Abstract
Glaucoma is a retinal disease caused due to increased intraocular pressure in the eyes. It is the second most dominant cause of irreversible blindness after cataract, and if this remains undiagnosed, it may become the first common cause. Ophthalmologists use different comprehensive retinal examinations such as ophthalmoscopy, tonometry, perimetry, gonioscopy and pachymetry to diagnose glaucoma. But all these approaches are manual and time-consuming. Thus, a computer-aided diagnosis system may aid as an assistive measure for the initial screening of glaucoma for diagnosis purposes, thereby reducing the computational complexity. This paper presents a deep learning-based disc cup segmentation glaucoma network (DC-Gnet) for the extraction of structural features namely cup-to-disc ratio, disc damage likelihood scale and inferior superior nasal temporal regions for diagnosis of glaucoma. The proposed approach of segmentation has been tested on RIM-One and Drishti-GS dataset. Further, based on experimental analysis, the DC-Gnet is found to outperform U-net, Gnet and Deep-lab architectures.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Glaucoma well-known as “silent theft of sight” was probably recognized as a disease in the initial years of the seventeenth century, when the Greek term glaukEoma meaning obscurity of the lens or cataract signifying lack of understanding of the disease came into existence. It begins due to an increase in the pressure of an eye, known as intraocular pressure leading to damage to the optic nerve. The aforementioned optic nerve carries visual information from the retina to the brain resulting in visualization of the outside world [1]. According to the World Health Organization (WHO), glaucoma is one of the leading causes of blindness, which has affected more than 60 million people to date across the globe and may rise to 80 million by 2020. Also, it is estimated to influence 12 million people in India around one-fifth of the total Indian population. Its common symptoms include sudden visual disturbance, severe eye pain, blurred vision, red eyes and halos around lights. This disease is normally found to affect people who are elder than 40 years [2]. Ophthalmologists use different comprehensive examinations such as ophthalmoscopy, tonometry, perimetry, gonioscopy and pachymetry to diagnose glaucoma. Tonometry is the measure of internal eye pressure, whereas ophthalmoscopy is the examination of the colour and shape of the optic nerve. Perimetry is the visual field analysis; pachymetry is the measure of the corneal thickness, and gonioscopy is the measure of the angle between the iris and cornea [3]. All these approaches are manual, consume time and may lead to biased decisions by different experts.
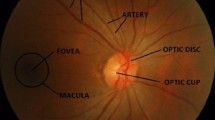
Thus, there is a need for a computer-aided diagnosis system (CAD) which can act as a second opinion for doctors. CADs for glaucoma use a retinal fundus imaging as an input to extract different types of features and classify the retinal image as “abnormal” or “normal” [4]. The retinal fundus image is acquired using a fundus camera which consists of a microscope attached with a flash-enabled camera to capture the image of the interior surface/backside of an eye. The fundus image comprises of the optic cup, optic disc, rim and inferior, superior nasal, temporal rim regions shown in Fig. 1. Here, the optic disc is the centric bright portion of retina where various optic nerves come together, and the optic cup is the depression of changeable size present on the optic disc. Glaucoma can be diagnosed by analysing the progression of the optic cup or thinning of rim [5].
The structural features used for diagnosis of glaucoma by CAD systems include cup-to-disc ratio (CDR), disc damage likelihood scale (DDLS) and inferior superior nasal temporal (ISNT) regions defined as:
- (i)
Cup-to-disc ratio (CDR) CDR is defined as the ratio of horizontal length, vertical length and area measurements of the optic cup to the optic disc, computed using Eqs. (1)–(3) for diagnosis of glaucoma.
$$ H_{\text{CDR}} = \frac{{H_{{{\text{optic}}\_{\text{cup}}}} }}{{H_{{{\text{optic}}\_{\text{disc}}}} }} $$(1)$$ V_{\text{CDR}} = \frac{{V_{{{\text{optic}}\_{\text{cup}}}} }}{{V_{{{\text{optic}}\_{\text{disc}}}} }} $$(2)$$ A_{\text{CDR}} = \frac{{A_{{{\text{optic}}\_{\text{cup}}}} }}{{A_{{{\text{optic}}\_{\text{disc}}}} }}. $$(3)Here, HCDR is the horizontal cup-to-disc ratio, VCDR is the vertical cup-to-disc ratio, and ACDR is the ratio of the area of the cup to the area of the disc. Further, Hoptic_cup, Voptic_cup and Aoptic_cup signify horizontal length, vertical length and area measurements of the optic cup, respectively. While Hoptic_disc, Voptic_disc and Aoptic_disc signify horizontal length, vertical length and area measurements of the optic disc, respectively. The average CDR value for a normal eye is considered to be less than 0.3 and that for a glaucomatous eye is considered to be more than 0.5, while the ones in the range 0.3–0.5 lies in suspect cases [6].
- (ii)
Disc damage likelihood scale (DDLS) DDLS is the ratio of the minimum rim width to the diameter of the disc. It is computed using Eq. (4).
$$ {\text{DDLS}} = \frac{{{\text{MinRim}}_{\text{width}} }}{{{\text{Disc}}_{\text{dia}} }}. $$(4)Here, MinRimwidth is the minimum width of the rim and Discdia is the diameter of the optic disc. DDLS value for a normal eye is more than 0.3, and that for a glaucomatous eye is less than 0.3 [7].
- (iii)
Inferior superior nasal temporal (ISNT) rule ISNT rule specifies the order of rim width in inferior, superior, nasal and temporal regions. The normal eye follows the order given in Eq. (5), while the abnormal eye does not follow the same.
$$ I < S < N < T. $$(5)Here, I, S, N and T represent the width of rim in inferior, superior, nasal and temporal regions [8].
