Abstract
Antimicrobial susceptibility test of 98 isolates of Salmonella was assayed from September 2003 to February 2004 using the guidelines of the National Committee for Clinical Laboratory Standards (NCCLS).The result revealed that 32.7% of Salmonella isolates were resistant to one or more of the 24 antimicrobials tested. Generally resistance for 13 different antimicrobial drugs was recognized. The most common resistance was to streptomycin (24/32, 75%), ampicillin (19/32, 59.4%), tetracycline (15/32, 46.9%), spectinomycin (13/32, 40.6%) and sulfisoxazole (13/32, 40.6%). All the three Salmonella Kentucky isolates showed resistance to at least 8 antimicrobials. Out of the 12 Salmonella Braenderup isolates, 10 (83.3%) showed multidrug resistance to ampicillin, spectinomycin, streptomycin, sulfisoxazole, sulfamethoxazole/trimethoprim, amoxicillin/clavulanic acid and trimethoprim. Among the 8 S. Hadar isolates 7 (86.5%) showed antimicrobial resistance. All the 6 S. Dublin isolates were resistant to carbadox (100%). All the 6 S. Haifa isolates were resistant for at least ampicillin, streptomycin and tetracycline. Up to ten different antimicrobial resistances pattern was observed. Multiple antimicrobial drug resistance was observed in 23 Salmonella isolates (23.5%). The level of antimicrobial resistance was significantly higher for isolates from chicken carcass (18/29, 62.1%) and pork isolates (5/22, 22.7%) (p = 0.003). The findings of the present study ascertain that significant proportion Salmonella isolates have developed resistance for routinely prescribed antimicrobial drugs and poses considerable health hazards to the consumers unless prudent control measures are instituted.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Antimicrobial-resistant salmonellae are increasing due to the use of antimicrobial agents in food animals, which are subsequently transmitted to humans usually through the food supply (White et al. 2001; Angulo et al. 2000; Fey et al. 2000; Mølback et al. 1999; Tollefson et al. 1998; D’Aoust 1989). Routine assessment of patterns of emerging antibiotic resistant Salmonella strains is of paramount importance because such information channeled to physicians and veterinarians help to timely redirect drug use so as to diminish the development and spread of resistance (Tellefson et al. 1998).
As serovars and phage types and, with them, antibiotic sensitivity patterns can vary annually (Van Duijkeren et al. 1994b); the choice of the drug for treatment of salmonellosis should always be based on sensitivity testing of the causative strain. However, since it takes two or three days before the result is available, blind therapy has to be started in severely ill animals. Therefore, susceptibility testing combined with knowledge of the pharmacokinetic and toxicologic data of the drug are essential in choosing an effective drug for antimicrobial therapy (Prescott and Baggot 1993).
The strains of S. Typhimurium known as definitive phage type 104 (DT 104) have become a worldwide health problem causing illness in humans and animals. It is usually resistant to five drugs: ampicillin, chloramphinicol, streptomycin, sulfonamides, and tetracyclines (White et al. 2001; Hohmann 2001; Lesser and Miller 2001; Cloeckaert et al. 2000; Mølback et al. 1999; Tellefson et al. 1998; Glynn et al. 1998). A report from the United Kingdom suggests that infections caused by this five-drug-resistant S. Typhimurium might be associated with greater morbidity and mortality than other Salmonella infections (Lesser and Miller 2001; Wall et al. 1994).
The emerging resistance in Salmonella is largely a consequence of the use of antimicrobial agents in animals (Acha and Szyfres 2001; Hohmann 2001; Olsen et al. 2001; Gorbach 2001) as well as the indiscriminate prescription-drug treatment of people and animals (Acha and Szyfres 2001). Resistance in Salmonella raise health care costs (Gorbach 2001) and limits the therapeutic options available to veterinarians and physicians in the treatment of certain cases of salmonellosis (Witte 1998).
