Abstract
The biosynthesis of aflatoxin in Aspergillus parasiticus is a complex process that involves the activities of at least 18 pathway enzymes. The distribution of these enzymes within fungal colonies and fungal cells is not clearly understood. The objective of this study was to investigate the distribution and subcellular location of Nor-1, Ver-1, and OmtA, which represent early, middle, and late enzymatic activities, respectively, in the aflatoxin biosynthetic pathway. The distribution of these three enzymes within A. parasiticus SU-1 was analyzed in time-fractionated, 72-h fungal colonies (fraction 1, 48–72 h; fraction 2, 24–48 h; fraction 3, 0–24 h). Western blot analysis and immunofluorescence microscopy demonstrated the highest abundance of Nor-1, Ver-1, and OmtA in colony fraction 2. Fungal tissues in this fraction were analyzed by immunoelectron microscopy. Nor-1 and Ver-1 were primarily localized to the cytoplasm, suggesting that they are cytosolic enzymes. OmtA was also detected in the cytoplasm. However, in cells located near the basal (substrate) surface of the colony, OmtA was predominantly detected in organelles tentatively identified as vacuoles. The role of this organelle in toxin biosynthesis is unclear. The relative distribution of OmtA to the cytoplasm or to vacuole-like organelles may depend on the age and/or physiological condition of the fungal cells.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Aflatoxins are highly toxic and carcinogenic secondary metabolites produced by several filamentous fungi, including Aspergillus parasiticus, A. flavus, A. nomius, and A. tamarii (Eaton and Groopman 1994; Payne and Brown 1998). Aflatoxins pose significant health and economic problems in the USA and many other locations throughout the world because they frequently contaminate food and feed crops, such as corn, peanuts, tree nuts and cottonseed (CAST 2003; Chang et al. 1998; Eaton and Groopman 1994).
The biosynthesis of aflatoxin is a complex process that involves at least 18 enzyme activities (Payne and Brown 1998). The distribution of these enzymes within fungal colonies and their location within fungal cells remain unknown. We hypothesized that aflatoxin enzymes are compartmentalized in a subcellular organelle(s) and the end-product, aflatoxin, is synthesized within this organelle; this could provide one mechanism to protect fungal cells from the potential deleterious effects of aflatoxin accumulation in the mycelium. To address this hypothesis, the distribution and subcellular location of representative enzymatic activities in the aflatoxin biosynthetic pathway were investigated. Toward this end, we focused attention on Nor-1 (Trail et al. 1994), Ver-1 (Liang et al. 1997) and OmtA (Yu et al. 1993), which catalyze early, middle, and late enzymatic steps in the aflatoxin biosynthetic pathway, respectively. Nor-1 and Ver-1 are NADPH-dependent keto-reductases involved in the conversion of norsolorinic acid (NA) to averantin (AVN), and versicolorin A (VERA) to demethylsterigmatocystin (DMST), respectively; OmtA is a methyltransferase that converts sterigmatocystin (ST) to O-methylsterigmatocystin (OMST).
Although a small number of reports have presented subcellular fractionation data, the main purpose of these studies was to purify and characterize the relevant enzyme activity (e.g. OmtA, OrdA, and VBS) (Cleveland et al. 1987; Watanabe and Townsend 1998; Yabe et al. 1999). However, the precise intracellular locations of these enzymes were not determined. In addition, the methods used in these experiments included grinding of mycelia under liquid nitrogen to obtain a cell extract followed by differential centrifugation. The potential drawbacks of vigorous cell disruption and time-consuming organelle isolation procedures were reported previously in studies focused on localization of penicillin enzymes in Penicillium chrysogenum. Müller et al. (1991) reported that organelles could be disrupted during grinding of mycelia and subsequent fractionation procedures, resulting in either release of their content into the cytoplasm or artifactual association of cytoplasmic proteins with the reformed organelles. Van der Lende et al. (2002) reported that, for isolation of protoplasts and organelles, the time-consuming cell-wall digestion procedure that is conducted under conditions of nutritional starvation can cause enzymes to relocate from the cytosol into the vacuole for protein degradation.
