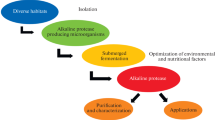
Abstract
Proteases, one of the largest groups of industrial enzymes occupy a major share in detergent industry. To meet the existing demands, proteases with efficient catalytic properties are being explored from bacteria residing in extreme habitats. Alkaline proteases are also considered as promising candidates for industrial sectors due to the activity and stability under alkaline and harsh environment. Therefore, a systematic review on experimental studies of bacterial proteases was conducted with emphasis on purification, characterization, cloning and expression and their suitability as detergent additive. Relevant searches using a combination of filters/keywords were performed in the online databases; PubMed, Science Direct, Scopus and Web of Science. Over thousands of research papers, 71 articles in Scopus, 48 articles in Science Direct, 18 articles in PubMed and 8 articles in Web of Science were selected with regard to bacterial extracellular proteases till date. Selected articles revealed majority of the studies conducted between the years 2015 and 17 and were focused on purification of proteases from bacteria. Among microbes, a total of 41 bacterial genera have been explored with limited studies from extreme habitats. Majority of the studies have reported the involvement of subtilisin-like serine proteases with effective properties for detergent industries. The studies revealed shifting of trend from purification to cloning to genetic engineering to meet the industrial demands. The present systematic review describes the proteases from extremophilic bacteria and use of biotechnological techniques such as site-directed mutagenesis and codon optimization to engineer enzymes with better hot spots in the active sites to meet industrial challenges.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Global market share of industrial enzymes was expected approximately $4.4 billion in 2015 http://bccresearch.blogspot). Among these industrial enzymes, proteases with over 60% share of worldwide market are being used as bioadditives in laundry, pharmaceutical, leather, food, agriculture and other industries (Gonzalez-Rabade et al. 2011; Sharma et al. 2016). Microbes from extreme environments in past decades serve as a centre of scientific spotlight which are being explored to meet existing industrial demands. The microbial enzymes from different habitats are continuously explored due to their better catalytic potential at low or moderate temperatures (Haki and Rakshit 2003). In present scenario and thrust to explore energy efficient industrial processes, attention on the use of enzymes from extremophiles has gained rapid attention keeping in view the importance of green processes. In comparison to mesophilic counterparts, extremophiles such as thermophiles and psychrophiles are being explored due to their better structural adaptability and enhanced specific activity of their enzymes at low temperature (Cowan et al. 1985; De Azeredo et al. 2003; Hajji et al. 2008; Zaferanloo et al. 2014; Boulkour et al. 2015). Bioadditives reported from psychrotrophs can interact with water, hydrophobic compounds, ice and gases at low temperature and hence offers a green sustainable and energy efficient processes and other biotechnological applications (Wu et al. 2004; Zhang et al. 2016; Yildirim et al. 2017).
Microbes from Arctic, Antarctic, and alpine zone containing ~ 6500 microbial strains have been explored for cold-active extracellular proteases, lipases, and exopolysaccharides production (Perfumo et al. 2018). Among proteases, alkaline proteases are the leading contributors and account for ~ 40% of the total global enzyme for industry (Kirk et al. 2002). Screening microbes from extreme environments can facilitate our thrust for energy efficient enzymes (Raval et al. 2014). Since the discovery of subtilisin Carlsberg, a first alkaline protease from Bacillus licheniformis for detergent industry (Lesney 2003), a number of alkaline proteases from Aeribacillus pallidus (Yildirim et al. 2017; Mechri et al. 2017), Bacillus circulans DZ100 (Benkiar et al. 2013), Bacillus pseudofirmus (Raval et al. 2014), Bacillus pumilus NRC21 (Tork et al. 2016), Bacillus sphaericus (Wati et al. 1997a, b), Bacillus sp. EMB9 (Sinha and Khare 2013) and WF146 (Wu et al. 2004), Bacillus subtilis AP-MSU6 (Maruthiah et al. 2013), Caldicoprobacter guelmensis (Bouacem et al. 2015), Geobacillus caldoproteolyticus (Chen et al. 2004), Geobacillus toebii Strain LBT 77 (Thebti et al. 2016), Planococcus sp. (Zhang et al. 2016), Pseudoalteromonas haloplanktis (dePascale et al. 2010), Serratia proteamaculans Gulmez et al. 2018), Shewanella arcticai (Gulmez et al. 2018), Stenotrophomonas maltophilia (Ribitsch et al. 2012), Streptomyces sp. AB1 (Jaouadi et al. 2010) and Streptomyces sp. strain AH4 (Touioui et al. 2015) have been characterized, cloned and explored for various biotechnological applications such as detergent, leather and feather degradation. Proteases from microbes isolated from extreme habitats with different features such as hot springs (Thebti et al. 2016), halophilic area, arctic or polar regions, deep sea, glacier ice and cold desert soils can be harnessed for efficient enzymes for various applications (Zhang et al. 2016). Limited studies has been done on the exploration of proteases for industry using culture independent techniques from extreme habitats (Qoura et al. 2015; Singh et al. 2015).
Application of proteases in detergents as green process with high activity at low temperature has brought a major shift from being minor additives to key ingredients. Proteases are considered as one of the most important industrial enzymes with applications in detergents, foods, textiles and leather, pharmaceuticals, silvery recovery from X-ray films and bioremediation processes. Presently, a number of Bacillus strains from alkalophilic habitats have been explored for detergents applications and first study was reported from Bacillus alcalophilus in 1934 by Vedder (Fritze et al. 1990; Gupta et al. 2002). Use of proteases in detergents constitutes ~ 25% of the total worldwide enzymes sales (Godfrey and West 1996). Among them, alkaline proteases or subtilisins of Bacillus sp. contribute to ~ 89% share of detergent enzymes and Novo Nordisk and Genencor International are the two major distributors covering ~ 95% global market of proteases (Gupta et al. 2002). A number of proteases with trademark such as Alcalase, BIO-40, and BIOTEX have been explored from Bacillus spp. (Lesney 2003). Industrial demands for proteases rely on a broad specificity for substrates, wide temperature and pH range and stability, and compatibility in chelating and oxidizing environments for the effective removal of stains. Proteases like kannases have been commercialized by Novo Nordisk Bioindustry, Japan.
Exploration of microbes from these extreme habitats offers access to novel microbes and robust enzymes which can meet industrial demands. Keeping in view the importance of proteases from extremophiles, an attempt has been made to summarise the literature published between 2000 till present based on the information collected from different databases.
