Abstract
Present study attempts in revealing taxonomic and functional diversity of microorganism from petroleum muck using metagenomics approach. Using Ion Torrent Personal Genome Machine, total of 249 Mb raw data were obtained which was analysed using MG-RAST platform. The taxonomic analysis revealed predominance of Proteobacteria with Gammaproteobacteria as major class and Pseudomonas stutzeri as most abundant organism. Several enzymes involved in aliphatic and aromatic hydrocarbon degradation through both aerobic and anaerobic routes and proteins related to stress response were also present. Comparison of our metagenome with the existing metagenomes from oil-contaminated sites and wastewater treatment plant indicated uniqueness of this metagenome taxonomically and functionally. Based on these results a hypothetical community model showing survival and syntrophy of microorganisms in hydrocarbon-rich environment is proposed. Validation of the metagenome data was done in three tiers by validating major OTUs by isolating oil-degrading microbes, confirmation of key genes responsible for hydrocarbon degradation by Sanger sequencing and studying functional dynamics for degradation of the hydrocarbons by the muck meta-community using GC–MS.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Petroleum popularly known as “Black Gold” has brought revolution in the history of mankind. According to the data obtained from Centre for Climate and Energy Solutions, petroleum contributes about 32.8 % of global energy use. Petroleum products are major drivers of transportation, energy, power, agriculture and industries. However, substantial quantity of petrol and other petroleum products enter into the environment as a result of accidental release. Annually, marine environment receives around 2–10 million tons of crude petroleum oil (Tyagi et al. 2011) which is highly toxic to animals, plants and microbes, thus creating a need of efficient remediation strategies (Lee and Vu 2010).
Remediation strategies which included sorption, volatilization and abiotic transformations are now being replaced with microbial remediation (Peixoto et al. 2011). However, one of the major constraints of microbial remediation is that our understanding of phylogenetic diversity of oil reservoir microbial community is very limited (Vartoukian et al. 2010) probably due to the limitation of the culturing techniques and inadequate understanding about microbial syntrophy (McInerney et al. 2009) for such special man-made environments.
Several molecular techniques have been developed to study organisms that are difficult to grow in the laboratory (Hugenholtz et al. 1998; MacNaughton et al. 1999; Kirk et al. 2004; Su et al. 2012). Metagenomics has emerged as a powerful method to study the diversity of both cultured and uncultured microbes (Satyanarayana et al. 2005; Kimes et al. 2013). It has been successfully employed to study the diversity and functional characteristics of oil spill affected mangrove area in Brazil (Andreote et al. 2012) and deep-sea sediments from Gulf of Mexico (Kimes et al. 2013). However, no attempt has been made to study the microbial population of petroleum pipelines. Such site assumes greater importance as compared to oil-contaminated sites because the indigenous population of this site will contain a population rich in microorganisms that are able to survive in that environment and in the process are resistant or able to degrade hydrocarbons, whereas the latter site will contain indigenous population of that site with increased population of microbes able to degrade/resist hydrocarbons.
Therefore, there is a need to study the metagenome of petroleum pipelines. In view of the above, it may be hypothesized that this metagenome would be different from that of other crude oil-contaminated sites in terms of taxonomic and functional diversity. Further on the basis of the unique environment mainly due to the presence of complex hydrocarbons, it was also hypothesized that microbial population engaged in aliphatic and aromatic hydrocarbon degradation may be present which once known can be cultured and used for evolving better remediation strategies.
This study provides first ever description of next generation sequencing (NGS)-based taxonomic and functional diversity of microbes from petroleum muck sample collected from petroleum pipeline. A comparative assessment of this metagenome with publically available metagenome of oil-contaminated and wastewater treatment plant is also provided. Validation of the metagenomics data obtained was done in three tier confirmatory tests using microbial isolation, gene sequencing and GC–MS. Based on the results obtained, hypothetical mechanistic model is also proposed for survival and syntrophy of microbes in hydrocarbon dominating environment.
Results and discussion
Metagenomic analysis
Metagenomic data of muck sample were analysed using MG-RAST platform using both assembled and unassembled data sets. In unassembled data set, total of 249 Mb data were obtained. Data set contained 2,228,423 sequences with an average length of 111 bps. After quality checking, 47,538 sequences (2.1 %) contained ribosomal RNA genes, 118,423 sequences (5.3 %) contained predicted proteins with known functions and 881,744 sequences (39.6 %) contained predicted proteins with unknown function. A total of 233,529 sequences (10.5 %) did not match with the databases of known rRNA genes or proteins.
Assembly of the sequences was done using MetaVelvet 1.13 from which 68,446 assembled sequences were submitted to MG-RAST for analysis. Around 4 % (2,926) sequences failed to pass the QC in pipeline hence omitted for further analysis. Analysis of the assembled contigs revealed 495 sequences (0.7 %) as ribosomal RNA genes and 46,824 sequences (68.4 %) contained predicted proteins with known functions. Around 19 % sequences contain proteins with unknown function, while 7.3 % sequences have no rRNA genes or predicted proteins. Further analysis of sequences with unknown functions and phylogeny may open newer avenues in understanding hidden diversity present in such hydrocarbon-rich environment.
Taxonomic analysis
Taxonomic analysis of both assembled and unassembled data resulted in similar taxonomic pattern. The data were compared with M5nr database using a maximum e value of 1e−5 with minimum identity and alignment length (basepairs for RNA databases) 60 % and 15 respectively. A holistic view of the petroleum muck diversity is shown in Fig. 1 in the form of a phylogenetic tree. The domain Bacteria (88.90 %) predominated, followed by Eukaryota (0.06 %) and Archaea (0.03 %). In the domain Bacteria, 99.09 % belonged to phylum Proteobacteria, 0.70 % Actinobacteria, 0.11 % Firmicutes and 0.75 % other phyla (Fig. 2a). Apart from predominance in the sample, phylum Proteobacteria is most phylogenetically (Silva et al. 2012) and metabolically diverse group (Spain et al. 2009), and members of this phylum are found to have major role in processes like hydrocarbon degradation (Barragán et al. 2008) and iron oxidation (Hedrich et al. 2011) and hence studied further.