Although many approaches have been used to date for glaucoma diagnosis using CDR, very few [9, 10] have utilized all the three structural features at the same time. Also, improving the accuracy of segmentation is of great significance for large scale clinical use, as better the performance of segmentation, the better is the diagnosis. Thus, this work presented an automated approach for diagnosis of glaucoma using CDR, DDLS and ISNT as a diagnostic feature. Here, the segmentation of optic disc and optic cup have been performed using DC-Gnet, and the performance is found to be better than the U-net, Gnet and Deep-lab. The proposed methodology comprises of:
- 1.
Pre-processing of input fundus images to remove outliers.
- 2.
Segmentation of optic disc and optic cup using DC-Gnet.
- 3.
Calculation of CDR, DDLS and ISNT.
Deep learning has played a major role in the detection and diagnosis of diseases such as cancer, glaucoma, etc. The approach that has been used here uses convolutional neural networks (CNNs) to constructively segment the optic disc and cup using downsampling and upsampling. CNNs are a class of deep neural networks that are primarily used for image processing. When an input is fed to a CNN, it is convolved with a feature map, i.e. convolution filter and the result is passed on to the next layer. The convolution operation is performed between the input and the feature maps thereby forming a network between each layer. The input to a CNN is an image of size (number of images) × (image width) × (image height) × (image depth), and after convolving with a feature map, i.e. a filter the output is an image of size (number of images) × (image width) × (image height) × (number of channels) [11].
Basic steps followed by CNN are as follows:
- (i)
Padding Padding prevents the size of the image from decreasing and thereby retaining the actual size of the image. It prevents the image from disappearing and allows the input size and output size to be the same, using Eq. (6)
$$ {\text{pad }} = (f - 1) \div 2. $$(6)Here, ‘f’ represents the size of the filter. It is used to perform the convolution of filter ‘f’ having size ‘f × f’ with the image portion of size ‘f × f’.
- (ii)
CNN output The output of a convolutional neural network is determined using Eq. (7)
$$ {\text{Output}} = ({\text{Input}} + 2 \times {\text{pad}} - f)/s + 1 $$(7)where ‘Input’ signifies input dimension, ‘pad’ is the padding size, ‘f’ is the filter dimension and ‘s’ is the stride size.
- (iii)
CNN layer output Finally, the output of a convolution layer is computed using Eqs. (8) and (9)
$$ z^{n } = W \cdot \left( {x^{n - 1} } \right) + b^{n} $$(8)$$ x^{n} = a^{n} ( z^{n} ) $$(9)where ‘W’ represents weights of the convolution layer, ‘\( x^{n - 1} \)’ is the intermediate value obtained from the previous layer, ‘b’ is the bias, ‘\( x^{n} \)’ signifies output of the convolution layer and ‘\( a^{n} \)’ is the activation function.
The approach presented in this paper involves modification in the architecture of U-net to improve the accuracy of the optic disc and optic cup segmentation. U-net is a CNN that includes upsampling layers instead of maxpooling layers to increase the resolution of the output. There is a large number of feature channels in the upsampling layers, which allows the propagation of context information to the higher resolution layer through the network, this gives the architecture a U-shaped structure. U-net is made up of two parts—a contracting and an expanding path. The contracting part consists of the typical convolutional network, i.e. convolution layers, rectified linear unit (ReLU) and maxpooling layers. The contracting part results in a decrease in the spatial dimensions and an increase in feature information. On the other hand, the expanding part consists of upsampling layers and concatenation layers. The expanding part involves the combining of the spatial and feature information and its concatenation with the high-resolution features from the contracting part [12].
The rest of the paper is further organized into Sect. 2 comprising of related work, Sect. 4 experimental setup and datasets used and Sect. 5 for the methodology. Further, Sect. 6 includes performance metrics, Sect. 7 includes training and testing parameters, experimental results and discussions followed by conclusion in Sect. 8.
2 Related work
Researchers across the globe have performed different approaches for segmentation of optic disc and optic cup using retinal fundus images. These approaches are categorized into two broad categories, i.e. the computer vision-based approach and deep learning-based approach. Computer vision-based approach is further categorized into thresholding, contouring and clustering-based approaches. Some of the commonly used approaches are as follows:
2.1 Computer vision-based approaches
2.1.1 Thresholding-based segmentation
Walter and Klein [13] applied thresholding followed by morphology to segment optic disc from input retinal fundus image, and the gradient-based traditional watershed was used to extract contours from the optic disc. But the approach was not able to perform optimally for images with low contrast.
Further, Pallawala et al. [14] used wavelet transformation and ellipse fitting to delineate optic disc from retinal fundus image. Daubechies wavelet was employed to get wavelet features and thresholding, followed by image subtraction to get the optic disc image.
Further, Abdel-Ghafar and Morris [15] gave thresholding-based approach for segmentation of optic disc from retinal fundus image. Sobel operator was initially applied to enhance the input image and thresholding followed by circular Hough transform was applied to achieve approximate boundaries of the optic disc with mean and variance as thresholds.
Liu et al. [16] segmented optic disc and optic cup to diagnose glaucoma using the CDR. Optic disc was segmented using the variational level set, and ellipse fitting was applied to smoothen the contour. Whereas, optic cup was segmented using thresholding followed by variational level set and ellipse fitting.
Zhang et al. [17] also worked on extraction of disc and cup using intensity-based information to analyse histogram and calculate threshold values for thresholding. The variational level set followed by ellipse fitting and the convex hull was applied for extraction disc and cup to calculate CDR.
Later, Sarkar and Das [18] suggested thresholding-based approach for segmentation of optic disc and optic cup. Median filter was initially applied to smoothen the image, and adaptive thresholding was applied to extract optic disc and optic cup. CDR was then calculated to diagnose glaucoma at an earlier stage.
Priyadharsini et al. [19] applied histogram equalization followed by connected components to extract segment optic disc. Circular Hough transform was thereafter applied to refine the segmentation and morphology followed by Otsu’s thresholding was used for segmentation of optic cup.
2.1.2 Contouring-based segmentation
Therefore, Osareh et al. [20] utilized colour spaces for localization of optic disc. Retinal vessels present in the input image were removed by applying morphology in the ‘L’ channel of ‘HLS’, ‘Lab’, and ‘Lch’ colour space. Gradient-based active contour was then employed for the segmentation of optic disc in case of low contrast images as well.