Database search and selection criteria
PubMed https://www.ncbi.nlm.nih.gov/pubmed/), ScienceDirect https://www.sciencedirect.com/), SCOPUS https://www.scopus.com/search/form) and Web of Science Clarivate Analytics version 1.0 were selected for online retrieval of the literature searches using a combination of keywords as filters, i.e. Protease) AND Peptidase) AND Proteolytic) AND Serine) AND Cloning, expression) AND Bacteria/bacterial/bacterium) AND Detergent/laundry). To limit the search, further terms like extracellular, purification, and recombinant were also included. Search was performed by the authors independently and data published till date was used for the present review. The literature included in the present study involved research articles exclusively and reviews articles were only studied and cited for supporting information on microbial proteases. Data extraction was conducted to establish reliability and avoiding any data entry errors.
A total of 4,97,201 articles were found with the keyword proteases in PubMed, 2,53,165, articles in Science Direct, 39,400 in Web of Science and 1, 82,895 in SCOPUS between the years 2000 to till present Fig. 1a). After applying all the keywords, a comprehensive list of abstracts was prepared to meet the inclusion criteria. Relevant studies based on titles and abstract were screened and selected for full-text evaluation. Duplicate entries from all the databases were screened and removed manually as well as by Mendeley. Studies highlighting purification and characterization, cloning and expression in heterologous host and application of purified protease as detergent additive and other industrial aspects were selected. After exclusion of irrelevant data, only 18 articles from PubMed, 48 articles from Science Direct, 8 articles from Web of Science and 537 from SCOPUS were retrieved and evaluated for the desired inclusion criteria (Fig. 1b). The full-text of selected research articles were selected and imported into Mendeley reference manager.
a Publications of research articles in (2000–till present) Scopus, Web of sciences, Pubmed and Science Direct using keywords (Protease) AND (Peptidase) AND (Proteolytic) AND (Serine) AND (Bacteria) AND (Detergent/laundry); b publications of research articles in Science Direct, Pubmed, Web of sciences and Scopus, using a keywords (Protease) AND (Peptidase) AND (Proteolytic) AND (Serine) AND (Detergent/laundry) from 2000 onwards to till present. The research articles evaluated for removal of duplicates, studies exactly not following the inclusion criteria and incomplete experimental research were removed; and c Venn diagram showing experimental studies on purification, expression and application of bacterial proteases as detergent additive based on research articles shared among Science Direct Scopus, PubMed and Web of Science. Triangle shows number of research articles on purification of proteases, circle shows number of research articles on expression of proteases in heterologous host and square shows number of articles on proteases with application as detergent additive
The experimental studies on proteases from extremophilic bacteria revealed 52 articles on purification and characterization of proteases, 19 articles on cloning and expression in heterologous host and 52 articles showing suitability of protease as detergent additive (Fig. 1b). Further analysis revealed 9 articles on purification of protease and expression in heterologous host, 6 articles on purification, expression and application as detergent additive, 39 articles on purification and application as detergent additive and only 6 articles on expression of protease in heterologous host with application as detergent additive (Fig. 1c).
Diversity of proteases producing bacteria residing in extreme environments
Evaluation of these research articles revealed 62% proteases from mesophiles, 13% from thermophiles, 11% from halophiles, 7% from psychrophiles and only 6% from alkaliphiles. Moreover, a total of 41 genera have been reported for protease production. Based on selected data used for comprehensive and systematic data analysis, 145 articles belonging to 41 bacterial genera were reported for purification, cloning and expression of proteases in heterologous host with majority of the studies from Bacillus spp. (Table 1). Different bacteria explored for industrial applications included Acinetobacter, Actinomadura, Aeribacillus, Aeromonas, Aeropyrum, Alcaligenes, Archaeoglobus, Bacillus, Brevibacillus, Burkholderia, Caldicoprobacter, Chryseobacterium, Dermatophilus, Dichelobacter, Fibrobacter, Lactobacillus, Lysinibacillus, Lysobacter, Micromonospora, Mycobacterium, Nocardiopsis, Oenococcus, Paenibacillus, Planococcus, Prevotella, Pseudoalteromonas, Pseudomonas, Salinicoccus, Serratia, Shewanella, Sphaerobacter, Stenotrophomonas, Streptococcus, Streptomyces, Teredinobacter, Thermoactinomyces, Thermoanaerobacter, Thermus, Treponema, Virgibacillus, Xanthomonas. Among different bacterial proteases, Bacillus and Streptomyces constituted the major part of literature (Table 1).
Among all publications, majority of the reported proteases have been reported as subtilisin-like serine proteases with limited reports on glutamyl-endopeptidase, Zn-metalloprotease/M14, proline specific prolyl endoprotease and kumamolysin. These proteases have been reported for effective wash performance in combination with detergents for suitability as additive in laundry (Table 1). Few reports on application of proteases in feather degradation, leather, fibrinolytic, pharmaceutical and biocidal activity have also been recorded. The research articles revealed purification of proteases using anion exchange, cation exchange, gel filtration, hydrophobic interaction and affinity, as well as FPLC and HPLC chromatography. The chromatographic based methods involve use of SP Sepharose, DEAE-cellulose, DEAE-Sepharose, Phenyl-Sepharose and superose, Sephadex G-100, Superdex-75, Sephacryl S-200, Superose and Ni–NTA column (Table 1). The molecular weight of the purified proteases ranged from 19.5 to 250 kDa and have optimum temperature range of 4–100 °C and pH range of 5–13 (Gabdrakhmanova et al. 2002, Trachuk et al. 2005; Snajder et al. 2015; Chin et al. 2007; Thebti et al. 2016). The presence of metal ions like Mg2+, Ca2+, Cu2+, Zn2+, Ba2+, Mn2+, Na+ and K+ stimulated enzyme activity (Kasana 2010). Regarding inhibition effect, majority of the proteases are inhibited in the presence of inhibitors PMSF and DFP, thus identified as subtilisin-like serine proteases. Other proteases were inhibited in the presence of inhibitors aprotinin, EDTA, EGTA, 1, 10-phenanthroline and phosphoramidon.
Structural analysis of bacterial extracellular S8 serine proteases
Representative proteases from thermophilic, mesophilic and psychrophilic bacteria were selected to compare their structural variation and evolutionary history. The gene sequences of serine proteases ranged from 1239 to 2371 bp have been identified as subtilisin like serine proteases belonging to family peptidase S8. Nine studies reported identification of proteases based on MALDI/TOF and N-terminal sequencing. Phylogenetic analysis of extracellular serine proteases was conducted using MEGA7 to analyse their association among each other. The analysis revealed close relatedness and clustering of proteases from thermophiles and psychrophiles as separate branches (Fig. 2).
