Abstract
We explored the molecular diversity and functional capabilities of cytochrome P450 monooxygenases (P450s) from the brown-rot basidiomycete Postia placenta. Using bioinformatic and experimental data, we found 250 genes of P450s in the whole genome, including 60 putative allelic variants. Phylogenetic analysis revealed the presence of 42 families, including 18 novel families. Comparative phylogenetic analysis of P450s from P. placenta and the white-rot basidiomycete Phanerochaete chrysosporium suggested that vigorous gene duplication and molecular evolution occurred after speciation of basidiomycetes. Among the 250 gene models, 184 were isolated as full-length cDNA and transformed into Saccharomyces cerevisiae to construct a functional library in which recombinant P450s were co-expressed with yeast NADPH-P450 oxidoreductase. Using this library, the catalytic potentials of P450s against a wide variety of compounds were investigated. A functionomic survey allowed the discovery of novel catalytic properties of P. placenta P450s. The phylogenetic diversity of the CYP53 family in P. placenta was clear, and CYP53D2 is capable of converting stilbene derivatives. This is the first report of this peculiar function of the CYP53 family. Our increased understanding of the molecular and functional diversity of P450s in this fungus will facilitate comprehension of metabolic diversity in basidiomycetes and has future biotechnology applications.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cytochromes P450 (P450s) comprise a superfamily of heme-containing monooxygenases, which are widely distributed in living organisms, and play numerous physiological roles in the metabolism of endogenous and exogenous compounds (Ortiz de Montellano 2005). The vast majority of P450s are thought to have emerged and diversified during the evolution of species, implying that a multitude of secondary metabolic systems are likely to be associated with a large-scale divergence of P450s. Within the last few years, a series of genome projects have accelerated sequence compilation of P450s, and as a result, the sequence database of P450s has greatly enlarged and continues to increase (Park et al. 2008; Nelson 2009). Thus, it becomes a challenging task to exploit the catalytic functions of numerous P450s to gain a better understanding of metabolic diversity in living organisms. In addition to the biological importance of P450s, it will be of great interest to utilize their catalytic potentials in biotechnology, because regio- and stereo-specific oxidations by P450s promise practical advantages (Guengerich 2002; Ro et al. 2006; Urlacher and Eiben 2006; Chang et al. 2007; Gillam 2008; Grogan 2011). Hence, increase in both molecular and functional information relating to P450s would facilitate advanced research into fungal biology and applied biotechnology (Lamb et al. 2002; Yadav et al. 2006; Sabbadin et al. 2010; Nazir et al. 2011; Hirosue et al. 2011).
Wood-rotting basidiomycetes, often categorized into white-rot and brown-rot basidiomycetes, are common inhabitants of forest litter and play crucial roles in the biospheric carbon cycle (Eriksson et al. 1990). Brown- and white-rot basidiomycetes employ independent strategies for the biodegradation of woody components, despite showing some phylogenetic relationship and taxonomic similarity. A significant ligninolytic potential, for example, is found in white-rot, but not in brown-rot basidiomycetes. However, the molecular mechanisms of brown-rot decay are poorly understood relative to those of white-rot decay (Eriksson et al. 1990; Kirk and Farrell 1987; Gold et al. 1989; Hammel and Moen 1991). There is great interest, however, in the biological performance of brown-rot basidiomycetes (Yelle et al. 2011; Wei et al. 2010; Niemenmaa et al. 2008). Recently, the whole genomes of the white-rot basidiomycete Phanerochaete chrysosporium (Martinez et al. 2004) and the brown-rot basidiomycete Postia placenta (Martinez et al. 2009) have been sequenced and are openly available (http://www.jgi.doe.gov/). Many researchers have taken advantage of these genomic projects and have carried out various “Omics” studies in the post genomic era (Matsuzaki et al. 2008; Sato et al. 2009; Vanden Wymelenberg et al. 2010). Based upon genome-wide comparison, it has been elucidated that the gene number of P450s in P. placenta (PpCYP) is larger than that of the P450s in P. chrysosporium (PcCYP) (Martinez et al. 2009; Hirosue et al. 2011). It could be hypothesized that brown-rot basidiomycetes have invested more heavily in the diversification of the P450 molecule than white-rot basidiomycetes. Thus, it would be of great interest to better understand the metabolic diversity and capabilities of brown-rot basidiomycetes through a comprehensive survey of PpCYPs.