To avoid potential artifacts introduced by vigorous cell disruption and time-consuming cell-wall digestion and organelle isolation procedures, we utilized another strategy for in situ localization of aflatoxin proteins, immunoelectron microscopy. To date, no in situ localization data for aflatoxin enzymes have been presented. In this study, a specific procedure was developed to detect the distribution of aflatoxin enzymes in solid-cultured fungal colonies (by colony fractionation, Western blot analysis, and immunofluorescence microscopy) and the intracellular distribution of aflatoxin enzymes (by freeze-substitution and immunoelectron microscopy). The in situ localization of aflatoxin enzymes was also conducted on fungal cells distributed vertically within a colony fraction. The results of this study demonstrated that Nor-1, Ver-1 and OmtA are found in highest abundance in colony fraction 2 (24- to 48-h-old) and Nor-1 and Ver-1 are found primarily in the cytoplasm. OmtA was also detected in the cytoplasm. However, in cells located near the basal (substrate) surface of colony fraction 2, OmtA was predominantly detected in organelles tentatively identified as vacuoles. Based on current data, we hypothesize that the relative distribution of OmtA to cytoplasm or vacuole-like organelles depends on the age and/or physiological condition of the fungal cells.
Materials and methods
Fungal strains and time-dependent fractionation of fungal colonies
A. parasiticus SU-1 (NRRL5862, ATCC 56775) is a wild-type, aflatoxin-producing strain. AFS10 (aflR via gene disruption) is a non-aflatoxin producing strain derived from A. parasiticus NR-1 (niaD), which in turn was derived from strain SU-1 (Cary et al. 2002). Asexual conidiospores (5 x105) of A. parasiticus SU-1 and AFS10 were inoculated onto the center of YES agar medium (2% yeast extract, 6% sucrose, pH 5.8) overlaid with sterile cellophane membranes, and incubated at 29 °C in the dark. After 72 h, colonies of SU-1 (S) and AFS10 (R) were fractionated into three concentric rings based on area covered at 24, 48, or 72 h of growth (fraction 1, 48–72 h; fraction 2, 24–48 h; fraction 3, 0–24 h) to generate fractions S1, S2, S3, and R1, R2, R3, respectively, as described by Lee et al. (2002). The harvested mycelia from appropriate sections of the colony were used in sample preparation for Western blot analysis and for fluorescence and electron microscopy.
Western blot analysis of proteins isolated from colony fractions
Sample preparation and Western blot analysis of A. parasiticus proteins isolated from colony fractions were conducted using methods described in Lee et al. (2002). Each lane on the 12% SDS-polyacrylamide gel contained 60 µg protein. Nor-1, Ver-1, and OmtA polyclonal antibodies were raised against maltose-binding-protein fusions generated for each aflatoxin protein (Liang et al. 1997; Zhou 1997; Lee et al. 2002). Anti-Nor-1 serum (rabbit no. 126) was generated in this study and used at a 1 to 5,000 dilution. Anti-Ver-1 IgG was used at 2 µg per ml.
Immunofluorescence labeling and confocal laser scanning microscopy
Preparation of paraffin-embedded fungal sections, immunolabeling, and confocal laser scanning microscopy (CLSM) analysis utilized methods described by Lee et al. (2002). The samples were probed with primary antibodies against Nor-1 (1:500 dilution), Ver-1 (20 µg/ml), or OmtA (20 µg/ml), followed by secondary antibody conjugated to a fluorescent probe [goat anti-rabbit IgG-Alexa 488 conjugate (5 µg/ml); absorbance 495 nm, emission 519 nm). Fluorescent image detection was carried out on a Meridian INSIGHT plus laser-scanning microscope (Meridian Instruments, Okemos, Mich., USA) with a 488-nm laser line. Alexa 488 green fluorescence was detected using a BP 530/30 barrier filter. All images were captured using a 40× Zeiss Plan-NEOFLUAR oil objective lens (N.A.=1.3) (Carl Zeis, Germany) with a 1× cool-charged detector (CCD) allowing analysis of a large number of cells in one 512×480 image. Fluorescence image analysis of strains SU-1 and AFS10 was conducted using the same instrument parameter settings. For all strain AFS10 samples, bright-field images were also generated to demonstrate the size and number of cells analyzed.
Quantitative fluorescence intensity analysis
Alexa 488 fluorescence was quantified using the IQ Master Program (V2.31) image analysis software that accompanied the Meridian Insight laser-scanning microscope (Meridian Instruments). The CLSM images were acquired under identical instrument settings for samples analyzed in the same immunolabeling experiments. For each colony fraction, 20 images were acquired for intensity analysis and the average pixel number was reported.
