Abstract
Effects of different flavonoids on various bacterial strains have been extensively reported; however, the mechanism(s) of their action on bacterial cells remain largely elusive. In this study, the antibacterial mechanism of soybean isoflavone (SI) on Staphylococcus aureus is systematically investigated using 4′6-diamidino-2-phenylindole (DAPI) staining, pBR322DNA decatenation experiment mediated by topoisomerase and agarose gel electrophoresis for direct decatenation. The results of fluorescence microscopy and fluorescence spectrophotometer indicated that DAPI was integrated in Staphylococcus aureus. Additionally, the quantity of both DNA and RNA reduced to 66.47 and 60.18%, respectively, after treated with SI for 28 h. Effects of SI on topoisomerase I and II were also investigated. SI completely inhibited the pBR322DNA unwinding mediated by topoisomerase I and topoisomerase II at the concentration of 6.4 mg/ml and could denature the plasmid DNA at the concentration of 12.8 mg/ml. These results indicate that topoisomerase I and II are the most important targets by SI to restrain bacterial cell division.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In recent years, there is a constant increase in the number of antibiotic-resistant bacteria due to extensive use of antimicrobial agents in domestic animals, which is then subsequently transmitted to humans through the food chain (Zewdu and Cornelius 2008; Angulo et al. 2000). The virulence and pathogenicity of a bacterium increases with increase in its antibiotic resistance (Levy 2005; Peterson 2005). Therefore, it is important to identify alternative drugs that can replace traditional antibiotics, thereby reducing development and spread of resistance. Additionally, elucidating the mechanism of action of these alternative compounds and the resistance of bacterium to these compounds will provide much needed information for the basic research on microbiology.
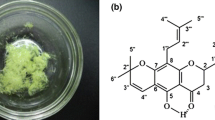
Soybean isoflavone (SI) is a secondary metabolite synthesized during the initial growth stages of soybeans. The SI is present mainly in the cotyledon and hypocotyl of soybean seeds, which is known to exhibit antibacterial properties. The SI is known to have low side effects, wide availability, low cost, less drug-fast characters and other beneficial properties. Therefore, extensive studies have been carried out to optimize the extraction of SI from plants. The SI surprisingly reveals a large spectrum of biological activities, such as anti-oxidation (Wang 2008), anti-tumor effect (Li et al. 2005; Sarkar and Li 2002), estrogen-like activities (An et al. 2001) and antiphlogistic function, and many studies have also investigated the antibacterial effect (Hyunki et al. 2006). To date, researchers pay much attention to the inhibition of the respiration and the synthesis of the cell wall and the cell membrane (Sabanero et al. 1995; Kubo and Taniguchi 1988), the protein (Haraguchi et al. 1996), DNA and RNA (Ulanowska et al. 2006), when developing novel antibacterial drugs. Our former studies on SI identified antimicrobial activity on Staphylococcus aureus, where SI affected the integrity of the cell wall and membrane and prevented the respiratory metabolism and protein synthesis of the bacteria (Wang et al. 2008). However, the nucleic acid metabolism plays a key role in the cell division of bacteria. Therefore, in this study, an attempt was made to investigate the effect of SI on DNA and RNA of Staphylococcus aureus.
DNA metabolism of bacteria, which includes the DNA replication, transcription, recombination and the delivery of genetic information, is related to the dynamic changes in their spatial configuration. DNA topoisomerase is a key enzyme controlling the dynamic changes of nucleic acids (McClendon and Osheroff 2007; Minkah et al. 2007; Wang 2002), and therefore, this enzyme has become one of the key targets for studying the antibacterial effect of naturally occurring antibacterial drugs. In this paper, antibacterial mechanism of SI on Staphylococcus aureus was investigated to provide a theoretical basis for the development of high-performance and low-toxicity antibacterial drugs.
Materials and methods
Bacterial strains
Staphylococcus aureus (ATCC26112) was obtained from the Chinese Medicine Bacteria Preservation Center. The bacteria were cultured in beef extract peptone medium at 37°C in shaking flasks to logarithmic growth phase.
Soybean isoflavone (SI)
Soybean isoflavone (SI), at 90% purity, was extracted from defatted soybean meal at the biology technology laboratory of Liaoning Normal University. The SI was dissolved in absolute ethanol, and concentrated solutions were added to bacterial cultures to keep the ethanol concentration in the cultures as low as possible.