We performed a genome-wide survey and functionomic investigation of PpCYPs. Using bioinformatic and experimental data, we identified 250 gene candidates of PpCYPs including 60 allelic variants. Using RT-PCR techniques, we confirmed gene expression in 210 species and isolated full-length cDNA from 184 species. Furthermore, we developed a functional screening system for PpCYPs in which 184 isoforms were co-expressed with yeast NADPH-P450 reductase in Saccharomyces cerevisiae. A functionomic survey resulted in the discovery of novel catalytic potentials of PpCYPs, providing new insight into their fascinating fungal biology and potential for biotechnology.
Materials and methods
Chemicals
Anthracene, carbazole, dibenzo-p-dioxin, 4-ethoxybenzoic acid, 7-ethoxycoumarin, pyrene, testosterone, and 3,5,4′-trimethoxy-trans-stilbene were purchased from Wako Pure Chemicals (Osaka, Japan). 7-Ethoxycoumarin was purified before use, using a silica gel column (hexane/ethyl acetate). 5-Aminolevulinic acid was purchased from Cosmo Bio Co. (Tokyo, Japan). DO supplement without Leu was purchased from TaKaRa Bio (Shiga, Japan). All other chemicals were reagent grade. Deionized water was obtained using a Milli-Q System (Millipore Japan Co Ltd., Tokyo, Japan).
Bioinformatic annotation of P450 from P. placenta
A possible coding sequence for PpCYPs was found in the US Department of Energy Joint Genome Initiative database, based upon sequence similarity to known P450s (http://genome.jgi-psf.org/Pospl1/Pospl1.home.html). To evaluate annotation accuracy, we identified the P450 signature sequence (F-x-x-G-x-x-x-C-x-G) in the heme-binding domain, the E-x-x-R motif in the K-helix, a conserved Thr in the center of the I-helix, and the hydrophobic transmembrane domain (TMD) at the N-terminal region. TMD sequences were analyzed using SOSUI (Hirokawa et al. 1998; http://bp.nuap.nagoya-u.ac.jp/sosui/). If candidates lacked sequences corresponding to these regions, their capability to encode P450 was judged by overall sequence similarity to known P450s.
Isolation of cDNA by RT-PCR
Postia placenta strain MAD-698 (ATCC 44394) was grown from hyphal inocula at 27°C in a stationary culture (10 mL medium) under aerobic conditions. The medium (pH 6.0) used in this study was, as previously described, utilizing 1% glucose and either 1.2 or 12 mM ammonium tartrate as the carbon and nitrogen sources (Kirk et al. 1978; Nazir et al. 2010). Total RNA was extracted individually from 10, 15, 20, 25, and 28-day-old mycelia using the acid guanidium-phenol–chloroform method (Sambrook and Russel 2001) and further purified using an RNeasy Plant Mini Kit (QIAGEN). The concentration of RNA was calculated from the absorbance at 260 nm. Equal quantities of RNA isolated from mycelia of the five different ages were then mixed and used for RT-PCR, as previously described (Nazir et al. 2010). Gene-specific primers used for PCR amplification were designed to anneal to 5′- and 3′-untranslated regions; to the 2–30 bp upstream or downstream flanking sequences from the putative start and stop codons (Table S1). Target cDNAs were cloned into pBluescript plasmid and sequenced using an automated DNA Sequencer (CEQ 8000; Beckman).
Sequence alignment and phylogenetic analysis
Multiple sequence alignment was carried out using the ClustalX program with a gap penalty of 10 and a gap extension penalty of 0.2 (Thompson et al. 1994). Our phylogenetic tree was constructed using the Unweighted Pair Group Method with Arithmetic Mean (UPGMA), with the Jones-Taylor-Thornton matrix using PHYLIP software (Felsenstein 1989), and visualized using the FigTree program (http://tree.bio.ed.ac.uk/).
