cyclic (simple or methylated) polyamines are not as good model compounds to derive C j,N shielding constants of cyclic triaza- and tetraaza-polycarboxylates as might be expected [52, 179, 185]. Here again, characteristic points of the 1H NMR titration curve of the cyclic polyaminocarboxylate under study can help deriving C j,N coefficients [52, 179, 186], which often turn out to be pH-dependent [52, 180, 185]. In triaza, oxatriaza, and tetraaza macrocycles, the first two associating protons attach to ring nitrogens to form hydrogen bonds with pendant carboxylates. The subsequent acid equivalents protonate almost exclusively those carboxylates that are not involved in such hydrogen bonds and the remaining ring nitrogens protonate only in strong acidic medium [52, 178, 179, 180, 185, 187, 188]. Thus, internal hydrogen bonds connecting the ammonium and the side chain carboxylate groups act as additional key factors to determine protonation sequences [186, 187, 188, 189, 190], to cause protonation shift anomalies [52, 179], to indicate redistribution of protons [190], to make C j,N coefficients pH-dependent [185], or to slow down the inversion of asymmetrically positioned nitrogen atoms [179]. Slow kinetics during the first protonation step, manifested in linewidth variations, was observed only in the case of macrorings containing amide groups [191]. Group constants of pendant carboxylates have been published for propionato analogues of DOTA (18) [192]. The impacts of protonation state and conformation on metal-binding characteristics for carboxylate and carbamoyl derivatives of cyclen and cyclam (12- and 14-membered tetraazamacrocycles) have been reviewed by Meyer et al. [193].
1H NMR–pH titrations of cyclic polyamino polyphosphonates and polyphosphinates [87, 90, 194, 195] revealed that the main factors determining their protonation sequence are the same as for polycarboxylates described above. The C j,P shielding constants proved to be pH-dependent, indicating conformational changes [87, 194].
The protonation sequence of piperazine-1,4-bis(N-methylacetohydroxamate) (19) has been characterized in terms of microconstants, which suggest highly overlapping protonation of ring nitrogens and hydroxamato side-groups [112]. In contrast, in the triaza analogue 1,5,9-triazacyclododecane-N,N′,N″-tris(N-methylacetohydroxamate) (20), the first proton coordinates to a ring nitrogen, followed by the independent, nearly statistical protonation of the three hydroxamato side-groups [177]. 1-(2-(9-Anthrylmethylamino)ethyl)-1,4,7,10-tetraazacyclo-dodecane was shown to take up the first two protons to ring nitrogens, followed by protonation at the pendant arm [196].
Dendrimers
Protonation behavior of poly(propylene imine) dendrimers has been studied by 15N NMR titration [26] and the microconstants have been calculated using the cluster expansion formalism [27, 28]. Repulsive nearest-neighbor pair-interactions have been shown to govern the protonation sequence, resulting in the typical odd–even shell protonation pattern of dendrimers [26].
Antibiotics, flavonoids, and other drugs
In an early application of the Sudmeier–Reilly approach, twelve microconstants of tetracycline were determined by 1H NMR–pH titration [197]. Group constants of individual NH2 groups have been determined by 15N NMR titration for the antibiotics tobramycin, apramycin [93], and neomycin B [94]. Szilágyi et al. have controlled the tobramycin basicity data by using 1H and 13C NMR titration as well as partially N-acetylated derivatives as model compounds [198].
Microconstants of phenylephrine have been determined by automatized, 13C NMR-controlled titration by Hägele and Ollig [199] and showed good agreement with those obtained by on-line UV titration [103].
13C NMR–pH titration yielded microconstants for the first deprotonation step of catechin and epicatechin [200] and the most acidic phenol group was found to coincide with the major site of metabolization.
Miscellaneous
Microconstants for the overlapping protonating phenolate groups of 3,4-dihydroxyphenylacetic acid have been deduced from 13C NMR–pH titration data, using the O-methylated and monohydroxy model compounds [201]. Microconstants of the diprotic aminobenzoic [202], nicotinic, and isonicotinic acids [203] have been determined by 13C NMR titration using methyl-4-aminobenzoate [202] or hydroxybenzoic acids [203] as model compounds.
Probing residue-specific basicity in polypeptides and proteins
No attempt will be made here to summarize the vast literature on the NMR–pH titration of protein residues; the reader is referred to separate monographs [23]. In the following discussion, a brief summary will be given, highlighting examples for unusual side-chain basicity and cooperativity as determined by NMR titrations.
Polypeptides and proteins can contain 10–100 ionizable residues. Beyond approximately 30 groups, a direct enumeration of all possible protonation states becomes prohibitive due to combinatorial reasons [20]. In theoretical calculations, titration curves of larger proteins are handled with special numerical methods [20, 204]. In practice, the residue-specific basicity is usually characterized in terms of group constants [12] and detailed microequilibrium treatments focus only on a few side-chains of special significance, which often coincide with the catalytic ones (some examples are discussed below).