In recent years, testing of Salmonella isolates has shown that an increasing proportion of isolates are resistant to several antimicrobial agents both in developing and developed countries. The issue of antimicrobial resistance is more complex in developing countries (Leegaard et al. 1996) like Ethiopia where Salmonella is not routinely isolated and resistance to commonly used antimicrobial drugs in veterinary and public health sector not regularly assessed. Therefore, the present study, which was a part of cross-sectional study of Salmonella from supermarket food items and personnel, was undertaken to investigate the susceptibility of Salmonella isolates to commonly used antimicrobial agents in Ethiopia for the treatment of bacterial diseases including salmonellosis.
Materials and methods
Isolation of salmonellae
A total of 1200 food samples i.e. chicken meat (208), pork (194), minced beef (142), mutton (212), cottage cheese (190), fish (128) and ice cream (126) and 68 stool samples collected from retail supermarkets, open markets and shops in Addis Ababa were examined for detection of Salmonella using the technique recommended by the International Organization for Standardization (ISO 1998) during the period from September 2003 to February 2004. Salmonella isolation was successful from all samples except ice cream. Ninety-eight Salmonella isolates identified into fourteen different serotypes were employed for the purpose of this study.
Antimicrobial susceptibility testing
The antimicrobial susceptibility test of all isolates of Salmonella was assayed in the Food Microbiology Laboratory, Laboratory Service Division, Animal Health Laboratory, University of Guelph, Guelph, Ontario; Canada. The National Committee for Clinical Laboratory Standards (NCCLS) (1999) guidelines was followed throughout the agar dilution testing procedure and interpretation of results as susceptible and resistant. Briefly, the isolates were grown to 0.5 – 1.0 McFarland density in Muller Hilton (MH) broth (Difco, Detroit, USA) and replica plated using a Cathra Replicator (Brown and Washington 1978) on to MH agar plates (Difco, Detroit, USA). The list of panel of antimicrobials utilized, their symbols and concentrations to classify an isolate as susceptible or resistant were shown on Table 1. An isolate was defined as resistant if it was resistant to one or more of the antimicrobial drugs tested whereas multiple resistance was defined as resistance to two or more antimicrobial drugs. Standard and reference strains were used following the recommendations of NCCLS (1999).
Fisher’s exact test was used to see the significance of antimicrobial resistance between food items. A difference will be statistically significant if the P-value is less than 0.05. Statistical analysis was performed using Intercooled Stata 6.0 soft ware package.
Results
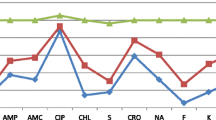
Of the 98 Salmonella serotypes subjected to antimicrobial susceptibility test, using a panel of 24 different antimicrobials (Table 1), 32 serotypes (32.7%) were found resistant to one or more of the antimicrobials used. A total of 66 (67.3%) Salmonella isolates belonging to S. Newport, S. Typhimurium, S. Infantis, S. Bovismorbificans, S. Anatum, S. Zanzibar, S. Kottbus, S. Saintpaul and S. I: 9, 12:- were found to be susceptible to all antimicrobials tested. However, 32 Salmonella isolates (32.7%) belonging to S. Braenderup, S. Hadar, S. Dublin, S. Haifa and S. Kentucky were resistant to one or more of the 24 antimicrobials tested (Tables 2 and 3). All Salmonella isolates belonging to S. Dublin (isolated from pork, mutton and minced beef) were resistant to carbadox and S. Haifa (isolated from pork and cottage cheese) were resistant to ampicillin, streptomycin and tetracycline. About 83% of S. Braenderup isolated from chicken carcass and 87.5% of S. Hadar isolated from chicken carcass and mutton were also resistant to one or more of the antimicrobials tested. In relation to the total Salmonella isolates tested, 24.5% were found resistant to streptomycin, while 19.4%, 15.3%, 13.3% and 13.3% were resistant to ampicillin, tetracycline, spectinomycin and sulfisoxazole, respectively.
With regards to source of the 32 resistant Salmonella isolates, chicken carcass accounted for 56.3% (18/32) while pork, mutton, minced beef and cottage cheese accounted for 21.9% (7/32), 9.4% (3/32), 9.4% (3/32) and 3.1% (1/32), respectively. Among Salmonella isolates from chicken carcass and pork samples 62.1% and 31.8% were resistant for one or more antimicrobials tested (Table 6). All Salmonella isolates from personnel and fish were susceptible to all antimicrobials tested (Tables 2 and 6).