Heterologous expression and functional screening of PpCYPs
The open reading frame of each PpCYP was re-amplified from the gene library (cloned in pBluescript vector) using gene-specific primers (Table S2). The resultant gene fragments were used for construction of expression plasmid, as previously described (Nazir et al. 2010). The expression plasmid harboring PpCYP was then transformed into S. cerevisiae AH22, using the Fast ™-Yeast Transformation Kit (G-Biosciences). Carbon monoxide (CO) difference spectra of the transformants were recorded on a UV–Vis spectrophotometer equipped with a head-on photomultiplier (Hitachi; U3900H) (Omura and Sato 1964; Nazir et al. 2011; Hirosue et al. 2011). To construct a functional library, S. cerevisiae harboring each expression plasmid was inoculated into 0.5 mL of synthetic dextrose liquid (SDL) medium (8% glucose, 2.68% yeast nitrogen base without amino acids, 0.1% DO supplement without Leu, and 0.5 mM 5-aminolevulinic acid), and these were simultaneously grown in a 96-deep-well plate. After 4 days incubation, transformants were harvested by centrifugation (1,300×g) and resuspended in 2 mL potassium phosphate (10 mM, pH 7.0) containing 10% glycerol. The 96-well plates accommodating transformants were stored at −80°C. For bioconversion, a 20 μl solution containing transformants was inoculated into 0.5 mL of SDL medium containing substrate (0.5 mM) and 5-aminolevulinic acid and incubated in a Micro Bio Shaker (TAITEC) at 28°C for 2 days. Reactions were stopped by the addition of methanol/acetone (0.5 mL), and metabolic products were analyzed by high-performance liquid chromatography (HPLC), after removal of cell debris by centrifugation (1,300×g) and filtration (0.45 mm, Whatman). If necessary, the metabolic products were extracted using ethyl acetate, purified by preparative HPLC, and analyzed by liquid chromatography electron spray ionization mass spectrometry (LC–ESI–MS) and/or 1H nuclear magnetic resonance (1H-NMR) spectrometry.
Instruments
HPLC analysis was carried out using a Prominence UFLC system (Shimadzu) consisting of two pumps (LC-20AD), an auto-injector (SIL-20AC HT), UV-detector (SPD-20A), and a column oven (CTO-20A). Chromatographic separation was performed using a Shima-pack XR-ODS II column (Shimadzu; 3.0 mm I.D. × 75 mm) with a column temperature of 40°C. The mobile phases for HPLC were (A) water with 0.05% phosphoric acid and (B) acetonitrile. The mobile phase gradient was as follows: 0–0.2 min, 10% B; 0.2–3.2 min, 10–40% B; 3.2–3.6 min, 40–100% B; 3.6–4.0 min, 100% B. The flow rate was 1.4 mL/min. An ultraviolet (UV) monitor was utilized for product detection. 1H-NMR (400 MHz) spectra were obtained using a JEOL JNM-AL400 spectrometer with the chemical shift expressed as parts per million downfield from an internal standard of tetramethylsilane. Samples were dissolved in deuterated methanol or chloroform.
Results and discussion
Genome-wide survey and molecular identification of PpCYPs
The whole genome of P. placenta strain MAD-698 has recently been sequenced (Martinez et al. 2009) and released by the US Department of Energy Joint Genome Initiative (http://genome.jgi-psf.org/Pospl1/Pospl1.home.html). In the current database, 17,173 gene models, over 300 PpCYPs, were predicted in the dikaryotic genome of this fungus; however, the gene model remains complicated due to the allelism of this strain and unidentified and/or unassembled segments. Moreover, it was presumed that gene annotation and prediction errors would be included in the current draft release, and in fact, several PpCYP candidates seem to have unexpected truncations at their N-and/or C-terminal region(s). We further refined the gene annotation accuracy, therefore, and selected gene candidates of PpCYP based on consideration of the conserved sequence feature in eukaryotic P450s (Ortiz de Montellano 2005; Nazir et al. 2010; Hirosue et al. 2011).
A bioinformatic survey was used to identify possible coding sequences of 242 PpCYPs from the whole genome sequence (Table 1; see also Figure S1, Figure S2, and Table S3). Among the 242 candidates, several gene pairs showed high sequence similarity; over 98% at amino acid level. Although this striking sequence homology implies allelic features, it could be expected that P. placenta possesses at least 190 haploidal PpCYP genes showing sequence identity of lower than 90%. In addition, we found several gene fragments showing significant homologies to P450s; however, we were not able to identify their full-length coding sequences because these sequences were connected to unassembled and/or uncharacterized segments. Although the dataset could be slightly enlarged after development of the database, we initiated further studies using the 242 PpCYP candidates.