Phylogenetic analysis of S8 family alkaline serine proteases showing clustering of psychrophilic and thermophilic proteases as different branches a WP 010369807 Pseudoalteromonas piscicida, WP 063374258; Pseudoalteromonas luteoviolacea, WP013051407; Shewanella violacea BAB61726, Pseudoalteromonas sp. AS-11, ABA60899 Serratia sp. GF96, WP 012197479 Shewanella baltica; b WP 094058666 Bacterioplanes sanyensis, WP 017819842 Vibrio alginolyticus, WP 015781271 Kangiella koreensis; c Thermophilic proteases WP 01083250 Nocardioides sp. CF8, WP 084100044 Knoellia flava; and d WP 027481108 Deinococcus pimensis; e WP 013704806 Marinithermus hydrothermalis, WP 013456966 Oceanithermus profundus, WP 082381964 Ardenticatena maritima; P08594 aqualysin-1, WP 053768483 Thermus aquaticus, WP 038035270 Thermus parvatiensis, WP 093005298 Thermus arciformis; f WP 010376775 Pseudoalteromonas piscicida, WP 039492850 Pseudoalteromonas elyakovii, WP 010605882 Pseudoalteromonas flavipulchrag AGV12715 Acinetobacter sp. IHB B 5011 (MN12). The phylogenetic tree was prepared in MEGA 7.0 using N–J method
The N-terminal signal peptide was predicted to determine the extracellular nature of proteases using SignalP Server 4.0 http://www.cbs.dtu.dk/services/SignalP/) (Petersen et al. 2011). The size of signal peptide is variable and ranges from 21 to 29 amino acid long among different proteases reported from extremophiles which facilitates secretion of prosubtilisin across cytoplasmic membrane. The structural analysis of representative proteases from thermophilic, mesophilic and psychrophilic bacteria was done using NCBI-CDD (http://www.ncbi.nlm.nih.gov/cdd) to predict active site residues, calcium binding sites and catalytic triad (Marchler-Bauer et al. 2005). The amino acid sequences of these proteases were also compared to predict the location of domains and inhibitor sites in SMART (Letunic et al. 2006). The domain analysis of representative proteases of psychrophilic Shewanella arctica revealed presence of inhibitor (41–120 aa), peptidase S8 (156–402 aa) and PPC domain (436–505 aa and 506–619 aa), thermophilic revealed the presence of signal peptide, preprosequence, peptidase S8 domain, inhibitors and PPC domain with variation in amino acid residues (Fig. 3). The domain analysis of thermophilic Thermus aquaticus revealed presence of signal peptide (1–25 aa), inhibitor (54–126 aa) and peptidase S8 domain (157–399 aa). The domain analysis of Pseudoalteromonas flavipulchra revealed signal peptide (1–27 aa), peptidase S8 (182–499 aa) and two PPC domain (524–594 aa and 632–701 aa). PPC domains are present only in psychrophilic bacteria and absent in thermophilic and mesophilic bacteria. The amino acid sequence of mesophilic B. clausii showed presence of signal peptide (1–29 aa), inhibitor (30–108) and peptidase S8 domain (131–374 aa). The domain analysis was also compared with previously reported alkaline serine protease of Acinetobacter sp. IHB B 5011 that showed the presence of signal peptide, pre-prosequence which guides correct folding of mature peptidase S8 domain (Salwan et al. 2018). No inhibitor and PPC domain were reported here. The present data revealed limited studies involving maturation of active proteases after N and C-terminal processing. Such autoprocessing has been reported in proteases of Bacillus spp., Burkholderia pseudomallei (Chin et al. 2007), Lysobacter sp. (Pereira et al. 2017), Streptococcus pneumonia (Romanello et al. 2006), Stenotrophomonas maltophilia Ribitsch et al. 2012) and Thermus aquaticus (Terada et al. 1990;). Ribitsch et al. 2012 reported truncation of the C-terminal domain of StmPr1 protease of Stenotrophomonas maltophilia which caused enhanced processing of N-terminal prosequences and hence production of active enzyme.
Domain analysis showing variation in position of signal peptide, inhibitor (if present) peptidase S8 and PPC domain in S8 proteases of a WP_010605882 Pseudoalteromonas flavipulchra; b WP_013051407 Shewanella violacea; c WP_053768483 Thermus aquaticus; d AGV12715 Acinetobacter sp. IHB B 5011 (MN12); e AGN91700 Bacillus circulans DZ100. The scanning of active sites was performed in ScanProsite tool in ExPASy
Homology modelling of peptidase S8 domain was done using Swiss-Model (Fig. 4) (http://swissmodel.expasy.org/) based on the template with PDB code 3TI7 Dichelobacter nodosus, 2B6N Serratia sp., 4DZT Thermus aquaticus and 3WHI Bacillus subtilis. The structure of peptidase S8 domain showing 66.48% sequence identity were prepared and aligned with the templates for quality assessment. All the model proteins showed root mean square deviation RMSD value below 1.5 Å which validates. All the model proteins were submitted to VERIFY-3D server http://servicesn.mbi.ucla.edu/Verify3d/) for validation and showed the acceptance of models with 95–100% passing score. PROCHECK analysis was done to check stereo-chemical quality properties of the model proteins which revealed over 90% of the residues in the most favourable regions with only 0.4–0.7% of the residues in disallowed regions.
Structural modeling of S8 alkaline proteases of psychrophilic aPseudoalteromonas flavipulchra using template 3TI7 of Dichelobacter nodosus); and bShewanella violacea using 2b6n of Serratia sp.); thermophilic, cThermus aquaticus using template 4dzt of Thermus aquaticus and mesophilic, dBacillus circulans DZ100 using template 3WHI of Bacillus subtilis. The graphics were prepared in PYMOL 2.1 for comparison. Catalytic triad (DHS) aspartate, histidine and serine residues are shown as spheres, active site residues as green sticks with labelled one letter code and calcium binding sites as sphere mesh in red
Engineering methods for enhanced production and effective catalytic behaviour
According to the literature, research articles were also searched for characterization of recombinant proteins, optimised expression of recombinant proteins and methods for engineering enzymes. Only 20 studies revealed the cloning and expression of proteases in heterologous host (Fig. 1b). The genes of varying size were successfully cloned in vectors like pHY300PLK, pMS470∆8, pET28a), pET19-b, pET32a-c +), pWEB-TNC, pBR322, pET22b, pET-30, pTrc99A, pTrc99A, pMCSGx, pWH1520, pD441-NH, pET-5a, pIJT02, pGEX-4T-2 and expressed in heterologous host such as Bacillus megaterium, B. subtilis AJ73, B. thuringiensis, E. coli BL21DE3, E. coli rosetta-gami DE3 and Streptomyces lividans. The recombinant proteins have also been purified using Ni–NTA affinity chromatography and characterized for substrate specificity, temperature and pH profile and utility in industrial applications.
