Abstract
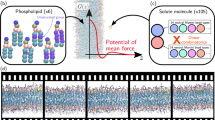
One of the major goals to improve drug discovery is to understand the molecular properties that influence oral bioavailability. Molecular dynamics-based methods have been used to understand passive diffusion with atomic detail and to predict diffusivities and permeabilities. In this paper, we explore the structural and dynamic behavior of piracetam, a nootropic drug, using umbrella sampling, as it crosses a 1,2-dioleoyl-sn-glycero-3-phosphocholine bilayer model. This is accompanied with a potential of mean force profile, showing the partition equilibrium within the bilayer. We discuss important molecular properties and their influence to the permeation of this important drug. Piracetam changes its solution-dominant conformation, modifying its polar surface area and forming an internal hydrogen bond, to facilitate penetration into the hydrophobic core. Rotation and orientation patterns, as piracetam diffuses across the membrane, were also analyzed, and we found out that in bulk water and at the membrane center piracetam shows no specific orientation as it rotates quickly and freely, whereas near the phospholipids’ polar head groups its polar atoms are much more oriented and the rotation is slowed down by more than one order of magnitude. Altogether, the study provides a very detailed view of the events mediating the permeation of this small and very polar drug.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Passive diffusion is the most common framework whereby drugs are absorbed when they are administered by an oral route [1, 2]. As a consequence, favorable membrane permeation has been recognized as a standard requirement for good oral bioavailability. While a high bioavailability allows us to reduce the amount of an administered drug necessary to achieve a desired pharmacological effect and the risks of side effects and toxicity, a low bioavailability, as the result of ineffective intestinal absorption and undesirable metabolic instability, has been considered as one of the major reasons why drug candidates fail to reach the market [1]. As a consequence, pharmaceutical companies have implemented the screening of absorption, distribution, metabolism, excretion and toxicity (ADME-T) properties at the initial stages of drug discovery, in order to avoid the downsides of poor pharmacokinetics.
A number of researchers, such as Lipinski [3, 4] or Veber [5], have defined rules that describe the most relevant molecular properties required for a molecule to be bioavailable. Apart from them, other researchers have also carried out analyses of the properties of marketed drugs and compounds under clinical investigation, and a summary of their works can be found in a review by Lajiness [6]. These rules had a huge impact on how the pharmaceutical industry evaluates a drug’s behavior: Computational methods, for instance, are commonly used to calculate molecular properties in order to predict intestinal absorption before embarking in the very large financial investment for developing a new drug. Quantitative structure–permeability relationship (QSPR) models are the simplest, more economical and the most used models to predict drugs’ permeability based on those molecular descriptors. However, apart from their low transferability, due to the limited datasets of compounds used in its training, QSPR models do not provide a detailed atomistic insight into the permeation process. Physics-based and time-/position-dependent methods, such as molecular dynamics (MD) simulations, have been increasingly applied to this field. They overcome the limitations of current QSPR-based methods and allow for the atomic-level understanding of the permeability process. However, they are still computationally demanding methods (comparing to QSPR approaches, for example), and large screening studies are still prohibitive without massive computational resources. The quality of the results is also dependent of the quality of the force field describing the system, and a common source of errors and disparities [6–9].
As many other chemical and biological processes, membrane permeation is a process that can be studied by following the free energy profile of the compounds as they cross membranes; so, the ability to predict these free energies profiles offers a great insight on how passive diffusion works. Accordingly, by using MD simulations we can estimate a drug’s free energy (ΔG) profile, known as the potential of mean force (PMF), ΔG(z), which describes the drug’s partition equilibrium (the relative concentration of a drug in different regions of the bilayer), as well as its diffusion coefficients, D(z), its local resistance, R, and its permeability, P [9]. Mathematically, these properties are related according to Eq. 1:
where β is the Boltzmann factor, z is the reaction coordinate (reflecting the position of the compound in relation to the axis perpendicular to the plane of the membrane), z 1 and z 2 are opposing points along the reaction coordinate, and as it can be noticed, permeability depends on the diffusion across the membrane (D(z)) and the solubility of the penetrant into the membrane (W(z)) [9, 10]. We have taken the log of the permeability because this quantity screens at least nine orders of magnitude among the known drugs, and the classification of a compound in terms of its permeability is based on the order of magnitude of their permeations.