Later, Chrastek et al. [21] presented an automatic approach for segmentation of optic nerve head for glaucoma diagnosis. Input retinal image was initially pre-processed using normalization and median filtering to estimate non-uniform illumination. Active contour modelling was applied to segment optic disc and circular Hough transform to refine boundaries.
Similarly, Blanco et al. [22] with few modifications presented another approach for segmentation of optic disc using gradient vector field (GVF) followed by fuzzy circular Hough transform, where the initial region of interest for GVF was extracted using canny edge detector.
Datt et al. [23] used red channel of input retinal fundus image to segment optic disc and green channel to segment optic cup. Retinal vessels present in input images were suppressed using morphological operations in respective channels. Canny edge detector followed by circular Hough transform was then employed for the extraction of initial contour, and active contour was applied for segmentation of optic disc. Medial axis was thereafter used for segmentation of optic cup by interpolation to avoid differences in intensity within cup region. Finally, the structural feature CDR was extracted to diagnose glaucoma using retinal fundus image.
Yin et al. [24] suggested median filtering of green channel input RGB fundus image for removal of vessels. Active shape modelling followed circular Hough transform and ellipse fitting was utilized to segment optic disc and optic cup for glaucoma diagnosis. The proposed approach was found to outperform level set by improving the accuracy of segmentation.
Xu et al. [25] gave reconstruction-based learning approach for localization of optic cup by codebook of reference images. Random sampling was used for the generation of a codebook, and cup descriptors identified were reconstructed with Hadamard product cost function. Lagrange multiplier with Gaussian distance and Kronecker product was used for minimization of the objective function, and the CDR was calculated for diagnosis of glaucoma.
Later, Cheng et al. [26] suggested dissimilarity-based approach for glaucoma screening. Three approaches, namely circular Hough transform, proceeded by ASM, superpixel classification and ellipse Hough transform to segment optic disc and optic cup for glaucoma diagnosis using CDR.
2.1.3 Clustering-based segmentation
Kavitha et al. [27] utilized morphologically closed green channel of retinal fundus image to segment optic cup by component analysis method. The value of CDR calculated thereafter using segmented regions was used for prediction of ‘normal’ and ‘abnormal’ glaucoma cases.
Khalid et al. [9] presented a clustering-based approach for segmentation of optic disc and optic cup from retinal fundus images. Morphological dilation and erosion were initially applied on the red channel and green channel of input images to remove the vessels for proper segmentation. Fuzzy C means clustering was then applied on pre-processed fundus image to segment optic disc and optic cup for glaucoma diagnosis using CDR.
Further, Mittapali and Kande [28] gave an optic cup and optic disc segmentation approach for diagnosis of glaucoma. Pre-processing was initially performed by replacing vessel pixels with non-vessel pixels followed by use of a median filter to smoothen the input image. Segmentation was then performed by spatial weighted fuzzy C means (SWFCM)-based thresholding using the distribution of pixels in a grayscale image. Finally, the structural features, namely CDR, ISNT and DDLS, were calculated for diagnosis of glaucoma.
Also, as per the survey conducted by Thakur and Juneja [10], it was observed that the approaches used emphasized more on the optic disc, while the diagnosis of glaucoma also involves the use of structural parameters such as CDR, DDLS and ISNT obtained using different segmentation approaches. Calculation of all these parameters is performed using the information of both optic disc and optic cup.
2.2 Deep learning-based approaches
Sevastopolsky [29] pre-processed the input retinal fundus image using contrast limited adaptive histogram equalization to improve the quality of the image. Segmentation of optic disc and optic cup was performed using modified U-net with a smaller number of filters to reduce the computational complexity. Thereafter, Fu et al. [30] presented a single-stage joint segmentation of optic disc and optic cup using convolutional neural network comprising of U-shaped convolutional network, multiscale input layer, multi-label loss function and side output layer. Also, Sun et al. [31] employed recurrent convolutional neural networks for the segmentation of optic disc from retinal fundus images and found that the performance of the proposed network was better than state of the art. Further, Chakravarty and Sivswamy [32] presented the joint segmentation of disc and cup using a multi-task convolutional neural network. Before segmentation, input images were cropped to extract the desired region of interest, and after segmentation thresholding followed by morphology was applied to refine the regions of segmented areas. Similarly, Edupuganti et al. [33] used a fully convolutional neural network (FCNN) to segment optic disc and optic cup. But the computational complexity of the network was very high. Thus, Wang et al. [34] recently in 2019 suggested U-net-based deep learning model for segmentation of optic disc from retinal images. Also, Juneja et al. [35] employed deep learning-based segmentation of optic disc and optic cup using Gnet and Kang et al. [36] used Deep-lab V3+ for segmentation of optic disc and optic cup .
3 Deep learning-based segmentation approaches
This section describes the implementation of various state-of-the-art approaches used for comparison with the proposed approach.
3.1 U-net
U-net is a CNN that was developed to segment biomedical images. U-net includes upsampling layers instead of maxpooling layers which increases the resolution of the output. There a large number of feature channels in the upsampling layers which allows the propagation of context information to the higher resolution layer through the network, this gives the architecture a U-shaped structure. U-net is made up of two parts—a contracting and an expanding path. The contracting part consists of the typical convolutional network, i.e. convolution layers, rectified linear unit (ReLU) and maxpooling layers. The contracting part results in a decrease in the spatial dimensions and an increase in feature information. On the other hand, the expanding part consists of upsampling layers and concatenation layers. The expanding part involves the combining of the spatial and feature information and concatenates with the high-resolution features from the contracting part [12].