Biotechnological techniques have seen evolutionary and cutting edge change in recent time. Systematic data analysis revealed emergence of alternative approaches from mining microbial diversity to recombinant protein engineering techniques such as site-directed mutagenesis involving substitution of one amino acid by other naturally occurring amino acids, codon optimisation and recombinant protein production by cloning gene in suitable vector and expression in heterologous systems to meet industrial demands. Engineering enzymes with residue-specific approach such as replacement of methionine by seleno-methionine residues is a common method of enzyme engineering for better catalytic properties. Various reports on modification of subtilisin type of proteases from Bacillus spp. using site-directed mutagenesis, DNA shuffling, cassette mutagenesis, error-prone PCR and loop removal are available (Gupta et al. 2002). However, engineering enzymes with modified unnatural amino acids and hot spots in the active sites has also been reported and continuously explored for improving enzymes with enhanced biocatalytic properties (Ravikumar et al. 2015; Balke et al. 2017). This strategy relies on the modification of active site residues (Bryan 2000). Heterologous expression of genes offers several advantages being fast to grow, higher and regulated protein production, easy to handle, and high versatility. However, expression of foreign genes of distant origin has several problems including modifications at posttranslational level, toxic or presence of rare codons in the host (Snajder et al. 2015). The associated challenges of expression in heterologous hosts limits the use in commercial scale preparation of recombinant proteins but can be addressed with development of engineered system (Wacker et al. 2002) and better regulation of expression host through promoters (Terpe 2006). Synthetic oligonucleotide shuffling is another method to facilitate the recombination of lower homology genes. Various wild-type protease genes were shuffled and this synthetic shuffling yielded diverse recombinants that displayed superior properties for use as additive in detergents (Yuan et al. 2005; Ness et al. 2002). Alternative to these methods, codon optimisation based on the modification of target gene sequence is a promising tool which has been explored for enhanced expression of serine protease of Aeropyrum pernix K1, human genes, and nucleopolyhedrovirus DNA polymerase of Bombyx mori in E. coli (Burgess-Brown et al. 2008; Song et al. 2014; Snajder et al. 2015). The study has reported codon optimisation as a key step for the enhanced expression of pernisine in E. coli by replacing the rare codons with the more frequently occurring amino acids while loss of activity occurs by doing changes in the active site residues (Mechri et al. 2017). Use of DNA sequence manipulation using codon-optimisation has resulted in optimised expression of proteins in heterologous hosts with considerably reduced production cost (Elena et al. 2014; Snajder et al. 2015).
Genomic and metagenomic approaches for discovering new proteases
Development in genome sequencing and annotation is rapidly growing but its application as an alternative approach for mining candidate genes for industrial application is in early stage. A comparative study of mesophiles, thermophiles, halophiles and psychrophiles provided an overview of richness of protease diversity and their organization in genomic map (Fig. 5; Table 2). Complete genome of representative mesophilic Bacillus clausii, psychrophilic Pseudoalteromonas flavipulchra and thermophilic Thermus aquaticus were retrieved from NCBI database and submitted to RAST annotation server. The genomes were analyzed and compared for genome size, GC content, presence of number of serine proteases (Table 2). The circular genome map of these selected bacteria was prepared in GView web server based on reference sequence of Pseudomonas aeruginosa PA103. The CDS were analysed according to the COGs functional classification categories. The comparative genomic analysis provides promising incidences in harnessing microbes for better and efficient protease production.
Circular genome map of extremophilic bacteria showing organization of various genes on forward and reverse strand, reference sequence of Pseudomonas aeruginosa PA103 (JARI01), genomes of mesophiles—Burkholderia pseudomallei AAHR02, Bacillus pumilus ATCC 7061 (ABRX01), genomes of thermophiles—Dermatophilus congolensis DSM 44180 (AUCS01), Caldanaerobacter subterraneus subsp. yonseiensis KB-1 (AXDC01), and archeal genome—Actinomadura viridilutea strain DSM 44433 (PVNI01) in different colors
Metagenomic based studies are useful and attractive alternate tools to discover uncharacterized enzymes from diverse ecological niches particularly for non-culturable microbes (Amann et al. 1995). Successful discoveries have been obtained using primers targeting conserved protein sequences (Morimoto and Fujii 2009; Fieseler et al. 2006), while activity-based screening is a more direct way to discover new enzymes. A variety of libraries have been reported for enzyme activity including lipases, β-lactamases, proteases, nitrilases, polysaccharide-modifying enzymes, oxidoreductases, and dehydrogenases (Ferrer et al. 2005; Hu et al. 2009; Jeon et al. 2009). Besides this, metagenomic approach not only provide information regarding single gene encoding activity but the entire operons can also be identified (Suenaga 2012; Uchiyama et al. 2005; Chung et al. 2008). The clones expressing a certain function are identified and indicating the genomic information and phylogeny of the genes (Handelsman 2005; Riesenfeld et al. 2004). This approach has recovered novel biocatalysts and expanded our knowledge on uncultured microorganisms (Daniel 2004, Riesenfeld et al. 2004). Despite the abundance of new enzymes isolated by metagenomic approaches and the industrial potential of proteases, only limited reports have been published concerning metagenome-derived proteases. Metalloproteases have been reported from a deep sea sediment metagenomic library (Lee et al. 2007) and soil metagenomic libraries (Waschkowitz et al. 2009). Similarly, serine proteases have been reported from metagenomic library of tannery activated sludge, saline habitat, yucatan underground water and oil-polluted mud-flat metagenome with suitability of protease as laundry additive (Apolinar-Hernández et al. 2016a, b; Ribitsch et al. 2010a, b; Purohit and Singh 2013; Devi et al. 2016; Gong et al. 2017). Several metagenomic studies have also reported screening of false-positive clones for proteolytic activities and has remained unsuccessful (Rondon et al. 2000; Jones et al. 2007).
Discussion
Proteases from diverse sources are extensively explored for various biotechnological and industrial applications as bioadditives in laundry, food, leather, textiles, pharmaceuticals and bioremediation processes (Kasana 2010), as well as in agricultural and basic research related to enzyme modification, photogenicity, complement system and apoptosis pathways (Rao et al. 1998; Mahajan and Badgujar 2010). Proteases are ubiquitous and are reported from a wide diversity of biological sources such as microorganisms, plants and animals. The inability of the plant and animal proteases to meet current world demands has led to an increased interest in microbial proteases. Proteases of bacterial origin are the most significant because of their wide biotechnological potential. Diverse species B. cereus, B. mojavensis, B. megaterium, B. stearothermophilus and B. subtilis have been exploited for extracellular production of proteases (Beg et al. 2002; Shumi et al. 2004; Ravishankar et al. 2012). Many proteases have been commercialized from these species and active under neutral and alkaline conditions. Many bacteria having ability to secrete serine, cysteine, metallo, glutamic and aspartic type of proteases have still remained uncharacterized.