In vivo nucleic acid staining
Staphylococcusaureus (final concentration 2%) was inoculated into 50 ml beef extract peptone medium with SI (final concentration 0.08%), and with ethanol for control group, and cultured at 37°C, in a rotary shaker (120 rpm) for 28 h, and the cells were then collected and resuspended in aquae sterilisata. An equal volume of diluted 4′6-diamidino-2-phenylindole (DAPI) (1:3, diluent, quarter-strength ringer’s solution) was added to obtain the resuspended bacterial culture. The cell samples were then placed on a microslide and kept in the dark for 10 min. The fluorescence of DAPI in cells was observed using inverted fluorescence microscope (OLYMPUS IX71).
Quantification of DNA and RNA
The nucleic acids were quantified using the DAPI staining method mentioned above. Eight hundred microliters of the sample solution (OD580 up to 0.6) treated with SI was added to an equal volume of diluted DAPI and kept in the dark for 10 min. The fluorescence density of DNA and RNA was separately estimated using fluorescence spectrophotometer (F-4500, Hitachi, Japan) with the excitation wavelength of 364 and 400 nm, respectively. Each experiment was repeated three times. Since standard deviation was lower than 15%, representative results are shown.
The extraction of DNA topoisomerase of Staphylococcus aureus
Staphylococcus aureus was collected by centrifugation at 6,000 rpm for 10 min and washed with cold PBS. The cells were then suspended in TMN buffer (1.0 mM Tris–HCl pH 7.5, 1.5 mM MgCl2, 10 mM NaCl) and then centrifuged at 6,000 rpm for 10 min at 4°C. The pellet was resuspended in 1 ml of crude enzyme preparation buffer (100 mM NaCl, 1 mM KH2PO4, 5 mM MgCl2, 5 mM EDTA, 10% glycerol, 0.5 mM PMSF, 250 mM saccharose, 1 mM DTT, pH 6.4) (Sulivan et al. 1986). The cells were then incubated on ice for 30 min and sonicated to enable cell lysis, and the Staphylococcus aureus suspension was then broken up using ultrasonic cell disrupter (SONICS & MATERIALS INC) for 10 s in order to the complete lysis for 15 cycles with 30-s interval between each cycle. Following sonication, the cells were centrifuged at 12,000 rpm for 15 min at 4°C. The supernatant was then transferred to a fresh tube and stored at −20°C. The supernatant fluid contains the crude extract of DNA topoisomerase used to measure the specific activity of DNA topoisomerase.
Determination of DNA topoisomerase activity
The supercoiling topoisomerase activity of DNA was determined by using 0.5 μg of pBR322 DNA. Various dilutions of DNA topoisomerase were added to 2.5 μl helicase buffer I (measuring the activity of topoisomerase I) or helicase buffer II (measuring the activity of topoisomerase II), with the final reaction volume made up to 20 μl with distilled water. The reactions were incubated for 30 min at 37°C. After incubation, 1 μl Proteinase K (10 mg/ml) and 2 μl 10% sodium dodecyl sulfate (SDS) were added and incubated for an additional 30 min at 37°C to stop enzyme activity. The samples were then loaded on 1% agarose gel and visualized using a transilluminator (Wang et al. 1991).
The effect of SI on DNA topoisomerase I/II activity
The complete conversion of per 0.5 μg double-stranded supercoiling pBR322 DNA to helicase DNA by cell extracts, which without/contain ATP, was defined as one activity unit of enzyme. One unit of topoisomerase I/II was added to a 20-μl reaction mix containing 2.5 μl helicase buffer I/II, 0.5 μg of pBR322 and 2 μl of SI or 100% ethanol (control). The reaction conditions were the same as mentioned above. The direct effect of SI on pBR322 DNA was determined by incubating 0.5 μg of pBR322 DNA in a 20-μl volume containing 2 μl SI or 100% ethanol control and 2.5 μl helicase buffer I, at 37°C for 30 min. The reaction was then loaded on 1% agarose gel and visualized using a transilluminator.