Residue-selective NMR–pH titration curves in proteins
Some protonating residues are buried in the interior of the protein molecule and are thus inaccessible to solvent. The nonzero spin nuclei in these “caves” usually exhibit no titration shifts unless they are influenced by proximate protonating residues. The residues situated on the protein surface or at the catalytic site are the main objects of protein NMR–pH titrations.
For larger peptides and proteins, it is customary to use Eq. (28) to elucidate the group constant of individual residues from NMR–pH profiles of appropriately selected “reporter” nuclei [205]. Rabenstein et al. have determined group constants for the pentadecapeptide FN-C/H II and a conotoxin G1-analogue tridecapeptide [44]. Both microconstants and group constants have been determined from 1H NMR titration of oxytocin, arginine vasopressin, and their derivatives (see the case study above, ref. [102]).
Since the aromatic 1H resonances of His and Tyr are well separated from the complicated aliphatic multiplets, earlier NMR–pH titrations focused mainly on these residues [206, 207, 208]. In fact, before the age of 2D NMR spectroscopy, protonation constants and 1H [208, 209, 210] and 13C [211] protonation shifts were used for peak assignment purposes in peptides and proteins, for example, serine proteases [208], hemerythrin [207], ribonuclease A [117, 209], myoglobins [211], and lysosime [212]. At present, pulse sequences are optimized specifically to ease protonation constant determination of particular residues [213].
1H NMR chemical shifts and group constants are especially useful to probe electrostatic and hydrophobic microenvironment of histidines and tyrosines, as demonstrated for azurin [216], bovine pancreatic ribonuclease A [123], apocytochrome c interacting with SDS micelles [214], hemoglobin [215], myoglobins [40, 41, 210, 216, 217], and subtilisin [218]. Histidine tautomerism, hydrogen bonding, protonation equilibrium, and kinetics in subtilisin BPN′ have recently been investigated by 1H, 13C, and 15N NMR titrations [95].
1H NMR–pH titration of phosphocarrier protein [219], class C β-lactamase [220] and the thermophilic protein Sso7d [45] contributed to a better understanding of the catalytic mechanism.
Group constants, when compared to “standard” basicities of the same residues in appropriately chosen model compounds are useful tools to identify salt bridges (hydrogen bonds). In this cases, logk of the more acidic residue decreases and that of the more basic increases as compared to the values in noninteracting form [12]. Changing one of the participating amino acids to a non-bridging one by site-directed mutagenesis restores the “normal” value of the remaining group constant. For a synthetic nonapeptide fragment of collagen, group constants from 1H NMR titration have lead to postulation of salt bridges, supported also by 13C protonation shifts of the corresponding residues [221]. In S-methylthio-papaine, the anomalous low basicity of His159 (logk=3.45) indicated the existence of a His–Cys ion pair [222]. Catalytic dyads have also been subject of NMR titrations, for example, His240–Asp77 in glucose 6-phosphate dehydrogenase [223]. Another interesting example of a perturbed logK value is that of the catalytic lysine in acetoacetate decarboxylase and polyamine enzyme models [224].
1H NMR titrations performed as 2D experiments to improve resolution lead to the determination of all (or nearly all) protonation constants of ribonuclease A [225], mouse epidermal growth factor [39], α-sarcin [226], and bovine β-lactoglobulin [227].
Composite NMR–pH titration curves in proteins
When a non-zero spin nucleus is influenced by several protonating groups, its NMR–pH profile often shows a biphasic (Fig. 2A) or even more complicated shape, which does not obey the sigmoidal run of a single protonation step as stated in Eq. (28). In these cases, the half-point of the titration curve, logk 1/2, has been used extensively in the past to characterize residue-specific basicity, which is, however, a qualitative feature, without matching any physically well-defined (group or micro) constants [25].
Earlier, the empirical Hill equation [228, 229, 230] was fitted instead of Eq. (28) to NMR–pH titration profiles not exhibiting the ideal, sigmoid shape:
Although the exact meaning of the Hill coefficient n could be given only recently [231], it has been used extensively as a model-free measure of cooperativity (n>1) or anticooperativity (n<1). The cooperative protonation of protein residues can be quantitated in a most straightforward way by microconstants [21, 24, 230] and interactivity parameters.
Positive cooperativity, namely the thermodynamically favored binding of the second proton upon binding the first, is rarely observed even with enzymes. For instance, two catalytic residues of fumarase bind hydrogen ions cooperatively when the enzyme is occupied by the competitive inhibitor l-tartarate [232, 233].