We have found hundreds of research articles related to purification and characterization of proteases as additive in laundry. Studies on cloning of candidate genes are limited and only 6 papers were found in which all the three aspects were discussed. Although a number of review articles are available on microbial proteases still systematic review are available only for proteases. Present review also highlights the scope of alterative tools for developing or engineering efficient enzymes. Although efforts have been placed for protein engineering, the recent advances are still not fully explored for developing efficient proteases. In an attempt to review serine proteases of microbes which is a major class of microbial proteases for detergent applications, systematic and meta-analysis was conducted. Present study revealed that majority of bacteria explored to industrial demands belongs to Bacillus spp. Most of the research has been done on the purification of enzymes with limited experimental efforts on cloning and exploration of candidate genes. Both the exploration of extremophiles for protease production is an attractive alternative and coupling of emerging techniques such as active hotspot manipulation of enzymes for broad substrates with enhanced properties can play big role in enzyme engineering. Studies using metagenomic approaches have shown potential alternate for mining unculturable diversity of microbes. Further studies on codon optimisation and use of unnatural amino acids for efficient enzymes engineering can be a boom to detergent industry. Advent of genomic approaches such as Hi seq 2500 Illumina and availability of complete genome of several bacteria from extreme habitat can help in developing environmental friendly and green sustainable processes. There are few studies that report identification of bacteria for protease activity, purification and characterization, cloning and expression in heterologous host with limited studies from extremophiles. Still exploring microbial systems for the protease production with enhanced efficacy, optimised expression system and higher production can ensure a boom in the enzymes.
Conclusions
Use of modern biotechnological techniques such as codon optimisation and other enzyme engineering tools can be promising for developing efficient enzymes to meet the demands for green and sustainable industrial processes. On the other hand, extremophiles such as halophiles, thermophiles and psychrophiles offer vast potential to tap their diversity to meet industrial demands. The present review article highlights the paradigmatic shift in trends of protein purification from native microbes to enzyme engineering and exploring whole genomes for mining protease diversity. Sequencing of complete genomes with cutting edge genomics techniques will speed-up the genome mining of proteases from bacteria residing in extreme habitats with potential industrial applications. Coupling of codon optimisation and site-directed mutagenesis can fasten our search and will contribute significantly to the next green revolution.
References
Amann RI, Ludwig W, Schleifer KH, Amann RI, Ludwig W (1995) Phylogenetic identification and in situ detection of individual microbial cells without cultivation. Microbiol Rev 591:143–169
Apolinar-Hernandez MM, Pera-Ramirez YJ, Perez-Rueda E, Canto-Canche BB et al (2016a) Identification and in silico characterization of two novel genes encoding peptidases S8 found by functional screening in a metagenomic library of yucatan underground water. Gene 5931:154–161
Apolinar-Hernandez MM, Peria-Ramirez YJ, Perez-Rueda E, Canto-Canche BB, De Los Santos-Briones C, Connor-Sanchez A (2016b) Identification and in silico characterization of two novel genes encoding peptidases S8 found by functional screening in a metagenomic library of yucatan underground water. Gene 5931:154–161
Balke K, Beier A, Bornscheuer UT (2017) Hot spots for the protein engineering of Baeyer–Villiger monooxygenases. Biotechnol Adv 361:247–263
Beg QK, Saxena RK, Gupta R (2002) Kinetic constants determination for an alkaline protease from Bacillus mojavensis using response surface methodology. Biotechnol Bioengg 783:289–295
Benkiar A, Jaouadi Z, Badis A, Rebzani F, Touioui B, Rekik H, Naili B, Zohra F (2013) Biochemical and molecular characterization of a thermo- and detergent-stable alkaline serine keratinolytic protease from Bacillus circulans strain DZ100 for detergent formulations and feather-biodegradation process. Int Biodeterior Biodegrad 83:129–138
Botos I, Melnikov EE, Cherry S, Kozlov S, Makhovskaya OV, Tropea JE, Gustchina A, Rotanova TV, Wlodawer A (2005) Atomic-resolution crystal structure of the proteolytic domain of Archaeoglobus fulgidus lon reveals the conformational variability in the active sites of lon proteases. J Mol Biol 3511:144–157
Bouacem K, Bouanane-Darenfed A, Laribi-Habchi H, Elhoul MB, Hmida-Sayari A, Hacene H, Ollivier B, Fardeau ML, Jaouadi B, Bejar S (2015) Biochemical characterization of a detergent-stable serine alkaline protease from Caldicoprobacter guelmensis. Int J Biol Macromol 81:299–307
Boulkour TS, Zarai JN, Boudjella H et al (2015) Purification and biochemical characterization of two detergent-stable serine alkaline proteases from Streptomyces sp. strain AH4. World J Microbiol Biotechnol 317:1079–1092
Bryan PN (2000) Protein engineering of subtilisin. Biochim Biophys Acta 1543:203–222
Burgess-Brown NA, Sharma S, Sobott F, Loenarz C, Oppermann U, Gileadi O (2008) Codon optimiza- tion can improve expression of human genes in Escherichia coli: a multi-gene study. Protein Expr Purif 59:94–102
Chen XG, Stabnikova O, Tay JH, Wang JY, Tay ST (2004) Thermoactive extracellular proteases of Geobacillus caldoproteolyticus sp. nov., from sewage sludge. Extremophiles. 86:489–498
Cheng Q, Xu F, Hu N, Liu X, Liu Z (2015) A novel Ca2+ dependent alkaline serine protease Bvsp) from Bacillus sp. with high fibrinolytic activity. J Mol Catal B Enzy 117:69–74
Chin CY, Othman R, Nathan S (2007) The Burkholderia pseudomallei serine protease MprA is autoproteolytically activated to produce a highly stable enzyme 40:370–377
Chung EJ, Lim HK, Kim JC, Choi GJ, Park EJ, Lee MH et al (2008) Forest soil metagenome gene cluster involved in antifungal activity expression in Escherichia coli. Appl Environ Microbiol 743:723–730
Cowan D, Daniel R, Morgan H (1985) Thermophilic proteases: properties and potential applications. Trends Biotechnol 33:68–72