Insight into PpCYP phylogenies
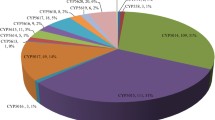
To better understand the molecular evolution of P450s within brown-rot basidiomycetes, we conducted a phylogenetic analysis using the 190 haploidal PpCYPs. The P450 nomenclature committee recommended that families should be considered to share greater than 40% identity, and subfamilies should be considered to share greater than 55% identity of amino acid sequences (Nelson et al. 1996). Based on sequence comparisons, PpCYPs were assigned to 42 families including 18 novel families. A comparative phylogenetic analysis of PpCYPs and PcCYPs revealed interesting aspects, including the revelation that CYP5027, CYP5350, and CYP5348 are large families in P. placenta but are absent from P. chrysosporium (Table 2). These results may highlight an evolutionary trajectory of vigorous gene duplication and molecular evolution within a short evolutionary period after the speciation of basidiomycetes, to meet the requirements of species. In fact, PpCYPs and PcCYPs shared few subfamilies (Table 2) and branched into unique clusters on our phylogenetic tree (Fig. S3). In addition, it was noteworthy that the gene number of the CYP53 family (widely distributed in the fungal kingdom and well known to catalyze benzoate hydroxylation) was significantly higher than in other organisms (Faber et al. 2001). Indeed, P. chrysosporium encodes one CYP53C2 in its genome (Martinez et al. 2004; Matsuzaki and Wariishi 2005). In P. placenta, the CYP53 family was shown to include orphan species assigned to the CYP53C subfamily, and 6 species assigned to a novel CYP53D subfamily. As shown in Fig. 1, PpCYPs CYP53D subfamily from P. placenta consisted of distinctive clusters showing significant phylogenetic distance from other subfamilies, suggesting a possibly unique evolutionary trajectory in the CYP53D subfamily. In addition, it should be noted that several PpCYPs were phylogenetically closer to P450s from ascomycetous fungi than to PcCYPs. P. placenta, for example, possesses a CYP537 family that is also found in Aspergillus species, but not in P. chrysosporium (Kelly et al. 2009; Nazir et al. 2010). PpCYPs assigned to the CYP5148 family showed higher sequence homology with CYP5148B1 from A. clavatus (74% identity) than to CYP5148A1 from P. chrysosporium (52% identity). Thus, one can assume that P. placenta possessed some evolutionary interaction such as gene transfers, at least in part, with ascomycetous fungi.
Phylogenetic tree of the CYP53 family. CYP number is represented by an abbreviated fungal species name. Ab Agaricus bisporus, Ac Aspergillus clavatus, Af Aspergillus flavus, Anid Aspergillus nidulans, Anig Aspergillus niger, Ao Aspergillus oryzae, At Aspergillus terreus, Cc Coprinus cinereus, Ci Coccidioides immitis, Cl Cochliobolus lunatus, Fg Fusarium graminearum, Fo Fusarium oxysporum, Fv Fusarium verticillioides, Lb Laccaria bicolor, Mf Mycosphaerella fijiensis, Mg Mycosphaerella graminicola, Nc Neurospora crassa, Nf Neosartorya fischeri, Nh Nectria haematococca, Pc Phanerochaete chrysosporium, Po Pleurotus ostreatus, Pp Postia placenta, Rm Rhodotorula minuta, Sr Sporobolomyces roseus, Ur Uncinocarpus reesii. P450s from Ab, Cc, Lb, and Po are able to be classified into the CYP53C subfamily but have not been named
cDNA isolation and heterologous expression of PpCYPs
In addition to bioinformatic studies, experimental approaches are compulsory to facilitate advanced research into basic biology and applied biotechnology. We performed isolation and characterization of full-length PpCYP cDNAs. Since several allelic P450s are known to exhibit different catalytic properties (Ingelman-Sundberg 2001; Yu et al. 2002), we aimed to isolate all possible PpCYPs found in the database, including allelic variants, and aimed for heterologous expression. We have previously demonstrated that the gene expression of fungal P450s is affected by cultural conditions (Ichinose et al. 1999, 2002; Chigu et al. 2010). In particular, it has been strongly suggested that the transcriptional regulation of fungal P450s may respond to nitrogen limitation or starvation (Ichinose et al. 1999; Nazir et al. 2010). Therefore, total RNA was extracted from P. placenta grown in a synthetic liquid medium; in which it has been shown that the white-rot basidiomycete P. chrysosporium and the ascomycetous fungus Aspergillus oryzae express a high number of P450s (Chigu et al. 2010; Nazir et al. 2010). Using RT-PCR technique, we isolated 167 species as full-length cDNA. Nevertheless, there are several nucleotide substitutions in some of the isolated P. placenta genes presumably attributable to polymorphisms. In addition, 43 species were amplified as immature cDNA whose open reading frames were shifted by illegal splicing events. During RT-PCR experiments, we obtained 8 allelic variants that could not be found in the current database, but that could share primer sequences with known PpCYPs. Eventually, we identified 250 possible sequence of PpCYPs and experimentation and isolated 175 species as full-length cDNA (Table 1; see also Table S3). cDNA and deduced amino acid sequences are listed in Figure S1 and Figure S2.