Site-directed mutagenesis and 13C NMR titration of isotopically labeled samples yielded microconstants for the active site Cys32, the Cys35, and the Asp26 residues of E. coli thioredoxin and its variants [234]. Previous 1H and 13C titrations revealed that Cys35 has an abnormally high thiolate basicity (logk=11.1) [235]. The microconstants of the Cys32 thiolate group bear significance on enzyme mechanism, since this residue initiates the catalysis by performing an intermolecular nucleophilic attack on the substrate. For the same reason, the nucleophilic Cys11 thiolate has an abnormally low basicity (logk=3.5; see, e.g., ref. [236]), stabilized by a Cys11–S...H–S–Cys14 hydrogen bridge. The non-exchanging bridging proton was directly observed by 1H NMR spectroscopy [237]. A correlation has been demonstrated between the site-specific basicity of the catalytic thiolate group and the redox potential of thiol-disulfide oxidoreductases [238].
Microconstants have been derived from the 13C NMR–pH titration of two catalytic, selectively 13C-labeled glutamyls of xylanase, the nucleophile Glu78 and the acid–base catalyst Glu172 [239]. 1H NMR–pH titration of wild and mutant ribonuclease A yielded microconstants for the catalytic imidazoles of His12 and His119 [240]. The microconstants quantitate that (a) the Asp121 of the catalytic site modulates only slightly the intrinsic basicity of His119 and (b) the negative cooperativity of His12 and His119 in the unliganded enzyme [241] changes to positive cooperativity upon binding the reaction product 3′-UMP [240].
References
Wishart DS, Sykes BD (1994) Methods Enzymol 239:363–392
Günther H (1992) NMR spectroscopy. Thieme, Stuttgart
Sudmeier JL, Evelhoch JL, Jonsson NBH (1980) J Magn Reson 40:377–390
Bányai I, Blixt J, Glaser J, Tóth I (1992) Acta Chem Scand 46:142–146
Zekri O, Boudeville P, Genay P, Perly B, Braquet P, Jouenne P, Burgot JL (1996) Anal Chem 68:2598–2604
Gutowsky HS, Saika A (1953) J Chem Phys 21:1688–1694
Grunwald E, Loewenstein A, Meiboom S (1957) J Chem Phys 27:641–642
Loewenstein A, Roberts JD (1960) J Am Chem Soc 82:2705–2710
King EJ (1965) Acid-base equilibria. Pergamon, Oxford
Bjerrum N (1923) Z Phys Chem 56:219–242
Noszál B, Burger K (1979) Acta Chim Acad Sci Hung 100:275–288
Noszál B (1986) J Phys Chem 90, 4104–4110
Noszál B (1990) Acid-base properties of bioligands. In: K Burger (ed) Biocoordination chemistry: coordination equilibria in biologically active systems. Ellis Horwood, Chichester
Noszál B, Sándor P (1989) Anal Chem 61:2631–2637
Edsall JT, Wyman J (1958) Biophysical chemistry. Academic Press, New York
Martell AE, Motekaitis RJ (1988) The determination and use of stability constants, Chap 7. VCH, New York
Szakács Z, Noszál B, J Math Chem 26, 139 (1999)
Noszál B, Szakács Z (2003) J Phys Chem B 107:5074–5080
Borkovec M, Koper GJM (1994) 98:6038–6045
Borkovec M, Jönsson B, Koper GJM (2000) Processes and proton binding in polyprotic systems: small molecules, proteins, interfaces and polyelectrolytes. In: Matijevic E (ed) Surface and colloid science, Vol 16. Plenum Press, New York
Shrager RI, Cohen JS, Heller SR, Sachs DH, Schechter AN (1972) Biochemistry 11:541–547
Rabenstein DL (1973) J Am Chem Soc 95:2797–2803