Sample preparation for immunoelectron microscopy
Circles of fungal tissue 3–5 mm in diameter were obtained from fraction 2 of colonies of strains SU-1 and AFS10 grown on YES agar plates. These circles were cut from the colony using sterile 200-µl pipette tips with the end removed with a sterile razor blade to generate a cutting edge with the correct diameter. Samples were immediately cryofixed at −190 °C in a commercial Jet-Freezer (RMC MF7200, Tucson, Ariz., USA) and then transferred to a tube containing acetone that had been frozen by immersion in liquid nitrogen. The following substitution procedure was modified from the method described by Xu and Mendgen (1994). Samples in frozen acetone were stored at −80 °C for 6 days. Samples were washed twice with fresh, ice-cold acetone (−20 °C) and then immersed in fresh, ice-cold acetone (−20 °C) containing 0.2% glutaradehyde and stored at −20 °C for 24 h. After washing with fresh, ice-cold acetone (−20 °C) three times, acetone in the samples was replaced with 100% ethanol by a graded series of ethanol/acetone [35, 50, 75, 90% (30 min each at −20 °C) and 100% (twice, 30 min each at −20 °C)]. Samples were then infiltrated with a graded series of LR White resin (Ted Pella, Redding, Calif., USA) ethanol: 5, 10, 20, 40, 60% (3 h each at −20 °C), 90% for 24 h, and 100% (3 changes) for 2 days at −20 °C. Polymerization was carried out under UV light (366 nm) at 4 °C for 48 h.
Immunogold labeling and electron microscopy
Fungal sections for immunogold labeling were cut to 90~100 nm thickness using an MTX ultramicrotome (RMC). Sections were collected onto Formvar-coated grids. Each grid contained approximately 10–20 thin sections. For each labeling experiment, two grids each from strain SU-1 and strain AFS10 tissues were used. For each antibody, at least three independent labeling experiments were done. First, grids with sections were incubated with blocking solution (1% BSA and 0.1% saponin in TBS) at 4 °C overnight. Primary antibody treatment (purified OmtA IgG, 50 µg per ml; anti-Ver1 IgG, 140 µg per ml; anti-serum: anti-SKL serum, 1:200; anti-Nor-1, 1:500) was carried out at room temperature for 4 h. Secondary antibody treatment (goat anti-rabbit conjugated with 10-nm gold; 30-fold dilution) (Ted Pella) was done at room temperature for 2 h. After each antibody incubation, grids were washed once with Tris-buffered saline (pH 7.5) containing 1% BSA and 0.1% saponin for 5 min followed by five washes with TBS for 25 min total. Finally, grids were washed with double-distilled water for 30 s and post-stained with 3% uranyl acetate for 20 min. Sections were observed using a JEOL 100CX II transmission electron microscope (Tokyo, Japan) at 100 kV.
Results
Both Western blot analysis and immunofluorescence detection by CLSM confirmed the specificity of the Nor-1 and Ver-1 polyclonal antibodies used in this study (Figs. 1 and 2). Western blot analysis of Nor-1and Ver-1 isolated from colony fractions 1, 2, and 3 of A. parasiticus strains SU-1 and AFS10 demonstrated that Nor-1 and Ver-1 appeared predominantly in full-length form in all strain SU-1 colony fractions (Fig. 1). In a similar analysis conducted previously on OmtA only, increasing quantities of proteolytically cleaved enzyme were detected in fractions 1 and 2 (Lee et al. 2002). Unlike the results obtained with OmtA, the proteolytically cleaved forms of Nor-1 and Ver-1 enzymes were not observed under the same culture conditions. The highest quantity of Nor-1 and Ver-1 was in fraction 2 of strain SU-1 (Fig. 1). Immunofluorescence detection byCLSM demonstrated that Nor-1 and Ver-1 were present in all colony fractions of strain SU-1 (Fig. 2a for Nor-1 and Fig. 2d for Ver-1). Very little signal was detected in strain AFS10 samples under the same instrument settings (Fig. 2b for Nor-1 and Fig. 2e for Ver-1). Therefore, bright-field images of the strain AFS10 colony fractions are shown in Fig. 2c, f to illustrate the typical size and numbers of cells analyzed. Quantitative analysis of the fluorescence intensity in the CLSM digital images confirmed that the highest quantity of these three aflatoxin enzymes occurred in fraction 2 (Table 1). Therefore, immuno-transmission electron microscopy (TEM) analysis focused on the proteins in this colony fraction.
