Abstract
A liquid chromatography method was developed for the determination of antifungal/antimicrobial proteins Rs-AFP1 and Dm-AMP1 in sandy loam soils. The extraction of these highly basic proteins was achieved by mechanical shaking with aqueous Tris buffer pH 9 containing guanidinium thiocyanate salt (4.1 M), EDTA and nonionic polyoxyethylene 20 cetyl ether, Brij-58 detergent. The extracts were cleaned up on Oasis HLB polymer solid-phase extraction cartridges and quantified by liquid chromatography fluorescence detection based on the fluorescence properties of the tryptophan content of these proteins. The detector response was linear for 0.3–10 μg mL−1. Procedural recoveries were tested in the range 10–100 mg kg−1. The limit of quantification was 10 mg kg−1 protein in the soil sample representing the lowest validated fortification level. The antifungal proteins were found to be stable in soil extract tested up to 9 days when stored at 4 °C.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Antifungal and antimicrobial proteins and polypeptides have been isolated from diverse groups of organisms, including plants, fungi, bacteria and insects. These proteins have a role in the host defence of plants by acting as natural defensins [1, 2, 3]. Seeds carrying fungicidal properties have been researched extensively to implant increased fungal resistance properties into commercial crops. Such proteins were identified from radish Raphanus sativus Rs-AFP1 and dahlia Dahlia merckii Dm-AMP1 [4], onion seeds Alium cepa Ace-AMP1 [5], Brassicaceae [6] and several other species [7]. Antifungal proteins are usually isolated and purified from the appropriate seeds or can be produced and isolated from biochemical fermentation of transgenic microorganisms such as yeast [8] and Pichia pastoris [9]. These active macromolecules are likely to be excreted into the soil at seed germination, thereby creating a microenvironment inhibiting fungal growth [1]. It is important to understand the fate of these pesticidal molecules in the soil environment, especially in circumstances where those compounds may be disposed in relatively large quantities from food processing plants. The objective of this study was to develop a method to monitor the fate of antifungal bioactive macromolecules in soil. Unlike for pesticides, limited publications are in the public domain related to the extraction, clean-up and determination of naturally occurring proteins [10] or pesticidal macromolecules [11]. Several papers reported investigations into the quantification and persistence of insecticidal proteins in soil derived from transgenic cotton that produces Bacillus thuringiensis var. kurstaki δ-endotoxin (Bt toxin 66 kDa) [12, 13]. The adsorption of Bt toxin to various soil constituents was discussed in several papers [14, 15, 16, 17, 18] but very little was published for quantitative assays of proteins in soil [19, 20]. The aim of this investigation was to establish an efficient extraction technique from soil and a quantification assay of antifungal protein Rs-AFP1 (5.5 kDa, calculated pI 8.3) and antimicrobial protein Dm-AMP1 (5.5 kDa, calculated pI 7.6) by using chromatographic techniques. Other techniques have been used to assess antifungal or antimicrobial proteins (e.g. enzyme-linked immunosorbent assays ELISA [21, 22]) but we have found none that quantitatively assess these proteins in soil by liquid chromatography. From aqueous solutions these proteins, especially Dm-AMP1, are susceptible to adsorption and other physicochemical interactions to various soil components, even to extraction vessels and pipettes; therefore, the handling of the extracts is itself a challenge. During the method development electrostatic and nonspecific interactions between proteins soil clay, silt, sand and humic organic matter were considered in order to achieve efficient extraction and keep the extracted proteins in solution. The limited availability of purified standards, no more than a few mg reference materials, forced the methodology towards miniaturisation in comparison with the study design and analysis of conventional pesticides. This limitation also restricted further evaluation of the specific interactions between protein soil constituents.
Experimental
Reagents and solvents
All reagents were analytical grade and solvents were HPLC grade. Guanidine thiocyanate, polyoxyethylene 20 cetyl ether (Brij-58), ethylenediaminetetraacetic acid disodium salt (EDTA), tris(hydroxymethyl)aminomethane (Trisma base), tris(hydroxymethyl)aminomethane hydrochloride (Trisma hydrochloride) were from Sigma–Aldrich Co. (Poole, Dorset, UK). Triethylamine (TEA) and trifluoroacetic acid (TFA) were from Fisher Scientific (Loughborough, Leicestershire, UK). Waters Oasis HLB extraction cartridges (60 mg in 3-mL cartridge) (Waters, Elstree, Hertfordshire, UK) were used. Deionised water was obtained from a Milli Q water purification system (Millipore Ltd., Watford, Hertfordshire, UK). The aqueous extraction solution comprised Tris buffer (32 mM) pH 9, Brij-58 (0.32%), EDTA (1.3 mM) and guanidine thiocyanate (4.1 M).