None of the Salmonella isolates showed resistance for the following antimicrobials: amikacin, apramycin, ceftriaxone, ceftiofur, cefoxitin, chloramphinicol, florfenicol, kanamycin, neomycin, nitrofurantoin and tobramycin.
Out of the 32 resistant Salmonella isolates, 23 (23.5%) were multidrug resistant (MDR) (Table 3). The proportion of MDR Salmonella isolates varied between sample types being highest in chicken carcass (65.2%, 15/23), cottage cheese (25%, 1/4) and pork 21.7%, 5/23). It is lowest in minced beef (8.3%, 1/12) and mutton (4.3%, 1/23) samples. Among MDR isolates resistance to streptomycin, spectinomycin, sulfisoxazole, ampicillin and tetracycline was most often observed (Table 5).
Serotypes isolated from chicken carcass (S. Braederup and S. Kentucky) showed resistance pattern for up to ten antimicrobials, while those isolates from pork (S. Haifa) showed resistance pattern for up to four antimicrobials. None of the mutton and cottage cheese isolates showed resistance for more than three antimicrobials and only one serotype from minced beef showed resistance for 8 antimicrobials. The most frequent combination of resistance was seen in S. Braenderup for the following antimicrobials: ampicillin, spectinomycin, streptomycin, sulfisoxazole, sulfamethoxazole/trimethoprim and trimethoprim. The three S. Kentucky serotypes isolated from exotic chicken carcass (one), local chicken carcass (one) and minced beef (one) samples were found to have MDR pattern for 10, 9 and 8 antimicrobials, respectively. Although 12 different antimicrobial resistance patterns were seen in this study, the two most common resistance patterns were Amp Spt Str Sul Sxt Tmp (9 isolates from chicken) and Amp Amc Str Tet (4 isolates from pork). Resistance to trimethoprim and sulfamethoxazole/trimethoprim was seen only in S. Braenderup and S. Kentucky isolated from chicken carcass while to carbadox was seen only among S. Dublin isolates from pork, mutton and minced beef (Tables 3 and 4).
Looking at individual antimicrobial drug, resistance to streptomycin was most frequently observed, followed by ampicillin, tetracycline, spectinomycin, and sulfisoxazole (Table 5). Isolates resistant to these antimicrobials were detected predominantly from chicken carcass and pork.
Discussion
Antimicrobial resistance recognizes no geographical boundaries and increasing rate of resistance of Salmonella isolates have been reported from developing and developed countries. Some of the antimicrobial drugs for which Salmonella serotypes/serogroups were resistant in our study have been reported earlier from Ethiopia (Molla et al. 2003; Alemayehu et al. 2004; Tibaijuka et al. 2003; Mache 2002; Molla et al. 1999b; Mache et al. 1997; Ashenafi and Gedebou 1985; Gedebou and Tassew 1981), other African countries (Leegaard et al. 1996; Adesiyun and Oni 1989; Hadfield and Monson 1985; Hummel 1979) and elsewhere (White et al. 2001; Gebreyes et al. 2000; Tellefson et al. 1998; Lee et al. 1993; D’Aoust et al. 1992).
The finding of 32.7% antimicrobial resistant Salmonella isolates from food samples examined was remarkable. It represents public health hazards due to the fact that food poisoning outbreaks would be difficult to treat and this pool of multi-drug resistant Salmonella in food supply represents a reservoir for transferable resistant genes (Diaz De Aguayo et al. 1992).