Western blot analysis of fungal protein extracts using Ver-1 and Nor-1 polyclonal antibodies. Proteins were extracted from time-fractionated colonies of Aspergillus parasiticus srtains SU-1 and AFS10 grown on YES agar medium and subjected to Western blot analysis with Ver-1 (A) and Nor-1 (B) polyclonal antibodies. At 72 h, colonies were fractionated into three concentric rings based on area covered at three time points: for S1 (strain SU-1) 1 and R1 (strain AFS10), 48–72 h; S2 and R2, 24–48 h; S3 and R3, 0–24 h. Lanes to the left and right contain molecular mass standards (M), which are marked at the right of A and B. The molecular masses of Ver-1 and Nor-1 (indicated by arrows at the left) are 28 and 31 kDa, respectively
Immunofluorescence confocal microscopy of Nor-1 and Ver-1 in A. parasiticus strains SU-1 and AFS10 (aflR knockout mutant) grown on YES agar for 72 h. Colonies of strains SU-1 and AFS10 were divided into three fractions (see Material and methods). The paraffin-embedded fungal sections were immunolabeled with anti-Nor-1 antiserum (A) (1:500) or anti-Ver-1 IgG (B) (20 μg/ml) followed by Alexa-488-conjugated goat anti-rabbit IgG. Fluorescence intensities of strain SU-1 sections immunolabeled with anti-Nor-1 and anti-Ver-1 are shown in rows a and d, respectively; and of strain AFS10 sections in rows b and e. The related bright-field images of the strain AFS10 colony fractions are shown in rows c and f . Bar 100 μm
Using our sample preparation, labeling, and imaging procedures, very few gold particles were observed on control sections of wild-type strain SU-1 treated with secondary antibodies only (Fig. 3A, B). Therefore, the intense signal (black dots representing 10-nm gold particles) on sections was interpreted as resulting from the interaction between primary antibodies and their target proteins immobilized within the cell. Use of the negative control strain AFS10 (no aflatoxin proteins produced) allowed non-specific labeling and cross-reactive binding to be recognized, and thus specific structures could be ruled out as the target organelles for aflatoxin proteins. For example, labeling of the cell wall, especially for anti-Ver-1, was occasionally detected in certain sections of both strain SU-1 and strain AFS10, leading to the conclusion that this was non-specific labeling. It was reported previously that non-specific labeling of the cell wall may result either from non-specific binding of primary antibodies or from cell-wall-reactive antibodies that arise in rabbits exposed to or infected by yeast or molds before or during antibody production (Atkin 1999; Binder et al. 1996). Scattered gold particles were found occasionally associated with the nucleus, mitochondria, and cytoplasm in strain AFS10 (for example, Fig. 3C), and with the nucleus and mitochondria in strain SU-1. This background “noise” could be largely eliminated by a further step of antibody purification using affinity subtraction as described in Lee et al. (2002). As shown in this study, very little labeling was noted in the cell wall, mitochondria, and nucleus using affinity-purified antibodies to OmtA (Fig. 5 and Table 1).
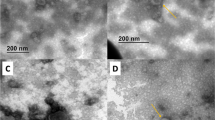
Immunogold labeling of Nor-1 in A. parasiticus strains SU-1 and AFS10. A, B Ultra-thin sections of fungal tissues of the aflatoxin producing strain SU-1 were prepared for transmission electron microscopy and labeled with secondary antibodies only. Ultra-thin sections of fungal tissues of strains AFS10 (C) and SU-1 (D) were labeled with primary antibodies against Nor-1, followed by 10-nm gold beads conjugated to goat anti-rabbit IgG secondary antibodies (1:30 dilution) as described in Materials and methods. cw Cell wall, mb microbodies, M mitochondria, N nuclei, V vacuole. Bars 500 nm (A), 250 nm (B–D)
When highly specific antibodies to Nor-1, Ver-1, and OmtA were used to label sections from fraction 2 of strain SU-1, high signal intensity was observed in the cytoplasm (Figs. 3D, 4E, and 5D). The gold particles frequently aggregated to form clusters (Figs. 3D and 5D); similar clusters were not seen in the cytoplasm of control sections obtained from strain AFS10 or when only gold-labeled secondary antibodies were used. The immunogold labeling results are in good agreement with immunofluorescence data in this and our previous study and may help explain the “patchy” appearance of the fluorescent signal observed (Fig. 2 and Lee et al. 2002).