Daniel R (2004) The soil metagenome—a rich resource for the discovery of novel natural products. Curr Opin Biotechnol 153:199–204
De Azeredo LAI, Castilho LR, Leite SGF, Freire DMG, Coelho RRR (2003) Protease from Streptomyces sp. isolated from Brazilian cerrado soil optimization of culture medium employing statistical experimental design. Appl Biochem Biotechnol 108:749–755
dePascale D, Giuliani M, De Santi C et al (2010) PhAP protease from Pseudoalteromonas haloplanktis TAC125: gene cloning, recombinant production in E. coli and enzyme characterization. Polar Sci 42:285–294
Devi SG, Fathima AA, Sanitha M, Iyappan S, Curtis WR, Ramya M, Devi et al (2016) Expression and characterization of alkaline protease from the metagenomic library of tannery activated sludge. J Biosci Bioeng 1226:694–700
Divakar K, Priya JDA, Gautam P (2010) Purification and characterization of thermostable organic solvent-stable protease from Aeromonas veronii PG01. J Mol Catal B Enzym 66:311–318
Dorra G, Ines K, Imen BS, Laurent C, Sana A, Olfa T et al (2018) Purification and characterization of a novel high molecular weight alkaline protease produced by an endophytic Bacillus halotolerans strain CT2. Int J Biol Macromol 111:342–351
Elena C, Ravasi P, Castelli ME, Peirú S, Menzella HG (2014) Expression of codon optimized genes in microbial systems: current industrial applications and perspectives. Front Microbiol 5:21
Elhoul BM, Zaraî Jaouadi N, Rekik H, Bejar W, Boulkour Touioui S, Hmidi M, Badis A, Bejar S, Jaouadi B (2015) A novel detergent-stable solvent-tolerant serine thiol alkaline protease from Streptomyces koyangensis TN650. Int J Biol Macromol 79:871–882
Elhoul BM, Zarai Jaouadi N, Rekik H, Omrane BM, Mechri S, Moujehed E, Kourdali SE, Hattab M, Badis A, Bejar S, Jaouadi B (2016) Biochemical and molecular characterization of new keratinoytic protease from Actinomadura viridilutea DZ50. Int J Biol Macromol 92:299–315
Ferrer M, Golyshina OV, Chernikova TN, Khachane AN, Reyes-Duarte D, Santos VA et al (2005) Novel hydrolase diversity retrieved from a metagenome library of bovine rumen microflora. Environ Microbiol 2005(712):1996–2010
Fieseler L, Quaiser A, Schleper C, Hentschel U (2006) Analysis of the first genome fragment from the marine sponge-associated, novel candidate phylum Poribacteria by environmental genomics. Environ Microbiol 8:612–624
Folio P, Ritt J, Alexandre H, Remize F (2008) Characterization of EprA, a major extracellular protein of Oenococcus oeni with protease activity. Int J Biol Macromol 127:26–31
Fritze D, Flossdorf J, Claus D (1990) Taxonomy of alkaliphilic Bacillus strains. Int J Syst Bacteriol 401:92–97
Gabdrakhmanova LA, Balaban NP, Sharipova MR, Kostrov SV, Akimkina TV, Rudenskaya GN, Leshchinskaya IB (2002) Optimization of Bacillus intermedius glutamyl endopeptidase production by recombinant strain of Bacillus subtilis and localization of glutamyl endopeptidase in Bacillus subtilis. Cells 31:256–263
Ghorbel S, Kammoun M, Soltana H, Nasri M, Hmidet N (2014) Streptomyces Flavogriseus HS1: isolation and characterization of extracellular proteases and their compatibility with laundry detergents. BioMed Res Int. https://doi.org/10.1155/2014/345980
Godfrey T, West S (1996) Industrial Enzymology, 2nd edn. Macmillan, Inc., New York, pp 03–04
Gong BL, Mao RQ, Xiao Y, Jia ML, Zhong XL, Liu Y, Xu PL, Li G (2017) Improvement of enzyme activity and soluble expression of an alkaline protease isolated from oil-polluted mud flat metagenome by random mutagenesis. Enzyme Microb Technol 106:97–105
Gonzalez-Rabade N, Badillo-Corona JA, Aranda-Barradas JS, del Carmen Oliver-Salvador M (2011) Production of plant proteases in vivo and in vitro—a review. Biotechnol Adv. 296:983–996
Gulmez C, Atakisi O, Dalginli KY, Atakisi E (2017) A novel detergent additive: organic solvent- and thermo-alkaline-stable recombinant subtilisin. Int J Biol Macromol 108:436–443
Gulmez C, Atakisi O, Dalginli KY, Atakisi E (2018) A novel detergent additive: organic solvent- and thermo-alkaline-stable recombinant subtilisin. Int J Biol Macromol 108:436–443
Gupta R, Beg QK, Lorenz P (2002) Bacterial alkaline proteases: molecular approaches and industrial applications. Appl Microbiol Biot. 59:15–32
Habbeche A, Saoudi B, Jaouadi B, Haberra S, Kerouaz B, Boudelaa M, Badis A, Ladjama A (2014) Purification and biochemical characterization of a detergent-stable keratinase from a newly thermophilic actinomycete Actinomadura keratinilytica strain Cpt29 isolated from poultry compost. J Biosci Bioeng. 117(4):413–421
Hajji M, Rebai A, Gharsallah N, Nasri M (2008) Optimization of alkaline protease production by Aspergillus clavatus ES1 in Mirabilis jalapa tuber powder using statistical experimental design. App Microbio Biotechnol. 79:915–923
Haki GD, Rakshit SK (2003) Developments in industrially important thermostable enzymes: a review. Bioresour Technol 891:17–34
Handelsman J (2005) Metagenomics: application of genomics to uncultured microorganisms. Microbiol Mol Biol Rev. 691:195
Hatanaka T, Yoshiko Uesugi JA, Iwabuchi M (2005) Purification, characterization cloning, and sequencing of metalloendopeptidase from Streptomyces septatus TH-2. Arch Biochem Biophys 434:289–298
Hu GQ, Guo JT, Liu YC, Zhu H (2009) MetaTISA: metagenomic translation initiation site annotator for improving gene start prediction. Bioinformatics 2514:1843–1845
Jankiewicz U, Larkowska E, Brzezinska MS (2016) Production, characterization, gene cloning, and nematocidal activity of the extracellular protease from Stenotrophomonas maltophilia N4. J Biosci Bioeng 1216:614–618
Jaouadi B, Abdelmalek B, Fodil D, Ferradji FZ, Rekik H, Zarai N, Bejar S (2010) Purification and characterization of a thermostable keratinolytic serine alkaline proteinase from Streptomyces sp. strain AB1 with high stability in organic solvents. Bioresour Technol. 10121:8361–8369
Jeon JH, Kim JT, Kim YJ et al (2009) Cloning and characterization of a new cold-active lipase from a deep-sea sediment metagenome. Appl Microbiol Biotechnol 815:865–874
Jones BV, Sun F, Marchesi JR (2007) Using skimmed milk agar to functionally screen a gut metagenomic library for proteases may lead to false positives. Lett Appl Microbiol 45:418–420
Kasana RC (2010) Proteases from psychrotrophs: an overview. Crit Rev Microbiol 362:134–145
Kirk O, Borchert TV, Fuglsang CC (2002) Industrial enzyme applications. Curr Opin Biotech 133:45