Among the important findings of the antimicrobial resistance testing was that 62.1% (18/29) of chicken carcass, 31.8% (7/22) of pork, 25% (1/4) of cottage cheese, 13% (3/23) of mutton and none of the fish and human isolates were antimicrobial resistant (Table 2). The level of resistance was significantly higher for chicken carcass and pork isolates (p = 0.003) (Table 6). Tibaijuka et al. (2003) also reported 60% antimicrobial drug resistance from chicken meat, which was similar to our findings. D’Aoust et al. (1992) also indicated a high antimicrobial resistance among poultry isolates as compared to Salmonella isolated from other sources. Out of the 32 resistant isolates 24 (75%), 19 (59.4%) and 15 (46.9%) were resistant for streptomycin, ampicillin and tetracycline, respectively (Table 5). The significantly high frequency of resistant salmonellae for these antimicrobials was probably an indication of their frequent usage both in livestock and public health sectors. The high prevalence of Salmonella isolates resistant to some of these relatively cheaper and commonly available antimicrobials is disturbing because of the limited access and high cost of newer cephalosporins and quinolone drugs (D’Aoust 1989) for poor citizens of developing countries like Ethiopia. Furthermore, systemic spread of such resistant isolates in human host could lead to serious complications or to a fatal outcome (D’Aoust 1991).
Our antimicrobial drug resistance result indicated that resistance to some extended spectrum cephalosporins (ceftriaxone, ceftiofur), aminoglycosides and newer quinolones was absent, perhaps due to their limited usage in veterinary and public health sectors of Ethiopia. On the other hand the occurrence of resistance to the quinolone (nalidic acid) and fluoroquinolone (ciprofloxacin) in 9.4% of resistant isolates from chicken carcass and minced beef or 3.1% of the total Salmonella isolates (S. Kentucky) was striking because development of resistance undermines the value of this first line drug (ciprofloxacin) for human systemic salmonellosis. Reasons for the emergence of resistance against these drugs were unknown and deserve investigation. However, introduction of resistant Salmonella with importation of food items and travelers was suspected (D’Aoust 1994). MDR was higher in Salmonella isolates from chicken carcass and pork. Thus, 65.2% of MDR isolates were S. Braenderup, S. Hadar and S. Kentucky from chicken carcass and 21.7% of MDR isolates from pork were S. Haifa. These all show that antimicrobial resistant Salmonella serotypes are widespread and more common particularly from chicken carcass, cottage cheese and pork samples as compared to mutton and minced beef. The isolation of susceptible S. Newport among supermarket butchery workers and food items examined indicate that the source of contamination could be either from reservoir animals or personnel. The reasons for the recovery of antimicrobial resistant Salmonella serotypes was most likely due to the indiscriminate use of antimicrobials (Guthrie 1992; WHO 1988), self-medication due to easy access to antibiotics without prescription (Acha and Szyfres 2001) in public health sector and the administration of sub-therapeutic dose of antimicrobials to livestock for prophylactic or nutritional purpose. Such agricultural practices introduce selective pressures that potentiate the emergence and distribution of resistant salmonellae in meats and other products (D’Aoust 1989). Therefore, attempts should be made to reduce the magnitude of the problem at various levels through prudent use of antimicrobials. The tendency of salmonellae for intra- and inter-generic exchange of cytoplasmic DNA (R plasmid) that encodes for single or multiple antimicrobial resistances is another contributing factor (D’Aoust et al. 1992; D’Aoust 1989, 1991; WHO 1988). Nonetheless, there is a need to relate the type and amount of antimicrobial drugs used in intensive farms with data from systematic survey of resistant Salmonella infection to monitor changing resistance and to determine if change in the frequency and pattern of resistance are related to specific pattern of antimicrobial usage (Lee et al. 1993).
None of the S. Typhimurium isolates were found resistant to any of the antimicrobial drugs used. In contrast, Alemayehu et al. (2004) detected MDR strain of S. Typhimurium phage type 2 and Molla et al. (1999b) reported 60% of S. Typhimurium isolates from chicken and minced beef to be MDR. Leegaard et al. (1996) also reported MDR S. Typhimurium. The absence of antimicrobial resistant Salmonella isolates from minced beef in Nyeleti et al. (2000) and 25% (3/12) resistant isolates in the present investigation was contrasting and suggests that antimicrobial resistant salmonellae from minced beef are emerging through time.
The present study demonstrated that supermarket meat samples particularly dressed chicken carcass and pork, were important sources of antimicrobial resistant Salmonella serotypes for consumers and stressed the need to regulate the ethical usage of antimicrobials and regular monitoring of antimicrobial resistance.