Within phylum Proteobacteria, class Gammaproteobacteria (51.31 %) was most abundant (Fig. 2b) followed by Alphaproteobacteria (41.99 %), Betaproteobacteria (6.45 %), unclassified (0.13 %), Deltaproteobacteria (0.11 %) and Epsilonproteobacteria (0.01 %). Similar observation of predominance of Gammaproteobacteria in oil-contaminated sites was reported by Hernandez-Raquet et al. (2006).
It is worth mentioning that the muck metagenome showed the presence of several xenobiotic and other complex hydrocarbon degraders. Among them Geobacillus thermoleovorans, G. stearothermophilus, G. anatolicus and Bacillus aeolius possess capacity for degradation of C26–C34 even carbon alkanes (Meintanis et al. 2006); Pseudomonas, Pedobacter, Micrococcus, Alcanivorax have been previously reported to possess oil biodegradation capacities (Raghavan and Vivekanandan 1999; Margesin et al. 2003; Khan and Singh 2011; Kostka et al. 2011); Corynebacterium sp. and Brevibacterium erythrogenes have been shown to degrade variety of compounds (Huang et al. 2008) including pristane (2,6,10,14-tetramethyl pentadecane) (Desai and Vyas 2006). Other hydrocarbon degraders included Acidovorax sp. JS42, Brevundimonas sp. BAL3, Erythrobacter, E. litoralis, Parvibaculum lavamentivorans and Sinorhizobium meliloti (Schleheck et al. 2011; Kryachko et al. 2012; Liu and Liu 2013).
In this metagenome, various methylotrophs such as Methylobacterium, Rhodopseudomonas, Xanthobacter, Methylococcus, Methylovorax and Methylophaga were observed. Methylotrophic bacteria have been reported to degrade different xenobiotic compounds such as MTBE (methyl tert butyl ether), dichloromethane, dichloroethane, dichlorobenzene, 1,2,4-trichlorobenzene, chlorotoluene (Janssen et al. 1985; Little et al. 1988; Van den Wijngaard et al. 1992; Dolfing et al. 1993; Piveteau et al. 2001; Chistoserdova et al. 2009) and are widely used in wastewater treatment plants. They also play an important role in carbon, nitrogen and sulphur cycle (Lidstrom 2006). Further, genera involved in sulphate reduction (Desulfotalea, Desulfotomaculum, Desulfococcus, Desulfurivibrio and Desulfobacterium), iron oxidation (Gallionella, Leptospirillum and Sulfolobus) and iron reduction (Geobacter, Desulfuromonas, Pelobacter and Shewanella) were also observed.
Among the different genera obtained, Pseudomonas stutzeri emerged as the most abundant species with maximum number of hits (22,309). Data obtained are supported by recruitment plot in which sequences of this metagenome were compared with reference genome of Pseudomonas stutzeri A1501 (Fig. 3). Recruitment plot showed genome coverage of 86.19 % with 19,104 sequences. Contig information for 3,541 unique sequences within metagenome for organism Pseudomonas stutzeri A1501 with a maximum e value of 0.001 is given in supplementary table S1. This abundance is explained on the basis of its reported potential to metabolize benzoate, cresol, naphthalene, xylene, toluene and phenol (Lalucat et al. 2006). The same species was also successfully isolated from the muck sample (NCBI Accession No. (KF939064, KF939066 and KF939075).
Recruitment plot of muck metagenome with reference genome of Pseudomonas stutzeri A1501 with scale of abundance log 2. The blue circle represents the bacterial contigs for the genome of interest, while the two black rings map genes on the forward and reverse strands. The inner graph consists of two stacked bar plots representing the number of matches to genes on the forward and reverse strands. The bars are colour coded according to the e value of the matches with red (<1e−30), orange (1e−30 to 1e−20), yellow (1e−20 to 1e−10), green (1e−10 to 1e−5) and blue (1e−5 to 1e−3) (color figure online)
Sequences affiliated to domain Archaea were only 0.03 % with major phyla Euryarchaeota (43.75 %), Thaumarchaeota (18.75 %) and Crenarchaeota (6.25 %). These data are supported by the reports that the presence of oil negatively affects abundance of Archaea (Röling et al. 2004). Even this minor level cannot be neglected as these Archaea may have developed a unique adaptive mechanism to resist oil toxicity. However, further studies on these Archaea will be needed to ascertain the above said proposition.
Functional analysis
Functional annotation of the raw as well as assembled sequences was done by MG-RAST platform using SEED and COG database. The data were compared to these systems using maximum e value of 1e−5, a minimum identity of 60 % and a minimum alignment length of 15 measured in amino acid for protein. Major functional categories obtained were related to metabolism of proteins, carbohydrates, RNA, DNA, amino acids, its derivatives, stress response and aromatic compounds. Sulphur, phosphorus and iron metabolism were also obtained. In case of iron metabolism, both iron oxidizers and reducers were obtained suggesting that microbial community within this environment might be involved in transition of iron from Fe(II) to Fe(III) and vice versa. It has been reported that some iron-reducing strains have ability to oxidize benzene, toluene, ethylbenzene, phenol, p-cresol and o-xylene (Kappler and Straub 2005).