Using the isolated cDNA, we performed heterologous expression of PpCYPs in S. cerevisiae. In addition to the 175 full-length cDNAs isolated by RT-PCR, 9 species were generated to encode a theoretical open reading frame by removing unexpected introns from their frame-shifted variants. Thus, 184 PpCYPs could be used for further experimentation. We have successfully expressed a number of fungal P450s from P. chrysosporium and A. oryzae in S. cerevisiae using the expression plasmid pGYR (Hirosue et al. 2011; Nazir et al. 2011). We therefore employed the pGYR vector system for heterologous expression of PpCYPs. To evaluate and optimize heterologous expression, we analyzed the CO difference spectra of transformants. A typical CO difference spectrum for P450s can be observed with an absorption maximum of around 450 nm (Omura and Sato 1964). We clearly demonstrated that an exogenous addition of heme precursor, 5-aminolevulinic acid, to liquid culture medium, can elevate protein concentration of several PpCYPs (Fig. 2); this supports the finding of earlier studies using a plant P450 (Jiang and Morgan 2004). Each transformant was therefore grown in culture medium supplemented with 5-aminolevulinic acid and was applied to spectroscopic analysis. We confirmed the substantial expression of 113 species based upon CO difference spectra. Furthermore, several PpCYPs showed catalytic activities even though we could not confirm their expression using CO difference spectra, due to low levels of expression. Combining the results of spectroscopic analysis and bioconversion, we concluded that at least 116 PpCYPs were functionally expressed in S. cerevisiae (Table 1).
Functional screening of PpCYPs
The expression plasmid pGYR is useful for screening of P450-dependent reactions because P450 is co-expressed with its redox partner cytochrome P450 oxidoreductase (Murakami et al. 1990; Sakaki et al. 1992, 2002; Hirosue et al. 2011; Nazir et al. 2011). To facilitate high-throughput screening, we constructed a functional library in which each transformant was separately inoculated into 0.5-mL culture medium and grown in 96 DeepWell plates. Transformants accommodated in the 96-well plates were easily replicated and used for functionomic surveys, in which each transformant was incubated with a wide variety of compounds, and the resultant metabolic products were comprehensively analyzed. The catalytic potentials of the PpCYPs revealed are summarized in Table 3.
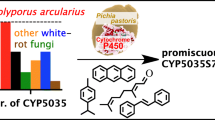
Through comprehensive functional screening, it was demonstrated that CYP53D2 exhibits O-demethylation activity against 3,5,4′-trimethoxy-trans-stilbene to produce 3-hydroxy-5,4′-dimethoxy-trans-stilbene and 3,5-dihydroxy-4′-methoxy-trans-stilbene (Fig. 3 and Table S4). O-demethylation activity was also shown from bioconversion experiments using 3,5-dimethoxy-trans-stilbene. In contrast, no reaction proceeded when trans-stilbene was used as substrate for CYP53D2. This may suggest that CYP53D2 recognizes the methoxyl group(s) in the stilbene derivatives. Although further investigation should be directed at understanding the mechanisms involved, it was noteworthy that substantial activities by CYP53D2 against stilbene derivatives were observed, because the CYP53 family has generally been considered to exhibit substrate specificity against benzoate, and the carboxyl group in benzoate is essential for enzyme-substrate binding (Matsuzaki and Wariishi 2005; Podobnik et al. 2008). To the best of our knowledge, this is the first report describing catalytic potentials of the CYP53 family against stilbene derivatives that seem structurally dissimilar to benzoic acid. However, we could not determine catalytic potentials of CYP53D2 against benzoic acid because S. cerevisiae metabolized benzoate without PpCYP. Thus, it would be of great interest to better understand biochemical aspects of CYP53D2 using purified enzymes. The finding of peculiar functions of CYP53D2 would also support unique evolutionary histories within the CYP53D2 subfamily (Fig. 1). These results should encourage further research to better understand the sequence–structure–function relationships of the CYP53 family. In addition to their biological roles, it would be of interest to utilize PpCYPs for biotechnological applications, such as the production of rare and/or value-added stilbenoids (Roupe et al. 2006; Rimando and Suh 2008). A wide variety of applied researches are now under way.