Results
Effect of SI on Staphylococcus aureus nucleic acid synthesis
DAPI is fluorescent dye that binds both DNA and RNA. The dye increases in its fluorescence with increase in the quantity of nucleic acids. The DAPI could be visualized using the inverted fluorescence microscope (OLYMPUS IX71), as the dye could penetrate into the bacteria cells and integrate with the nucleic acids. Staphylococcus aureus cells treated with SI for 28 h were incubated with DAPI for 10 min, and the fluorescence intensities of the bacteria were recorded. The fluorescence intensities of the cells treated with DAPI-SI were significantly lower compared to the control group (Fig. 1), which implies that SI could prevent nucleic acid synthesis of Staphylococcus aureus, thereby reducing the nucleic acid content. The fluorescence spectrophotometer measurements indicated that the DNA synthesis was significantly reduced up to 66.47% and that RNA synthesis was also significantly reduced up to 60.18%, compared to the control group (Fig. 2), which indicated that SI had an adverse effect on nucleic acid synthesis. The analysis of the effect of this compound on macromolecular synthesis pathways as revealed by the incorporation of DAPI-labeled precursors showed that SI inhibits nucleic acid synthesis in Staphylococcus aureus. Statistical analysis indicated that compared to the control group, the difference in significant level of SI group was up to the most notable level (P < 0.01).
Effect of SI on the synthesis of nucleic acid in Staphylococcus aureus for 28 h by using fluorescence spectrophotometer. a Changes in DNA of Staphylococcus aureus effected by SI, control group (filled bar), SI group (unfilled bar). b Changes in RNA of Staphylococcus aureus effected by SI, control group (filled bar), SI group (unfilled bar). Error bars show the mean ± SEM (standard error of the mean), n = 3, *P < 0.05, **P < 0.01
Effect of SI on DNA topoisomerase activity
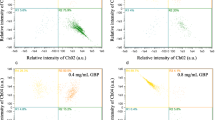
The DNA topoisomerase could alter the supercoiling of double-stranded DNA, and it is a key enzyme that controls the dynamics and synthesis of nucleic acids in bacteria. To understand the possible mechanism of action involved in the inhibition of nucleic acid synthesis, the effect of SI on the DNA topoisomerase activity was investigated using pBR322 DNA decatenation experiment (Fig. 3). The results indicated that with the increase in the concentrations of SI, the quantity of supercoiled DNA (Form I) increased and that of open circular DNA and linear DNA (Form II) decreased, compared to the no effect in the absolute ethanol (negative control). The activity of topoisomerase I was significantly (P < 0.01) inhibited by SI at 6.4 mg/ml. Moreover, SI was relatively less active on topoisomerase II where the inhibition of activity was observed at 6.4 mg/ml (Fig. 4). This phenomenon was similar to the effect on topoisomerase I activity. The results mentioned above suggested that SI acted on both topoisomerase I and II, thereby affecting nucleic acid synthesis and inhibiting bacterial growth.
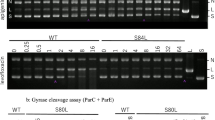
Effect of SI on topoisomerase I in Staphylococcus aureus by using pBR322 DNA decatenation experiment and agarose gel electrophoresis. a The electrophoretogram. b, c The IOD of the bands (Form II: open circular DNA and linear DNA; Form I: supercoiled DNA; a: pBR322 DNA; b: 1 U topoisomerase I; c: 100% ethanol, negative control; d–k: 0.2, 0.4, 0.8, 1.6, 3.2, 6.4, 12.8, 25.6 mg/ml SI). Error bars show the mean ± SEM (standard error of the mean), n = 3, **P < 0.01 (Compared with the negative control)
Effect of SI on topoisomerase II in Staphylococcus aureus by using pBR322 DNA decatenation experiment and agarose gel electrophoresis. a The electrophoretogram. b, c The IOD of the bands (Form II: open circular DNA and linear DNA; Form I: supercoiled DNA; a: pBR322 DNA; b: 1 U topoisomerase II; c: 100% ethanol, negative control; d–k: 0.2, 0.4, 0.8, 1.6, 3.2, 6.4, 12.8, 25.6 mg/ml SI). Error bars show the mean ± SEM (standard error of the mean), n = 3, *P < 0.05, **P < 0.01 (Compared with the negative control)
To investigate the direct cleavage effect of SI on DNA, pBR322 DNA was incubated with different concentrations of SI. The quantity of supercoiled DNA (Form I) was decreased, when the SI concentration was more than 3.2 mg/ml and at this concentration, the quantity of open circular DNA and linear DNA (Form II) increased (Fig. 5), which indicated that SI could cleave pBR322 DNA.
Discussion
The SI is found to have many biological activities, and it is extensively investigated (Wang 2008; Hyunki et al. 2006; Li et al. 2005; Sarkar and Li 2002; An et al. 2001). It has also been known that SI has antibacterial activity with limited information on the mechanisms of its action, especially in the facet of nucleic acid synthesis. This study investigated the effects and mode of action of SI on DNA and RNA quantity of Staphylococcus aureus cells. Incubating S. aureus with SI for 28 h, significantly, affected the quantities of DNA and RNA by reducing to 66.47 and 60.18%, respectively, compared to the control group (without SI). This observation indicates that SI prevents nucleic acid synthesis in Staphylococcus aureus.