In this paper, we intend to show how passive diffusion works in atomistic detail for a very hydrophilic small drug, piracetam, by simulating its diffusion through a membrane model. Piracetam, a nootropic drug known to improve a whole series of mental activities, is an orally administered drug with a bioavailability of almost 100%; its plasma peak concentration is reached 30–60 min after being administered [11, 12]. We chose piracetam mainly because it is a small molecule (with a molecular weight of 142.15 g mol−1) with only two rotatable bonds. The fact that it is very hydrophilic (logP of −3.51) adds interest to the understanding of how it achieves its high bioavailability. Table 1 summarizes the main molecular properties that have already been marked as important descriptors for the bioavailability and drug-like character of compounds. Thus, the system we propose to study is small and it has few variables to study, which allows us to follow all the process more accurately and makes the understanding easier. In addition, piracetam is able to form an internal hydrogen bond, which consequently makes it to be able to tune its polar surface area and its internal flexibility by forming the bond. This is a relevant issue, since Veber et al. [5] proposed that the permeation rate is entropically favored by a low molecular flexibility.
Apart from the permeability and diffusion coefficients, we analyzed several of piracetam’s properties during its permeation through the bilayer: the flexibility, the polar surface area (PSA) and the formation of the internal hydrogen bond. Moreover, we also analyzed its orientation and rotation dynamics as it was crossing the membrane. This study highlights the capabilities of MD-based methods in attaining a level of detail over the permeation process of compounds through lipid bilayers. Ultimately, these tools could definitely have a direct impact on drug design campaigns, as these could be easily integrated in drug design efforts in order to understand the pharmacokinetics of drug candidates.
2 Computational details
The lipid membrane system was built using CHARM-GUI, a membrane builder Web-based graphical user interface [14–17]. The 1,2-dioleoyl-sn-glycero-3-phosphocholine (DOPC) glycerophospholipid was carefully chosen because it has been one of the most employed lipids for constructing bilayer system for in silico models [18–22]. DOPC has a transition temperature of −16.5 °C, which means that it is in the fluid phase state at room temperature [23]. The Slipids force field [24–26] was used for describing the DOPC molecule, whereas the GAFF force field was used for piracetam. Water molecules were described with the TIP3P water model—the same water model employed in the validation of Slipids [27]. The topology parameters of piracetam were generated using the ANTECHAMBER suite [28] and the GAUSSIAN 03 [29] program. Firstly, the molecule’s geometry was optimized at the HF/6-31G(d) level of theory in gas phase; secondly, a RESP approach was used to generate the molecule’s point charges at the same level of theory, also in gas phase [30]. The structure and topology files were converted to GROMACS’ compatible files using the ACPYPE tool [31]. The complete system consisted of 72 DOPC molecules, 1 piracetam molecule, 3399 water molecules and 21 molecules of Na+ and Cl− (0.154 M) to mimic physiological conditions. The system had 20,195 atoms in total.
Before starting the umbrella sampling simulations, the system was equilibrated for 40 ns. Membranes have a complex phase behavior, and to have a realistic description of their structural parameters, good-quality force fields (FF) are required. Many studies have shown that some improved FF like Slipids, the one we use in this work, represent accurately those structural parameters [24–26, 32]. This FF was specially developed to reproduce lipid bilayers’ structural and dynamic data and, according to Paloncyova et al. [32], it proved to be the best to calculate the partition coefficients than other all-atomic force fields like CHARMM36 and GAFFlipids.
All MD simulations were carried out using the GROMACS program, version 4.6.7 [33]. The system was equilibrated through a steepest descent minimization algorithm with a gradient of 10 kJ mol−1 nm−1, an NVT stage of 250 ps, a NPT stage of 1 ns and an MD production under the NPT ensemble over 40 ns. The integration of Newton’s equations of motion was carried out using a leap-frog MD integrator with a time step of 2 fs. Short-range non-bonded interactions were calculated using the Verlet cutoff scheme [34] at a distance of 1.2 nm, and long-range electrostatics was calculated using the PME algorithm. The Verlet cutoff scheme was required to calculate the non-bonded interactions on GPU accelerators, whereas bonded and non-bonded interactions were calculated on standard CPU hardware. Dispersion corrections were applied to energy and pressure terms to account for truncation of the van der Waals terms. Periodic boundary conditions were applied in all directions. Bonds involving hydrogen atoms were constrained using the LINCS algorithm [35]. Temperature coupling was handled by the Nosé–Hoover [36, 37] thermostat with a time constant of 0.5 ps and a reference temperature of 310 K, whereas semi-isotropic (separate pressure coupling in xy and z) pressure coupling was handled by the Parrinello–Rahman [38] barostat with a time constant of 10.0 ps. A reference pressure of 1 bar and an isothermal compressibility of 5.0 × 10−5 bar−1 were used. Next, to verify whether the system was well equilibrated several structural parameters for the bilayer were analyzed: the deuterium order parameters of the acyl chains (Scd), the electron density profiles of the system relative to the axis perpendicular to the plane of the membrane, the area per lipid (A L) and the bilayer’s thickness (D HH).