We have provided a brief overview showing the phylogenetic relationships and functions of PpCYPs. Figure 4 shows a phylogenetic tree combined with functional information relating to PpCYPs. A bioconversion experiment revealed catalytic activities of CYP5139 family against aromatic compounds such as 7-ethoxycoumarin, carbazole, phenanthrene, and/or stilbene (Fig. 4 and Table 3). The ability of the CYP5139 family to convert 7-ethoxycoumarin has been previously demonstrated using CYP5139A1 from P. chrysosporium; this is the orphan PcCYP categorized into the CYP5139 family (Hirosue et al. 2011). These results would suggest that CYP5139 family recognize aromatic ring(s) for substrate binding. CYP512N6v1, CYP512N6v2, and CYP512P showed catalytic activity against the provisional substrates, testosterones. Several PcCYPs assigned into the CYP512 family but belonging to different subfamilies (CYP512C1, E1, F1, and G1) have been shown to convert steroid compounds. Combining the catalytic activities of PpCYPs and PcCYPs, it can be thus hypothesized that the natural substrate(s) of the CYP512 family may be structurally related to steroid compounds. The catalytic potentials of CYP512N6v1, CYP512N6v2, and CYP512P to convert both testosterone and dehydroabietic acid highlight the structural similarity of steroids and abietane diterpenoids. In addition, CYP5150D1, CYP5027B1, and CYP5350B2v1 showed multifunctional properties against a series of polycyclic aromatic hydrocarbons (PAHs), such as anthracene, carbazole, phenanthrene, and pyrene. The versatile functions would play important roles, at least in part, in fungal metabolic systems involved in xenobiotic detoxification. Although further investigation should aim to identify reaction products, our functionomic survey provides novel insights into the catalytic potentials of PpCYPs that will open the door for advanced fungal biology and biotechnology.
Phylogenetic relationships and functions of PpCYPs. Multiple alignment of PpCYPs was carried out using the ClustalW program. The phylogenetic tree was constructed using the Unweighted Pair Group Method with Arithmetic Mean, with the Jones-Taylor-Thornton matrix using PHYLIP software and visualized using the FigTree program. Catalytic potentials of PpCYPs are represented on the concentric circles
In conclusion, we elucidated the molecular diversity of PpCYPs using bioinformatic and experimental approaches. A genome-wide survey highlighted the unique evolutionary histories of P450s in basidiomycetes. In addition, we constructed a functional library that is potentially useful for comprehensive functional screening. A functionomic approach resulted in characterization of novel catalytic properties of PpCYPs. A compilation of both molecular and functional information relating to PpCYPs will help to facilitate the study of fungal biology and applied biotechnology.
References
Chang MCY, Eachus RA, Trieu W, Ro D-K, Keasling JD (2007) Engineering Escherichia coli for production of functionalized terpenoids using plant P450s. Nat Chem Biol 3:274–277
Chigu NL, Hirosue S, Nakamura C, Teramoto H, Ichinose H, Wariishi H (2010) Cytochrome P450 monooxygenases involved in anthracene metabolism by the white-rot basidiomycete Phanerochaete chrysosporium. Appl Microbiol Biotechnol 87:1907–1916
Eriksson K-EL, Blanchette RA, Ander P (1990) Microbial and enzymatic degradation of wood and wood components. Springer, Berlin
Faber BW, Van Gorcom RFM, Duine JA (2001) Purification and characterization of benzoate-para-hydroxylase, a cytochrome P450 (CYP53A1), from Aspergillus niger. Arch Biochem Biophys 394:245–254
Felsenstein J (1989) PHYLIP—phylogeny inference package (version 3.2). Cladistics 5:164–166
Gillam EMJ (2008) Engineering cytochrome P450 enzymes. Chem Res Toxicol 21:220–231
Gold MH, Wariishi H, Valli K (1989) Extracellular peroxidases involved in lignin degradation by the white rot basidiomycete Phanerochaete chrysosporium. In: Whitaker JR, Sonnet PE (eds) Biocatalysis in agricultural biotechnology, ASC symposium series 389. American Chemical Society, Washington D.C, pp 127–140