3.2 Gnet
Gnet is a modified U-net architecture which can perform better than the traditional U-net in segmenting the optic disc and optic cup. The key feature of Gnet is the increase in the filter size from (3, 3) to (4, 4), in maxpool layer from (2, 2) to (4, 4) and in upsampling layer from (2, 2) to (4, 4). This leads to an increase in the total number of filters used in Gnet to 256, which is double the number of filters used in U-net. This increase in filter size leads to an increase in the accuracy of segmenting the optic disc and optic cup [35].
3.3 Deep-lab
Deep-lab is a model developed for semantic segmentation of the optic cup and optic disc. Deep-lab uses atrous convolutions to extract features from the images along with batch normalization layers to facilitate an efficient training process. The model also uses atrous spatial pyramid pooling to extract features from the images with long-range information. Deep-lab has an encoder–decoder structure, in which the encoder performs the above task and decoder refines the model results along with the boundaries. The output from the encoder–decoder structure is passed through a binary component analysis technique to get the final result. The final result is then used to calculate CDR [36].
4 Experimental setup and datasets used
This section presents the details of the environmental setup employed for the performance of experimentation of the proposed approach and state of the art for the comparison. Also, it emphasizes the details of the dataset used for analysis in the study.
4.1 Experimental setup
The segmentation approaches of the optic disc and optic cup in retinal fundus images used in this study were implemented on python version 3.7 and TensorFlow version 1.13.1. Furthermore, the execution was performed on the following hardware setup:
CPU: Two Intel® Xeon® CPU E5-2650 v4 @2.20 GHz
GPU: nVidia Titan Xp 12 GB GDDR5X
Memory: 256 GB ECC DDR4 RAM (32 GB × 8)
OS: KDE Neon
Nvidia Drivers v430 and CUDA Toolkit 10.1
4.2 Datasets
-
(i)
DRISHTI-GS The dataset comprises 101 retinal fundus images with 30 normal images and 71 glaucomatous images acquired using a retinal fundus camera. The ground truth for comparison of implemented approaches comprises of the ‘normal/abnormal’ labels and soft segmented maps of ‘disc/cup’ generated by the researchers of the IIIT Hyderabad in alliance with Aravind eye hospital in Madurai, India. It also includes a.txt file for each retinal image comprising of CDR values, which is a significant diagnostic parameter for glaucoma. Further, the images in the data repository are gathered from people of varying age groups visiting the hospital, with images acquired under varying brightness and contrast [37].
-
(ii)
RIM-One This dataset comprises of 166 retinal fundus images with labels “glaucoma” or “non-glaucoma” followed by ground truth of cup and disc. The images in the dataset are acquired under different brightness and contrast using Nidek AFC-210 retinal fundus camera. It includes 92 normal images and 74 abnormal images (glaucoma/suspect) [38].
5 Methodology
The approach used for the detection of glaucoma relies on precise segmentation of optic disc and optic cup. The fundus images available in the dataset were too large to be directly fed into the convolutional network; hence, cropping was used to reduce the size of the image. Further, as the training of network requires a large number of data, augmentation was performed to increase the number of images in the dataset. The images were then split into red, green and blue channels. Red channel was used for the segmentation of the optic disc due to better visibility of optic disc in the red channel as compared to green and blue channels. Whereas the blue channel was used for the segmentation of the optic cup due to the highest contrast of optic cup boundaries in this channel as compared to green and red channels. Thereafter, the images were fed into the CNN model as input and were trained to segment the optic disc and optic cup while passing through the different convolutional layers. Finally, the values of CDR, DDLS and ISNT were computed for diagnosis. Figure 2 shows the diagrammatic representation of the methodology used for the detection of glaucoma.
The detailed explanation of the procedural steps is as follows:
5.1 Pre-processing
The pre-processing was done to remove irrelevant details from the input data. Diagnosis of glaucoma with structural parameters such as CDR, DDLS and ISNT in retinal imaging involves an analysis of the pale yellowish region in the centre called the optic disc and depth of variable size called an optic cup. The acquired image comprises of extra irrelevant details leading to an increase in the size of an image, as a result of which computation time is increased. Also, the presence of retinal vessels results in the reduction in segmentation accuracy. Thus, the pre-processing of the retinal images included cropping to remove irrelevant details and reduce the size of the image, augmentation to increase the number of images in the dataset for training convolution network followed by separation of channels for clear visibility of optic disc and optic cup suppressing the retinal vessels.
5.2 Segmentation using DC-Gnet
The retinal fundus images input to the CNN model were initially cropped to 512 × 512 pixels and then were resized to 256 × 256 pixels, as the original size of the images were quite large, and CNN was not able to train due to the high-power computation. The images were then further normalized for easier computation. The DC-Gnet with 28 layers comprising of 2D convolutional layers, pooling layers, dropout layers and upsampling layers was used for segmentation of optic disc and optic cup from pre-processed images. The convolutional layers extracted features from the resized images using the convolutional filters. The images were then downsampled up to some extent without significant loss of data using a low pass filter in the convolutional layer. The use of low pass filter before downsampling in the convolutional layer reduces the high-frequency components, which creates attenuation and retains only the significant details for further processing. The model then built a binary mask using the features extracted by upsampling using the upsampling 2D layer. The model gives a binary image as an output of the input image. The logic output per pixel is itself combined by the model to give a binary image as an output. Also, the ground truth available in the datasets used for experimentation is in the form of a binary image.
Further, the training is performed in such a way that output is a binary mask, and thus the model outputs a binary image. The upsampling 2D layer maintained the connectivity in the model by doubling the dimensions of input from convolution 2D layer, thereby attaining the shape of the input image. Pooling layers and dropout layers were employed to drop features while training to prevent the model from overfitting. A dropout of 0.18 was used after every two convolutional 2D layers and after every upsampling layer. Also, the value of dropout was varied, but the best results were observed in this case. The layers were then activated by ReLU activation function. The output obtained was a binary mask of size 256 × 256 pixels.