Immunogold labeling using anti-SKL and anti-Ver-1 in A. parasiticus strains AFS10 and SU-1. Ultra-thin sections of fungal tissues obtained from fraction 2 were labeled with primary antibodies to PST-1 (anti-SKL) (A–C) or primary antibodies to Ver-1 (D–G) followed by gold-labeled secondary antibodies as described in Materials and methods. A, B, D, F, G Ultra-thin sections prepared from fungal tissues of strain AFS10; C, E ultra-thin sections prepared from fungal tissues of strain SU-1. cw Cell wall, mb microbodies, wb woronin body, st septum, M mitochondria, V vacuole. Bars 250 nm
Immunogold labeling of OmtA in A. parasiticus strains AFS10 and SU-1. Ultra-thin sections of fungal tissues of strains AFS10 (A, B) and SU-1 (C, D) were labeled with primary antibodies against OmtA followed by gold-labeled secondary antibodies as described in Materials and methods. E The colony fractionation procedure and the cell distribution in a typical thin section. The specific location of OmtA in cells near the basal surface of the colony (gray) is also shown. cw Cell wall, M mitochondria, N nuclei, V vacuole. Bars 500 nm (A, C, E), 250 nm (B, D)
The gross ultrastructure of cells observed in thin sections suggested that our fixation procedures successfully preserved the integrity of organelles bounded by double (nuclei and mitochondria) and single (vacuoles, microbodies, Woronin bodies) membranes. Although the data appeared quite convincing, it was possible that disruption of organelles occurred during sample preparation and was in part or fully responsible for the observed cytoplasmic location of Nor-1, Ver-1, or OmtA. To minimize this possibility, we demonstrated that the ultrastructure of Woronin bodies (derived from microbodies; Valenciano et al. 1998) and microbodies, and the antigenicity of their content proteins were preserved using our sample preparation methods. Woronin bodies and microbodies were intensely labeled by anti-SKL (peroxisomal targeting signal 1) (Fig. 4A–C) but not by antibodies to Nor-1, Ver-1, and OmtA in strains AFS10 and SU-1 (Fig. 4D, E), which provided a good positive control for sample preparation. Some labeling of Woronin bodies did occur with anti-Ver-1 (Fig. 4F, G). However, the observed labeling occurred in both strain AFS10 (control) and strain SU-1, allowing us to interpret this as non-specific labeling possibly due to high antibody concentration or the binding of antibodies to cross-reactive proteins located in this organelle. This interpretation was likely because disruption of aflR in A. parasiticus (such as in strain AFS10) results in loss of nor-1, ver-1, and omtA gene transcripts (Cary et al. 2000, 2002) and proteins (this study). Therefore, it is unlikely that Ver-1 exists at any location in strain AFS10. The ultrastructure of double-membrane-bound organelles (nuclei and mitochondria) as well as other single-membrane-bound organelles (vacuoles) were also maintained in this study. With the exception of vacuole-like organelles in the basal region of fraction 2, these organelles did not label with antibodies against the aflatoxin enzymes (Figs. 3, 4 and 5). These results suggest that Nor-1, Ver-1, and OmtA do not localize to nuclei, mitochondria, Woronin bodies, or microbodies.
Of particular interest, most cells located near the basal surface of fraction 2 were closely packed and contained one to several large organelles heavily labeled with OmtA antibodies (Figs. 5E and 6). Because the absence of signal in organelles (nuclei, mitochondria, Woronin bodies, or microbodies) does not appear to be an artifact arising from organelle breakage (Figs. 3,4 and 5), these data strongly support the specificity of labeling of the vacuole-like organelles observed in the basal region of fraction 2 (Figs. 5E and 6). In some cells, the large and small organelles appeared to fuse together (Fig. 6). The highly dynamic structure of these organelles is consistent with a model for vacuolar development in Aspergillus (Ohsumi et al. 2002) and the vacuolation phenomena observed by Paul et al. (1994). The morphology and apparent developmental origin of the labeled organelles in fraction 2 prompted us to tentatively identify them as vacuoles (Amor et al. 2000). The labeling intensity suggested a high concentration of OmtA in these organelles. In contrast, antibodies against Nor-1 or Ver-1 did not label these organelles in sections from the same location (basal portion of fraction 2) (Figs. 7 and 8). Cells located above the basal region contained smaller organelles that were sporadically labeled by OmtA and not at all by antibodies to Nor-1 and Ver-1. Similar organelles in AFS10 did not label with polyclonal antibodies to any aflatoxin enzyme. We attempted to label these organelles using antibodies against yeast vacuolar aminopeptidase I (kindly provided by Dr. Klionsky (Division of Biology, California Institute of Technology, Pasadena Calif., USA). However, no specific binding was observed (data not shown). It is possible that antibody against yeast aminopeptidase I is not suitable for A. parasiticus because the proteins structures are not sufficiently conserved.