Sarneski JE, Reilly CN (1977) The determination of proton binding sites by NMR-titrations. In: E. Wänninen (ed) Essays on analytical chemistry. Pergamon, Oxford
Onufriev A, Case DA, Ullmann GM (2001) Biochemistry 40:3413–3419
Ullmann GM (2003) J Phys Chem B 107:1263–1271
Koper GJM, van Genderen MHP, Elissen-Roman C, Baars MWPL, Meijer EW, Borkovec M (1997) J Am Chem Soc 119:6512–6521
Borkovec M, Koper GJM (2000) Anal Chem 72:3272-3279
Borkovec M, Koper GJM (2002) Chimia 56:695–701
Glasoe PK, Long FA (1960) 64:188–190
Baucke FGK (1998) J Phys Chem B 102:4835–4842
Wiberg KB (1966) Physical organic chemistry, Chaps 2–7. Wiley, New York
Paabo M, Bates RG (1969) Anal Chem 41:283–285
Bundi A, Wüthrich K (1979) 18:285–297
Delgado R, Da Silve JJRF, Amorim MTS, Cabral MF, Chaves S, Costa J (1991) Anal Chim Acta 245:271–282
Hore PJ (1983) J Magn Reson 55:283–300
Hore PJ (1989) Methods Enzymol 176:64–77
Larive CK, Rabenstein DL (1991) Magn Reson Chem 29:409-417
Price WS (1999) Annu Rep NMR Spectrosc 38:289–354
Kohda D, Sawada T, Inagaki F (1991) Biochemistry 30:4896–4900
Cocco MJ, Kao YH, Phillips AT, Lecomte JTJ (1992) Biochemistry 6481–6491
Bashford D, Case DA, Dalvit C, Tennant L, Wright PE (1993) Biochemistry 32:8045–8056
Kesvatera T, Jönsson B, Thulin E, Linse S (1996) J Mol Biol 259:828–839
Chen HA, Pfuhl M, McAlister MSB, Driscoll PC (2000) Biochemistry 39:6814–6824
Rabenstein DL, Hari SP, Kaerner A (1997) Anal Chem 69:4310–4316
Consonni R, Arosio I, Belloni B, Fogolari F, Fusi P, Shehi E, Zetta L (2003) Biochemistry 42:1421–1429
Wang C, Gao H, Gaffney BL, Jones RA (1991) J Am Chem Soc 113:5486–5488
Holmes DL, Lightner DA (1995) Tetrahedron 51:1607–1622
Holmes DL, Lightner DA (1996) Tetrahedron 52:5319–5338
Koeberg-Telder A, Cerfontain H (1975) J Chem Soc Perkin Trans 2 226–229
De Marco A (1977) J Magn Res 26:527–528
Rabenstein DL, Greenberg MS, Evans CA (1977) Biochemistry 16:977-981
Amorim MTS, Ascenso JR, Delgado R, Faústo da Silva JJR (1990) J Chem Soc Dalton Trans 3449–3455
Szakács Z (2002) PhD thesis, Heinrich-Heine-University Düsseldorf
Ollig J, Hägele G (1995) Computers Chem 19:287–294
G Hägele (1994) NMR controlled titrations of phosphorus-containing acids and bases in protolysis and complex formation. In: Quin LD, Verkade JG (eds) Phosphorus-31P-NMR spectral properties in compound characterization and structural analysis. VCH-Verlag, Weinheim
Hägele G, Szakács Z, Ollig J, Hermens S, Pfaff C (2000) Heteroatom Chem 11:562–582
Uhlemann CE, Pfaff CG, Hägele G (2002) Magn Reson Chem 40:573–580
Hermens S (2002) PhD thesis, Heinrich-Heine-University Düsseldorf
Rabenstein DL, Mariappan SVS (1993) J Org Chem 58:4487–4489
Perrin CL, Fabian MA, Armstrong KB (1994) J Org Chem 59:5246–5253
Perrin CL, Fabian MA (1996) Anal Chem 68:2127–2134
Pehk T, Kiirend E, Lippmaa E, Ragnarsson U (1996) J Chem Soc Perkin Trans 2 2351–2357
Perrin CL, Rivero IA (1999) Rev Sci Instr 70:2173–2174
Allred AL, Rochow EG (1957) J Am Chem Soc 79:5361–5365
Karplus M, Pople JA (1963) J Chem Phys 38:2803–2807
Buckingham AD (1960) Can J Chem 38:300–307
Perczel A, Császár A (2001) Chem Eur J 7:1069–1083
Sterk H, Holzer H (1974) Org Magn Reson 6:133–143
Hagen R, Roberts JD (1969) J Am Chem Soc 91:4504–4506
Horsley WJ, Sternlicht H (1968) J Am Chem Soc 90:3738–3748
Keire DA, Robert JM, Rabenstein DL (1992) J Org Chem 57:4427–4431