For our equilibrated system, we found a bilayer thickness (D HH) value of 3.74 nm and an area per lipid (A L) value of 0.659 nm2. Both values are reasonably close to experimental data [39, 40]. Moreover, we also verified whether the membrane had entered into gel phase, by determining the deuterium order parameters (Scd) of the DOPC’s acyl chains. The equilibrated system showed Scd values smaller than 0.30 (see SI, Fig. S1), which is characteristic of a disordered hydrocarbon region [39].
After the system energy was equilibrated, 60 umbrella sampling windows were obtained from a pull simulation conducted using the cylinder geometry (with a radius of 1.0 nm). A force constant for pulling of 100 kJ mol−1 nm−2 and a pull rate of 0.1 nm ps−1 were applied. The pull simulation was performed until a complete crossing through the membrane had occurred (reaching a total of 3-µm-long simulation), applying the same simulation conditions as in the membrane equilibration stage. Subsequently, frames for defining the umbrella sampling windows, spaced out every 0.1 nm, were extracted from the pull’s trajectory files. These frames were submitted to a NPT stage of 1 ns and then to an MD production under a NPT ensemble of 50 ns. The simulation parameters were the same as above, apart from the ‘pull rate’, which was set to zero to hinder the configuration’s movement during the exploration of the reaction coordinate, and the harmonic restraint force constant of 1000 kJ mol−1 nm−2. Finally, the PMF profile was obtained using the weighted histogram analysis method (WHAM), and the statistical errors were estimated with bootstrap analysis (200 bootstraps were defined) [41].
The diffusion coefficient of a solute dissolved in a specific solvent can be estimated through the calculation of the mean squared displacement. However, this approach is not applicable to a heterogeneous membrane environment. To estimate the position-dependent diffusion coefficient, D(z), we used the method described by Hummer, which was specially designed for calculating D(z) from umbrella sampling (US) simulations [9, 42]. In this method, D(z) is estimated within each window according to Eq. 2:
where var(z) is the coordinate variance in each i sampled window, and τ i is the characteristic time of its autocorrelation function given by Eq. 3:
However, calculating D(z) from the same MD simulations used to estimate W(z) can be inaccurate: The harmonic potential used to calculate W(z) is commonly too large and, if so, the coordinate variance may not be valid [9]. Although, in less fluid environments, such as the membrane’s polar head groups, correlation functions require extensive sampling (µm to ms length simulations), they can be calculated collecting time series from those long simulations [18].
Hence, to calculate the z-dependent diffusion coefficients, D(z), we performed MD simulations of 500 ps, starting from the last structure of the 50-ns simulation of each slab, with a harmonic potential of 10 kJ mol−1 nm−2. These simulations were repeated five times, with new velocities for each one, to get the standard error of the mean.
The analysis of piracetam’s structural and dynamical properties as it crossed the bilayer system was carried out using some of the GROMACS software integrated tolls. Visual inspection of the simulations and image rendering were performed with the visual molecular dynamics (VMD) software [43].
3 Results and discussion
3.1 Potential of mean force for bilayer crossing
Frequently, PMF profiles found in the literature are obtained by simulating the permeation process until the drug has reached the bilayer’s center, and then, the profile is plotted symmetrically to get the profile over the full bilayer. Despite being a reasonable approach, in this work we simulated the full permeation process, starting from the water phase on one side of the membrane and finishing when the drug has reached the water phase interacting with the opposing lipid layer. In this way, it was possible to verify the convergence of the PMF for piracetam, in particular in the center of the membrane. This is expected to happen when full symmetry is observed around 0.0 nm. Several WHAM analyses with different time distributions of the MD simulations were performed, and, as expected, the PMF convergence depends both on the amount of data that is analyzed and on the total simulation time (see SI, fig. S2). From all WHAM analysis, we observed that the profile based on the first 40 ns of each umbrella sampling window almost completely overlaps the total PMF profile. This means that with only 40-ns simulation in each US window, the profile was already converged (see SI, fig. S3). We also evaluated the histograms configurations, which showed that the neighboring windows were overlapped (see SI, fig. S4).