Conclusion
Antimicrobial susceptibility testing of Salmonella isolates from food items and food handling personnel is of considerable importance in attempt of supplying sound and safe food for community people. Antimicrobial resistance of Salmonella isolates from food items may suggest the possible existence of antimicrobial resistance at farm level among food animals. Significant proportions of Salmonella isolates were resistant for antimicrobials (32.7%), of which 23.5% were MDR. This could make treatment of humans’ clinical salmonellosis and other bacterial diseases difficult should food poisoning by similar resistant Salmonella serotype ensue. Among MDR serotypes, Salmonella Kentucky was resistant to up to ten antimicrobials. The findings also suggest the need for developing educational program to address issues related to the consumption of raw animal products that might contain antimicrobial resistant Salmonella.
References
Acha, P.N. and Szyfres, B. (2001): Zoonoses and Communicable Diseases Common to Man and Animals. 3rd ed., Washington DC: Pan American Health Organization, Vol.1. Pp 233–246.
Adesiyun, A.A., and Oni, O.O. (1989): Prevalence and Antibiograms of salmonellae in slaughter cattle, slaughter areas and effluents in Zaria abattoir, Nigeria. Journal of Food Protection, 52 (4): 232–235.
Alemayehu, D., Molla, B., and Muckle, A. (2004): Prevalence and antimicrobial resistance pattern of Salmonella isolates from apparently healthy slaughtered cattle in Ethiopia. Tropical Animal Health and Production, 35: 309–319. DOI 10.1023/A:1025189204496.
Angulo, F., Johnson, K., Tauxe, R., and Cohen, M. (2000): Significance and source of antimicrobial-resistant nontyphoidal Salmonella infection in the United States. Microbial Drug resistance, 6 (1): 77–83.
Ashenafi, M., and Gedebou, M. (1985): Salmonella and Shigella in adult diarrhea in Addis Ababa – prevalence and antibiograms. Transaction of the Royal Society of Tropical Medicine and Hygiene, 79: 719–721 Medline. DOI 10.1016/0035-9203(85)90201-9.
Cloeckaert, A., Boumedine, K.S., Flaujac, G., Imberechts, H., D’Hooghe, I., Chaslus-Dancla, E. (2000): Occurrence of a Salmonella enterica serovar typhimurium DT104-like antibiotic resistance gene cluster including the floR gene in S. enterica serovar agona. Antimicrobial Agent Chemotherapy, 44 (5): 1350–13461. DOI 10.1128/AAC.44.5.1359-1361.2000.
D’Aoust, J.-Y. (1989): Salmonella. In: Doyle, M.P. (ed). Foodborne Bacterial Pathogens. New York Marcel Dekker Inc. Pp 327–445.
D’Aoust, J.-Y. (1994): Salmonella and the international food trade. International Journal of Food Microbiology, 24: 11–31 Medline. DOI 10.1016/0168-1605(94)90103-1.
D’Aoust, J.-Y. (1991): Pathogenecity of foodborne Salmonella. International Journal of Food Microbiology, 12: 17–40 Medline. DOI 10.1016/0168-1605(91)90045-Q.
D’Aoust, J.-Y., Sewell, A.M., Daley, E., and Greco, P. (1992): Antibiotic resistance of agricultural and foodborne Salmonella isolated in Canada: 1986–1989. Journal of Food Protection, 55 (6), 428–434.
Diaz De Aguayo, M.E., Duarte, A.B.L., and Montes De Oca Canastillo, F. (1992): Incidence of multiple antibiotic resistant organisms isolated from retail milk products in Hermosillo, Mexico. Journal of Food Protection, 55 (5): 370–373.
Fey, P.C., Safranek, T.J., Rupp, M.E., Dunne, E.F., Ribot, E., Iwen, P.C., Bradford, P.A., Angulo, F.J., and Hinrichs, S.H. (2000): Ceftriaxone-resistant Salmonella infection acquired by a child from cattle. New England Journal of Medicine, 342 (17): 1242–1249 Medline. DOI 10.1056/NEJM200004273421703.