Aliphatic hydrocarbon degradation
Several enzymes related to methane metabolism like methane monooxygenase (1.14.13.25), methanol dehydrogenase (EC 1.1.1.244), formaldehyde dehydrogenase (EC 1.2.1.46), formate dehydrogenase (EC 1.2.1.2), methenyl tetrahydromethanopterin cyclohydrolase (EC 3.5.4.27), glycine hydroxymethyl transferase (EC 2.1.2.1) and S-formyl glutathione hydrolase (EC 3.1.2.12) were present in the metagenome. Similar results were obtained with contig level analysis. The former two enzymes, with broad substrate specificities, are responsible for co-oxidation of C1–C8 alkanes to their corresponding 1- and 2-alcohols (Anthony 1982). Rubredoxin-bearing alkane 1-monooxygenase and cytochrome P-450 monooxygenase found in this metagenome are reported to be involved in degradation of higher chain alkanes (Rojo 2009; Nie et al. 2011).
Aromatic hydrocarbon degradation
Although aromatic hydrocarbons were not detected by GC–MS, SEED subsystem annotation revealed the presence of several enzymes involved in both aerobic and anaerobic degradation of aromatic hydrocarbons such as benzoate, toluene, ethylbenzene. Figure 4 shows three categories of aromatic hydrocarbon degradation obtained in this metagenome. Several enzymes of the aerobic metabolism such as benzoate dioxygenase (EC 1.14.12.10), biphenyl dioxygenase (EC 1.14.12.18), toluene dioxygenase (EC 1.14.12. 11), phenol hydroxylase (EC 1.14.13.7), catechol 1,2-dioxygenase (EC 1.13.11.1) and homogentisate 1,2-dioxygenase (EC 1.13.11.5) were present. Enzymes related to anaerobic metabolism included acetyl-CoA acetyl transferase, benzoyl CoA reductase and benzylsuccinate synthase. Apart from this, contig analysis also showed the presence of several other enzymes in this metagenome like muconate cycloisomerase (EC 5.5.1.1), muconolactone delta-isomerase (EC 5.3.3.4), 3-oxoadipate enol-lactonase (EC 3.1.1.24), aryl alcohol dehydrogenase (EC 1.1.1.90), benzyldehyde dehydrogenase (EC 1.2.1.28) 3-oxo-3 hexenedioate decarboxylase (EC 4.1.1.77) indicating the presence of near-complete pathway for aromatic hydrocarbon degradation. The KEGG pathway depicting the hydrocarbon degradation is shown in supplementary figure S1. Occurrence of aerobic degradation of aromatic compounds is supported by the presence of genera Pseudomonas, Burkholderia and Ralstonia (Shinoda et al. 2004) and that of anaerobic by the presence of Desulfatibacillum and Geobacter (Kimes et al. 2013) from this metagenome.
Stress response
Majority of proteins found were related to oxidative stress and detoxification followed by osmotic stress, heat shock and acid stress (Fig. 5). Proteins involved in oxidative stress were catalase, peroxidase, superoxide dismutase, glutathione and glutaredoxins which are known to be upregulated during metabolism of toxic compound and polycyclic aromatic hydrocarbons (Seo et al. 2009). Further, among the detoxification strategies proteins related to formaldehyde detoxification were most common (55.56 %) followed by Nudix proteins (32.48 %).
The presence of formaldehyde at this site could be correlated with the metabolism of microorganisms such as methylotrophs, which produces formaldehyde as an intermediate during metabolism of methylamine, methanol, etc. Its toxicity is mainly due to formation of cross-links with proteins and nucleic acids (Gonzalez et al. 2006), and hence, detoxification is necessary. Further, members of Nudix proteins act as “housecleaning” enzymes to destroy potentially deleterious compounds such as damaged nucleotides, metabolizing intermediates or signalling molecules (Makarova et al. 2001; Ooga et al. 2009). The presence of many complex aromatic and aliphatic hydrocarbons and scarcity of water has created a stressful environment for the species living in the pipeline; hence, the presence of these stress proteins is well explained.
Comparative metagenomics
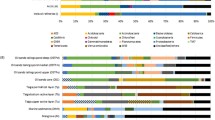
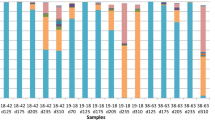
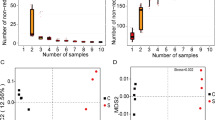
To test our hypothesis that a petroleum pipeline metagenome may be different from that of oil-contaminated sites, this metagenome was compared with six other publically available metagenomes. Principle component analyses (PCA) using MG-RAST M5nr database for taxonomy and subsystem for functional classification revealed that muck metagenome did not cluster with any of the publically available oil-contaminated metagenomes indicating its uniqueness (Fig. 6a, b). The scatter plots shown in Fig. 7a–d explain the uniqueness of muck metagenome. Predominance of Gammaproteobacteria and Alphaproteobacteria in oil-contaminated as well as muck samples suggests that they may act as indicator of oil contamination (Popp et al. 2006; Alonso-Gutiérrez et al. 2009; Nouira et al. 2012). The difference between oil-contaminated sites and muck sample was prominent at order level. Within Gammaproteobacteria, Pseudomonadales was more abundant in muck as compared to deep-sea sediment group which contained higher number of reads for Alteromonadales. Even in Pseudomonadaceae, difference in the species was observed with P. stutzeri and P. mendocina being more common in muck, whereas deep-sea sediment contained mainly P. aeruginosa, P. fluorescens and P. syringae.
Scatter plot showing relative proportion of sequences determined using STAMP software, a and b phylum level distribution; c and d level 1 functional distribution (1 carbohydrate metabolism, 2 protein metabolism, 3 RNA metabolism, 4 respiration; 5 membrane transport; 6 stress response, 7 metabolism of aromatic compounds, 8 nitrogen metabolism, 9 regulation and cell signalling)
Functionally, sequences affiliated to aromatic hydrocarbon degradation, stress responses and membrane transport were more abundant in muck metagenome as compared to other two metagenomic groups (Fig. 7c, d) which is well explained in terms of higher content of hydrocarbons in muck sample.