In Fig. 1, we present the full and the symmetrized PMF for piracetam, considering 50-ns simulation per US window. This profile was symmetrized around 0.0 nm considering the mean of equidistant points from the bilayer’s center (0.0 nm). The reference value of the free energy in water was set (in water) to 0.0 kcal mol−1. The PMF shows a typical behavior of an amphiphilic drug as it crosses the DOPC bilayer system [44]. As piracetam goes from bulk water to the DOPC membrane, the free energy decreases to a minimum of −0.90 kcal mol−1 at 1.71 nm from the membrane’s center. This minimum, located in the high head-group density region, represents the most favorable position of the molecule inside the bilayer, which was expected due to the polar character of piracetam. However, before the energy reaches its minimum, it seems there is a slight increase in energy (ca. 0.3 kcal mol−1) just before piracetam enters the membrane. Miguel et al. [45] observed that the transfer of a drug into the head-group region is driven by an entropic increase, and they have attributed this behavior to the energy required for the drug desolvation. In fact, we observed a decrease of 66% in water molecules of the piracetam’s first solvation shell as it reaches the high head-group density region. As piracetam goes deeper into the membrane, the free energy increases, showing a maximum value of 6.65 kcal mol−1 at the membrane center (0.0 nm).
Piracetam’s potential of mean force (PMF) profile before (red) and after (black) being symmetrized around 0.0 nm considering the mean of equidistant points from the bilayer’s center (0.0 nm) (top) and system’s densities (bottom). In addition to the full system density (along the axis perpendicular to the plane of the membrane, the z axis), separated densities of particular groups (water, polar head groups and acyl chains) are also presented. Background colors represent the pure water phase (blue), and the four region-model phases (shades of red and orange) that from left to right correspond to the low head-group density, high head-group density, high tail density and low tail density [46]. Also depicted is the head-to-head distance (D HH) considering the maximum densities of the system (3.74 nm); the water/lipid free energy barrier ∆G wat; and the bilayer center penetration free energy barrier, ∆G pen
The affinity of the drug to the polar heads in comparison with the water environment can be evaluated from the water/lipid barrier, ΔG wat. We obtained a value of −0.61 kcal mol−1 for ΔG wat, which only translates a slight affinity of piracetam to the bilayer polar region. Similarly, the permeability of drugs depends on the bilayer center penetration barrier, ΔG pen, i.e., the free energy barrier that the drug should overcome to cross the whole membrane [7, 44]. The ΔG pen is determined by the difference between the minimum and maximum values of the PMF profile, having a value of 7.55 kcal mol−1 for piracetam.
3.2 Diffusion coefficients and permeability
According to Eq. 1, the permeability can be obtained if the equilibrium thermodynamics is integrated. In Fig. 2, we present the diffusion profile across the bilayer. It shows a fast diffusion in bulk water, in the low head-group and low tail density regions, and lower diffusivity in the high head-group and high tail density regions. Indeed, the diffusion values in water (1.31 × 10−5 cm2 s−1) are three times larger than in the hydrophilic head-group region and in the dense region of the membrane (0.41 × 10−5 cm2 s−1). The values in the low density region of the bilayer (0.93 × 10−5 cm2 s−1) are twice the values in the head-group region or in the dense region of the membrane. This picture is consistent with the scenario (and the values) found for other similar size molecules [8, 47]. Equation 1 provides a permeability (logP m) value of −2.74 ± 0.09, which lies within the range of experimental values: from −6 to +3 [48].
Piracetam’s diffusion profile before (red) and after (black) being symmetrized around 0.0 nm considering the mean of equidistant points from the bilayer’s center (0.0 nm), which they had previously been averaged in blocks of three points (the error bars represent the standard error of the mean). Background colors represent the pure water phase (blue), and the four region-model phases (shades of red and orange) that from left to right are: low head-group density, high head-group density, high tail density and low tail density [46]
Although no experimental logP m value for piracetam was found in the literature for comparison, it was shown that the permeability of small compounds can be robustly estimated with this method [9]. In fact, Lee et al. [9] tested several methods to predict logP m, and the errors were about 1–2 units, which was considered to be acceptable [9]. Permeability coefficients estimates using MD-based methods have a tremendous value for predicting the bioavailability of compounds, which is a key of importance for drug design and toxicology [9].
3.3 Rotatable bonds, internal H-bond and PSA
Apart from the PMF and the permeability estimates of piracetam, we have analyzed important properties that lead piracetam’s permeation across the DOPC bilayer. Firstly, we have studied the piracetam’s flexibility. This property is related to the number of rotatable bonds, which can be defined as any single non-ring bond attached to a non-terminal heavy atom. According to the Veber’s rules, the number of rotatable bonds in a drug has to be 10 or less for the drug to achieve a bioavailability larger than 20–40% upon oral administration in rats [5]. Piracetam has two rotatable bonds (see Table 1), and for each one, the dihedral angle population was analyzed in function of the distance from the bilayer center. Figure 3 shows the results. The chosen dihedrals are also represented (panel a). We see that the D1 dihedral (formed by atoms 1–4) shows high flexibility in bulk water, as it oscillates approximately between 50° and 150° (see Fig. 3b). Similarly, the D2 dihedral angle (formed between atoms 2–5) also presents high flexibility in bulk water. This can oscillate between 0° and 50°, or 150° and 180° (see Fig. 3c). However, it is clear that as piracetam penetrates deeply into the membrane, both dihedral angles become more constrained, which makes piracetam stiffer. The population of dihedral angles becomes less and less broad and concentrates much more around the minimum as the molecule penetrates into the membrane. In the light of this result, we can see the logic of the empiric Veber rule: Molecular rigidity facilitates the diffusion process because less entropy is lost along the membrane penetration, due to conformational restriction [46, 49].