Grogan G (2011) Cytochromes P450: exploiting diversity and enabling application as biocatalysts. Curr Opin Chem Biol 15:241–248
Guengerich FP (2002) Cytochrome p450 enzymes in the generation of commercial products. Nat Rev Drug Discov 1:359–366
Hammel KE, Moen MA (1991) Depolymerization of a synthetic lignin in vitro by lignin peroxidase. Enzyme Microb Technol 13:15–18
Hirokawa T, Boon-Chieng S, Mitaku S (1998) SOSUI: classification and secondary structure prediction system for membrane proteins. Bioinformatics 14:378–379
Hirosue S, Tazaki M, Hiratsuka N, Yanai S, Kabumoto H, Shinkyo R et al (2011) Insight into functional diversity of cytochrome P450 in the white-rot basidiomycete Phanerochaete chrysosporium: involvement of versatile monooxygenase. Biochem Biophys Res Commun 407:118–123
Ichinose H, Wariishi H, Tanaka H (1999) Biotransformation of recalcitrant 4-methyldibenzothiophene to water-extractable products using lignin-degrading basidiomycete Coriolus versicolor. Biotechnol Prog 15:706–714
Ichinose H, Wariishi H, Tanaka H (2002) Identification and characterization of novel cytochrome P450 genes from the white-rot basidiomycetes Coriolus versicolor. Appl Microbiol Biotechnol 58:97–105
Ingelman-Sundberg M (2001) Implications of polymorphic cytochrome p450-dependent drug metabolism for drug development. Drug Metab Dispos 29:570–573
Jiang H, Morgan JA (2004) Optimization of an in vivo plant P450 monooxygenase system in Saccharomyces cerevisiae. Biotechnol Bioeng 85:130–137
Kelly DE, Krasevec N, Mullins J, Nelson DR (2009) The CYPome (Cytochrome P450 complement) of Aspergillus nidulans. Fungal Genet Biol 46:S53–S61
Kirk TK, Farrell RL (1987) Enzymatic combustion—the microbial-degradation of lignin. Annu Rev Microbiol 41:465–505
Kirk TK, Schultz E, Connors WJ, Lorenz LF, Zeikus JG (1978) Influence of culture parameters on lignin metabolism by Phanerochaete chrysosporium. Arch Microbiol 117:277–285
Lamb DC, Skaug T, Song HL, Jackson CJ, Podust LM, Waterman MR et al (2002) The cytochrome P450 complement (CYPome) of Streptomyces coelicolor A3(2). J Biol Chem 277:24000–24005
Martinez D, Larrondo LF, Putnam N, Gelpke MD, Huang K, Chapman J et al (2004) Genome sequence of the lignocellulose degrading fungus Phanerochaete chrysosporium strain RP78. Nat Biotechnol 22:695–700
Martinez D, Challacombe J, Morgenstern I, Hibbett D, Schmoll M, Kubicek CP et al (2009) Genome, transcriptome, and secretome analysis of wood decay fungus Postia placenta supports unique mechanisms of lignocellulose conversion. Proc Natl Acad Sci USA 106:1954–1959
Matsuzaki F, Wariishi H (2005) Molecular characterization of cytochrome P450 catalyzing hydroxylation of benzoates from the white-rot fungus Phanerochaete chrysosporium. Biochem Biophys Res Commun 334:1184–1190
Matsuzaki F, Shimizu M, Wariishi H (2008) Proteomic and metabolomic analyses of the white-rot fungus Phanerochaete chrysosporium exposed to exogenous benzoic acid. J Proteome Res 7:2342–2350
Murakami H, Yabusaki Y, Sakaki T, Shibata M, Ohkawa H (1990) Expression of cloned yeast NADPH-cytochrome P450 reductase gene in Saccharomyces cerevisiae. J Biochem 108:859–865
Nazir KHMNH, Ichinose H, Wariishi H (2010) Molecular characterization and isolation of cytochrome P450 genes from the filamentous fungus Aspergillus oryzae. Arch Microbiol 192:395–408
Nazir KHMNH, Ichinose H, Wariishi H (2011) Construction and application of a functional library of cytochrome P450 monooxygenases from the filamentous fungus Aspergillus oryzae. Appl Environ Microbiol 77:3147–3150
Nelson DR (2009) The cytochrome p450 homepage. Hum Genomics 4:59–65
Nelson DR, Koymans L, Kamataki T, Stegeman JJ, Feyereisen R, Waxman DJ et al (1996) P450 superfamily: update on new sequences, gene mapping, accession numbers and nomenclature. Pharmacogenetics 6:1–42
Niemenmaa O, Uusi-Rauva A, Hatakka A (2008) Demethoxylation of O14CH3-labelled lignin model compounds by the brown-rot fungi Gloeophyllum trabeum and Poria (Postia) placenta. Biodegradation 19:555–565
Omura T, Sato R (1964) The carbon monoxide-binding pigment of liver microsomes, II. Solubilization, purification, and properties. J Biol Chem 239:2379–2385