Plant defensin proteins
Samples of Rs-AFP1 and Dm-AMP1 were available as reference standards of known purity. These were prepared and characterised in-house by Syngenta Discovery and Analytical Sciences. Samples of reference materials were dissolved in sterile water for fortification and analysis.
Chromatographic system
Chromatography column
Jupiter C5, 5 μm, 30-nm pore size, 150 mm×4.6 mm coupled to a SecurityGuard column containing two 4 mm×3 mm wide-pore C5 guard cartridges (Phenomenex UK Ltd., Macclesfied, Cheshire, UK).
Mobile phase and gradient
Solvent A was water + 0.5% TEA + 0.4% TFA pH 2.2; solvent B was acetonitrile + 0.5% TEA + 0.4% TFA. Gradient: initial 12% B then linear gradient to 47% B at 8.75 min, 47% B at 8.76 min, linear gradient to 95% B at 20 min, 99% B from 21.1–25 min and kept. Column re-equilibration time was 5 min at 12% B. The flow rate was 1 mL min−1. Injection volumes were 14 μL.
Liquid chromatograph
Agilent 1100 system equipped as follows: quaternary pump (G1311A), degasser (G13322A), column heater (G1316A) at 40 °C, chilled auto sampler at 4 °C (G1313A), fluorescence detector (1046A) excitation 278 nm, emission 350 nm, UV detector (G1314A) at 280 nm and ChemStation instrument control and data processor (Rev. A.05.01).
Sample preparation
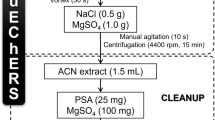
Three different representative European soils (sandy loam, sandy clay loam and loamy sand) of 10–15 kg were sampled. The upper 5 cm soil containing the turf was removed and it was sampled in 5–20 cm depth. Each sample was thoroughly mixed, air dried, stones and root debris were removed and sieved through a 2-mm sieve to achieve homogeneity. A sub-sample was sent for soil characterisation and biomass determination. The major soil characteristics were in the range pH 6.2–7.4, organic matter content 2.8–4.6%, cation exchange capacity 5.3–12.9 meq/100 g. These soil types were selected because they are routinely used for regulatory soil degradation studies in our laboratories. A representative sub-sample of soil (1 g) was weighed into a screw cap vial (15 mL). The water content of the soil was adjusted to its 40% moisture holding capacity, homogenised by shaking and equilibrated for 30 min prior to fortification and/or extraction. Samples were extracted with extraction solution (6 mL) [Tris buffer (32 mM) pH 9, Brij-58 (0.32%), EDTA (1.3 mM) and guanidine thiocyanate (4.1 M)] by agitating for 60 min on a laboratory wrist shaker. The extract was separated from solid debris by centrifugation at 3,500 rpm for 5 min. The soil pellet was re-extracted with the buffer (4 mL) as above. The combined turbid extracts were further centrifuged at 3,500 rpm for 10 min before clean-up.
Sample clean-up was performed on Oasis HLB solid-phase extraction (SPE) cartridges. The cartridges were conditioned with acetonitrile (2.5 mL) containing TEA (15 mM) and TFA (0.22%) and then washed with water (2.5 mL) containing TEA (15 mM) and TFA (0.22%). Sample aliquots (0.25 g in 2.5 mL extract) were applied onto the column and percolated through under gravity at approximately 1 drop s−1. The column was washed first with water (3×1 mL) containing TEA (15 mM) and 0.22% TFA followed by 10% acetonitrile in water (2×0.5 mL) containing TEA (15 mM) and TFA (0.22%) under gravity at a speed of approximately 1–2 drops s−1 ensuring that the sorbent was always kept moist. The proteins were eluted with 30% acetonitrile in water (2×0.55 mL) containing TEA (15 mM) and TFA (0.22%) pH 2.2 at a slow speed of 1 drop s−1 and collected into a pre-weighed vial (1.5-mL size). The last drops of the solution retained by the SPE sorbent were collected by applying gentle vacuum or pressure. The samples were concentrated to 0.4–0.6 mL using a stream of nitrogen at ≤45 °C. The sample concentration was calculated by weight assuming a density of 1 g mL−1.