Immunogold labeling of OmtA in A. parasiticus strain SU-1. Sections cut from the basal surface of the colony were labeled with primary antibodies against OmtA. A The gold-labeled vacuole-like organelles inside the cells were numbered (1–8). B–E Higher magnification images of these organelles. cw Cell wall, M mitochondria, N nuclei, V vacuole-like organelle. Bars 2 μm (A), 1 μm (B–E)
Discussion
Our goal was to analyze the distribution and subcellular localization of the aflatoxin enzymes Nor-1, Ver-1, and OmtA in A. parasiticus grown on YES agar medium. Immunofluorescence microscopy and Western blot analysis of protein extracts from time-fractionated colonies demonstrated that fraction 2 (cells 24- to 48-h-old) contained the highest concentration of each target protein. In addition, proteolytic cleavage of OmtA occurred to a greater extent in “older” fungal tissues (fractions 1 and 2) (Lee et al. 2002) but this was not observed for Nor-1 and Ver-1 under these culture conditions. It was reported previously that OmtA purified from A. parasiticus is a 40-kDa protein whereas the cDNA encodes a 45-kDa protein (Keller et al. 1993; Liu et al. 1993; Yu et al. 1993). We speculate that loss of the 5 kDa of N-terminal sequence may be associated with localization to the organelle and possibly the function of the enzyme. The specificity and timing of expression of the proteolytic enzymes and the potential role of cleavage in activation/inactivation of aflatoxin enzymes is clearly of interest for follow-up studies.
To our knowledge, this is the first study to conduct in situ localization of aflatoxin enzymes and to localize an aflatoxin enzyme to a specific cellular organelle. We demonstrated that Nor-1 and Ver-1 were primarily localized to the cytoplasm, suggesting that they are cytosolic enzymes. OmtA was also detected in the cytoplasm. However, in cells located near the basal (substrate) surface of the colony, OmtA was predominantly detected in organelles tentatively identified as vacuoles. Vacuoles are reported to associate with several biological functions in fungi, including degradation or recycling of proteins and whole organelles (Amor et al. 2000), storage of metabolites, ions and amino acids, enzyme maturation (e.g. aminopeptidase I) and pH homeostasis (Klionsky et al. 1990; Thumm 2000). At specific stages of appressoria formation in Magnaporthe grisea, vacuoles serve as a compartment for lipid degradation and glycerol production (Weber et al. 2001). In yeast, the biological sulfonium compound S-adenosyl-methionine (AdoMet), accumulates to high levels in vacuoles (Schwencke et al. 1976; Svihla et al. 1969). S-adenosyl-methionine is an important cofactor and provides the methyl group in the reaction catalyzed by OmtA. Based on the available data, it is difficult to determine whether the localization of OmtA to vacuole-like organelles in A. parasiticus occurs for protein recycling, enzyme activation, or some other purpose .
Two observations from this study are particularly noteworthy. First, intensely labeled, weakly labeled, and unlabeled cells could be found adjacent to each other in the same thin section analyzed by TEM. These data suggest that the extent and timing of aflatoxin gene expression varies from cell to cell, even in an area of the colony that is presumably the same age. This variation may be related to the local concentration of available nutrients or the relative age of fungal cells (Calvo et al. 2002; Chiou et al. 2002; Luchese et al. 1993). Second, OmtA appeared in the cytoplasm in certain cells and in vacuole-like organelles in other cells in the same colony fraction. The large and small vacuole-like organelles were fused together, similar to the vacuolation process, suggesting that these cells were under nutrient limitation (Paul et al. 1994). We propose that a specific set of changes in cell age, physiological conditions, or nutrient availability triggers the transport of OmtA from cytoplasm into the vacuole-like organelle via a pathway similar to the yeast cytoplasm-to-vacuole targeting (cvt) pathway. In Saccharomyces, the cvt pathway is regulated by the availability or limitation of specific nutrients (e.g. glucose or ethanol) (Klionsky et al. 1990; Klionsky 1997; Thumm 2000). To date, no pathway similar to cvt has been reported in filamentous fungi.
Based on data from this and previous studies, it is reasonable to assign two potential alternative roles for the vacuole-like organelle in aflatoxin synthesis: (1) as a means to reduce or limit aflatoxin synthesis, OmtA is transported to and inactivated in the vacuole-like organelle via proteolytic cleavage; (2) as a means to shield the cell from the potential toxic effects of aflatoxin in the mycelium (Saxena et al. 1991), OmtA is transported to the vacuole-like organelle together with the late aflatoxin pathway intermediate ST. Here, OmtA is activated and ST is converted to OMST and further converted to AFB1 by OrdA (which catalyzes the final step of aflatoxin biosynthesis) (Prieto and Woloshuk 1997; Yu et al. 1998) localized in or on the same vacuole. These hypotheses form the basis for our future studies on aflatoxin synthesis.