Lee DG, Jeon JH, Jang MK, Kim NY, Lee JH, Lee JH et al (2007) Screening and characterization of a novel fibrinolytic metalloprotease from a metagenomic library. Biotechnol Lett 293:465–472
Lesney M (2003) For more and more industrial applications enzymes, natural and engineered are replacing traditional chemistry. Todays chemist at work. Biocatalysis on Board, pp 21–30. http://www.tcawonline.org
Letunic I, Copley RR, Pils B, Pinkert S, Schultz J, Bork P (2006) SMART 5: domains in the context of genomes and networks. Nucleic Acids Res 34:D257–D260
Mahajan RT, Badgujar SB (2010) Biological aspects of proteolytic enzymes: a review. Pharma Res 39:2048–2068
Marchler-Bauer A, Anderson JB, Cherukuri PF, DeWeese-Scott C, Geer LY, Gwadz M, He S, Hurwitz DI, Jackson JD et al (2005) CDD: a conserved domain database for protein classification. Nucleic Acids Res 33:192–196
Marquart ME, Caballero AR, Chomnawang M, Thibodeaux BA, Twining SS, O’Callaghan RJ (2005) Identification of a novel secreted protease from Pseudomonas aeruginosa that causes corneal erosions. Invest Ophthalmol Vis Sci 4610:3761–3768
Martinez R, Jakob F, Tu R, Siegert P, Maurer KH, Schwaneberg U (2013) Increasing activity and thermal resistance of Bacillus gibsonii alkaline protease BgAP) by directed evolution. Biotechnol Bioeng 1103:711–720
Maruthiah T, Esakkiraj P, Prabakaran G, Palavesam A, Immanuel G (2013) Purification and characterization of moderately halophilic alkaline serine protease from marine Bacillus subtilis AP-MSU 6. Biocatal Agric Biotechnol 2:116–119
Mechri S, Kriaa M, Ben A, Berrouina E, Benmrad MO, Jaouadi NZ, Rekik H et al (2017) Optimized Production and characterization of a detergent-stable protease from Lysinibacillus Fusiformis C250R. Int J Biol Macromol 101:387–393
Mikhailova AG, Khairullin RF, Demidyuk IV, Kostrov SV, Grinberg NV, Burova TV et al (2014) Cloning, sequencing, expression, and characterization of thermostability of oligopeptidase B from Serratia proteamaculans, a novel psychrophilic protease. Protein Expr Purif 93:63–76
Moradian F, Khajeh K, Naderi-Manesh H, Sadeghizadeh M (2009) Isolation, purification and characterization of a surfactants—laundry detergents—and organic solvents-resistant alkaline protease from Bacillus sp. HR-08. Appl Biochem Biotechnol 1591:33–45
Morimoto S, Fujii T (2009) A new approach to retrieve full lengths of functional genes from soil by PCR-DGGE and metagenome walking. Appl Microbiol Biotechnol 832:389–396
Ness JE, Kim S, Gottman A, Pak R, Krebber A, Borchert TV, Govindarajan S, Mundorff EC, Minshull J (2002) Synthetic shuffling expands functional protein diversity by allowing amino acids to recombine independently. Nat Biotechnol 20:1251–1255
Pereira JQ, Ambrosini A, Passaglia LMP, Brandelli A (2017) A new cold-adapted serine peptidase from Antarctic Lysobacter sp. A03: insights about enzyme activity at low temperatures. Int J Biol Macromol 103:854–862
Perfumo A, Banat I, Marchant R (2018) Going green and cold: biosurfactants from low-temperature environments to Biotechnology applications. Trends Biotechnol 363:277–289
Petersen TN, Brunak S, vonHeijne G, Nielsen H (2011) SignalP 4.0: discriminating signal peptides from transmembrane regions. Nat Methods 8:785–786
Prakash M, Banik RM, Koch-Brandt C (2005) Purification and characterization of Bacillus cereus protease suitable for detergent industry. Appl Biochem Biotechnol 1273:143–155
Purohit MK, Singh SP (2013) A metagenomic alkaline protease from saline habitat : cloning, over-expression and functional attributes. Int J Biol Macromol 53:138–143
Qoura F, Kassab E, Reiße S, Antranikian G, Brueck T (2015) Characterization of a new, recombinant thermo-active subtilisin-like serine protease derived from Shewanella arctica. J Mol Catal B Enzym 116:16–23
Rao MB, Tanksale AM, Ghatge MS, Deshpande VV (1998) Molecular and biotechnological protease. Microbiol Mol Biol Rev 623:597–635
Raval VH, Pillai S, Rawal CM, Singh SP (2014) Biochemical and structural characterization of a detergent—stable serine alkaline protease from sea water haloalkaliphilic bacteria. Process Biochem 496:955–962
Ravikumar Y, Nadarajan SP, Hyeon Yoo T, Lee CS, Yun H (2015) Unnatural amino acid mutagenesis-based enzyme engineering. Trends Biotechnol 338:462–470
Ravishankar K, Kumar PS, Jacob B, Saravanan K, Kumar MA, Jacob A (2012) Optimal conditions for production of extracellular alkaline protease from a newly isolated Bacillus subtilis strain AKRS3. Res Biotechnol 35:45–54
Ribeiro-Guimaraes ML, Marengo EB, Tempone AJ, Amaral JJ, Klitzke CF, da Silveira EKX et al (2009) Cloning, expression and characterisation of an HtrA-like serine protease produced in vivo by Mycobacterium leprae. Mem Inst Oswaldo Cruz 1048:1132–1138
Ribitsch D, Karl W, Birner-Gruenberger R, Gruber K, Eiteljoerg I, Remler P et al (2010a) C-terminal truncation of a metagenome-derived detergent protease for effective expression in E. coli. J Biotechnol 150:408–416
Ribitsch D, Karl W, Birner-gruenberger R, Gruber K, Eiteljoerg I, Remler P, Wieland S (2010b) C-Terminal truncation of a metagenome-derived detergent protease for effective expression in E. coli. J Biotechnol 1503:408–416
Ribitsch D, Heumann S, Karl W et al (2012) Extracellular serine proteases from Stenotrophomonas maltophilia: Screening, isolation and heterologous expression in E. coli. J Biotechnol 1571:140–147
Riesenfeld CS, Schloss PD, Handelsman J (2004) Metagenomics: genomic analysis of microbial communities. Annu Rev Genet 381:525–552
Riffel A, Brandelli A (2007) Purification and characterization of a keratinolytic metalloprotease from Chryseobacterium sp. J Biotechnol 1283:693–703
Romanello V, Marcacci M, Dal Molin F, Moschioni M, Censini S, Covacci A, Baritussio AG, Montecucco C, Tonello F (2006) Cloning, expression, purification, and characterization of Streptococcus Pneumoniae IgA1. Protease 45:142–149. https://doi.org/10.1016/j.pep.2005.07.015
Rondon MR, August PR, Bettermann AD et al (2000) Cloning the soil metagenome: a strategy for accessing the genetic and functional diversity of uncultured microorganisms. Appl Environ Microbiol 666:2541–2547
Salwan R, Kasana RC (2013) Purification and characterization of extracellular low temperature active and alkaline stable serine peptidases from psychrotrophic Acinetobacter sp. MN 12 MTCC 10786. Indian J Microbiol 531:63–69