Results and discussion
During the method development careful consideration was given to the handling of protein macromolecules during preparation and analysis. The major parameters considered during the optimisation of extraction were salt, pH, organic additives, surfactants and temperature. At pH values below the pI of the antifungal proteins there is a potential for strong electrostatic interactions with negatively charged components in the soil and extraction vessels. Therefore various salts (e.g. potassium chloride, guanidium chloride, calcium nitrate and guanidium thiocyanate) were investigated to compete with these interactions. Guanidium thiocyanate was found to be the most effective over a concentration range of 0.1–4.1 M, while the others caused precipitation at higher salt concentration and posed various other technical limitations. This supports earlier observations of the stabilising effect of guanidium thiocyanate upon solutions of peptides and proteins [20]. Organic additives for example ethanol, isopropanol, acetonitrile and various nonionic (Tween 20, Brij-58, Brij-35) and ionic detergents (sodium dodecyl sulfate and hexadecyltrimethylammonium bromide) were explored to minimise the secondary, van der Waals, interactions to nonpolar surfaces. Organic additives failed to prevent the "stickiness" of the AFPs and restricted the solubility of the proteins. The ionic detergents would not be compatible with the liquid chromatography; therefore a nonionic monodisperse detergent polyoxyethylene 20 cetyl ether was selected (Brij-58). Various buffers were also tested in the pH range 2–11. The extraction efficiency was mainly controlled by the relatively high concentration (4.1 M) of guanidium thiocyanate. The pH had little effect on the efficiency: the alkaline range pH 8–11 was more favourable; therefore pH 9 Tris buffer was selected, since this was compatible with proteins. The antifungal proteins are heat-stable at 60–80 °C [4], so extraction at 60 °C was also tried. No benefit was found at this elevated temperature versus ambient temperature; however, the robustness of the method with regards to temperature was established. EDTA was added to the extraction system as a precautionary measure to complex polyvalent ions rather than letting them strongly interact with the proteins. Eventually the extraction mixture containing Tris buffer (32 mM) pH 9, Brij-58 (0.32%), EDTA (1.3 mM) and guanidine thiocyanate (4.1 M) proved to be suitable for the extraction of antifungal proteins from three different textural soil classifications, sandy loam, sandy clay loam and loamy sand soils. The high salt and other lipophilic co-extractives were separated on polymer-based solid-phase extraction cartridges (polyvinyl pyrrolidine polystyrene co-polymer Oasis HLB) with a separation profile mimicking the reversed-phase liquid chromatography. It was found that the TEA additive was essential to minimise the "stickiness" of these compounds while maintaining the overall pH of the systems at pH 2.2 using TFA in excess.
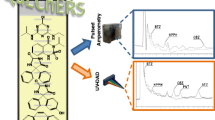
The analysis was performed on several Jupiter C5 columns using fluorescence detection. Variation of peak tailing was observed from column to column when conventional mobile phases containing 0.1% TFA only were used. It was found that tailing could be avoided by the addition of 0.5% TEA in the mobile phase while maintaining the overall acidic pH by use of a slight excess of TFA.
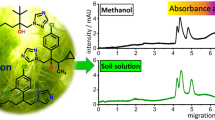
The detector response was calibrated by using a standard curve and shown to be linear over the range 0.3–10 μg mL−1. A typical equation for a calibration line would be y=29.01x−3.751, R 2=0.9955 where y is the peak area (mAUs) and x is the concentration of Dm-AMP1 (μg mL−1). The limit of quantification is defined as the lowest validated fortification level giving a S/N=10 and was found to be 10 mg kg−1 protein in soil sample (Figs. 1 and 2).
Fortified samples were used to establish the procedural recoveries of Rs-AFP1 and Dm-AMP1. Soils were re-equilibrated with water to a moisture holding capacity of 40% prior to fortification followed by a 0.5 h equilibration before subjecting them to the whole procedure. Three soil types were tested, which were known to be microbiologically active soils with normal soil-born bacterial colonies capable metabolising organic substances. One of the soils, sandy clay loam, was sterilised by γ-ray irradiation and the Rs-AFP1 recoveries were compared with that of generated on viable soil. The results were similar (Table 1) demonstrating that there were no artefacts due to quick metabolisation during the extraction step. The overall mean recoveries were 87.2% (RSD 7.9%, n=17) and 78.3% (RSD 8.9%, n=23) for Rs-AFP1 and Dm-AMP1, respectively (Table 1). The physicochemical properties of soils were determined by a standard method and are detailed in Table 2.
Whilst extended stability tests were not performed, the Dm-AMP1 levels were found to be unchanged in sandy loam soil extracts after 9 days storage at 4 °C (Table 3), thereby indicating that extracts are likely to be stable for a considerably longer period of time.