Validation of metagenomic data
In the fast developing era of genomics, techniques of metagenome analysis has emerged as potential tool for prediction of total taxonomic and functional diversity, studying community modelling and predicting ecological indicators but at the same time it completely relies on the analysis revealed by bioinformatic tools. Hence, establishing link between in silico and in vitro laboratory data is essential and may conclude about the validity of the metagenomic data. In this paper, attempt has been made to validate the metagenome data in three tier system. The approach for taxonomic validation involved, isolation of oil-degrading microbes while the approach to attest the function revealed in metagenome data involved PCR amplification and Sanger sequencing of certain representative genes responsible for hydrocarbon degradation. Apart from this, hydrocarbon-degrading potential of the microbial community present in the muck sample was also studied using GC–MS.
Microbial isolation
In view of the taxonomy revealed by the metagenomic data, microbial isolation was attempted. In silico analysis clearly revealed Pseudomonas stutzeri as the most abundant organism. Therefore, an attempt was made to isolate this organism from muck along with other hydrocarbon degraders. It was possible to successfully isolate three strain of Pseudomonas stutzeri strain BAB-3460, strain BAB-3462 and strain BAB-3471. Apart from this, other known hydrocarbon degraders such as Diaphorobacter sp. BAB-3478, Burkholderia sp. BAB-3479, Pseudomonas sp. BAB-3481, Micrococcus sp. BAB-2954, Bacterium close to Lysinibacillus BAB-2943, Pseudomonas pseudoalcaligenes strain BAB-3067, Pseudomonas aeruginosa strain BAB-3476 and Bacillus pichinotyi strain BAB-3066 were also isolated. Accession numbers of these cultures are KF939064, KF939066, KF939075, KF939082, KF939083, KF939085, KF889300, KF889318, KF960966, KF939080 and KF960965. Phylogeny of these cultures is provided in supplementary figure S2. All these organisms are proven hydrocarbon degraders (Piveteau et al. 2001; Ghazali et al. 2004; Lalucat et al. 2006; Khan and Singh 2011; Jin et al. 2011). Successful isolation of these organisms from muck sample provide attestation of major OTUs from metagenome, and since these are known hydrocarbon degraders, the presence of hydrocarbon degradation pathways as revealed in analysis of metagenome is also verified.
PCR amplification and sequencing of selected genes
The ability of organism to degrade hydrocarbon gives clear evidence for the presence of genes responsible for degradation in that organism. The enzymes used in this study, benzoate dioxygenase (BD), toluene 1,2-dioxygenase (T12D), catechol 1,2-dioxygenase (C12D), are known to play a role in aromatic hydrocarbon degradation, whereas methanol dehydrogenase (MDH) has been reported to be involved in aliphatic hydrocarbon degradation.
It was possible to amplify these genes from the pure isolate of Pseudomonas stutzeri as well as from metagenome. The genes encoding all the enzymes used in this study gave amplification with expected size (BD: 331 bp; T12D: 356 bp; C12D: 213 bp and MDH: 358 bp). The amplified PCR products of Pseudomonas stutzeri were sequenced and submitted to GenBank accession numbers KJ778618, KJ778619, KJ778620. The sequences reconfirmed the presence of key genes encoding hydrocarbon-degrading pathway enzyme in metagenome.
GC–MS analysis
Hydrocarbon analysis of the muck sample
GC–MS analysis of petroleum muck sample revealed the presence of several higher chain alkanes as shown in Table 1. However, except for phenol 2 chloro short chain alkanes and aromatic hydrocarbons were not obtained. This might be due to volatile nature of short chain alkanes (C6–C16) and aromatics (C8–C21) and easy degradation of short and intermediate chain length alkanes (C10–C24) (Kostka et al. 2011). Moreover, only major peaks obtained were analysed by GC–MS.
This muck was then used to assess the hydrocarbon-degrading potential of the microbial community present in it. For this, GC–MS analysis was performed for control (media + crude oil) and experimental sample (media + crude oil + muck) after 30 days incubation. The control sample showed the presence of (C7–C22) linear as well as certain branched alkanes. In the experimental sample, a significant reduction in the hydrocarbon concentration was observed. Most significant degradation was observed in case of undecane (C11), where 84.63 % alkane was removed (Fig. 9). Apart from this, 67 to 84 % reduction was observed in short- and middle-chain (C8–C17) aliphatic compounds. Higher chain alkane such as docosane (C22) showed 51.09 % degradation (Fig. 9). Around 55 % degradation was observed in branched chain alkanes like dodecane, 2-methyl- and dodecane, 2,6,11-trimethyl. This result attests the presence of hydrocarbon degradation pathway as observed in the metagenomic analysis.
Hypothetical model
Based on the metagenomic data, a hypothetical model is proposed to depict community dynamics, survival and syntrophy of microorganisms in petroleum muck (Fig. 8). The central cell in the model depicts the in general mechanism of hydrocarbon uptake and utilization by cells capable of hydrocarbon degradation. In this mechanism, biosurfactants internalize the hydrocarbon, and this entire complex enters the cell by a process similar to pinocytosis (Cameotra and Singh 2009). This is supported by the data generated during functional annotation of the metagenome showing the presence of quorum sensing molecules including autoinducers such as acyl-homoserine lactone (AHL) and Pseudomonas quinolone signal (PQS) which regulate biosurfactant production (Dusane et al. 2010). After uptake, hydrocarbons are degraded by various catabolic pathways and finally enter into central metabolism for energy generation and biomass formation. The model also depicts a synergistic community dynamics between some hydrocarbon degraders with additional phenomenon such as methylotrophy, methanogenesis, denitrification, sulphur and iron reduction and oxidation. It is known that methanogens uses CO2 and H2 to produce methane which in turn is used by methylotrophs generating formate as an intermediate. During oxidation of formate to CO2, electrons generated are transferred to nitrate reductase of its own cell or of other denitrifiers (Ingeldew and Poole 1984). Methane so generated is not only used by methylotrophs but by sulphate reducers as well, and reduced form of sulphur is oxidized by sulphur-oxidizing bacteria. Iron reducers degrade hydrocarbons and at the same time reduce Fe(III) to Fe(II) which in turn is utilized by iron oxidizers. The balance between iron oxidizers and iron reducers is important as imbalance may lead to corrosion of the pipeline. Sulphur-oxidizing and iron-oxidizing bacteria are chemolithoautotrophs utilizing CO2 as a carbon source (Fig. 9).