Piracetam’s rotatable bonds and internal hydrogen bond analysis. a Studied dihedral angles (D1 and D2), together with the internal hydrogen bond that is formed in piracetam (dashed line). b Population for the D1 angle along the axis perpendicular to the plane of the membrane. c Population for the D2 angle along the axis perpendicular to the plane of the membrane. d Internal H-bond distance (r) and angle (θ)
The number of hydrogen bonds is also a relevant parameter that directly influences the drug lipophilicity and consequently its permeability. This parameter was included in the Veber’s rules, in which an upper limit of 12 donors and acceptors, or a PSA of 140 Å2 was established. However, these single parameters are insufficient to predict the permeability of a drug. Indeed, apart from the fact that their calculation does not take into account the molecular conformation, there are class of drugs like renin and HIV protease inhibitors, or even the immunosuppressant cyclosporine A, which are in disagreement with these rules and still show acceptable oral bioavailability [50].
Molecular flexibility allows for intramolecular H-bonds between pairs of functional groups to be formed as the medium changes [51, 52]. When piracetam reaches the membrane center, as no relevant electrostatic interactions are formed with the predominately apolar environment, piracetam tends to favor the formation of an internal hydrogen bond. This bond is established between the oxygen of pyrrolidone and the amide group. This H-bond was studied throughout the permeation process, through the analysis of the angle of the vectors connecting the acceptor–donor atoms and the donor–hydrogen bond (which should be less than or equal to 35° to fulfill the criterion of a hydrogen bond interaction) [53] and intramolecular distances (distance between the donor and the acceptor atoms, which should be less than 3.25 Å) [53].
In Fig. 3 (see panel D), we present both quantities along the axis perpendicular to the plane of the membrane. The formation of this internal hydrogen bond is an important structural feature, with an impact on piracetam’s polar character, as a decrease of 7% in PSA is observed when the drug penetrates into the membrane. Moreover, the major contribution from the H-bond for that decrease is important to highlight. Piracetam becomes less polar when the internal H-bond is formed: The hydrogen bond donor and hydrogen bond acceptor are no longer available to form H-bonds with the surrounding environment. This hydrogen bond also has an important role in stabilizing both donors and acceptors when desolvation takes place, reducing the cost for desolvation. This ability to be more water soluble when the internal H-bond is not formed and more lipid soluble when H-bond is formed probably increases the drug’s permeation [51].
3.4 Rotation and orientation
The rotation of a molecule is influenced by its neighboring molecules; thus, owing to the density of the lipid bilayer environment, a slow rotational diffusion of drugs inside the membrane is expected. To explore the rotational dynamics of piracetam, the rotational autocorrelation functions (ACFs) were calculated. These functions indicate the degree to which the orientation of a molecule at time t + ∆t is correlated with its orientation at time t, and its value will decay to zero over time. Applying a single exponential fitting to these functions, the relaxation times, τ, were obtained [52, 54]. We used the ACF embedded on GROMACS using the first Legendre polynomial and the perpendicular vector to the plane containing the atoms 1, 2 and 3 (see Fig. 3a). In Fig. 4, we present the results of the piracetam’s center-of-mass rotation correlation time. In bulk water, as piracetam is able to rotate very freely, we see a small correlation time. As piracetam penetrates the bilayer, the correlation time increases sharply, reaching its maximum value near the polar head-group and high tails density region, which means that rotation is severely slowed down by more than one order of magnitude (63 ± 3 ps in water vs. 740 ± 107 ps in the head groups). This result may be due to the loss of freedom as the solute moves into the membrane, a hypothesis proposed by Bemporad et al. [52]. The larger τ values are located in the same region where PMF reaches its minimum value, which means that the slower rotation is due to the very strong interactions between piracetam and the head-group region of DOPC. As these interactions are electrostatic by nature (polar and ionic hydrogen bonds), and the head groups are highly immobilized (in terms of rotation) by the lipophilic chains, the strict directionality of the hydrogen bonds ‘freezes’ rotationally the piracetam molecules, with a consequent entropic penalty, obviously overcome by the enthalpic gains of these interactions.