Salwan R, Sharma V, Pal M, Kasana RC, Yadav SK, Gulati A (2018) Heterologous expression and structure-function relationship of low-temperature and alkaline active protease from Acinetobacter sp. IHB B 5011MN12. Int J Biol Macromol 107(Part A):567–574
Sharma V, Salwan R, Sharma PN (2016) Differential response of extracellular proteases of Trichoderma Harzianum against fungal phytopathogens. Curr Microbiol 733:419–425
Sharmila GR, Halami PM, Venkateswaran G (2018) Identification and characterization of a calcium dependent bacillopeptidase from Bacillus subtilis CFR5 with novel kunitz trypsin inhibitor degradation activity. Food Res Int 103:263–272
Shetty R, Vestergaard M, Jessen F, Hagglund P, Knorr V, Koehler P et al (2017) Discovery, cloning and characterisation of proline specific prolyl endopeptidase, a gluten degrading thermo-stable enzyme from Sphaerobacter thermophiles. Enzy Microbial Technol 107:57–63
Shumi W, Hossain T, Anwar MN (2004) Proteolytic activity of a bacterial isolate Bacillus fastidiosus den Dooren de Jong. J Biol Sci 43:370–374
Singh R, Chopra C, Gupta VK (2015) Purification and characterization of CHpro1, a thermotolerant, alkali-stable and oxidation-resisting protease of chumathang hotspring. Sci Bull 60:1252–1260
Sinha R, Khare SK (2013) Characterization of detergent compatible protease of a halophilic Bacillus sp. EMB9: differ-ential role of metal ions in stability and activity. Bioresour Technol 145:357–361
Snajder M, Mihelic M, Turk D, Ulrih NP (2015) Codon optimisation is key for pernisine expression in Escherichia coli. PLoS One 104:1–16
Song H, Li G, Mai W, Huang G, Chen K, Zhou J et al (2014) Codon optimization enhances protein ex- pression of Bombyx mori nucleopolyhedrovirus DNA polymerase in E. coli. Curr Microbiol 68:293–300
Suenaga H (2012) Targeted metagenomics: a high-resolution metagenomics approach for specific gene clusters in complex microbial communities. Environ Microbiol 141:13–22
Tatineni R, Doddapaneni KK, Potumarthi RC et al (2008) Purification and characterization of an alkaline keratinase from Streptomyces sp. Bioresour Technol 996:1596–1602
Terada I, Kwon ST, Miyata Y, Matsuzawa H, Ohta T (1990) Unique precursor structure of an extracellular protease, aqualysin I, with NH2- and COOH-terminal pro-sequences and its processing in Escherichia coli. J Biol Chem 26512:6576–6581
Terpe K (2006) Overview of bacterial expression systems for heterologous protein production: from mo- lecular and biochemical fundamentals to commercial systems. Appl Microbiol Biotechnol 72:211–222
Thebti W, Riahi Y, Belhadj O (2016) Purification and characterization of a new thermostable, haloalkaline, solvent stable and detergent compatible serine protease from Geobacillus toebii strain LBT 77. BioMed Res Int 2016:8
Tork SE, Shahein YE, El-Hakim AE, Abdel-Aty AM, Aly MM (2016) Purification and partial characterization of serine-metallokeratinase from a newly isolated Bacillus pumilus NRC21. Int J Biol Macromol 86:189–196
Touioui SB, Jaouadi NZ, Boudjella H, Ferradji FZ, Belhoul M, Rekik H, Badis A, Bejar S, Jaouadi B (2015) Purification and biochemical characterization of two detergent-stable serine alkaline proteases from Streptomyces sp. strain AH4. World J Microbiol Biotechnol 31:1079–1092
Trachuk LA, Shcheglov AS, Milgotina EI, Chestukhina GG (2005) In vitro maturation pathway of a glutamyl endopeptidase precursor from Bacillus licheniformis. Biochimie 876:529–537
Uchiyama T, Abe T, Ikemura T, Watanabe K (2005) Substrate-induced gene-expression screening of environmental metagenome libraries for isolation of catabolic genes. Nature Biotechnol 231:88–93
Wacker M, Linton D, Hitchen PG, Nita-Lazar M, Haslam SM, North SJ et al (2002) N-Linked glycosylation in Campylobacter jejuni and its functional transfer into E. coli. Science 298:1790–1793
Wani AH, Sharma M, Salwan R, Singh G, Chahota R, Verma S (2016) Cloning, expression, and functional characterization of serine protease Aprv2 from virulent isolate Dichelobacter nodosus of Indian origin. Appl Biochem Biotechnol 180:576–587
Waschkowitz T, Rockstroh S, Daniel R (2009) Isolation and characterization of metalloproteases with a novel domain structure by construction and screening of metagenomic libraries. Appl Environ Microbiol 758:2506–2516
Wati MR, Thanabalu T, Porter AG (1997a) Gene from tropical Bacillus sphaericus encoding a protease closely related to subtilisins from Antarctic bacilli. Biochim Biophy Acta Gene Struct Exp 13521:56–62
Wati MR, Thanabalu T, Porter AG (1997b) Gene from tropical Bacillus Sphaericus encoding a protease closely related to subtilisins from Antarctic Bacilli. Biochim Biophys Acta Gene Struct Exp 13521:56–62
Wu J, Bian Y, Tang B, Chen X, Shen P, Peng Z (2004) Cloning and analysis of WF146 protease, a novel thermophilic subtilisin-like protease with four inserted surface loops. FEMS Microbiol Lett 2302:251–258
Yildirim V, Baltaci MO, Ozgencli I, Sisecioglu M, Adiguzel A, Adiguzel G (2017) Purification and biochemical characterization of a novel thermostable serine alkaline protease from Aeribacillus pallidus C10: a potential additive for detergents. J Enzyme Inhib Med Chem 321:468–477
Yuan L, Kurek I, English J, Keenan R (2005) Laboratory-directed protein evolution. Microbiol Mol Biol Rev 693:373–392
Zaferanloo B, Quang TD, Daumoo S, Ghorbani MM, Mahon PJ, Palombo EA (2014) Optimization of protease production by endophytic fungus, Alternaria alternata, isolated from an Australian native plant. World J Microbiol Biotechnol 306:1755–1762
Zhang H, Mu H, Mo Q, Sun T, Liu Y, Xu M, Wang H, Dai Y, Lu F (2016) Gene cloning, expression and characterization of a novel cold-adapted protease from Planococcus sp. J Mol Catal B Enzym 130:1–8
Acknowledgements
The authors are thankful to SEED Division, Department of Science and Technology, India for providing financial support under the project SP/YO/125/2017. The authors also acknowledge Chandigarh University, Gharuan for providing necessary infrastructure for the successful completion of this article.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Communicated by Erko Stackebrandt.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Salwan, R., Sharma, V. Trends in extracellular serine proteases of bacteria as detergent bioadditive: alternate and environmental friendly tool for detergent industry. Arch Microbiol 201, 863–877 (2019). https://doi.org/10.1007/s00203-019-01662-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-019-01662-8