Since the microbial diversity of the muck metagenome contains microbes such as diazotrophs which cannot utilize hydrocarbon as carbon source (Moreno et al. 1986), it is further proposed that intermediate products formed during hydrocarbon degradation may be used by them. These diazotrophs in turn provides fixed nitrogen to hydrocarbon degraders thereby depicting syntrophy. This finds support from the work of Onwurah and Nwuke (2004) who reported growth of diazotroph, Azotobacter vinlandii, in the presence of oil by utilizing by-products of petroleum hydrocarbon compounds formed by Pseudomonas.
The model further depicts that the organism are living in stress conditions and hence shows expression of various stress response proteins. One of the responses worth mentioning here is that quorum sensing also regulates biofilm formation which is supposed to be a protection mechanism against stressful conditions (Kang and Park 2010). Thus, the model typically depicts a community dynamics prevailing in the petroleum pipeline and association of various microflora found in the metagenome in the form of syntrophy.
Conclusions
Thus, taxonomic and functional metagenomic analysis revealed the presence of both aliphatic and aromatic hydrocarbon degraders as well as enzymes associated with hydrocarbon degradation. Through a hypothetical model of community dynamics, an effort is made to depict microbial syntrophy prevailing in the petroleum pipeline. Further successful metagenome validation using microbial isolation, gene amplification and sequencing and hydrocarbon degradation using GC–MS suggest that the muck community may fulfil the need for efficient removal of huge amount of hydrocarbons which enters in the environment due to accidental oil spillage.
Experimental procedures
Sampling
Petroleum muck sample was kindly provided by Indian Oil Corporation (IOC), Kandla, as a part of pigging (maintenance exercise) for pipeline which was ideal for 6 years. Immediately after pigging, the muck was collected in sterile container and preserved at 4 °C till transported and further processed for metagenome analysis. The sample collection was as per NACE guidelines (NACE Standard TM0194-2004); GPS co-ordinates of this sample are 23°01′53.12″N and 70°12′54.61″E.
Metagenomic DNA isolation and sequencing
Metagenomic DNA extraction was carried out using the Power Soil DNA Isolation kit (Mo BioLaboratories Inc., Carlsbad, CA, USA). DNA was isolated from 0.25 g of muck samples using bead beating method following the manufacturer’s protocol. Qubit® 2.0 Fluorometer (Invitrogen, USA) was used to obtain an accurate quantitation of DNA.
Whole-genome sequencing of the metagenomic DNA was done with a high-throughput Ion Torrent Personal Genome Machine with Ion Torrent Server (Torrent suite version 3.2). Library was prepared using Ion Express Plus Fragment library kit. Metagenomic DNA was sheared into blunt-ended fragments by enzymatic lysis using Ion Shear Plus Reagents. The fragmented DNA was ligated to ion-compatible adapters, followed by nick repair to complete the linkage between adapters and DNA inserts. Sequencing was performed using Ion Express Template 300 chemistry on 318 chip, quality filtered and then exported in FastQ format. Raw reads obtained by Ion Torrent are submitted to SRA with accession SRX314771.
Assembly and annotation
Sequencing data set in FastQ format was uploaded to the Metagenome Rapid Annotation using Subsystem Technology (MG-RAST) server (http://metagenomics.nmpdr.org/) (Meyer et al. 2008) under submission ID 4517794.3. In MG-RAST, sequences were subjected to quality control (QC), which includes dereplication (removing artificial replicate sequences produced by sequencing artefacts), ambiguous base filtering (removing sequences with >5 ambiguous base pairs) and length filtering (removing sequences with a length of >2 standard deviation from mean). Artificial duplicate reads (ADRs) are analysed using duplicate read inferred sequencing error estimation (DRISEE). Near-exact matches to the genomes of fly, mouse, cow and human were also filtered by MG-RAST. After filtering of the data, gene calling is done using FragGeneScan to predict coding regions. Clusters of proteins are build using uclust in QIIME followed by BLAT analysis. The annotation mapping is done against M5nr which provides nonredundant integration of many data sets such as Genbank, SEED, IMG, UniProt, KEGG and eggNOGs.
In order to check contig level analysis, assembly was done with MetaVelvet 1.13 assembler (Namiki et al. 2012) using max k-mer length as 51. Expected coverage was kept as auto and the expected distance between two paired reads was kept 150 in velvetg. The assemblies were uploaded to MG-RAST with submission ID 4556373.3.
Enrichment and isolation of microorganisms
Microorganisms capable of degrading hydrocarbons were enriched in sterile mineral salt medium (MSM) (Teli et al. 2013) supplemented with hydrocarbons as sole carbon source (1 % petrol, diesel, crude oil or engine oil). Approximately 0.1 g of muck sample was inoculated in 10 ml sterile medium and allowed to incubate at 37 °C for 5 days. Cells were subcultured twice in fresh sterile medium followed by its isolation on sterile MSM agar plates. Pure cultures were streaked and maintained on nutrient agar plates.