Conclusions
An understanding of protein–soil interaction dynamics allowed efficient recovery of antifungal proteins from soil matrices. The critical parameter for the effective extraction was the use of guanidium thiocyanate at high concentration (4.1 M) in the extraction buffer. This, in combination with conventional solid-phase extraction (SPE) clean-up and analytical techniques, showed that the protein extraction and analysis from soil can be as robust as conventional chemical techniques used in the analysis of trace-level xenobiotics in soils. Inclusion of triethylamine as a competitive agent in the LC and SPE mobile phases was beneficial to the chromatographic performance (e.g. minimisation of peak tailing and recoveries, respectively). This study demonstrated that the solid-phase extraction procedures conventionally used for quantitative pesticide residue analysis can also be applied to proteins extracted from complex matrices such as soils.
References
Epple P, Apel K, Bohlmann H (1997) FEBS Lett 400:168–172
Selitrennikoff CP (2001) Appl Environ Microbiol 67(7):2883–2894
Terras FRG, Schoofs HME, De Bolle MFC, Van Leuven F, Rees SB, Vanderleyden J, Cammue BPA, Broekaert WF (1992) J Biol Chem 267 (22):15301–15309
Francois IEJA, De Bolle MFC, Dwyer G, Goderis IJWM, Woutors PFJ, Verhaert PD, Proost P, Schaaper MMW, Cammue BPA, Broekaert WF (2002) Plant Phys 128:1346–1358
Tassin S, Broekaert WF, Marion D, Acland DP, Ptak M, Vovelle F, Sodano P (1998) Biochemistry 37(11):3623–3637
Terras FRG, Torrekens S, Van Leuven F, Osborn RW, Vanderleyden J, Cammue BPA, Broekaert WF (1993) FEBS Lett 316:233–240
Osborn RW, Genoveva W, De Samblanx GW, Thevissen K, Goderis I, Torrekens S, Van Leuven F, Attenborough S, Rees SB, Broekaert WF (1995) FEBS Lett 368:257–262
Alves AL, De Samblanx GW, Terras RR, Cammue BP, Broekaert WF (1994) FEBS Lett 348:228–232
Kristensen AK, Brunsted J, Nielsen JW, Mikkelsen JD, Roepstorff P, Nielsen KK (1999) Protein Expr Purif 16:377–387
Ogunseitan OA (1997) J Microbiol Methods 28:55–63
Palm CJ, Donegan K, Harris D, Seidler RJ (1994) Molecular Ecology 3:145–151
Palm CJ, Schaller DL, Donegan K, Seidler RJ (1996) Can J Microbiol 42:1258–1262
Saxena D, Flores P, Stotzky G (1999) Nature 402:480–481
Tapp H, Calamai L, Stotzky G (1994) Soil Biol Biochem 26:663–679
Crecchio C, Stotzky G (1998) Soil Biol Biochem 30:463–470
Koskella J, Stotzky G (1997) Appl Environ Microbiol 63:3561–3568
Stotzky G (2000) J Environ Quality 29:691–705
Tapp H, Stotzky G (1998) Soil Biol Biochem 30:471–476
Sims SR, Holden LR (1996) Environ Entomology 25:659–664
Moffatt F, Senkans P, Ricketts D (2000) J Chrom A 891:235–242
François IEJA, Dwyer GI, De Bolle MFC, Goderis JWM, Van Hemelrijck W, Proost P, Wouters P, Broekaert F, Cammue BPA (2002) Plant Physiol Biochem 40:871–879
Penninckx IAMA, Eggermont K, Terras FRG, Thomma BPHJ, De Samblanx GW, Buchala A, Métraux J-P, Manners JM, Broekaert WF (1996) Plant Cell 8:2309–2323
Walkley A, Black IA (1934) Soil Sci 37:29–38
Anderson JPE, Domsch KH (1978) Soil Biol Biochem 10:215–221
Chapman HD (1965) In: Black CA (ed) Methods of soil analysis. Am Soc Agronomy, Wisconsin pp 891–901
Acknowledgement
The authors wish to express their gratitude to Mike Duffin and Peter Massey (Syngenta Analytical Sciences Jealott's Hill) for providing characterised test substances and Sui Kuet (Syngenta Ecological Risk Assessment) for providing characterised soil samples.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Bolygo, E., Ricketts, D., Moffatt, F. et al. Determination of antifungal proteins in soil by liquid chromatography. Anal Bioanal Chem 376, 701–705 (2003). https://doi.org/10.1007/s00216-003-1955-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-003-1955-5