The binary image was then masked and ANDed with the original cropped RGB fundus image. Thus, the two stages of segmentation, i.e. for optic disc and optic cup, were performed. Two-stage segmentation used the two different U nets for optic disc and optic cup, and the masking was used to improve the segmentation outputs. Also, it reduced the number of unwanted pixels and offered better visibility of the optic cup in the image achieved. Thus, masking helps to reduce the computation and removes the background pixels which do not contribute to the prediction of the optic cup. The original cropped RGB fundus image was then again split into red, green and blue channels. The blue channel was then used for the segmentation of optic cup since, due to clear visibility in this channel. The model was thus made to perform the same operations on the image as it did to get the binary mask of the optic disc, but this time the region of interest for the model was the optic cup. The binary image of the optic cup was then ANDed with the cropped RGB fundus image. Masking the original cropped fundus image with a binary mask of the optic cup creates a new RGB image. This RGB image is split into RGB channels, and the green channel is used to train another model to get the binary mask of the optic cup. This resulted in a binary image of the optic cup of size 256 × 256 pixels.
In conventional U-net, all conv2d layers had a filter size of (3, 3), whereas, in modified U-net the downsampling conv2d has a filter size of (4, 4). The upsampling layer also has a filter size of (2, 2) in conventional layers, whereas, in modified U-net, it has a filter size of (4, 4). The number of filters in the first conv2d layer is 32, 64 in the next 2 layers, and 128 in the further 2 layers. Maxpool has a pool size of (2, 2) in the conventional U-net, whereas, (4, 4) in the modified U-net. Larger filter size in modified U-net helps to extract more features from the image. As each layer adds a large number of parameters, a larger size of maxpool helps to downsample effectively. The dropout layer in the middle has been removed in the modified U-net as it leads to loss of information which further leads to underperformance of the model. Thus, only a single dropout was used in such a way that loss of data is minimal, and no significant data are lost. The use of low pass filter before downsampling in the convolutional layer reduces the high-frequency components, which creates attenuation and thus retains the significant details for further processing. Also, the larger filter size of the upsampling layer than the conventional U-net was used to get better performance. The DC-Gnet used for segmentation with the arrangement of different layers is shown in Fig. 3, and the block diagram for the same is given in Fig. 4.
5.3 Calculation of CDR, DDLS and ISNT
The predicted binary mask of disc and cup was then used to calculate the structural parameters CDR, DDLS and ISNT. These masks were then individually traversed either vertically or horizontally and width measures were noted. After getting the maximum or minimum width for each mask in each traversal, the axis of traversal was rotated by an angle equal to the value given by the precision angle (more is the precision angle, more is the accuracy) and the same process was repeated till the rotation of the image to 360 degrees. These measures were then inserted into Eqs. (1)–(5) to get values of CDR, DDLS and ISNT.
6 Performance metrics
The performance of the segmentation approaches used for an optic disc and optic cup was analysed using metrics such as dice similarity, Jaccard index and accuracy derived from the values of true positive (TP), false positive (FP), true negative (TN) and false negative (FN).
TP is defined as the region of image segmented as disc/cup and also belongs to disc/cup.
FP is defined as region of image segmented as disc/cup, but does not belong to disc/cup.
TN is defined as region of image not segmented as disc/cup, and also does not belong to disc/cup.
FN is defined as region of image not segmented as disc/cup, but belongs to disc/cup.
Here, the values of TP, FP, TN and FN are derived using background area (BA) of the segmented region, desired area (DA) of ground truth and segmented area (SA) of segmented region using the calculations in Table 1.
6.1 Dice similarity
Dice similarity (DS) is a measure of similarity between the segmented image and the ground truth. Higher is the value of DS, better is the performance of segmentation [39]. DS is calculated using Eq. (10):
6.2 Jaccard index
Jaccard index (JI) is the measure of dissimilarity between the segmented image and the ground truth. Lower value of JI signifies the better performance of segmentation [40]. JI is calculated using Eq. (11):
6.3 Accuracy
Accuracy (Acc) is the measure of preciseness or correctness of segmented image with respect to the ground truth. Higher is the value of Acc, better is the performance of segmentation [9]. Acc is calculated using Eq. (12):
7 Experimental results and discussions
This section emphasizes the performance of deep learning-based optic disc and optic segmentation for diagnosis of glaucoma using structural parameters such as CDR, DDLS and ISNT. The performance of DC-Gnet is compared with the U-net [9], Gnet [35] and Deep-lab [36] for segmentation. Performance metrics such as Dice similarity, Jaccard index and accuracy are used for comparison of the approaches.
7.1 Training and testing parameters
The experimental evaluation was performed on the total number of 267 images combined from both datasets. The original retinal images in the dataset were of dimensions 2896 × 1944 which were quite large to be given directly as an input to the deep neural networks. The input image was initially cropped to 512 × 512 pertaining targeted portion, i.e. optic disc in the centre and was further resized to 256 × 256 to reduce computational complexity. Cropping was performed manually in such a way that it included the region comprising of the optic disc and the outer boundaries within a distance of 1 cm approx. Cropping thus removed data irrelevant for glaucoma diagnosis, reduced the computation time and raised the accuracy. Computational time was reduced as the image without cropping was of bigger dimension, while the cropping reduced the dimension of the image. The approximate training time for the U-net is observed to be 9 min, Gnet is 12 min, Deep-lab is 15 min and DC-Gnet is 10 min.
On the other hand, segmentation accuracy was increased as the cropped image included only the desired region of interest (ROI) due to which better was the segmentation accuracy, and correct was the prediction. Also, it increased the visibility of ROI as a result of which segmentation was improved. While the image without cropping may include false regions as well which may result in inaccurate prediction. Figure 5 shows the original and the input cropped image in Rim-One and Drishti-GS dataset used in the study. The reduced dimension of the input image does not affect the optic disc region. The desired region of interest remains in the same proportion as that in the input image. The objective function of the training process is logarithmic dice error which calculates the loss of data while the optimization algorithm used to decrease the loss in data is adadelta.