Values of the rotational autocorrelation function (ACF) (bullet points), plotted against the electron density profiles of the bilayer system, and bilayer system groups (water, polar head groups, lipid tails and glycerol molecular framework). The ACF values shown were symmetrized around 0.0 nm considering the mean of equidistant points from the bilayer’s center (0.0 nm), which had previously been averaged in blocks of two points (the error bars represent the standard error of the mean)
Since the ability of piracetam to rotate around its center is greatly diminished when it is located at the polar heads groups of the membrane and the kind of interactions established are highly directional, it is probable that piracetam adopts a narrow range of orientations in this region. In Fig. 5, we present two snapshots representing the orientation behavior of piracetam at opposing sites inside the membrane. A visual analysis of the MD trajectories revealed a full isotropic orientation in bulk water, but as piracetam moves into the membrane it tends to align its polar atoms toward the water phase and polar heads, as expected (see Fig. 5a). In the center of the bilayer, the drug shows a full isotropic motion, like in water, because the DOPC tails are hydrophobic and homogeneous; thus, there is no reason for a particular orientation to be favored. Piracetam leaves the membrane with the opposite orientation as it has entered: In the polar region of the opposing membrane leaflet, the piracetam’s polar atoms tend to be aligned toward the water phase again (see Fig. 5b).
In brief, piracetam enters the membrane with a specific orientation, and after reaching the membrane center it leaves the membrane with the inverse orientation, that is to say that its behavior is symmetric. Bemporad et al. [52] had already observed this preferential orientation; however, like piracetam, the molecules they studied were small (≤266 g mol−1), and this behavior might not hold in more complex molecules.
4 Conclusions
In this article, the passive diffusion of piracetam through a DOPC membrane model was studied in detail by using MD-based methods, in particular by employing the umbrella sampling technique. The piracetam’s PMF profile showed a typical behavior for an amphiphilic drug. As the molecule enters the membrane, the free energy decreases, reaching a minimum at the lipids polar head groups. Then, as we approach the bilayer center, the free energy begins to rapidly increase, exhibiting a maximum value at the membrane center.
As piracetam diffuses through the membrane, a stiffening of its molecular structure was observed: As it goes deeper into the membrane, its dihedrals become rigidified, indicative also of the formation of an internal hydrogen bond. In turn, the internal hydrogen bond makes piracetam less polar, as it can be observed by a decreasing PSA value as the compound reaches the center of the bilayer. This observation is fully consistent with the Veber’s rules and provides a physical reasoning for its empirically derived observations about the influence of the number of rotatable bonds over bioavailability.
The rotation dynamics and the orientation that piracetam assumes as it diffuses across the membrane were also explored. While a full isotropic movement in bulk water and at the membrane center is observed, when piracetam is at the lipids’ polar head-group region, it rotates more than an order of magnitude slower, tending to align its polar groups toward the water phase, with a consequent entropic penalty precisely in the region where the free potential of mean force has a minimum, evidencing a perfect enthalpic/entropic (over)compensation.
Piracetam is an interesting case study, as it is a very small and highly hydrophilic molecule that exhibits good permeation. The extent to which these observations will hold in very large and highly hydrophobic drugs is still to be studied. Nonetheless, the influence of its conversion of an external to an internal hydrogen bond along the membrane crossing its permeability is undoubtedly interesting, as this feature may be more rationally used in drug screening campaigns.
With this work, we have shown the potential of MD-based methods to study in atomistic detail the permeation of compounds through the bilayer. This is an important field as it is intimately linked with drug design campaigns: Poor pharmacokinetics will undoubtedly influence drugs disposition in the organisms and consequently its efficacy.