Molecular identification of bacterial isolates
Genomic DNA of bacterial isolates was extracted as described by Sambrook and Russel (2001) and was PCR amplified using 16S rDNA universal primers 8f (5′-AGAGTTTGATCCTGGCTCAG-3′) and 1492r (5′-ACGGCTACCTTGTTACGACTT-3′). All reactions were carried out in 20 µl volumes with2x ReadyMix™Taq PCR mix (Sigma-Aldrich). PCR was performed in Applied BiosystemVeriti® Thermal Cycler with following reaction conditions: 5 min denaturation at 94 °C, followed by 35 cycles of 30 s denaturation at 94 °C, 2 min annealing at 55 °C, 1.5 min extension at 72 °C and final extension of 5 min at 72 °C. PCR products were purified using affymatrix USB ExoSap-IT PCR product cleanup kit and cycle sequencing was done using BigDye® Terminator v3.1 Cycle Sequencing Kit followed by purification with BigDyeXTerminator® Purification Kit in order to remove unincorporated dye terminators and free salts from postsequencing reactions. Products were analysed using ABI PRISM 3500 XL genetic analyser. The obtained sequences of each isolate were compared with sequences of known bacteria in GenBank database.
Partial amplification of genes involved in hydrocarbon degradation
Four different enzymes were used to confirm the presence of genes involved in hydrocarbon degradation, viz. benzoate 1,2 dioxygenase, catechol 1,2 dioxygenase, toluene 1,2 dioxygenase and methanol dehydrogenase. Primers were designed based upon gene sequences of Pseudomonas stutzeri DSM 10701 in GenBank. Primer details are listed in Table 2. PCR conditions used were stage 1, 94 °C (1 min), stage 2, 94 °C (30 s), 60 °C (2 min), 72 °C (50 s) (35 cycles), stage 3, 72 °C (5 min), 4 °C (∞).
GC–MS analysis of muck sample
In order to check the presence of hydrocarbons in muck sample, equal volume of n-hexane was used to extract hydrocarbons. The n-hexane-soluble fraction was analysed by gas chromatography–mass spectrophotometry (GC–MS) (Autosystem XL GC+). Column used was PE-5MS column (30 m × 0.250 mm × 0.250 µm) with helium as carrier gas. Injection volume was 1 µl with temperature maintained at 250 °C. Oven temperature was programmed to rise from 80 to 280 °C at 10 °C min−1 increment and held at 280 °C for 20 min. Initial holding time was 5 min at 80 °C. Electron ionization temperature was 220 °C. The MS had a mass range of 20–600 atomic mass unit.
GC–MS analysis to assess hydrocarbon degradation potential of muck community
Hydrocarbon degradation study was conducted by inoculating 0.1 g muck sample in MSM containing 0.25 % crude oil. Inoculated broth was maintained at 37 °C and 150 rpm for 30 days. MSM containing 0.25 % crude oil was kept as negative control. After 30 days of incubation period, broth was centrifuged at 10,000 rpm for 2 min. Hydrocarbons were extracted from supernatant with equal volume of n-hexane. n-hexane fraction was analysed using GC–MS (Shimadzu QP2010 Ultra) fitted with a flame ionization detector and Rtx-5 (Restek) column of dimension 30 m × 0.25 mm × 0.25 µm. Parameters for GC–MS were same except for mass range which was 40–700 amu. Per cent hydrocarbon degradation was calculated by comparing peak areas of sample with that of control.
Statistical analysis
For comparative metagenomics, STAMP (Statistical Analyses of Metagenomic Profiles) software v2.0 (Parks and Beiko 2010) was used in which data were uploaded from MG-RAST. Seven metagenomes from different ecosystems were divided into three groups. First group contained muck metagenome. Second group contained metagenomes of deep-sea sediments from Gulf of Mexico after 2010 deepwater horizon oil spill [GoM deep-sea sediment 023 (4465489.3), GoM deep-sea sediment 278 (4465490.3) and GoM deep-sea sediment 315 (4465491.3)]. Third group contained three metagenomes of wastewater treatment plants in which metagenome Beer_CE_Day 44 (4480719.3) examined anaerobic digester sludge whereas M8_META (4494854.3) and M9_META (4494855.3) examined activated sludge. For principle component analysis (PCA), ANOVA and Tukey–Kramer with p value filter (>0.05) were used as statistical hypotheses test and as post hoc test, respectively. High level of variance was explained with 83.8 and 81.1 % (x axis) and 9.1 % and 14.6 % (y axis) for taxonomic and functional plot, respectively. Two-sided Welch’s test was used for scatter plot.