References
Amor C, Dominguez AI, Lucas JRD, Laborda F (2000) The catabolite inactivation of Aspergillus nidulans isocitrate lyase occurs by specific autophagy of peroxisomes. Arch Microbiol. 174:59–66
Atkin AL (1999) Preparation of yeast cells for confocal microscopy. Methods Mol Biol 122:131–139
Binder M, Hartig A, Sata T (1996) Immunogold labeling of yeast cells: an efficient tool for the study of protein targeting and morphological alterations due to overexpression and inactivation of genes. Histochem Cell Biol 106:115–130.
Calvo AM, Wilson RA, Bok JW, Keller NP (2002) Relationship between secondary metabolism and fungal development. Microbiol Mol Biol Rev 66:447–459
Cary JW, Ehrlich KC, Wright MS, Chang P-K, Bhatnagar D (2000) Generation of aflR disruption mutants of Aspergillus parasiticus. Appl Microbiol Biotechnol. 53:680–684
Cary JW, John MD, Ehrlich KC, Wright MS, Liang S-H, Linz JE (2002) Molecular and functional characterization of a second copy of the aflatoxin regulatory gene, AflR, from Aspergillus parasiticus. Biochim Biophys Acta 1576:316–323
CAST (2003) Council for Agricultural Science and Technology. Mycotoxins: risks in plant, animal, and human systems. Report 139. Council for Agricultural Science and Technology, Ames, Iowa
Chang P-K, Yu J, Bhatnagar D, Cleveland TE (1998) Abstract. The USDA-ARS Aflatoxin Elimination Workshop, October 25–27, 1998, St. Louis, Missouri
Chiou C-H, Miller M, Wilson DL, Trail F, Linz J (2002) Chromosomal location plays a role in regulation of aflatoxin gene expression in Aspergillus parasiticus. Appl Environ Microbiol 68:306–315
Cleveland TE, Lax AR, Lee LS, Bhatnagar D (1987) Appearance of enzyme activities catalyzing conversion of sterigmatocystin to aflatoxin B1 in late-growth-phase Aspergillus parasiticus cultures. Appl Environ Microbiol 53:1711–3
Eaton DL, Groopman JD (eds.) (1994) The toxicology of aflatoxins: human health, veterinary, and agricultural significance. Academic, San Diego, California
Keller NP, Dischinger HC, JR, Bhatnagar D, Cleveland TE, Ullah AHJ (1993) Purification of a 40-kDa methyltransferase activity in the aflatoxin biosynthetic pathway. Appl Environ Microbiol 36:270–273
Klionsky DJ (1997) Protein transport from the cytoplasm into vacuole. J Membr Biol 157:105–115
Klionsky DJ, Herman PK, EMR SD (1990) The fungal vacuole: composition, function, and biogenesis. Microbiol Rev 54:266–292
Lee L-W, Chiou C-H, Linz JE (2002) Function of native OmtA in vivo and expression and distribution of this protein in colonies of Aspergillus parasiticus. Appl Environ Microbiol 68:5718–5727
Liang S-H (1996) The function and expression of the ver-1 gene and localization of the Ver-1 protein involved in aflatoxin B1 biosynthesis in Aspergillus parasiticus. PhD Dissertation, Michigan State University, East Lansing
Liang S-H, Wu T-S, Lee R, Chu FS, Linz JE (1997) Analysis of mechanism regulating expression of the ver-1 gene, involved in aflatoxin biosynthesis. Appl Environ Microbiol 63:1058–1065
Liu BH, Keller NP, Bhatnagar D, Cleveland TE, Chu FS (1993) Production and characterization of antibodies against sterigmatocystin O-methyltransferase. Food Agric Immunol 5:155–164
Luchese RH, Harrigan WF (1993) Biosynthesis of aflatoxin- the role of nutritional factors. J Applied Bacteriol 74:5–14
Müller WH, van der Krift TP, Knoll G, Smaal EB, Verkleij AJ (1991) A preparation method of specimens of the fungus Penicillium chrysogenum for ultrastuctural and immuno-electron microscopical studies. J Microsc 164:29–41
Ohsumi K, Arioka M, Nakajima H, Kitamoto K (2002) Cloning and characterization of a gene (avaA) from Aspergillus nidulans encoding a small GTPase involved in vacuole biogenesis. Gene 291:77–84
Paul GC, Kent CA, Thomas CR (1994) Hyphal vacuolation and fragmentation in Penicillium chrysogenum. Biotechnol Bioeng 44:655–660
Payne GA, Brown MP (1998) Genetics and physiology of aflatoxin biosynthesis. Annu Rev Phytopathol 36:329–362