Gebreyes, W.A., Davies, P.R., Morrow, W.E.M., Funk, J.A. and Altier, C. (2000): Antimicrobial resistance of Salmonella isolated from swine. Journal of Clinical Microbiology, 38 12: 4633–4635 Medline.
Gedebou, M. and Tassew, A. (1981): Antimicrobial resistance and R factor of Salmonella isolates from Addis Ababa. Ethiopian Medical Journal, 19: 77–85 Medline.
Glynn, M.K., Bopp, C., Dewitt, W., Dabney, P., Mokhtar, M., and Angulo, F.J. (1998): Emergence of multidrug-resistant Salmonella enterica serotype Typhimurium DT104 infection in the United States. New England Journal of Medicine, 338 (19): 1333–1339 Medline. DOI 10.1056/NEJM199805073381901.
Gorbach, S.L. (2001): Antimicrobial use in animal feed – time to stop. New England Journal of Medicine, 345 (16): 1202–1203 Medline. DOI 10.1056/NEJM200110183451610.
Guthrie, R.K. (1992): Salmonella. CRS Press, USA. Pp 23–156.
Hadfield, T.L. and Monson, M.H. (1985): An outbreak of antibiotic-resistant Salmonella Enteritidis in Liberia, West Africa. Journal of Infectious Disease, 151 (5): 790–795 Medline.
Hohmann, E.L. (2001): Nontyphoidal salmonellosis. Clinical Infectious Disease, 32: 253–269. DOI 10.1086/318457.
Hummel, P.H. (1979): Antibiotic resistance among salmonellae isolated from animals in Tanzania. Bulletin of Animal Health and Production in Africa, 27: 113–121.
International Organization for Standardization 6579 (1998): Microbiology of food and animal feeding stuff-horizontal method for the detection of Salmonella, ISO, Geneva.
Lee, L.A., Threatt, V.L., Puhr, N.D., Levine, P., Ferris, K., Tauxe, R.V. (1993): Antimicrobial–resistant Salmonella spp isolated from healthy broiler chickens after slaughter. Journal of American Veterinary Medical Association, 202 (5): 752–755 Medline.
Leegaard, T.M., Van Gestel, M.H., Petit, P.L.C., and Van De Klundert, J.A.M. (1996): Antibiotic resistance mechanisms in Salmonella species causing bacteraemia in Malawi and Kenya. Acta Pathologica Microbiologica et Immunologica Scandinavica, 104: 302–306.
Lesser, C.F., and Miller, S.I. (2001): Salmonellosis. In: Harrison’s Principles of Internal Medicine. Braunwald, E., Fauci, A.S., Kasper, D.L., Hauser, S.L., Longo, D.L., and Jameson, J.L. (eds.) 15 th ed. Vol.1, The McGraw – Hill Companies, Inc. India. Pp 973–975.
Mache, A. (2002): Salmonella serogroups and their antibiotic resistance patterns isolated from diarrhoeal stools of padiatric out-patients in Jimma Hospital and Jimma Health Center, South West Ethiopia. Ethiopian Journal of Health Science, 12 (1): 37–45.
Mache, A., Mengistu, Y., and Cowley, S. (1997): Salmonella serogrouups identified from adult diarrhoeal out-patients in Addis Ababa, Ethiopia: Antibiotic resistance and plasmid profile analysis. East African Medical Journal, 74 (3): 183–186 Medline.
Mølback, K., Baggesen, D.L., Aarestrup, F.M., et al., (1999): An outbreak of multi-drug resistant, quinolone-resistant Salmonella enterica serotype Typhimurium DT 104. New England Journal of Medicine, 19: 1420–1425. DOI 10.1056/NEJM199911043411902.
Molla, B.,Kleer, J. and Sinell, H.-J. (1999a): Occurrence, distribution and level of Salmonella in selected food items in Addis Ababa (Ethiopia). Fleischwirtschaftlisch International, 4: 37–39.
Molla, B., Kleer, J. and Sinell, H.-J. (1999b): Antibiotic resistance pattern of foodborne Salmonella isolates in Addis Ababa (Ethiopia). München Tierärztlisch Wochenschr 112:41–43.