References
Alonso-Gutiérrez J, Figueras A, Albaigés J, Jiménez N, Viñas M, Solanas AM, Novoa B (2009) Bacterial communities from shoreline environments (Costa da Morte, Northwestern Spain) affected by the Prestige oil spill. Appl Environ Microbiol 75:3407–3418
Andreote FD, Jiménez DJ, Chaves D et al (2012) The microbiome of Brazilian mangrove sediments as revealed by metagenomics. PLoS ONE 7:e38600
Anthony C (1982) The biochemistry of methylotrophs. Academic Press, London
Barragán V, Aveiga I, Trueba G (2008) Microbial community composition in petroleum-contaminated and uncontaminated soil from Francisco de Orellana, in the northern Ecuadorian Amazon. Int Microbiol 11:121–126
Cameotra SS, Singh P (2009) Synthesis of rhamnolipid biosurfactant and mode of hexadecane uptake by Pseudomonas species. Microb Cell Fact 8:16–22
Chistoserdova L, Kalyuzhnaya MG, Lidstrom ME (2009) The expanding world of methylotrophic metabolism. Ann Rev Microbiol 63:477
Desai A, Vyas P (2006) Petroleum and hydrocarbon microbiology. Applied Microbiology. Report of Department of Microbiology. MS University of Baroda, Vadodara, pp 1–22
Dolfing J, van den Wijngaard AJ, Janssen DB (1993) Microbiological aspects of the removal of chlorinated hydrocarbons from air. Biodegradation 4:261–282
Dusane DH, Zinjarde SS, Venugopalan VP, Mclean RJ, Weber MM, Rahman PK (2010) Quorum sensing: implications on rhamnolipid biosurfactant production. Biotechnol Genet Eng Rev 27:159–184
Ghazali FM, Rahman RNZA, Salleh AB, Basri M (2004) Biodegradation of hydrocarbons in soil by microbial consortium. Int Biodeterior Biodegrad 54:61–67
Gonzalez CF, Proudfoot M, Brown G, Korniyenko Y, Mori H, Savchenko AV, Yakunin AF (2006) Molecular basis of formaldehyde detoxification characterization of two s-formyl glutathione hydrolases from Escherichia coli, FrmB and YeiG. J Biol Chem 281:14514–14522
Hedrich S, Schlömann M, Johnson DB (2011) The iron-oxidizing Proteobacteria. Microbiology 157:1551–1564
Hernandez-Raquet G, Budzinski H, Caumette P, Dabert P, Le Ménach K, Muyzer G (2006) Molecular diversity studies of bacterial communities of oil polluted microbial mats from the Etang de Berre (France). FEMS Microbiol Ecol 58:550–562
Huang Y, Zhao KX, Shen XH, Jiang CY, Liu SJ (2008) Genetic and biochemical characterization of a 4-hydroxybenzoate hydroxylase from Corynebacterium glutamicum. Appl Microbiol Biotechnol 78:75–83
Hugenholtz P, Goebel BM, Pace NR (1998) Impact of culture-independent studies on the emerging phylogenetic view of bacterial diversity. J Bacteriol 180:4765–4774
Ingeldew WJ, Poole RK (1984) The respiratory chains of Escherichia coli. Microbiol Rev 48:222–271
Janssen DB, Scheper A, Dijkhuizen L, Witholt B (1985) Degradation of halogenated aliphatic compounds by Xanthobacter autotrophicus GJ10. Appl Environ Microbiol 49:673–677
Jin D, Wang P, Bai Z, Bo J, Lv Z, Wei D, Zhuang G (2011) Biodegradation of di-n-butyl phthalate by a newly isolated Diaphorobacter sp. strain QH-6. Afr J Microbiol Res 5:1322–1328
Kang YS, Park W (2010) Contribution of quorum-sensing system to hexadecane degradation and biofilm formation in Acinetobacter sp. strain DR1. J Appl Microbiol 109:1650–1659
Kappler A, Straub KL (2005) Geomicrobiological cycling of iron. Rev Mineral Geochem 59:85–108
Khan JA, Singh S (2011) Evaluation of oil degradation potential of Micrococcus varians. Int J Appl Microbiol Pharm Technol 2:75–80
Kimes NE, Callaghan AV, Atkas D et al (2013) Metagenomic analysis and metabolite profiling of deep-sea sediments from the Gulf of Mexico following the Deepwater Horizon oil spill. Front Microbiol 4:1–17
Kirk JL, Beaudette LA, Hart M, Moutoglis P, Klironomos JN, Lee H, Trevors JT (2004) Methods of studying soil microbial diversity. J Microbiol Methods 58:169–188
Kostka JE, Prakash O, Overholt WA et al (2011) Hydrocarbon-degrading bacteria and the bacterial community response in Gulf of Mexico beach sands impacted by the Deepwater Horizon oil spill. Appl Environ Microbiol 77:7962–7974
Kryachko Y, Dong X, Sensen CW, Voordouw G (2012) Compositions of microbial communities associated with oil and water in a mesothermic oil field. Antonie Van Leeuwenhoek 101:493–506
Lalucat J, Bennasar A, Bosch R, García-Valdés E, Palleroni NJ (2006) Biology of Pseudomonas stutzeri. Microbiol Mol Biol Rev 70:510–547
Lee BK, Vu VT (2010) Sources, distribution and toxicity of polyaromatic hydrocarbons (PAHs). In: Villanyi V (ed) Particulate matter, air pollution. ISBN 978-953-307-143-5, InTech. Available from http://www.intechopen.com/books/air-pollution/sources-distribution-and-toxicity-of-polyaromatic-hydrocarbonspahs-in-particulate-matter
Lidstrom ME (2006) Aerobic methylotrophic prokaryotes. In: Dworkin M, Falkow S, Rosenberg E, Schleifer KH, Stackebrandt E (eds) The prokaryotes. Springer, New York, pp 618–634
Little CD, Palumbo AV, Herbes SE, Lidstrom ME, Tyndall RL, Gilmer PJ (1988) Trichloroethylene biodegradation by a methane-oxidizing bacterium. Appl Environ Microbiol 54:951–956
Liu Z, Liu J (2013) Evaluating bacterial community structures in oil collected from the sea surface and sediment in the northern Gulf of Mexico after the Deepwater Horizon oil spill. Microbiol Open 2:492–504
MacNaughton SJ, Stephen JR, Venosa AD, Davis GA, Chang YJ, White DC (1999) Microbial population changes during bioremediation of an experimental oil spill. Appl Environ Microbiol 65:3566–3574
Makarova KS, Aravind L, Wolf YI, Tatusov RL, Minton KW, Koonin EV, Daly MJ (2001) Genome of the extremely radiation-resistant bacterium Deinococcus radiodurans viewed from the perspective of comparative genomics. Microbiol Mol Biol Rev 65:44–79