References
Li AP (2001) Screening for human ADME/Tox drug properties in drug discovery. Drug Discov Today 6:357–366
Chawla G, Gupta P, Koradia V, Bansal A (2003) A means to address regional variability in intestinal drug absorption. Pharm Technol 27:50–68
Lipinski CA (2000) Drug-like properties and the causes of poor solubility and poor permeability. J Pharmacol Toxicol Methods 44:235–249
Lipinski CA, Lombardo F, Dominy BW, Feeney PJ (2001) Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv Drug Deliv Rev 46:3–26
Veber DF, Johnson SR, Cheng HY (2002) Molecular properties that influence the oral bioavailability of drug candidates. J Med Chem 45:2615–2623
Lajiness MS, Vieth M, Erickson J (2004) Molecular properties that influence oral drug-like behavior. Curr Opin Drug Discov Dev 7:470–477
Meng F, Xu W (2013) Drug permeability prediction using PMF method. J Mol Model 19:991–997
Swift RV, Amaro RE (2013) Back to the future: can physical models of passive membrane permeability help reduce drug candidate attrition and move us beyond QSPR? Chem Biol Drug Des 81:61–71
Lee CT, Comer J, Herndon C et al (2016) Simulation-based approaches for determining membrane permeability of small compounds. J Chem Inf Model 56:721–733
Jämbeck J, Lyubartsev AP (2013) Exploring the free energy landscape of solutes embedded in lipid bilayers. J Phys Chem Lett 4:1781–1787
Gouliaev AH, Senning A (1994) Piracetam and other structurally related nootropics. Brain Res Brain Res Rev 19:180–222
Hitzenberger G, Rameis H, Manigley C (1998) Pharmacological properties of piracetam. CNS Drugs 9:19–27
Viswanadhan VN, Ghose AK, Revankar GR, Robins RK (1989) Atomic physicochemical parameters for three dimensional structure directed quantitative structure-activity relationships. 4. Additional parameters for hydrophobic and dispersive interactions and their application for an automated superposition of certain naturally occurring nucleoside antibiotics. J Chem Inf Model 29:163–172
Jo S, Kim T, Im W (2007) Automated builder and database of protein/membrane complexes for molecular dynamics simulations. PLoS ONE 2:e880
Jo S, Kim T, Iyer VG, Im W (2008) CHARMM-GUI: a web-based graphical user interface for CHARMM. J Comput Chem 29:1859–1865
Jo S, Lim JB, Klauda JB, Im W (2009) CHARMM-GUI Membrane Builder for mixed bilayers and its application to yeast membranes. Biophys J 97:50–58
Wu EL, Cheng X, Jo S et al (2014) CHARMM-GUI Membrane Builder toward realistic biological membrane simulations. J Comput Chem 35:1997–2004
Neale C, Bennett WFD, Tieleman DP, Pomès R (2011) Statistical convergence of equilibrium properties in simulations of molecular solutes embedded in lipid bilayers. J Chem Theory Comput 7:4175–4188
Tieleman DP (2006) Computer simulations of transport through membranes: passive diffusion, pores, channels and transporters. Clin Exp Pharmacol Physiol 33:893–903
MacCallum JL, Tieleman DP (2006) Computer simulation of the distribution of hexane in a lipid bilayer: spatially resolved free energy, entropy, and enthalpy profiles. J Am Chem Soc 128:125–130
Parisio G, Stocchero M, Ferrarini A (2013) Passive membrane permeability: beyond the standard solubility-diffusion model. J Chem Theory Comput 9:5236–5246
Carpenter TS, Kirshner DA, Lau EY, Wong SE (2014) A method to predict blood–brain barrier permeability of drug-like compounds using molecular dynamics simulations. Biophys J 107:630–641
Ulrich AS, Sami M, Watts A (1994) Hydration of DOPC bilayers by differential scanning calorimetry. BBA Biomembr 1191:225–230
Jämbeck JPM, Lyubartsev AP (2012) Derivation and systematic validation of a refined all-atom force field for phosphatidylcholine lipids. J Phys Chem B 116:3164–3179
Jämbeck J, Lyubartsev AP (2012) An extension and further validation of an all-atomistic force field for biological membranes. J Chem Theory Comput 8:2938–2948
Jämbeck J, Lyubartsev AP (2012) Another piece of the membrane puzzle: extending slipids further. J Chem Theory Comput 9:774–784
Jorgensen WL, Chandrasekhar J, Madura JD et al (1983) Comparison of simple potential functions for simulating liquid water. J Chem Phys 79:926–935
Wang J, Wang W, Kollman PA, Case DA (2005) Antechamber: an accessory software package for molecular mechanical calculations. J Comput Chem 25:1157–1174
Frisch MJ, Trucks GW, Schlegel HB et al (2004) Gaussian 03, Revision C.02. Gaussian Inc, Wallingford
Bayly CI, Cieplak P, Cornell WD, Kollman PA (1993) A well-behaved electrostatic potential based method using charge restraints for deriving atomic charges—the RESP model. J Phys Chem 97:10269–10280
Sousa da Silva AW, Vranken WF (2012) ACPYPE—antechamber python parser interface. BMC Res Notes 5:367