Tribolet R, Sigel H (1987) Eur J Biochem 163:353–363
Blindauer CA, Holy A, Dvorakova H, Sigel H (1997) J Chem Soc Perkin 2 2353–2363
Felemez M, Marwood RD, Potter BVL, Spiess B (1999) Biochem Biophys Res Comm 266:334–340
Cistola DP, Small DM, Hamilton JA (1982) Biochemistry 23:795–799
Batchelor JG, Prestegard JH, Cushley RJ, Lipsy SR (1973) J Am Chem Soc 95:6358-6364
Batchelor JG, Feeney J, Roberts GCK (1975) J Magn Reson 20:19–38
Quirt AR, Lyerla JR Jr, Peat IR, Cohen JS, Reynold WR, Freedman MH (1974) J Am Chem Soc 96:570–574
Batchelor JG (1975) J Am Chem Soc 97:3410-3415
Rabenstein DL, Sayer TL (1976) J Magn Reson 24:27–39
Surprenant HL, Sarneski JE, Key RR, Byrd JT, Reilly CN (1980) J Magn Reson 40:231–243
Nelson DJ, Yeagle PL, Miller TL, Martin RB (1976) Bioinorg Chem 5:353–358
Sarneski JE, Surprenant HL, Molen FK, Reilley CN (1975) Anal Chem 47:2116–2124
Dagnall SP, Hague DN, McAdam ME (1984) J Chem Soc Perkin Trans 2, 435–440
Letcher JH, Van Wazer JR (1966) J Chem Phys 45:2926–2930
Moedritzer I (1967) Inorg Chem 5:936–939
Geraldes CFGC, Sherry AD, Cacheris WP (1989) Inorg Chem 28:3336–3341
Jaffe EK, Kohn M (1978) Biochemistry 17:652–657
Gillard RD, Newman PD (1989) Polyhedron 8:2077–2086
Rohovec J, Kyvala M, Vojtisek P, Hermann P, Lukes I (2000) Eur J Inorg Chem 195–203
Levy GC, Lichter RL (1979) Nitrogen-15 nuclear magnetic resonance spectroscopy. Wiley, New York
Witanowski M, Stefaniak L, Webb GA (1986) Annu Rep NMR Spectrosc 18:1–756
Dorman DE, Paschal JW, Merkel KE (1976) J Am Chem Soc 98:6885–6888
Botto RE, Coxon B (1983) J Am Chem Soc 150:1021–1028
Day RM, Thalhauser CJ, Sudmeier JL, Vincent MP, Torchilin EV, Sanford DG, Bachovchin CW, Bachovchin WW (2003) Protein Sci 12:794–810
Gertl JA, Demou PC, Mehdi S (1982) 104:2848–2856
Gerlt JA, Reynolds MA, Demou PC, Kenyon GL (1983) 105:6469–6474
Reynolds MA, Gerlt JA, Demou PC, Oppenheimer NJ, Kenyon GL (1983) 105:6475–6481
Gajda T, Henry B, Delpuech JJ (1994) J Chem Soc Perkin Trans 2, 157-164
Forsyth DA, Yang JR (1986) J Amer Chem Soc 108:2157-2161
Harper JL, Smith RAJ, Bedford JJ, Leader JP (1997) Tetrahedron 53:8211–8224
Noszál B, Guo W, Rabenstein DL (1992) J Org Chem 57:2327–2334
Arendt C, Hägele G (1995) Comput Chem 19:263–268
Burger K, Sipos P, Véber M, Horváth I, Noszál B, Lőw M (1988) Inorg Chim Acta 152:233–239
Rabenstein DL, Sayer TL (1976) Anal Chem 48:1141–1146
Sudmeier JL, Reilly CN (1964) Anal Chem 36:1698-1706
Sayer TL, Rabenstein DL (1976) Can J Chem 54:3392-3403
Jameson RF, Hunter G, Kiss T (1980) J Chem Soc Perkin Trans 2 1105–1110
Kiss T, Tóth B (1982) Talanta 29:539–544
Farkas E, Kiss T, Kurzak B (1990) J Chem Soc Perkin Trans 2 1255–1257
Benesch RE, Benesch R (1955) J Am Chem Soc 77:5877–5881
Santos MA, Esteves MA, Vaz MC, Fraústo da Silva JJR, Noszál B, Farkas E (1997) J Chem Soc Perkin Trans 2 1977–1983
D’Angelo JC, Collette TW (1997) 69:1642–1650
Mernissi-Arifi K, Schmitt L, Schlewer G, Spiess B (1995) Anal Chem 67:2567–2574
Sørensen MD, Led JJ (1994) Biochemistry 33:13727–13733
Jameson RF, Hunter G, Kiss T (1978) J Chem Soc Chem Comm 768–769
Bradbury JH, Scheraga HA (1966) J Am Chem Soc 88:4240–4246
Gajda T, Henry B, Aubry A, Delpuech JJ (1996) Inorg Chem 35:586–593
Walters DB, Leyden DE (1974) Anal Chim Acta 72:275–283