DNA topoisomerase is necessary for DNA replication, facilitating short-term separation of single-stranded or double-stranded DNA (McClendon and Osheroff 2007; Minkah et al. 2007; Wang 2002). It has been reported that the efficiency of transcription of rRNA coding genes depends on DNA topology (Hraiky et al. 2000; Rochman et al. 2004). Recent studies found that some drugs exhibit activities by affecting DNA topoisomerase, thereby interfering with DNA replication, gene expression and recombination. Oppegard et al. (2009) found that some novel acridine-based compounds inhibited the proliferation of pancreatic cancer cell lines in vitro and were indicated to be catalytic inhibitors, not poisons, of human topoisomerase II by a series of biochemical assays. In addition, Liu and Liu (2010) showed that nitidine chloride can inhibit hepatic carcinoma growth in nude mice, and the anti-tumor mechanism is probably related to the inhibitory effect on topoisomerase. Some researches, at the same time, showed that genistein, the main component of SI, was a typical topoisomerase inhibitor (Chang et al. 1995; Snyder and Gillies 2002). Although most of these studies were performed using eukaryotic enzymes, considering evolutionary conservation of their active domains, it is likely that SI could influence prokaryotic DNA topology significantly (at least those coming from some species). This could in turn lead to strong negative effects on DNA replication and the reduction in the quantities of DNA. To understand the possible mode of action involved in the inhibition of nucleic acid synthesis, the effect of SI on the activity of S. aureus DNA topoisomerase was investigated. The result showed that SI can inhibit both DNA topoisomerase I and II activity significantly, with complete inhibition of topoisomerase I and II activity at 6.4 mg/ml of SI. Therefore, from this observation, it is inferred that DNA topoisomerase could be one of the direct targets for antibacterial action of SI.
How can SI influence activity of topoisomerase I and II? Recent studies found that the mechanism of topoisomerase inhibitors was to produce drug–topoisomerase–DNA cleavable complexes or interfere with binding of topoisomerase to DNA (Raspaglio et al. 2005); Boege et al. 1996). Otherwise, some researches on the quinolone resistance showed that quinolone-resistant mutants exhibited an alteration in ParC (the subunit of topoisomerase IV) and GyrA (the subunit of DNA gyrase) at a few proteins, suggesting that the ParC subunits and the GyrA subunits are primary topoisomerase targets (Morrissey and George 1999; Pan et al. 1996). Accordingly, it is speculated that the inhibitory effect of SI on topoisomerase may be related to the interaction of the drug, topoisomerase and DNA, and the primary target may be related to the subunit of topoisomerase. However, the specific mode of action and the exact targets involved in the inhibition of DNA topoisomerase would be expected to study further.
References
An J, Tzagarakis-foster C, Scharschmid TC (2001) Estrogen receptor beta-selective trans-criptional activity and recruitment of coregulators by phytoestrogens. J Biol Chem 276:17–808
Angulo F, Johnson K, Tauxe R, Cohen M (2000) Significance and source of antimicrobial-resistant nontyphoidal Salmonella infection in the United States. Microb Drug Resist 1:77–83
Boege F, Straub T, Kehr A, Boesenberg C, Christiansen K, Andersen A, Jakob F, Köhrle J (1996) Selected novel flavones inhibit the DNA binding or the DNA religation step of eukaryotic topoisomerase I. Biol Chem 271:2262–2270
Chang YC, Nair MG, Nitiss JL (1995) Metabolites of daidzein and genistein and their biological activities. J Nat Prod 58:1901–1905
Haraguchi H, OikeS MuroiH, Kubo I (1996) Mode of antibacterial action of totarol, diterpene from Podocarpus nagi. J Planta Med 62:122–125
Hraiky C, Raymond MA, Drolet M (2000) RNase H overproduction corrects a defect at the level of transcription elongation during rRNA synthesis in the absence of DNA topoisomerase I in Escherichia coli. J Biol Chem 275:11257–11263