Paloncýová M, Fabre G, DeVane RH et al (2014) Benchmarking of force fields for molecule-membrane interactions. J Chem Theory Comput 10:4143–4151
van der Spoel D, Lindahl E, Hess B et al (2005) GROMACS: fast, flexible, and free. J Comput Chem 26:1701–1718
Verlet L (1967) Computer experiments on classical fluids. I. Thermodynamical properties of Lennard–Jones molecules. Phys Rev 159:98
Hess B, Bekker H, Berendsen H, Fraaije J (1997) LINCS: a linear constraint solver for molecular simulations. J Comput Chem 18:1463–1472
Nosé S (1984) A unified formulation of the constant temperature molecular dynamics methods. J Chem Phys 81:511–519
Hoover WG (1985) Canonical dynamics: equilibrium phase-space distributions. Phys Rev A Gen Phys 31:1695–1697
Parrinello M, Rahman A (1981) Polymorphic transitions in single-crystals—a new molecular-dynamics method. J Appl Phys 52:7182–7190
Rosso L, Gould IR (2008) Structure and dynamics of phospholipid bilayers using recently developed general all-atom force fields. J Comput Chem 29:24–37
Kucerka N, Nagle JF, Sachs JN et al (2008) Lipid bilayer structure determined by the simultaneous analysis of neutron and X-ray scattering data. Biophys J 95:2356–2367
Hub JS, De Groot BL (2010) g_wham—a free weighted histogram analysis implementation including robust error and autocorrelation estimates. J Chem Theory Comput 6:3713–3720
Hummer G (2005) Position-dependent diffusion coefficients and free energies from Bayesian analysis of equilibrium and replica molecular dynamics simulations. New J Phys 7:34
Humphrey W, Dalke A, Schulten K (1996) VMD: visual molecular dynamics. J Mol Graph 14:33–38
Paloncýová M, DeVane R, Murch B (2014) Amphiphilic drug-like molecules accumulate in a membrane below the head group region. J Phys Chem B 118:1030–1039
Miguel V, Defonsi Lestard ME, Tuttolomondo ME et al (2016) Molecular view of the interaction of S-methyl methanethiosulfonate with DPPC bilayer. Biochim Biophys Acta 1858:38–46
Bemporad D, Luttmann C, Essex JW (2004) Computer simulation of small molecule permeation across a lipid bilayer: dependence on bilayer properties and solute volume, size, and cross-sectional area. Biophys J 87:1–13
Boggara MB, Krishnamoorti R (2010) Partitioning of nonsteroidal antiinflammatory drugs in lipid membranes: a molecular dynamics simulation study. Biophys J 98:586–595
Marrink SJ, Berendsen H (1996) Permeation process of small molecules across lipid membranes studied by molecular dynamics simulations. J Phys Chem 100:16729–16738
Rezai T, Bock JE, Zhou MV et al (2006) Conformational flexibility, internal hydrogen bonding, and passive membrane permeability: successful in silico prediction of the relative permeabilities of cyclic peptides. J Am Chem Soc 128:14073–14080
Navia MA, Chaturvedi PR (1996) Design principles for orally bioavailable drugs. Drug Discov Today 1:179–189
Alex A, Millan DS, Perez M et al (2011) Intramolecular hydrogen bonding to improve membrane permeability and absorption in beyond rule of five chemical space. Med Chem Commun 2:669–674
Bemporad D, Luttmann C, Essex JW (2005) Behaviour of small solutes and large drugs in a lipid bilayer from computer simulations. Biochim Biophys Acta 1718:1–21
PasenkiewiczGierula M, Takaoka Y, Miyagawa H et al (1997) Hydrogen bonding of water to phosphatidylcholine in the membrane as studied by a molecular dynamics simulation: location, geometry, and lipid-lipid bridging via hydrogen-bonded water. J Phys Chem A 101:3677–3691
Leach AR (2001) Molecular modelling: principles and applications, 2nd edn. Pearson Education, Upper Saddle River
Acknowledgements
This work received financial support from the following institutions: European Union (FEDER funds POCI/01/0145/FEDER/007728) and National Funds (FCT/MEC, Fundação para a Ciência e Tecnologia and Ministério da Educação e Ciência) under the Partnership Agreement PT2020 UID/MULTI/04378/2013; NORTE-01-0145-FEDER-000024, supported by Norte Portugal Regional Operational Programme (NORTE 2020), under the PORTUGAL 2020 Partnership Agreement, through the European Regional Development Fund (ERDF); JTSC further thanks the FCT for Grant SFRH/BD/87434/2012.
Author information
Authors and Affiliations
Corresponding author
Additional information
Published as part of the special collection of articles derived from the 10th Congress on Electronic Structure: Principles and Applications (ESPA-2016).
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ribeiro, R.P., Coimbra, J.T.S., Ramos, M.J. et al. Diffusion of the small, very polar, drug piracetam through a lipid bilayer: an MD simulation study. Theor Chem Acc 136, 46 (2017). https://doi.org/10.1007/s00214-017-2073-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00214-017-2073-3