Noszál B, Rabenstein DL (1991) J Phys Chem 95:4761–4765
Tanokura M, Tasumi M, Miyazawa T (1976) Biopolymers 15:393–401
Tanokura M (1983) Biochim Biophys Acta 742:576–585
Tanokura M (1983) Biochim Biophys Acta 742:586–596
Tanokura M (1983) Biopolymers 22:2563–2576
Kawano K, Kyogoku Y (1975) Chem Lett 1305–1308
Blomberg F, Maurer W, Rüterjans H (1977) J Am Chem Soc 99:8149–8159
Roberts JD, Yu C, Flanagan C, Birdseye TR (1982) 104:3945–3949
Reynolds WF, Peat IR, Freedman MH, Lyerla JR Jr (1973) J Amer Chem Soc 95:328–331
Deslauriers R, McGregor WH, Sarantakis D, Smith ICP (1974) Biochemistry 13:3443–3448
Wasylishen RE, Tomlinson G (1977) Can J Biochem 55:579–582
London RE (1978) J Chem Soc Chem Comm 1070–1071
Friedrich JO, Wasylishen RE (1986) Can J Chem 64:2132–2138
Henry B, Tekely P, Delpuech JJ (2002) J Am Chem Soc 124:2025–2034
Martin RB (1985) Acc Chem Rev 18:32–38
Barbarella G, Bertoluzza A, Tugnoli V (1987) Magn Reson Chem 25:864–868
Sowers LC, Fazakerly GV, Kim H, Dalton L, Goodman MF (1986) Biochemistry 25:3983–3988
Crisponi G, Nurchi V, Casu M, Lai A (1993) Spectrochim Acta A 49:1643–1649
Pilatus U, Mayer A, Offermann W, Leibfritz D (1987) Biochim Biophys Acta 926:106–113
Wang C, Gao H, Gaffney BL, Jones RA (1991) 113:5486–5488
Szpoganicz B, Martell AE (1984) 106:5513–5521
Berridge MJ (1993) Nature 361:315–325
Worley PF, Baraban JM, Supattapone S, Wilson VS, Snyder SH (1987) J Biol Chem 262:12132–12136
Schmitt L, Bortmann P, Schlewer G, Spiess B (1993) J Chem Soc Perkin Trans 2 2257–2263
Mernissi-Arifi K, Ballareau S, Schlewer G, Spiess B, Zenkouar M (1996) New J Chem 20:1087–1092
Mernissi-Arifi K, Schlewer G, Spiess B (1998) 308:9–17
Felemez M, Schlewer G, Jenkins DJ, Correa V, Taylor CW, Potter BVL, Spiess B (1999) Carbohydr Res 322:95–101
Felemez M, Bernard P, Schlewer G, Spiess B (2000) J Am Chem Soc 122:3156–3165
Dozol H, Blum-Held C, Guédat P, Maechling C, Lanners S, Schlewer G, Spiess B (2002) J Mol Struct 643:171–181
Pasini A, Perego P, Balconi M, Lupatini M (1995) J Chem Soc Dalton Trans 575–578
Bebot-Brigaud A, Dange C, Fauconnier N, Gérard C (1999) J Inorg Biochem 75:71–78
Bencini A, Bianchi A, Garcia-Espana E, Micheloni M, Ramirez JA (1999) Coord Chem Rev 188:97–156
Dagnall SP, Hague DN, McAdam ME (1984) J Chem Soc Perkin Trans 2, 1111–1114
Kimberly MM, Goldstein JH (1981) 53:789–793
Aikens DA, Bunce SC, Onasch OF, Schwartz HM, Hurwitz C (1983) J Chem Soc Chem Commun 43-45
Hague DN, Moreton AD (1994) J Chem Soc Perkin Trans 2 265–270
Onasch F, Aikens D, Bunce S, Schwartz H, Nairn D, Hurwitz C (1984) Biophys Chem 19:245–253
Takeda Y, Samejima K, Nagano K, Watanabe M, Sugeta H, Kyogoku Y (1983) Eur J Biochem 130:383-389
Jaroszewski JW, Matzen L, Frolund B, Krogsgaard-Larsen P (1996) J Med Chem 39:515–521
Newton GL, Dwyer TJ, Kim T, Ward JF, Fahey RC (1992) Radiat Res 131:143–151
Aikens DA, Bunge SC, Onasch OF, Parker R, Hurwitz C, Clemans S (1983) Biophys Chem 17:67–74
Aguilar JA, Garcia-Espana E, Guerrero JA, Luis SV, Llinares JM, Ramírez JA, Soriano C (1996) Inorg Chim Acta 246:287–294
Letkeman P, Martell AE (1979) Inorg Chem 18:1284–1289
Fujiwara Y, Reilly CN (1968) Anal Chem 40:890–894
Sawyer DT, Tackett JE (1963) J Am Chem Soc 85:314–316
Chapman D, Lloyd DR, Prince RH (1963) J Chem Soc 3645–3658