Hyunki H, Michael R, Landauer M, Foriska G, David L (2006) Antibacterial activity of the soy isoflavone genistein. Basic Microbiol 46:329–335
Kubo I, Taniguchi M (1988) Polygodial, an antifungal potentiator. J Nat Prod 51:22–29
Levy SB (2005) Antibiotic resistance—the problem intensifies. Adv Drug Del Rev 57:1443–1450
Li YZ, Lin QL, Xiao HQ (2005) Research of features progress about soy isoflavones. J Food Nutr China 10:21–22
Liu LM, Liu HG (2010) Anti-hepatoma activity of nitidine chloride and its effect on topoisomerase. Chin Pharmacol Bull 26:497–500
McClendon A, Osheroff N (2007) DNA Topoisomerase II, genotoxicity, and cancer. Mutat Res 623:83–97
Minkah N, Hwang Y, Perry K, Duyne GD, Hendrick RC, Lefkowitz EJ, Hannenhalli S, Bushman FD (2007) Variola virus topoisomerase: DNA cleavage specificity and distribution of sites in poxvirus genomes. Virology 365:60–69
Morrissey I, George J (1999) Activities of fluoroquinolones against Streptococcus pneumoniae Type II topoisomerases purified as recombinant proteins. Antimicrob Agents Chemother 43:2579–2585
Oppegard LM, Ougolkov AV, Luchini DN, Schoon RA, Goodell JR, Kaur H, Billadeau DD, Ferguson DM, Hiasa H (2009) Novel acridine-based compounds that exhibit an anti-pancreatic cancer activity are catalytic inhibitors of human topoisomerase II. Eur J Pharmacol 602:223–229
Pan XS, Ambler J, Mehtar S, Fisher LM (1996) Involvement of topoisomerase IV and DNA gyrase as ciprofloxacin targets in Streptococcus pneumoniae. Antimicrob Agents Chemother 40:2321–2326
Peterson LR (2005) Squeezing the antibiotic balloon: the impact of antimicrobial classes on emerging resistance. Clin Microb Infect 11:4–16
Raspaglio G, Ferlini C, Mozzetti S (2005) Thiocolchicine dimers: a novel class of topoisomerase-I inhibitors. Biochem Pharmacol 69:113–121
Rochman M, Blot N, Dyachenko M, Glaser G, Travers A, Muskhelishvili G (2004) Buffering of stable RNA promoter activity against DNA relaxation requires a far upstream sequence. Mol Microbiol 53:143–152
Sabanero M, Quijano L, Rios T, Trejo R (1995) Encelin: a fungal growth inhibitor. J Planta Med 61:185–186
Sarkar FH, Li Y (2002) Mechanisms of cancer chemoprevention by soy isoflavone genistein. Cancer Metastasis Rev 21:265–280
Snyder RD, Gillies PJ (2002) Evaluation of the clastogenic, DNA intercalative, and topoisomerase II-interactive properties of bioflavonoids in Chinese hamster V79 ells. Environ Mol Mutagen 40:266–276
Sulivan DM, Glisson BS, Hodges PK (1986) Proliferation dependence of topoisomerase II mediated drug action. J Biochem 5:2248–2256
Ulanowska K, Tkaczyk A, Konopa G, Wegrzyn G (2006) Differential antibacterial activity of genistein arising from global inhibition of DNA, RNA and protein synthesis in some bacterial strains. J Arch Microb 14:271–278
Wang JC (2002) Cellular roles of DNA topoisomerases: a molecular perspective. Nat Rev Mol Cell Biol 3:430–440
Wang HB (2008) The study of antioxidant activity of soyasaponin and isoflavone. J Food Res Devel 29:9–12
Wang LG, Liu XM, Ji XJ (1991) Determination of DNA topoisomerase II activity from L1210 cells-a target for screening antitumor agent. J Acta Pharmacol Sinica 12:108–114
Wang HT, Shi SS, Li YX, Li HQ, Xie MJ (2008) Study on anti-microbial activity of genistein and its mechanism. Acta Nutr Sinica 30: 403–406, 409
Zewdu E, Cornelius P (2008) Antimicrobial resistance pattern of Salmonella serotypes isolated from food items and personnel in addis ababa, Ethiopia. Trop Anim Health Prod 41:241–249
Acknowledgments
This work was supported by the Natural Science Foundation of Liaoning Province under grants 20072153 and the Liaoning Provincial Department of Education research project under grants L2010236.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Jose membrillo-Hernandez.
Rights and permissions
About this article
Cite this article
Wang, Q., Wang, H. & Xie, M. Antibacterial mechanism of soybean isoflavone on Staphylococcus aureus . Arch Microbiol 192, 893–898 (2010). https://doi.org/10.1007/s00203-010-0617-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-010-0617-1