Ortiz de Montellano PR (2005) Cytochrome P450: structure, mechanism, and biochemistry, 3rd ed. Kluwer Academic/Plenum Publishers, New York
Park J, Park B, Jung K, Jang S, Yu K, Choi J et al (2008) CFGP: a web-based, comparative fungal genomics platform. Nucleic Acids Res 36:D562–D571
Podobnik B, Stojan J, Lah L, Krasevec N, Seliskar M, Rizner TL et al (2008) CYP53A15 of Cochliobolus lunatus, a target for natural antifungal compounds. J Med Chem 51:3480–3486
Rimando AM, Suh N (2008) Biological/chemopreventive activity of stilbenes and their effect on colon cancer. Planta Med 74:1635–1643
Ro DK, Paradise EM, Ouellet M, Fisher KJ, Newman KL, Ndungu JM et al (2006) Production of the antimalarial drug precursor artemisinic acid in engineered yeast. Nature 440:940–943
Roupe KA, Remsberg CM, Yáñez JA, Davies NM (2006) Pharmacometrics of stilbenes: seguing towards the clinic. Curr Clin Pharmacol 1:81–101
Sabbadin F, Hyde R, Robin A, Hilgarth EM, Delenne M, Flitsch S et al (2010) LICRED: a versatile drop-in vector for rapid generation of redox-self-sufficient cytochrome P450s. Chembiochem 11:987–994
Sakaki T, Akiyoshi-Shibata M, Yabusaki Y, Ohkawa H (1992) Organella-targeted expression of rat liver cytochrome P450c27 in yeast: genetically engineered alteration of mitochondrial P450 into a microsomal form creates a novel functional electron transport chain. J Biol Chem 267:16497–16502
Sakaki T, Shinkyo R, Takita T, Ohta M, Inouye K (2002) Biodegradation of polychlorinated dibenzo-p-dioxins by recombinant yeast expressing rat CYP1A subfamily. Arch Biochem Biophys 410:91–98
Sambrook J, Russel DW (2001) Extraction, purification, and analysis of mRNA from eukaryotic cells. In: Argentine J (ed) Molecular cloning: a laboratory manual, 3rd edn, vol 1. Cold spring harbor laboratory press, New York
Sato S, Feltus FA, Iyer P, Tien M (2009) The first genome-level transcriptome of the wood-degrading fungus Phanerochaete chrysosporium grown on red oak. Curr Genet 55:273–286
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Urlacher VB, Eiben S (2006) Cytochrome P450 monooxygenases: perspectives for synthetic application. Trends Biotechnol 24:324–330
Vanden Wymelenberg A, Gaskell J, Mozuch M, Sabat G, Ralph J, Skyba O et al (2010) Comparative transcriptome and secretome analysis of wood decay fungi Postia placenta and Phanerochaete chrysosporium. Appl Environ Microbiol 76:3599–3610
Wei D, Houtman CJ, Kapich AN, Hunt CG, Cullen D, Hammel KE (2010) Laccase and its role in production of extracellular reactive oxygen species during wood decay by the brown rot basidiomycete Postia placenta. Appl Environ Microbiol 76:2091–2097
Yadav JS, Doddapaneni H, Subramanian V (2006) P450ome of the white rot fungus Phanerochaete chrysosporium: structure, evolution and regulation of expression of genomic P450 clusters. Biochem Soc Trans 34:1165–1169
Yelle DJ, Wei D, Ralph J, Hammel KE (2011) Multidimensional NMR analysis reveals truncated lignin structures in wood decayed by the brown rot basidiomycete Postia placenta. Environ Microbiol 13:1091–1100
Yu A, Kneller BM, Rettie AE, Haining RL (2002) Expression, purification, biochemical characterization, and comparative function of human cytochrome P450 2D6.1, 2D6.2, 2D6.10, and 2D6.17 allelic isoforms. J Pharmacol Exp Ther 303:1291–1300
Acknowledgments
We are deeply grateful to Dr. Osamu Gotoh (Kyoto University, Japan) for technical support of our bioinformatic gene prediction and to Dr. David Nelson (University of Tennessee, USA) for assistance with the CYP naming. This research was supported in part by a Grant-in-Aid (no. 21688013) for Young Scientists (A) from the Ministry of Education, Culture, Sports, Science and Technology, Japan (to H.I.).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Axel Brakhage.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ide, M., Ichinose, H. & Wariishi, H. Molecular identification and functional characterization of cytochrome P450 monooxygenases from the brown-rot basidiomycete Postia placenta . Arch Microbiol 194, 243–253 (2012). https://doi.org/10.1007/s00203-011-0753-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-011-0753-2