Kula RJ, Sawyer DT (1963) Inorg Chem 3:458–458
Day RJ, Reilly CN (1964) Anal Chem 36:1073–1076
Geraldes CFGC, Urbano AM, Alpoim MC, Sherry AD, Kuan KT, Rajagopalan R, Maton F, Muller RN (1995) Magn Res Imag 13:401–420
Imura H, Choppin GR, Cacheris WP, de Learie LA, Dunn TJ, White DH (1997) Inorg Chim Acta 258:227–236
Uggeri F, Aime S, Anelli PL, Botta M, Brocchetta M, de Haen C, Ermondi G, Grandi M, Paoli P (1995) Inorg Chem 34:633–642
Alderighi L, Bianchi A, Biondi L, Calabi L, De Miranda M, Gans P, Ghelli S, Losi P, Paleari L, Sabatini A, Vacca A (1999) J Chem Soc Perkin 2 2741–2745
Lammers H, van der Heijden AM, van Bekkum H, Geraldes CFGC, Peters JA (1998) 277:193–201
Letkeman P, Westmore JB (1971) Can J Chem 49:2096–2102
Rizkalla EN, Choppin GR (1983) Inorg Chem 22:1478–1482
Sroczynski D, Grzejdziak A, Nazarski RB (1999) J Incl Phenom Macro 35:251–260
Bell TW, Choi HJ, Harte H (1986) J Am Chem Soc 108:7427–7428
Esteves MA, Vaz MCT, Goncalves MLSS, Farkas E, Santos MA (1995) J Chem Soc Dalton Trans 2565–2573
Desreux JF, Merciny E, Loncin MF (1981) Inorg Chem 20:987–991
Geraldes CFGC, Sherry AD, Marques MPM, Alpoim MC, Cortes S (1991) J Chem Soc Perkin Trans 2 137–146
Geraldes CFGC, Alpoim MC, Marques MPM, Sherry AD, Singh M (1985) Inorg Chem 24:3876–3881
Ciampolini M, Micheloni M, Nardi N, Paoletti P, Dapporto P, Zanobini F (1984) J Chem Soc Dalton Trans 1357–1362
Bencini A, Bianchi A, Garcia-Espana E, Fusi V, Micheloni M, Paoletti P, Ramirez JA, Rodriguez A, Valtancoli B (1992) J Chem Soc Perkin Trans 2 1059–1065
Andres A, Bazzicalopi C, Bencini A, Bianchi A, Fusi V, Garcia-Espana E, Giorgi C, Nardi N, Paoletti P, Ramirez JA, Valtancoli B (1994) J Chem Soc Perkin Trans 2 2367–2373
Nazarski RB (2003) Magn Reson Chem 41:70–74
Ascenso JR, Delgado R, Fraústo da Silva JJR (1985) J Chem Soc Perkin 2 781–788
Geraldes CFGC, Brücher E, Cortes S, Koenig SH, Sherry AD (1992) J Chem Soc Dalton Trans 2517–2521
Kumar K, Chang CA, Francesconi LC, Dischino DD, Malley MF, Gougoutas JZ, Tweedle MF (1994) Inorg Chem 33:3567–3575
Kumar K, Jin T, Wang X, Desreux JF, Tweedle MF (1994) Inorg Chem 33:3823–3829
Wambeke DM, Lippens W, Herman GG, Goeminne AM, Van De Vondel D, Van Der Kelen GP (1992) Polyhedron 11:1305–1313
Aime S, Botta M, Crich SG, Giovenzana GB, Jommi G, Pagliarin R, Sisti M (1997) Inorg Chem 36:2992–3000
Inoue MB, Oram P, Inoue M, Fernando Q (1995) Inorg Chim Acta 232:91–98
Keire DA, Jang YH, Li L, Dasgupta S, Goddard WA, Shively JE (2001) Inorg Chem 40:4310–4318
Meyer M, Dahaoui-Gindrey V, Lecomte C, Guilard L (1998) Coord Chem Rev 178:1313–1405
Clegg W, Iveson PB, Lockhart JC (1992) J Chem Soc Dalton Trans 3291–3298
Lukes I, Kotek J, Vojtisek P, Hermann P (2001) 216:287–312
Aoki S, Kaido S, Fujioka H, Kimura E (2003) Inorg Chem 42:1023–1030
Rigler NE, Bag SP, Leyden DE, Sudmeier JL, Reilly CN (1965) Anal Chem 872–875
Szilágyi L, Pusztahelyi SzZ, Jakab S, Kovács I (1993) Carbohydr Res 247:99-109
Hägele G, Holzgrabe U (1999) pH-dependent NMR measurements. In: Holzgrabe U, Wavez I, Diehl B (eds) NMR spectroscopy in drug development and analysis.Wiley, Weinheim
Cren-Olivé C, Wieruszeski JM, Maes E, Rolando C (2002) Tetrahedron Lett 43:4545–4549
Ishimitsu T, Fujiwara Y, Hirose S (1979) Talanta 26:67–69