In order to increase the number of images, the images in the dataset, as well as the binary masks, were augmented manually using custom augmentor. Several augmentation techniques such as zooming, width shifting, height shifting, rotation, vertical flipping and horizontal flipping were used. The images and the binary masks were zoomed in a range of 0.9–1.0, thereby increasing the size of the dataset. Then, the dataset size was further increased by shifting the width and height by 0.1. Furthermore, the images and binary masks were rotated by − 10° to 10°. Figure 6 shows the augmentation of images performed to increase the number of input images for the training of the proposed CNN model.
The cropped and augmented images were then split into three channels viz. red, green and blue. The red channel was then used for disc segmentation, as the disc is more clearly visible in the red channel as compared to other channels. While the blue channel was used for cup segmentation, due to clear most visibility of cup in this channel. The clear visibility of the optic disc and cup enables the neural network to extract maximum and the significant features which further leads to an increase in the performance of segmentation accuracy. Figure 7 shows the red, green and blue channels of the input RGB image.
Thus, the total number of 267 images combined from both the datasets was further increased to 1332 for experimental evaluation of deep learning approaches. The number of images was increased to 1332 as the deep learning-based approaches require training at large scale for correct prediction. Data augmentation increases the number of images for training, as a result of which “better is the training improved the accuracy of testing”. Thus, augmentation plays a significant role in the case of a smaller number of samples in the dataset. The training was performed on RIM-One dataset and testing on Drishti dataset. The same set of pre-processing comprising of cropping, augmentation and channel separation was applied on U-net, Gnet, Deep-lab and DC-Gnet to avoid biases in performance evaluation. Further, the learning rate of 0.41, batch size of 8, 60 number of epochs, ReLU activation for hidden layers and sigmoid activation for output layers with ada_delta optimizer was used.
7.2 Performance analysis of optic disc segmentation
Segmentation of optic disc has been performed using deep learning-based approaches such as U-net, Gnet, Deep-lab and DC-Gnet. Figure 8 shows the outputs of disc segmentation using different approaches followed by its quantitative analysis in Table 2 using metrics such as DS, JI derived from DA and SA, and accuracy derived from TP, FP, TN, FN (where TP, FP, TN, FN are further derived from BA, DA and SA).
Based on the experimental analysis, values of DS, JI, accuracy for optic disc segmentation using U-net are observed to be 0.77, 0.36, 0.81 for Drishti dataset with 0.63 TP, 0.36 FP, 1 TN, 0 FN and 0.94, 0.13, 0.96 for RIM-One with 1 TP, 0.02 FP, 0.97 TN, 0 FN. Also, for the segmentation of optic disc using Gnet the DS of 0.80, JI of 0.32 and accuracy of 0.83 is observed for Drishti dataset with 0.67 TP, 0 FP, 1 TN, 0.32 FN and the DS of 0.96, JI of 0.04 and accuracy of 0.98 is observed for RIM-One dataset with 0.95 TP, 0 FP, 1 TN, 0.04 FN. Further, the values of DS, JI, accuracy for optic disc segmentation using Deep-lab are found to be 0.73, 0.42, 0.78 for Drishti dataset with 0.57 TP, 0 FP, 1 TN, 0.42 FN and 0.91, 0.15, 0.93 for RIM-One with 1 TP, 0 FP, 0.8 TN, 0.13 FN. Finally, for the optic disc segmentation using DC-Gnet, DS of 0.88, JI of 0.12, accuracy of 0.93 is observed for Drishti dataset with 0.87 TP, 0 FP, 1 TN, 0.12 FN and DS of 0.98, JI of 0.03, accuracy of 0.99 is observed for RIM-One dataset with 1 TP, 0.007 FP, 0.99 TN, 0 FN.
7.3 Performance analysis of optic cup segmentation
Similar to optic disc segmentation, the optic cup is also extracted using deep learning-based approaches such as U-net, Gnet, Deep-lab and Modified U-net. Figure 9 shows the outputs of cup segmentation using different approaches followed by its quantitative analysis in Table 3 using metrics such as DS, JI derived from DA and SA, and accuracy derived from TP, FP, TN, FN (where TP, FP, TN, FN are further derived from BA, DA and SA).
Similar to the experimental analysis of optic disc, values of DS, JI, accuracy for optic cup segmentation using U-net are observed to be 0.63, 0.62, 0.79 for Drishti dataset with 1 TP, 0.25 FP, 0.99 TN, 0 FN and 0.79, 0.38, 0.943 for RIM-One with 1 TP, 0.01 FP, 0.98 TN, 0 FN. Also, for the segmentation of optic cup using Gnet the DS of 0.73, JI of 0.41 and accuracy of 0.81 is observed for Drishti dataset with 0.58 TP, 0 FP, 1 TN, 0.41 FN and the DS of 0.89, JI of 0.26 and accuracy of 0.96 is observed for RIM-One dataset with 0.96 TP, 0.03 FP, 1 TN, 0 FN. Further, the values of DS, JI, accuracy for optic cup segmentation using Deep-lab are found to be 0.42, 0.72, 0.69 for Drishti dataset with 0.23 TP, 0 FP, 1 TN, 0.76 FN and 0.69, 0.41, 0.94 for RIM-One with 1 TP, 0.11 FP, 0.88 TN, 0 FN. Finally, for the optic cup segmentation using DC-Gnet, DS of 0.84, JI 0.36, accuracy of 0.90 is observed for Drishti dataset with 0.63 TP, 0 FP, 1 TN, 0.36 FN and DS of 0.91, JI of 0.19, accuracy of 0.978 is observed for RIM-One dataset with 1 TP, 0.002 FP, 0.91 TN, 0.19 FN.
Further, the approximate execution time for U-net is observed to be 2.8 s, Gnet is 3.2 s, Deep-lab is 3.5 s and DC-Gnet is 3 s. It can be observed from Figs. 8 and 9 that U-net performs over-segmentation, while Gnet performs under-segmentation and the Deep-lab gives coarse/incomplete segment boundaries. The inaccuracies in the state-of-the-art models were observed due to overfitting. To overcome this, the DC-Gnet increased the size of kernel in case of downsampling and decreased in case of upsampling to avoid overfitting of the model. Also, the size of the pool was increased in case of maxpooling layer for refined edges of disc and cup. Thus, it can be stated both qualitatively and quantitatively that the DC-Gnet outperforms U-net, Gnet and Deep-lab for segmentation of optic disc and optic cup with highest values of DS, accuracy and least value of JI.