Margesin R, Spröer C, Schumann P, Schinner F (2003) Pedobacter cryoconitis sp. nov., a facultative psychrophile from alpine glacier cryoconite. Int J Syst Evol Microbiol 53:1291–1296
McInerney MJ, Sieber JR, Gunsalus RP (2009) Syntrophy in anaerobic global carbon cycles. Curr Opin Biotechnol 20:623–632
Meintanis C, Chalkou KI, Kormas KA, Karagouni AD (2006) Biodegradation of crude oil by thermophilic bacteria isolated from a volcano island. Biodegrad 17:3–9
Meyer F, Paarmann D, D’Souza M et al (2008) The metagenomics RAST server–a public resource for the automatic phylogenetic and functional analysis of metagenomes. BMC Bioinf 9:386
Moreno J, Gonzalez-Lopez J, Vela GR (1986) Survival of Azotobacter spp. in dry soils. Appl Environ Microbiol 51:123–125
Namiki T, Hachiya T, Tanaka H, Sakakibara Y (2012) MetaVelvet: an extension of Velvet assembler to de novo metagenome assembly from short sequence reads. Nucleic Acids Res 40:e155
National Association of Corrosion Engineers, International Standard Test Method, NACE Standard TM0194-2004, item no. 21224
Nie Y, Liang J, Fang H, Tang YQ, Wu XL (2011) Two novel alkane hydroxylase-rubredoxin fusion genes isolated from a Dietzia bacterium and the functions of fused rubredoxin domains in long-chain n-alkane degradation. Appl Environ Microbiol 77:7279–7288
Nouira IZ, Mosbahi DS, Abdelghani S, Bakhrouf A, Rouabhia M (2012) Crude oil metagenomics for better bioremediation of contaminated environments. In: Zerón LR (ed) Introduction to enhanced oil recovery (eor) processes and bioremediation of oil-contaminated sites. In Tech, pp 261–290
Onwurah IN, Nwuke C (2004) Enhanced bioremediation of crude oil-contaminated soil by a Pseudomonas species and mutually associated adapted Azotobacter vinelandii. J Chem Technol Biotechnol 79:491–498
Ooga T, Ohashi Y, Kuramitsu S, Koyama Y, Tomita M, Soga T, Masui R (2009) Degradation of ppGpp by nudixpyrophosphatase modulates the transition of growth phase in the bacterium Thermusthermophilus. J Biol Chem 284:15549–15556
Parks DH, Beiko RG (2010) Identifying biologically relevant differences between metagenomic communities. Bioinformatics 26:715–721
Peixoto RS, Vermelho AB, Rosado AS (2011) Petroleum-degrading enzymes: bioremediation and new prospects. Enzyme Res 2011:1–7
Piveteau P, Fayolle F, Vandecasteele JP, Monot F (2001) Biodegradation of tert-butyl alcohol and related xenobiotics by a methylotrophic bacterial isolate. Appl Microbiol Biotechnol 55:369–373
Popp N, Schlömann M, Mau M (2006) Bacterial diversity in the active stage of a bioremediation system for mineral oil hydrocarbon-contaminated soils. Microbiology 152:3291–3304
Raghavan P, Vivekanandan M (1999) Bioremediation of oil-spilled sites through seeding of naturally adapted Pseudomonas putida. Int Biodeterior Biodegrad 44:29–32
Rojo F (2009) Degradation of alkanes by bacteria. Environ Microbiol 11:2477–2490
Röling WF, de Brito IRC, Swannell RP, Head IM (2004) Response of archaeal communities in beach sediments to spilled oil and bioremediation. Appl Environ Microbiol 70:2614–2620
Sambrook J, Russel DW (2001) Molecular cloning-A laboratory manual, 3rd edn. Cold spring harbor laboratory press, New York
Satyanarayana T, Raghukumar C, Shivaji S (2005) Extremophilic microbes: diversity and perspectives. Curr Sci 89:78–90
Schleheck D, Weiss M, Pitluck S et al (2011) Complete genome sequence of Parvibaculum lavamentivorans type strain (DS-1T). Stand Genomic Sci 5:298–310
Seo JS, Keum YS, Li QX (2009) Bacterial degradation of aromatic compounds. Int J Environ Res Public Health 6:278–309
Shinoda Y, Sakai Y, Uenishi H et al (2004) Aerobic and anaerobic toluene degradation by a newly isolated denitrifying bacterium, Thauera sp. strain DNT-1. Appl Environ Microbiol 70:1385–1392
Silva CC, Hayden H, Sawbridge T et al (2012) Phylogenetic and functional diversity of metagenomic libraries of phenol degrading sludge from petroleum refinery wastewater treatment system. AMB Express 2:1–13
Spain AM, Krumholz LR, Elshahed MS (2009) Abundance, composition, diversity and novelty of soil Proteobacteria. Int Soc Microb Ecol J 3:992–1000
Su C, Lei L, Duan Y, Zhang KQ, Yang J (2012) Culture-independent methods for studying environmental microorganisms: methods, application, and perspective. Appl Microbiol Biotechnol 93:993–1003
Teli N, Verma D, Gavankar R, Bhalerao S (2013) Isolation, characterization and identification of diesel engine oil degrading bacteria from garage soil and comparison of their bioremediation potential. Int Res J Environ Sci 2:48–52
Tyagi M, da Fonseca MMR, de Carvalho CC (2011) Bioaugmentation and biostimulation strategies to improve the effectiveness of bioremediation processes. Biodegrad 22:231–241
Van den Wijngaard A, Van Der Kamp K, Van der Ploeg J, Pries F, Kazemier B, Janssen DB (1992) Degradation of 1, 2-dichloroethane by Ancylobacter aquaticus and other facultative methylotrophs. Appl Environ Microbiol 58:976–983
Vartoukian SR, Palmer RM, Wade WG (2010) Strategies for culture of ‘unculturable’ bacteria. FEMS Microbiol Lett 309:1–7
Acknowledgments
This work was supported by the Department of Science and Technology, Government of Gujarat, under Project-151, a joint initiative of Gujarat Biodiversity Gene Bank, Gujarat Genomics Initiative and the Virtual Institute of Bioinformatics. We are thankful to Indian Oil Corporation, Kandla, Gujarat, India, for providing muck sample.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Erko Stackebrandt.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Joshi, M.N., Dhebar, S.V., Dhebar, S.V. et al. Metagenomics of petroleum muck: revealing microbial diversity and depicting microbial syntrophy. Arch Microbiol 196, 531–544 (2014). https://doi.org/10.1007/s00203-014-0992-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-014-0992-0