Zhang XX, Oscarson JL, Izatt RM, Schuck PC, Li D (2000) J Phys Chem B 104:8598–8605
Schwartz LM, Gelb RI, Mumford-Zisk J, Laufer DA (1987) J Chem Soc Perkin 2 453–460
Sandberg L, Edholm O (1999) Proteins Struct Funct Genet 36:474–483
Giralt E, Viladrich R, Pedroso E (1983) Org Magn Res 21:208–213
Hill HA, Smith BE (1979) J Inorg Biochem 11:79–93
York JL, Millett FS, Minor LB (1980) Biochemistry 19:2583–2588
Chen HA, Pfuhl M, McAlister MSB, Driscoll PC (2000) Biochemistry 39:6814–6824
Markley JL (1975) Biochemistry 14:3546–3554
Carver JA, Bradbury JH (1984) Biochemistry 21:4890–4905
Wilbur DJ, Allerhand A (1976) J Biol Chem 251:5187–5194
Shindo H, Cohen JS (1976) Proc Natl Acad Sci USA 73:1979–1983
Tollinger M, Forman-Kay JD, Kay LE (2002) J Amer Chem Soc 124:5714–5717
Snel MME, Kaptein R, de Kruijff B (1991) Biochemistry 30:3387–3395
Russu IM, Ho NT, HO C (1982) Biochemistry 21:5031–5043
Ugurbil K, Norton RS, Allerhand A, Bersohn R (1977) Biochemistry 5:886–894
Bradbury JH, Carver JA (1984) Biochemistry 23:4905–4913
Jordan F, Polgar L, Tous G (1985) Biochemistry 24:7711–7717
Rösch P, Kalbitzer HR, Schmidt-Aderjan U, Hengstenberg W (1981) Biochemistry 20:1599–1605
Kato-Toma Y, Iwashita T, Masuda K, Oyama Y, Ishiguro M (2003) Biochem J 371:175–181
Gervais M, Commenges G, Laussac JP (1987) Magn Reson Chem 25:594–599
Johnson FA, Lewis SD, Shafer JA (1981) Biochemistry 20:44–48
Cosgrove MS, Loh SN, Ha JH, Levy HR (2002) Biochemistry 41:6939–6945
Highbarger LA, Gerlt JA, Kenyon GL (1996) Biochemistry 35:41-46
Baker WR, Kintanar A (1996) Arch Biochem Biophys 327:189–199
Pérez-Canadillas JM, Campos-Olivas R, Lacadena J, del Pozo AM, Gavilanes JG, Santoro J, Rico M, Bruix M (1998) Biochemistry 37:15865–15876
Fogolari F, Ragona L, Licciardi S, Romagnoli S, Michelutti R, Ugolini R, Molinari H (2000) Proteins Struct Funct Genet 39:317–330
Hill AV (1910) J Physiol (London) 40:4–7
Markley J (1973) Biochemistry 12:2245–2249
Markley J (1975) Acc Chem Res 8:70–80
Acerenza L, Mizraji E (1997) Biochim Biophys Acta 1339:155–166
Wigler PW, Alberty RA (1960) 82:5482–5488
Alberty RA (2000) J Phys Chem B 104:9929–9934
Chivers PT, Prehoda KE, Volkman BF, Kim BM, Markley JL, Raines RT (1997) Biochemistry 36:14985–14991
LeMaster DM (1996) Biochemistry 35:14876–14881
Foloppe N, Sagemark J, Nordstrand K, Berndt KD, Nilsson L (2001) J Mol Biol 310:449–470
Nordstrand K, Aslund F, Meunier S, Holmgren A, Otting G, Berndt KD (1999) FEBS Lett 449:196–200
Grauschopf U, Winther JR, Korber P, Zander T, Dallinger P, Bardwell JC (1995) Cell 83:947–955
McIntosh LP, Hand G, Johnson PE, Joshi MD, Körner M, Plesniak LA, Ziser L, Wakarchuk WW, Withers SG (1996) Biochemistry 35:9958–9966
Quirk DJ, Raines RT (1999) Biophys J 76:1571–1579
Markley JL, Finkenstadt WR (1975) Biochemistry 14:3562–3566
Acknowledgements
This work was supported by the grant OTKA T43579. Z. Sz. is indebted to Prof Gerhard Hägele (Heinrich Heine University Düsseldorf) for invaluable mentoring during the DAAD scholarship years.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Szakács, Z., Kraszni, M. & Noszál, B. Determination of microscopic acid–base parameters from NMR–pH titrations. Anal Bioanal Chem 378, 1428–1448 (2004). https://doi.org/10.1007/s00216-003-2390-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-003-2390-3