8 Conclusion
The paper presented a deep learning-based DC-Gnet for segmentation of optic disc and optic cup, followed by extraction of structural features namely cup-to-disc ratio (CDR), disc damage likelihood scale (DDLS) and inferior superior nasal temporal (ISNT) regions for diagnosis of glaucoma. As, the traditional approaches employed for diagnosis were manual time-consuming, and offers diagnosis often on the values CDR values. This approach offers improved automated diagnosis, additionally based on DDLS and ISNT. Based on the experimental evaluations, the performance of DC-Gnet is found to outperform other deep learning-based models such as U-net, Gnet and Deep-lab with DS of 0.84, JI of 0.36, accuracy of 0.90 for Drishti dataset with 0.63 TP, 0 FP, 1 TN, 0.36 FN and DS of 0.91, JI of 0.19, accuracy of 0.978 for RIM-One dataset with 1 TP, 0.002 FP, 0.91 TN, 0.19 FN. As the performance of segmentation on Drishti dataset was less remarkable than RIM-One due to training solely in RIM-One dataset. Future efforts could be to train the proposed model on varied images to improve the performance on other datasets and make it generalize for all fundus images.
References
Foster, P.J., Buhrmann, R., Quigley, H.A., Johnson, G.J.: The definition and classification of glaucoma in prevalence surveys. Br. J. Ophthalmol. 86, 238–242 (2002)
Saxena, R., Singh, D., Vashist, P.: Glaucoma: an emerging peril. Indian J. Community Med. 38, 135–137 (2013)
Glaucoma Research foundation: Five Common Glaucoma Tests. https://www.glaucoma.org/glaucoma/diagnostic-tests.php. (Available 4 April 2019). (Accessed 10 May 2019)
Yamada, S., Komatsu, K., Ema, T., Inventors; Toshiba Corp, Assignee: Computer-aided diagnosis system for medical use. United States Patent US 5,235,510 (1993)
Abràmoff, M.D., Garvin, M.K., Sonka, M.: Retinal imaging and image analysis. IEEE Rev. Biomed. Eng. 3, 169–208 (2010)
Garway-Heath, D.F., Ruben, S.T., Viswanathan, A., Hitchings, R.A.: Vertical cup/disc ratio in relation to optic disc size: its value in the assessment of the glaucoma suspect. Br. J. Ophthalmol. 82(10), 1118–1124 (1998)
Harizman, N., Oliveira, C., Chiang, A., Tello, C., Marmor, M., Ritch, R., Liebmann, J.M.: The ISNT rule and differentiation of normal from glaucomatous eyes. Arch. Ophthalmol. 124(11), 1579–1583 (2006)
Henderer, J.D., Liu, C., Kesen, M., Altangerel, U., Bayer, A., Steinmann, W.C., Spaeth, G.L.: Reliability of the disk damage likelihood scale. Am. J. Ophthalmol. 135(1), 44–48 (2003)
Khalid, N.E.A., Noor, N.M., Ariff, N.M.: Fuzzy c-means (FCM) for optic cup and disc segmentation with morphological operation. Procedia Comput. Sci. 42, 255–262 (2014)
Thakur, N., Juneja, M.: Survey on segmentation and classification approaches of optic cup and optic disc for diagnosis of glaucoma. Biomed. Signal Process. Control 1(42), 162–189 (2018)
Gentle Dive into Math Behind Convolutional Neural Networks: https://towardsdatascience.com/gentle-dive-into-math-behind-convolutional-neural-networks-79a07dd44cf9. (Available 13 April 2019). (Accessed 17 July 2019)
Ronneberger, O., Fischer, P., Brox, T.: U-net: convolutional networks for biomedical image segmentation. In: International Conference on Medical Image Computing and Computer-Assisted Intervention, pp. 234–241. Springer, Cham (2015)
Walter, T., Klein, J.C.: Segmentation of color fundus images of the human retina: detection of the optic disc and the vascular tree using morphological techniques. In: Springer Proceedings of International Symposium on Medical Data Analysis, pp. 282–287 (2015)
Pallawala, P.M.D.S., Hsu, W., Lee, M.L., Eong, K.G.A.: Automated optic disc localization and contour detection using ellipse fitting and wavelet transform. In: Springer Proceedings of European Conference on Computer Vision, pp. 139–151 (2004)
Abdel-Ghafar, R.A., Morris, T.: Progress towards automated detection and characterization of the optic disc in glaucoma and diabetic retinopathy. Med. Inform. Internet Med. 32(1), 19–25 (2007)
Liu, J., Wong, D.W.K., Lim, J.H., Jia, X., Yin, F., Li, H., Xiong, W., Wong, T.Y.: Optic cup and disk extraction from retinal fundus images for determination of cup-to-disc ratio. In: IEEE Proceedings of 3rd Conference on Industrial Electronics and Applications, pp. 1828–1832 (2008)
Zhang, Z., Liu, J., Wong, W.K., Tan, N.M., Lim, J.H., Lu, S., Li, H., Liang, Z., Wong, T.Y.: Neuro-retinal optic cup detection in glaucoma diagnosis. In: IEEE Proceedings of 2nd International Conference on Biomedical Engineering and Informatics, pp. 1–4 (2009)
Sarkar, D., Das, S.: Automated glaucoma detection of medical image using biogeography based optimization. In: Proceedings of Advances in Optical Science and Engineering, pp. 381–388. Springer, Singapore (2017)
Priyadharsini, R., Beulah, A., Sharmila, T.S.: Optic disc and cup segmentation in fundus retinal images using feature detection and morphological techniques. Curr. Sci. 115(4), 748 (2018)
Osareh, A., Mirmehdi, M., Thomas, B., Markham, R.: Comparison of colour spaces for optic disc localisation in retinal images. In: Proceedings of 16th International Conference on Pattern Recognition, pp. 743–746 (2002)