Molla, B, Mesfin, A, and Alemayehu, D. (2003): Multiple antimicrobial–resistant Salmonella serotypes isolated from chicken carcass and giblets in Debre Zeit and Addis Ababa, Ethiopia. Ethiopian Journal of Health Development, 17 (2): 131–149.
National Committee for Clinical Laboratory Standards (1999): Performance standards for antimicrobial disc and dilution susceptibility tests for bacteria isolated from animals and humans. Approved Standard. NCCLS Document M31– A, NCCLS, Villanova, PA.
Nyeleti, C., Molla, B., Hilderbandt, G., and Kleer, J. (2000): The prevalence and distribution of Salmonella in slaughter cattle, slaughterhouse personnel and minced beef in Addis Ababa, Ethiopia. Bulletin of Animal Health and Production in Africa, 48: 19–24.
Olsen, S.J., DeBess, E.E., McGivern, T.E., Marano, N., Eby, T., Mauvais, S., Balan, V.K. Zirnstein, G. Cieslak, P.R. and Angulo, F.J. (2001): A nosocomial outbreak of fluoroquinolone-resistant Salmonella infection. New England Journal of Medicine, 344 (21): 1572–1579 Medline. DOI 10.1056/NEJM200105243442102.
Prescott, J.P. and Baggot, J.D. (1993): Antimicrobial susceptibility and drug dosage prediction. Sulfonamides,trimethoprim, ormetoprim and their combinations. Miscellaneous antibiotics: Ionophores, nitrofurans, nitroimidazoles, rifampin, and others. In: J.P. Prescott and J.D. Baggot (eds.) Antimicrobial Therapy in Veterinary Medicine. II Iowa State University Press; Pp 11–20.
Tellefson, L., Angulo, F., and Fedorka-Cray, P. (1998): National surveillance for antibiotic resistance in zoonotic enteric pathogens. Veterinary Clinics of North America and Food Animal Practice, 14 (1): 141–150 Medline.
Tibaijuka, B., Molla, B., Hilderbrandt, G. and Kleer, J. (2003): Occurrence of salmonellae in retail raw chicken products in Ethiopia. Berlin München Tieräztlisch Wschenschr 116: 55–58.
Van Duijkeren, E., Slot Van Oldruitenborgh Oosterbaan, M.M., Houwers, D.J., Van Leeuwen, W.J. and Kalsbeek, H.C. (1994b): A study of equine salmonellosis in a Dutch veterinary teaching hospital. Veterinary Record, 135: 248–250 Medline.
Wall, P.G., Morgan, D., and Lamden, K. (1994): A case control study of infection with an epidemic strain of multi-resistant Salmonella Typhimurium DT 104 in England and Wales. Communicable Disease Report Review, 4: 130–135.
White, D.G., Zhao, S., Sudler, R., Ayers, S., Friedman, S., Chen, S., McDermott, S., Waner, D.D., Meng, J. (2001): The isolation of antibiotic-resistant Salmonella from retail ground meats. New England Journal of Medicine, 345 (16): 1147–1154 Medline. DOI 10.1056/NEJMoa010315.
Witte, W. (1998): Medical consequences of antibiotic use in agriculture. Science, 279 (5353): 996–997 Medline. DOI 10.1126/science.279.5353.996.
World Health Organization (1988): Salmonellosis Control: The Role of Animal and Product Hygiene, Technical Report Series 774, World Health Organization, Geneva.
Acknowledgements
I am extremely grateful to Dr. Anne Muckle, head of Laboratory for Foodborne Zoonoses, Health Canada, and Dr. Cornelius Poppe and Ms. Linda Cole at the Office International des Epizooties (OIEì), Reference Laboratory for Salmonellosis, in Guelph, Ontario, Canada, for their excellent endeavors in serotyping, phage typing and antimicrobial susceptibility testing of Salmonella isolates. The financial assistance of Addis Ababa University is highly acknowledged.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zewdu, E., Cornelius, P. Antimicrobial resistance pattern of Salmonella serotypes isolated from food items and personnel in Addis Ababa, Ethiopia. Trop Anim Health Prod 41, 241–249 (2009). https://doi.org/10.1007/s11250-008-9181-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11250-008-9181-y