Prieto R, Woloshuk CP (1997) ord1, an oxidoreductase gene responsible for the conversion of O-methylsterigmatocystin to aflatoxin in Aspergillus flavus. Appl Environ Microbiol 63:1661–1666
Saxena M, Allameh A, Mukerji KG, Raj HG (1991) Epoxidation of aflatoxin B1 by Aspergillus flavus microsomes in vitro: interaction with DNA and formation of aflatoxin B1-glutathione conjugate. Chem Biol Interactions 78:13–22
Schwencke J, De Robichon-Szulmajster H (1976) The transport of S-adenosyl-L-methionine in the isolated yeast vacuoles and spheroplasts. Eur J Biochem 65:49–60
Svihla G, Dainko JL, Schlenk F (1969) Ultraviolet micrography of penetration of exogenous cytochrome c into the yeast cell. J Bacteriol 100:498–504
Taylor KM, Kaplan CP, Gao X, Baker A (1996) Localization and targeting of isocitrate lyases in Saccharomyces cerevisiae. Biochem J 319:255–262
Thumm M (2000) Structure and function of the yeast vacuole and its role in autophagy. Microsc Res Tech 51:563–572
Trail F, Chang P-K, Cary J, Linz JE (1994) Structural and functional analysis of the nor-1 gene involved in the biosynthesis of aflatoxin by Aspergillus parasiticus. Appl Environ Microbiol 60:3315–3320
Valenciano S, De Lucas JR, Van der Klei I, Veenhuis M, Laborda F (1998) Characterization of Aspergillus nidulans peroxisomes by immunoelectron microscopy. Arch Microbiol 170:370–376
van der Lende TR, van de Kamp M, Berg M, Sjollema K, Bovenberg RA, Veenhuis M, Konings WN, Driessen AJ (2002) δ-(l-α-aminoadipyl)-l-cysteinyl-d-valine sythetase, that mediates the first committed step in penicillin biosynthesis, is a cytosolic enzyme. Fungal Genet Biol 37:49–55
Watanabe CMH, Townsend CA (1998) The in vitro conversion of norsolorinic acid to aflatoxin B1. An improved method of cell-free enzyme preparation and stabilization. J Am Chem Soc 120:6231–6239
Weber RWS, Wakley GE, Thines E, Talbot NJ (2001) The vacuole as element of the lytic system and sink for lipid droplets in maturing appressoria of Magnaporthe grisea. Protoplasma 216:101–112
Xu H, Mendgen K (1994) Endocytosis of 1,3-ß-glucans by broad bean cells at the penetration site of the cowpea rust fungus (haploid stage). Planta 195:282–290
Yabe K, Nakamura M, Hamasaki T (1999) Enzymatic formation of G-group aflatoxins and biosynthetic relationship between G- and B-group aflatoxins. Appl Environ Microbiol 65:3867–3872
Yu, J, Cary JW, Bhatnagar D, Cleveland TE, Keller NP, Chu FS (1993) Cloning and characterization of a cDNA from Aspergillus parasiticus encoding an O-methyltransferase involved in aflatoxin biosynthesis. Appl Environ Microbiol 59:3564–71
Yu J, Chang P-K, Ehrlich KC, Cary JW, Bhatnagar D, Cleveland TE (1998) Characterization of the critical amino acids of Aspergillus parasiticus cytochrome P-450 monooxygenease encoded by ordA that is involved in the biosynthesis of aflatoxin B1, G1, B2 and G2. Appl Environ Microbiol 64:4834–4841
Zhou R (1997) The function, accumulation, and localization of the Nor-1 protein involved in aflatoxin biosynthesis; the function of the fluP gene associated with sporulation in Aspergillus parasiticus. PhD Dissertation, Michigan State University, East Lansing
Acknowledgements
We thank Dr. Shirley Owens for help in quantification of fluorescence intensity and Dr. Xudong Fan in operating the jet-freezer at the Center of Advanced Microscopy at Michigan State University. This work was supported by the Michigan Agricultural Experiment Station (MSU), the National Food Safety and Toxicology Center (MSU), the Center for Environmental Toxicology (MSU), and the National Institutes of Health (RO1 CA52003-12).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Lee, LW., Chiou, CH., Klomparens, K.L. et al. Subcellular localization of aflatoxin biosynthetic enzymes Nor-1, Ver-1, and OmtA in time-dependent fractionated colonies of Aspergillus parasiticus . Arch Microbiol 181, 204–214 (2004). https://doi.org/10.1007/s00203-003-0643-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-003-0643-3