Chrastek, R., Wolf, M., Donath, K., Niemann, H., Paulus, D., Hothorn, T., Lausen, B., Lämmer, R., Mardin, C.Y., Michelson, G.: Automated segmentation of the optic nerve head for diagnosis of glaucoma. Med. Image Anal. 9(4), 297–314 (2005)
Blanco, M., Penedo, M.G., Barreira, N., Penas, M., Carreira, M.J.: Localization and extraction of the optic disc using the fuzzy circular Hough transform. In: Proceedings of International Conference on Artificial Intelligence and Soft Computing, pp. 712–721 (2006)
Datt, J.G., Sivaswamy, J., Krishnadas, S.R.: Optic disk and cup segmentation from monocular color retinal images for glaucoma assessment. IEEE Trans. Med. Imaging 30(6), 1192–1205 (2011)
Yin, F., Liu, J., Wong, D.W.K., Tan, N.M., Cheung, C., Baskaran, M., Aung, T., Wong, T.Y.: Automated segmentation of optic disc and optic cup in fundus images for glaucoma diagnosis. In: Proceedings of 25th International Symposium on Computer-Based Medical Systems, pp. 1–6 (2012)
Xu, Y., Lin, S., Wong, D.W., Liu, J., Xu, D.: Efficient reconstruction-based optic cup localization for glaucoma screening. In: Proceedings of International Conference on Medical Image Computing and Computer-Assisted Intervention, pp. 445–452 (2013)
Cheng, J., Yin, F., Wong, D.W., Tao, D., Liu, J.: Sparse dissimilarity-constrained coding for glaucoma screening. IEEE Trans. Biomed. Eng. 62(5), 1395–1403 (2015)
Kavitha, S., Karthikeyan, S., Duraiswamy, K.: Early detection of glaucoma in retinal images using cup to disc ratio. In: Proceedings of International Conference on Computing Communication and Networking Technologies, pp. 1–5 (2010)
Mittapalli, P.S., Kande, G.B.: Segmentation of optic disk and optic cup from digital fundus images for the assessment of glaucoma. Biomed. Signal Process. Control 24, 34–46 (2016)
Sevastopolsky, A.: Optic disc and cup segmentation methods for glaucoma detection with modification of U-Net convolutional neural network. Pattern Recognit. Image Anal. 27(3), 618–624 (2017)
Fu, H., Cheng, J., Xu, Y., Wong, D.W., Liu, J., Cao, X.: Joint optic disc and cup segmentation based on multi-label deep network and polar transformation. IEEE Trans. Med. Imaging 37(7), 1597–1605 (2018)
Sun, X., Xu, Y., Zhao, W., You, T., Liu, J.: Optic disc segmentation from retinal fundus images via deep object detection networks. In: 2018 40th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), pp. 5954–5957. IEEE (2018)
Chakravarty, A., Sivswamy, J.: A deep learning based joint segmentation and classification framework for glaucoma assesment in retinal color fundus images. arXiv preprint arXiv:1808.01355 (2018)
Edupuganti, V.G., Chawla, A., Kale, A.: Automatic optic disk and cup segmentation of fundus images using deep learning. In: 2018 25th IEEE International Conference on Image Processing (ICIP), pp. 2227–2231. IEEE (2018)
Wang, L., Liu, H., Zhang, J., Chen, H., Pu, J.: Automated segmentation of the optic disc using the deep learning. In: Medical Imaging 2019: Image Processing, vol. 10949, p. 1094923. International Society for Optics and Photonics (2019)
Juneja, M., Singh, S., Agarwal, N., Bali, S., Gupta, S., Thakur, N., Jindal, P.: Automated detection of glaucoma using deep learning convolution network (G-net). Multimed. Tools Appl. (2019). https://doi.org/10.1007/s11042-019-7460-4
Kang, H., Wang, K., Guo, S., Gao, Y., Li, N., Weng, J., Li, X., Li, T.: Pixel quantification for robust segmentation of optic cup (2019). https://grand-challenge-public.s3.amazonaws.com/f/challenge/229/b14c03ff-a260-4272-b5ee-67e4b695f14a/REFUGE-NKSG-slides.pdf
Sivaswamy, J., Krishnadas, S.R., Joshi, G.D., Jain, M., Tabish, A.U.: Drishti-GS: retinal image dataset for optic nerve head (ONH) segmentation. In: 2014 IEEE 11th International Symposium on Biomedical Imaging (ISBI), pp. 53–56. IEEE (2014)
Fumero, F., Alayón, S., Sanchez, J.L., Sigut, J., Gonzalez-Hernandez, M.: RIM-One: an open retinal image database for optic nerve evaluation. In: 2011 24th International Symposium on Computer-Based Medical Systems (CBMS), pp. 1–6. IEEE (2011)
Jackson, D.A., Somers, K.M., Harvey, H.H.: Similarity coefficients: measures of co-occurrence and association or simply measures of occurrence. Am. Nat. 133(3), 436–453 (1989)
Ferdous, R.: An efficient k-means algorithm integrated with Jaccard distance measure for document clustering. In: 2009 First Asian Himalayas International Conference on Internet, pp. 1–6. IEEE (2009)
Acknowledgements
The authors are also grateful to the Ministry of Human Resource Development (MHRD), Govt. of India for funding this Project (17-11/2015-PN-1) under Design Innovation Centre (DIC) sub-theme Medical Devices & Restorative Technologies.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Juneja, M., Thakur, S., Wani, A. et al. DC-Gnet for detection of glaucoma in retinal fundus imaging. Machine Vision and Applications 31, 34 (2020). https://doi.org/10.1007/s00138-020-01085-2
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00138-020-01085-2